Abstract
Mesenchymal stem cells (MSCs) represent a promising source of cell-based therapies for treatment of a wide variety of injuries and diseases. Their tropism and migration to the damaged sites, which are elicited by cytokines secreted from tissues around pathology, are the prerequisite for tissue repair and regeneration. Better understanding of the elicited-migration of MSCs and discovering conditions that elevate their migration ability, will help to increase their homing to pathologies and improve therapeutic efficacy. It is increasingly recognized that microRNAs are important regulators of cell migration. Here we summarize current understanding of the microRNA-regulated migration of MSCs.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Small non-coding RNAs (miRNAs), typically 21~22 nucleotide-long and evolutionarily conserved, are predicted to regulate about 30% of protein-coding genes by binding to the 3′untranslated region (3′UTR) of target mRNAs particularly at the 5′end seed region, resulting in translational suppression or degradation of target transcripts, thereby participating in the regulation of almost every cellular process and emerging as master regulators of cell function [1,2,3].
By global profiling, miRNAs have been demonstrated to selectively express in MSCs of different origins [4,5,6,7] or under differentiation [8,9,10,11,12], and those play critical roles in MSC self-renewal, proliferation, and differentiation have been extensively and well reviewed [13,14,15,16,17,18]. However, miRNAs that regulate the chemotactic migration of MSCs have not been summarized.
Cell migration is a highly regulated multi-step process, initiated by extracellular growth factors or chemokines, and followed by the continuous formation and turnover of cell-to-matrix adhesions, the persistent protrusion, retraction, and rearrangement of cytoskeleton [19]. MiRNAs have been found to regulate the migration of a wide variety of cell types through direct modulation of extracellular matrix (ECM), cell adhesion, cytoskeleton rearrangement, and cell signaling cascades [20]. Here, we review current knowledge of miRNA’s regulatory roles in MSC migration and the underlying mechanisms.
MiRNAs Regulate Cell Migration by Targeting Cytokine/Receptor Axis
Cytokines secreted from tissues of pathology, including growth factors and chemokines, have been demonstrated to stimulate the migration of MSCs in vitro and induce their pathotropism in vivo by interacting with their receptors and thereby activating downstream migration-related signaling cascades. Stromal cell-derived factor (SDF)-1 and its receptor chemokine receptor type 4 (CXCR4) play important role in stem cell migration and mobilization. Knockdown of either SDF-1α or CXCR4 or inhibition of CXCR4 with antagonist significantly decreases the migration ability of MSCs [7, 21,22,23]. Multiple miRNAs, including miR-27b, miR-27a, miR-146a-5p and miR-886-3p, have been reported to affect the migration of MSCs through targeting SDF-1α/CXCR4 axis.
MiR-27b and miR-27a were found to be significantly downregulated in burned skin of bone marrow-chimeric mice and in CCl4-injured livers of mice, while the expression of their potential target SDF-1α continuously increased after injury. Dual luciferase reporter assay demonstrated the direct binding of miR-27b and miR-27a with SDF-1α 3′UTR, suggesting that SDF-1α is a direct target of miR-27b and miR-27a. Transwell migration assay indicated that fewer MSCs migrated to the lower well containing suspensions of miR-27b- or miR-27a-overexpressing MSCs, in which the content of SDF-1α was significantly decreased [24, 25]. These results suggest that downregulation of miR-27b and miR-27a upon injury, leading to the increase of SDF-1α, may contribute to the mobilization of MSCs, thereby promoting wound healing and tissue repair.
MiR-146a-5p was significantly overexpressed and had higher abundance in human umbilical cord Wharton’s jelly derived MSCs (hWJ-MSCs), which showed poorer motility yet better proliferation compared to bone marrow derived MSCs (hBM-MSCs). A 50% downregulation of miR-146a-5p in hWJ-MSCs resulted in over 3 folds increase of cell migration, while overexpression of miR-146a-5p in hBM-MSCs resulted in significant decrease of cell motility. CXCL12 (SDF-1) and SIKE1, an I-kappa-B kinase epsilon (IKKε) suppressor that inhibits the activity of NFκB, were direct targets of miR-146a-5p, and knockdown of either CXCL12 or SIKE1 significantly decreased the migration of hWJ-MSCs. Moreover, expression of miR-146a-5p was induced by binding of NFκB to its promoter, but was inhibited by CXCL12. These results suggest a positively self-regulated expression of miR-146a-5p in hWJ-MSCs through targeting CXCL12 and SIKE1 [7].
In two human marrow stromal cell lines HS5 and HS27A distinguished by function and expression of SDF-1 (CXCL12, CXCL12+ or CXCL12−), miR-886-3p was found to express >40 fold more in CXCL12− cells than in CXCL12+ cells, and overexpression of miR-886-3p in CXCL12+ cells resulted in as much as 85% decreased CXCL12 expression and loss of CXCL12-directed chemotaxis, suggesting that miR-886-3p netatively regulates the chemotactic migration of hMSCs through inhibition of SDF-1/CXCR4 axis [26].
In addition to regulate cell migration by targeting cytokines/receptors upstreamly, miRNAs are also found to affect cell migration at downstream of cytokines/receptors. Upon stimulation with cytokines, some miRNAs differentially express and regulate cell migration through diverse mRNA targets [27,28,29,30,31]. Using a miRNA microarray containing 679 oligonucleotide probes, we detected 18 miRNAs exhibited ≥2 fold upregulation and 8 miRNAs exhibited >2 fold downregulation in rat bone marrow derived MSCs (rBM-MSCs) stimulated with hepatocyte growth factor (HGF) at 25 ng/ml for 24 h (Table 1); further, we demonstrated that miR-221 and miR-26b, two HGF-upregulated miRNAs, promoted the migration of MSCs [31], while miR-375, which was downregulated by HGF, negatively regulated the migration of MSCs [34].
Interferon-γ (IFN-γ) treatment of human umbilical cord derived MSCs led to the upregulation of 5 miRNAs and downregulation of 3 miRNAs (Table 1), and bioinformatics analysis indicated a majority of targets involved in signal transduction and cell migration [32]. IFN-γ downregulated the expression of miR-335 in human MSCs derived from bone marrow, adipose tissue and articular cartilage, and downregulation of miR-335 promoted the migration of MSCs, but the direct targets involved require further elucidation [29].
Overnight treatment of foreskin derived MSCs with a pro-inflammatory cytokine cocktail containing IL-1β (25 ng/ml), TNF-α (50 ng/ml), IFN-α (3000 U/ml or 10 ng/ml) and IFN-γ (1000 U/ml or 50 ng/ml) upregulated 5 miRNAs and downregulated 20 miRNAs (Table 1), and DIANA-miRPath v.3.0 analysis revealed significantly enriched Kyoto encyclopedia of genes and genomes (KEGG) pathways targeted by miR-27a-3p, miR-145, miR-199a-5p, miR-194-5p, miR-221-3p, miR-423-5p, miR-485-5p, and miR-155, suggesting their strongly regulatory roles in inflammation-primed cell processes, including cell migration [33]. Yet how these miRNAs regulate MSC migration and the direct transcript targets involved require further study.
MiRNAs Regulate Cell Migration by Targeting Components of Cell Signaling Pathway
Cytokines/receptors stimulate cell migration by activating signaling pathways, such as the mitogen-activated protein kinases (MAPKs), phosphatidylinositol 3-kinase (PI3K)/Akt, and Wnt/β-catenin [35, 36]. A variety of miRNAs have been demonstrated to regulate the migration of MSCs by targeting signaling components [30, 31, 37].
MiRNAs and PI3K/Akt Signaling Pathway
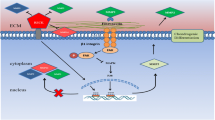
Upon activation by cytokine/receptor interaction (Fig. 1a), PI3K converts phosphatidylinositol 4, 5-biphosphate (PIP2) into phosphatidylinositol 3, 4, 5-triphosphate (PIP3), which binds both Akt and 3-phosphoinositide-dependent protein kinase 1 (PDK1), allowing PDK1 to phosphorylate Akt at Thr308 [38]. Full activation of Akt also requires its phosphorylation at Ser473, which is catalyzed by mammalian target of rapamycin complex 2 (mTORC2). Phosphorylation defect of either Thr308 or Ser473 will decrease the activity of Akt [39,40,41,42]. Phosphatase and tensin homologue deleted on chromosome ten (PTEN) opposes the function of PI3K by dephosphorylating PIP3, leading to the inactivation of Akt [43].
MiRNAs regulate the migration of MSCs through targeting cytokines/receptors and components of PI3K/Akt signaling pathway or interaction with MAPK signaling pathways. a Interaction of cytokines and receptors activates PI3K, which converts PIP2 into PIP3. PIP3 binds both PDK1 and Akt, allowing PDK1 to phosphorylate Akt at Thr308. Full activation of Akt also requires its phosphorylation at Ser473, which is catalyzed by mTORC2. PTEN inhibits the activation of Akt through dephosphorylating PIP3 and opposing the action of PI3K. b Interaction of cytokines and receptors induces a kinase cascade, sequentially MAP3Ks (MAPKKKs) and MAP2Ks (MAPKKs), which culminates the activation of three major groups of MAPKs, SAPK/JNK, p38MAPK, and ERK1/2, and promotes cell migration
Akt activation is required for HGF-induced migration of neural differentiating MSCs, and inhibition of PI3K/Akt signaling pathway by LY294002 significantly impairs directional migration of MSCs [44]. MiR-221 and miR-26b, two HGF-upregulated miRNAs, promoted the migration of MSCs by targeting PTEN and leading to the activation of Akt [31]. MiR-375, downregulated by HGF for about 70% [31], negatively regulated MSC migration through decreasing the expression of PDK1 and leading to the inactivation of Akt [34]. Although ERK1/2, SAPK/JNK, and p38MAPK, three major groups of MAPKs, are also reported to mediate HGF-elicited migration of neural differentiating MSCs [44], it seems less likely that they are involved in miR-26b-, miR-221-, and miR-375-regulated migration of MSCs [31, 34]. The above studies suggest that, upon HGF stimulation, downregulation of miR-375 and upregulation of miR-26b and miR-221 coordinate the activation of Akt signaling pathway, thereby promoting the chemotactic migration of MSCs.
MiRNAs and MAPK Signaling Pathways
Conventional MAPKs (Fig. 1b), including extracellular signal-reugalted kinase 1 and 2 (ERK1/2), c-Jun N-terminal kinases (JNK)/stress activated protein kinases (SAPK), and the p38 isoforms (p38MAPK), are activated by upstream kinases, referred to as MAPK kinases (MAPKKs or MAP2Ks). MAPKKs are, in turn, activated by their upstream kinases, referred to as MAPKKKs or MAP3Ks. MAPK signaling pathways have been demonstrated to mediate the chemotactic migration of MSCs [44], however, their interactions with miRNAs remain largely unkwown.
Precondition of MSCs with multiple myeloma cell lines was found to elevate cell migration in a JNK/SAPK- and ERK1/2-dependent manner, and downregulation of miR-199b and miR-125a by preconditioning also contributed to the elevated migration of MSCs [45]. However, the interrelationship between activation of MAPKs and downregulation of miR-199b/miR-125a in mediating the migration of MSCs remain further elusive.
MiRNAs and Wnt/β-catenin Signaling Pathway
Wnt signaling pathways, including the canonical Wnt/β-catenin signaling pathway (Fig. 2) and the non-canonical Wnt signaling pathways that use downstream effectors other than β-catenin, are important regulators of cell migration [46,47,48,49,50,51,52,53]. The canonical Wnt pathway is triggered by the interaction of Wnt with its receptor Frizzled (FZD) and coreceptor low-density lipoprotein receptor-related protein (LRP5 or LRP6). Upon activation, the receptor inhibits the activity of the destruction complex and allows the accumulation of β-catenin, which then translocates to the nuclear, where it associates with the transcription factor T cell factor/lymphoid enhancer factor (TCF/LEF) and activates the transcription of downstream target genes, thereby regulating cell migration. The destruction complex consists of the cytosolic scaffold protein axin, adenomatous polyposis coli (APC), casein kinase 1α (CK1α) and glycogen synthase kinase 3β (GSK3β), and acts as negative regulator of canonical Wnt pathway through phosphorylation of β-catenin at Ser45 and Ser33/Ser37/Thr41 sequentially by CK1α and GSK3β, which subsequently leads to the ubiquitination and degradation of β-catenin and keeps the free β-catenin in the cytosol at very low level (Fig. 2). Accumulating studies demonstrate that components of canonical Wnt/β-catenin signaling pathways are involved in miRNA-regulated cell migration [30, 37, 54].
MiRNAs regulate the migration of MSC through targeting the components of canonical Wnt/β-catenin signaling pathway. a Inactivated Wnt/β-catenin signaling. β-catenin is tracted by the destruction complex, which consists of the cytosolic scaffold protein axin, adenomatous polyposis coli (APC), casein kinase 1α (CK1α) and glycogen synthase kinase 3β (GSK3β), and is phosphorylated at Ser45 and Ser33/Ser37/Thr41 sequentially by CK1α and GSK3β. Subsequently the phospho-β-catenin is ubiquitinated by β-TrCP and degraded, keeping the free β-catenin in cytosol at very low level. Therefore, CK1α and GSK3β act as negative regulators of Wnt/β-catenin signaling pathway. b Wnt/β-catenin signaling pathway is triggered by interaction of Wnt with its receptor Frizzled and coreceptor LRP5/LRP6, which allows β-catenin to accumulate in cytosol through inhibiting the activity of destruction complex. β-catenin then translocates to the nuclear, where it associates with transcription factor T cell factor/lymphoid enhancer factor (TCF/LEF) and activates the transcription of downstream target genes, and regulates cell migration
MiR-124 has been reported to suppress the migration of MSCs toward HGF through inhibition of Wnt/β-catenin signaling pathway, and FZD4/LRP6, the receptor/co-receptor of Wnt, are direct targets of miR-124 involved in this process [30]. Overexpression of miR-124 significantly suppressed the activity of Wnt/β-catenin signaling, as evidenced by the decrease of Wnt3a-induced TOPflash activity, the level of active β-catenin, the nuclear translocation of β-catenin, and the transcription of downstream target genes c-Myc and RUNX2. In contrary, inhibition of endogenous miR-124 increased the level of active β-catenin and enhanced the nuclear translocation of β-catenin, indicating the activation of Wnt/β-catenin signaling. The decreased chemotactic migration of MSCs by miR-124 could be rescued by LiCl and the constitutively active β-catenin mutant (ΔN89 β-catenin), which ensures the activation of Wnt/β-catenin signaling, while the enhanced chemotactic migration of MSCs by miR-124 inhibitor could be abolished by FH535, the specific inhibitor of Wnt/β-catenin signaling. These results clearly demonstrate that miR-124 suppresses the chemotactic migration of MSCs through inhibition of Wnt/β-catenin signaling pathway.
MiR-9-5p has recently been reported to promote the chemotactic migration of MSCs toward HGF through activating β-catenin signaling pathway, and CK1α and GSK3β, two negative regulators of active β-catenin, are direct targets of miR-9-5p involved in this process [37]. The enhanced chemotactic migration of MSCs by miR-9-5p was significantly decreased by inhibition of β-catenin signaling with FH535, while the suppressed chemotactic migration of MSCs by miR-9-5p inhibitor was rescued by activation of β-catenin signaling with LiCl. These results clearly suggest that miR-9-5p promotes the chemotactic migration of MSCs through activating β-catenin signaling. In addition to act through β-catenin signaling pathway, direct inhibition of GSK3β by inhibitors LiCl, SB-415286, and AR-A014418 is also found to enhance the transmembrane migration of MSCs through increasing the expression of migration-related proteins including CXCR4 and phospho-β-PAK-interacting exchange factor (PIX) [55]. Moreover, GSK3β, as one of the few master switch kinases with multiple functions, has been found to influence cell migration through direct regulation of the spatiotemporally controlled dynamics of actin cytoskeleton, microtubules, and cell-to-matrix adhesions [52, 56,57,58]. It would be interesting to interpret how GSK3β, β-catenin signaling pathway, and other migration-related signaling proteins such as phospho-β-PIX and CXCR4 coordinate the regulation of cytoskeletons and cell-to-matrix adhesions, and affect cell migration.
In addition to affect the activity of Wnt/β-catenin signaling pathway upstreamly, miRNAs might also regulate cell migration at downstream of Wnt/β-catenin. LiCl or Wnt3a, which activates the canonical Wnt signaling pathway, upregulated the expression of miR-335 in human MSCs, while DKK1, a specific inhibitor of canonical Wnt signaling, downregulated the expression of miR-335 in the presence of exogenous Wnt3a [29]. Upregulation of miR-335 was found to inhibit the migration of human MSCs [29], consistent with the finding that Wnt3a inhibited the invasion of murine MSCs [59], but inconsistent with the reports that activation of canonical Wnt/β-catenin signaling pathway by LiCl and Wnt3a enhanced the migration [37, 55] and invasion [60] of human and rat MSCs. These discrepancies suggest a complex and delicate function of Wnt/β-catenin signaling pathway in MSC migration, which requires further investigation.
The best-characterized noncanonical Wnt signaling pathways are the planar cell polarity (PCP) pathway and the Wnt/Ca2+ pathway [48]. In the PCP pathway, FZD receptors activate a cascade consisting of the small GTPases Rac1, cdc42, Rho, and downstream protein kinases JNK or Rho kinase, thereby regulating cytoskeleton rearrangement and cell migration. In the Wnt/Ca2+ pathway, FZD receptors activate heterotrimeric G proteins, which in turn activate phospholipase C (PLC) and trigger Ca2+ release from intracellular stores, thereby activating effectors controlling cell migration. However, little is known at present about the function of noncanonical Wnt signaling pathways in MSC migration and their interactions with miRNAs during this process.
MiRNAs Regulate MSC Migration through Affecting Cell-to-Matrix Adhesions
Cell-to-matrix adhesions (Fig. 3), also termed focal adhesions (FAs), are formed by a surprisingly diverse and large number of proteins, including transmembrane proteins (e.g. integrins and E-cadherins), cytoskeletal proteins (e.g. paxillin, tensin, vinculin, α-actinin, and talin), tyrosine kinases such as focal adhesion kinase (FAK) and Src, modulators of small GTPases, tyrosine phosphatases, and other enzymes such as PI3K and the protease calpain II [61, 62]. Among these molecules, FAK is considered a key regulator of the assembly and disassembly (turnover) of FAs depending on its tyrosine phosphorylation status especially at Tyr397 [63]. Reduced tyrosine phosphorylation of FAK results in larger and more stable FAs, thereby slowing down the turnover of FAs and suppressing cell migration [63, 64]. Accumulating evidence indicates that miRNAs regulate cell migration through affecting the turnover of FAs [20].
MiRNAs regulate the migration of MSCs through affecting cell-matrix adhesions and actin cytoskeleton. Cell-matrix adhesions are formed by a large number of proteins, including transmembrane proteins such as integrins, cytoskeletal proteins such as paxillin and α-actinin, and tyrosine kinases such as FAK and Src, among which FAK is an essential regulator of the turnover of cell-matrix adhesions depending on its tyrosine phosphorylation status
In MSCs, upregulation of miR-26b, miR-221, and miR-9-5p promoted the formation of dot-like and peripherally distributed FAs [31, 37], which are more dynamic than larger and centrally distributed FAs, indicating that these miRNAs might promote cell migration through increasing the dynamics of FAs. In contrary, upregulation of miR-124 and miR-375 decreased the number and peripheral distribution of FAs in MSCs, suggesting that miR-124 and miR-375 might suppress cell migration through inhibiting the assembly and remodeling of FAs [30, 34]. FAK phosphorylation at Tyr397 was upregulated by miR-26b and miR-221 [31], but downregulated by miR-124 and miR-375 [30, 34], consistent with the aforementioned studies about FAK’s regulation of FA turnover and cell migration [63, 64]. Treatment of MSCs overexpressing miR-221 or miR-26b with LY294002, the well-known inhibitor of PI3K/Akt signaling pathway, decreasesd the phosphorylation of FAK at Tyr397 and the formation of dot-like FAs at cell periphery, indicating the crosstalk between PI3K/Akt and FAK signaling pathways in miR-221- and miR-26b-regulated migration of MSCs [31], yet the detailed mechanism requires further elucidation.
MiR-10b was reported to promote the migration of mouse bone marrow-derived MSCs by targeting E-cadherin [65], a mediator of cell-cell adhesions involved in cell migration [66], suggesting that miR-10b might promote MSC migration through reducing cell-cell adhesions.
Conditioned medium collected from rBM-MSCs under hypoxic condition for 12 h significantly downregulated the expression of miR-221 in MSCs, leading to the upregulation of its direct target intercellular adhesion molecule-1 (ICAM-1), and induced the transfilter migration of MSCs and promoted their adhesion and spreading [67]. These results indicate that miR-221 negatively regulate the migration of MSCs toward hypoxic conditioned medium, which is contrary to our previous report that miR-221 promoted the migration of MSCs toward HGF [31]. The inconsistency may be related to the difference of factors that induce cell migration, suggesting that a miRNA may lead to distinct migration behavior of MSCs under different conditions.
MiRNAs Regulate MSC Migration through Affecting Cytoskeleton Remodeling
The migration of cells is driven by continuous reorganization of actin cytoskeleton, which generate the forces of push at the front part by actin polymerization and the forces of contraction at the rear part by interacting with myosin, thereby facilitating cell migration [19]. Besides, microtubule network plays pivotal role in the establishment and maintenance of front-rear polarization, and is also indispensable for directional cell migration [68,69,70]. Accumulating studies show that miRNAs affect cell migration through regulation of cytoskeleton remodeling [20].
MiR-9 suppresses the migration of human embryonic stem cell-derived neural progenitor cells (hNPCs) by directly targeting stathmin, a cytosolic phosphoprotein with catastrophe-promoting activity of microtubule-depolymerization, thereby affecting the dynamics of microtubule network [70]. In MSCs, miR-9-5p was found to promote cell migration through regulating F-actin reorganization and/or cell polarization. Overexpression of miR-9-5p helped MSCs to form one large lamellipodium in response to HGF, while control MSCs tended to form more and smaller lamellipodia and were less polarized after HGF treatment [37].
MiRNAs from MSC-Derived Exosomes Regulate Cell Migration
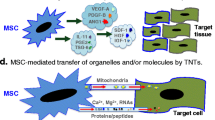
Recently, MSC-derived exosomes are found to deliver the mRNAs, miRNAs and proteins they contained to recipient cells, affecting the function of recipient cells and displaying significant therapeutic benefit as MSCs against a wide variety of diseases and injuries [71,72,73,74,75,76,77,78,79,80,81]. MiR-494 in exosomes derived from human bone marrow stem cells promoted myogenesis and the migration activity of endothelial cells [79]. MiR-140-5p in exosomes derived from miR-140-5p-overexpressing MSCs promotes the migration and proliferation of chondrocytes without side-effect of damaging extracellular matrix secretion, and contributes to prevention of osteoarthritis [80]. These results suggest that miRNAs contained in exosomes may also regulate the migration of MSCs via exosome-mediated delivering.
Conclusions
In summary, a range of miRNAs is reported to regulate the migration of MSCs from diverse aspects including cytokine-receptor interaction, intracellular signaling cascade, cytoskeleton rearrangement, and focal adhesion turnover (Table 2). Regarding the cytokine- or conditioned medium-induced migration of MSCs, it’s reasonable that a wide variety of miRNAs are activated or inhibited upon stimulation with a certain kind of cytokines or conditioned medium, which coordinate downstream signaling transduction and lead to certain locomotive behavior. Future efforts are required to analyze genome-wide miRNA content and address miRNAs that play a fundamental role in the elicited migration of MSCs, find out their direct targets and interactions with signaling pathways, and elucidate how they affect the cytoskeleton reorganization and focal adhesion turnover, thereby regulating cell migration.
Moreover, since each miRNA can simultaneously regulate potentially hundreds or even thousands of mRNA transcripts, it is possible that miRNA regulates cell migration through multiple downstream targets, leading to different locomotive behavior in different cells or in the same kind of cells under different conditions, such as miR-9-5p [37, 70] and miR-221 [31, 67]. It’ll be interesting to understand why and how a specific mRNA is “targeted” under certain conditions, and to find out the downstream mechanism that MSCs migrate to certain pathologies or injuries.
References
Filipowicz, W., Bhattacharyya, S. N., & Sonenberg, N. (2008). Mechanisms of post-transcriptional regulation by microRNAs: Are the answers in sight? Nature Reviews Genetics, 9, 102–114.
Hausser, J., & Zavolan, M. (2014). Identification and consequences of miRNA-target interactions - beyond repression of gene expression. Nature Reviews Genetics, 15, 599–612.
Huntzinger, E., & Izaurralde, E. (2011). Gene silencing by microRNAs: Contributions of translational repression and mRNA decay. Nature Reviews Genetics, 12, 99–110.
Lindsay, S. L., Johnstone, S. A., McGrath, M. A., Mallinson, D., & Barnett, S. C. (2016). Comparative miRNA-based fingerprinting reveals biological differences in human olfactory mucosa-and bone-marrow-derived mesenchymal stromal cells. Stem Cell Reports, 6, 729–742.
Bellayr, I. H., Kumar, A., & Puri, R. K. (2017). MicroRNA expression in bone marrow-derived human multipotent stromal cells. BMC Genomics, 18, 605–617.
Ali, N. M., Boo, L., Yeap, S. K., et al. (2016). Probable impact of age and hypoxia on proliferation and microRNA expression profile of bone marrow-derived human mesenchymal stem cells. Peer J, e1536, 4.
Hsieh, J. Y., Huang, T. S., Cheng, S. M., Lin, W. S., Tsai, T. N., Lee, O. K., & Wang, H. W. (2013). MiR-146a-5p circuitry uncouples cell proliferation and migration, but not differentiation, in human mesenchymal stem cells. Nucleic Acids Research, 41, 9753–9763.
Baglio, S. R., Devescovi, V., Granchi, D., & Baldini, N. (2013). MicroRNA expression profiling of human bone marrow mesenchymal stem cells during osteogenic differentiation reveals Osterix regulation by miR-31. Gene, 527, 321–331.
Chang, C. C., Veno, M. T., Chen, L., et al. (2018). Global microRNA profiling in human bone marrow skeletal-stromal or mesenchymal-stem cells identified candidates for bone regeneration. Molecular Therapy, 26, 593–605.
Cui, L. N., Zhou, X. M., Li, J. G., et al. (2012). Dynamic microRNA profiles of hepatic differentiated human umbilical cord lining-derived mesenchymal stem cells. PLoS One, 7, e44737.
Huat, T. J., Khan, A. A., Abdullah, J. M., Idris, F. M., & Jaafar, H. (2015). MicroRNA expression profile of neural progenitor-like cells derived from rat bone marrow mesenchymal stem cells under the influence of IGF-1, bFGF and EGF. International Journal of Molecular Sciences, 16, 9693–9718.
Gothelf, Y., Kaspi, H., Abramov, N., & Aricha, R. (2017). MiRNA profiling of NurOwn (R): Mesenchymal stem cells secreting neurotrophic factors. Stem Cell Research & Therapy, 8, 249–257.
Clark, E. A., Kalomoiris, S., Nolta, J. A., & Fierro, F. A. (2014). Concise review: microRNA function in multipotent mesenchymal stromal cells. Stem Cells, 32, 1074–1082.
Guo, L., Zhao, R. C. H., & Wu, Y. J. (2011). The role of microRNAs in self-renewal and differentiation of mesenchymal stem cells. Experimental Hematology, 39, 608–616.
Peng, S. P., Gao, D., Gao, C. D., Wei, P. P., Niu, M., & Shuai, C. J. (2016). MicroRNAs regulate signaling pathways in osteogenic differentiation of mesenchymal stem cells (review). Molecular Medicine Reports, 14, 623–629.
Huang, C., Geng, J. N., & Jiang, S. W. (2017). MicroRNAs in regulation of osteogenic differentiation of mesenchymal stem cells. Cell and Tissue Research, 368, 229–238.
Kang, H., & Hata, A. (2015). The role of microRNAs in cell fate determination of mesenchymal stem cells: Balancing adipogenesis and osteogenesis. BMB Reports, 48, 319–323.
Hamam, D., Ali, D., Kassem, M., Aldahmash, A., & Alajez, N. M. (2015). MicroRNAs as regulators of adipogenic differentiation of mesenchymal stem cells. Stem Cells and Development, 24, 417–425.
Parsons, J. T., Horwitz, A. R., & Schwartz, M. A. (2010). Cell adhesion: Integrating cytoskeletal dynamics and cellular tension. Nature Reviews Molecular Cell Biology, 11, 633–643.
Huang, S. L., & He, X. H. (2010). MicroRNAs: Tiny RNA molecules, huge driving forces to move the cell. Protein & Cell, 1, 916–926.
Chen, D., Xia, Y. L., Zuo, K., et al. (2015). Crosstalk between SDF-1/CXCR4 and SDF-1/CXCR7 in cardiac stem cell migration. Scientific Reports, 5, 16813–16821.
Liu, X. L., Duan, B. Y., Cheng, Z. K., et al. (2011). SDF-1/CXCR4 axis modulates bone marrow mesenchymal stem cell apoptosis, migration and cytokine secretion. Protein & Cell, 2, 845–854.
Son, B. R., Zhao, D. L., Marquez-Curtis, L. A., Shirvaikar, N., Ratajczak, M. Z., & Janowska-Wieczorek, A. (2004). SDF-1-CXCR4 and HGF-c-met axes regulate mobilization/recruitment to injured tissue of human mesenchymal stem cells. Blood, 642a, 104.
Lu, M. H., Hu, C. J., Chen, L., et al. (2013). miR-27b represses migration of mouse MSCs to burned margins and prolongs wound repair through silencing SDF-1a. PLoS One, 8, e68972.
Lu, M. H., Li, C. Z., Hu, C. J., et al. (2012). MicroRNA-27b suppresses mouse MSC migration to the liver by targeting SDF-1 alpha in vitro. Biochemical and Biophysical Research Communications, 421, 389–395.
Pillai, M. M., Yang, X., Balakrishnan, I., Bemis, L., & Torok-Storb, B. (2010). MiR-886-3p down regulates CXCL12 (SDF1) expression in human marrow stromal cells. PLoS One, 5, e14304.
Li, J. N., Li, L., Li, Z. X., et al. (2015). The role of miR-205 in the VEGF-mediated promotion of human ovarian cancer cell invasion. Gynecologic Oncology, 137, 125–133.
Susuki, D., Kimura, S., Naganuma, S., Tsuchiyama, K., Tanaka, T., Kitamura, N., Fujieda, S., & Itoh, H. (2011). Regulation of microRNA expression by hepatocyte growth factor in human head and neck squamous cell carcinoma. Cancer Science, 102, 2164–2171.
Tome, M., Lopez-Romero, P., Albo, C., et al. (2011). MiR-335 orchestrates cell proliferation, migration and differentiation in human mesenchymal stem cells. Cell Death and Differentiation, 18, 985–995.
Yue, Q., Zhang, Y., Li, X. Y., et al. (2016). MiR-124 suppresses the chemotactic migration of rat mesenchymal stem cells toward HGF by downregulating Wnt/beta-catenin signaling. European Journal of Cell Biology, 95, 342–353.
Zhu, A., Kang, N., He, L., Li, X., Xu, X., & Zhang, H. (2016). MiR-221 and miR-26b regulate chemotactic migration of MSCs toward HGF through activation of Akt and FAK. Journal of Cellular Biochemistry, 117, 1370–1383.
Chi, Y., Cui, J., Wang, Y., du, W., Chen, F., Li, Z., Ma, F., Song, B., Xu, F., Zhao, Q., Han, Z., & Han, Z. (2016). Interferongamma alters the microRNA profile of umbilical cord-derived mesenchymal stem cells. Molecular Medicine Reports, 14, 4187–4197.
Fayyad-Kazan, H., Fayyad-Kazan, M., Badran, B., Bron, D., Lagneaux, L., & Najar, M. (2017). Study of the microRNA expression profile of foreskin derived mesenchymal stromal cells following inflammation priming. Journal of Translational Medicine, 15, 10.
He, L. H., Wang, X. Y., Kang, N. X., et al. (2018). MiR-375 inhibits the hepatocyte growth factor-elicited migration of mesenchymal stem cells by downregulating Akt signaling. Cell and Tissue Research, 372, 99–114.
Neth, P., Ries, C., Karow, M., Egea, V., Ilmer, M., & Jochum, M. (2007). The Wnt signal transduction pathway in stem cells and cancer cells: Influence on cellular invasion. Stem Cell Reviews, 3, 18–29.
Ryu, C. H., Park, S. A., Kim, S. M., Lim, J. Y., Jeong, C. H., Jun, J. A., Oh, J. H., Park, S. H., Oh, W. I., & Jeun, S. S. (2010). Migration of human umbilical cord blood mesenchymal stem cells mediated by stromal cell-derived factor-1/CXCR4 axis via Akt, ERK, and p38 signal transduction pathways. Biochemical and Biophysical Research Communications, 398, 105–110.
Li, X. Y., He, L. H., Yue, Q., et al. (2017). MiR-9-5p promotes MSC migration by activating beta-catenin signaling pathway. American Journal of Physiology-Cell Physiology, 313, C80–C93.
Sotsios, Y., & Ward, S. G. (2000). Phosphoinositide 3-kinase: A key biochemical signal for cell migration in response to chemokines. Immunological Reviews, 177, 217–235.
Garcia-Martinez, J. M., Moran, J., Clarke, R. G., et al. (2009). Ku-0063794 is a specific inhibitor of the mammalian target of rapamycin (mTOR). Biochemical Journal, 421, 29–42.
Mora, A., Davies, A. M., Bertrand, L., Sharif, I., Budas, G. R., Jovanović, S., Mouton, V., Kahn, C. R., Lucocq, J. M., Gray, G. A., Jovanović, A., & Alessi, D. R. (2003). Deficiency of PDK1 in cardiac muscle results in heart failure and increased sensitivity to hypoxia. The EMBO Journal, 22, 4666–4676.
Mora, A., Lipina, C., Tronche, F., Sutherland, C., & Alessi, D. R. (2005). Deficiency of PDK1 in liver results in glucose intolerance, impairment of insulin-regulated gene expression and liver failure. Biochemical Journal, 385, 639–648.
Sarbassov, D. D., Guertin, D. A., Ali, S. M., & Sabatini, D. M. (2005). Phosphorylation and regulation of Akt/PKB by the rictor-mTOR complex. Science, 307, 1098–1101.
Song, M. S., Salmena, L., & Pandolfi, P. P. (2012). The functions and regulation of the PTEN tumour suppressor. Nature Reviews Molecular Cell Biology, 13, 283–296.
Zheng, B., Wang, C., He, L., Xu, X., Qu, J., Hu, J., & Zhang, H. (2013). Neural differentiation of mesenchymal stem cells influences chemotactic responses to HGF. Journal of Cellular Physiology, 228, 149–162.
Dabbah, M., Attar-Schneider, O., Zismanov, V., Matalon, S. T., Lishner, M., & Drucker, L. (2016). Multiple myeloma cells promote migration of bone marrow mesenchymal stem cells by altering their translation initiation. Journal of Leukocyte Biology, 100, 761–770.
Aman, A., & Piotrowski, T. (2008). Wnt/beta-catenin and FGF signaling control collective cell migration by restricting chemokine receptor expression. Developmental Cell, 15, 749–761.
Asad, M., Wong, M. K., Tan, T. Z., Choolani, M., Low, J., Mori, S., Virshup, D., Thiery, J. P., & Huang, R. Y. J. (2014). FZD7 drives in vitro aggressiveness in stem-a subtype of ovarian cancer via regulation of non-canonical Wnt/PCP pathway. Cell Death & Disease, 5, e1346.
Gomez-Orte, E., Saenz-Narciso, B., Moreno, S., & Cabello, J. (2013). Multiple functions of the noncanonical Wnt pathway. Trends in Genetics, 29, 545–553.
Montcouquiol, M., Crenshaw, E. B., & Kelley, M. W. (2006). Noncanonical Wnt signaling and neural polarity. Annual Review of Neuroscience, 29, 363–386.
Song, J. L., Nigam, P., Tektas, S. S., & Selva, E. (2015). MicroRNA regulation of Wnt signaling pathways in development and disease. Cellular Signalling, 27, 1380–1391.
Ueno, K., Hirata, H., Hinoda, Y., & Dahiya, R. (2013). Frizzled homolog proteins, microRNAs and Wnt signaling in cancer. International Journal of Cancer, 132, 1731–1740.
Wu, X. Y., Shen, Q. T., Oristian, D. S., et al. (2011). Skin stem cells orchestrate directional migration by regulating microtubule-ACF7 connections through GSK3 beta. Cell, 144, 341–352.
Köhler, A., Schambony, A., & Wedlich, D. . (2007). Cell migration under control of Wnt-signaling in the vertebrate embryo. Advances in Developmental Biology, 17, 159–201.
Zhang, J., Han, C., & Wu, T. (2012). MicroRNA-26a promotes cholangiocarcinoma growth by activating beta-catenin. Gastroenterology, 143, 246–256.e248.
Kim, Y. S., Noh, M. Y., Kim, J. Y., Yu, H. J., Kim, K. S., Kim, S. H., & Koh, S. H. (2013). Direct GSK-3beta inhibition enhances mesenchymal stromal cell migration by increasing expression of beta-PIX and CXCR4. Molecular Neurobiology, 47, 811–820.
Lapid, K., Itkin, T., D'Uva, G., et al. (2013). GSK3 beta regulates physiological migration of stem/progenitor cells via cytoskeletal rearrangement. Journal of Clinical Investigation, 123, 1705–1717.
Sun, T., Rodriguez, M., & Kim, L. (2009). Glycogen synthase kinase 3 in the world of cell migration. Development, Growth & Differentiation, 51, 735–742.
Yucel, G., & Oro, A. E. (2011). Cell migration: GSK3 beta steers the cytoskeleton's tip. Cell, 144, 319–321.
Karow, M., Popp, T., Egea, V., Ries, C., Jochum, M., & Neth, P. (2009). Wnt signalling in mouse mesenchymal stem cells: Impact on proliferation, invasion and MMP expression. Journal of Cellular and Molecular Medicine, 13, 2506–2520.
Neth, P., Ciccarella, M., Egea, V., Hoelters, J., Jochum, M., & Ries, C. (2006). Wnt signaling regulates the invasion capacity of human mesenchymal stem cells. Stem Cells, 24, 1892–1903.
Romer, L. H., Birukov, K. G., & Garcia, J. G. N. (2006). Focal adhesions - paradigm for a signaling nexus. Circulation Research, 98, 606–616.
Zamir, E., & Geiger, B. (2001). Molecular complexity and dynamics of cell-matrix adhesions. Journal of Cell Science, 114, 3583–3590.
Hamadi, A., Bouali, M., Dontenwill, M., Stoeckel, H., Takeda, K., & Ronde, P. (2005). Regulation of focal adhesion dynamics and disassembly by phosphorylation of FAK at tyrosine 397. Journal of Cell Science, 118, 4415–4425.
Illc, D., Furuta, Y., Kanazawa, S., et al. (1995). Reduced sell motility and enhanced focal adhesion contact formation in cells from FAK-deficient mice. Nature, 377, 539–544.
Zhang, F. X., Jing, S. H., Ren, T. M., & Lin, J. T. (2013). MicroRNA-10b promotes the migration of mouse bone marrow-derived mesenchymal stem cells and downregulates the expression of E-cadherin. Molecular Medicine Reports, 8, 1084–1088.
Vogelmann, R., Nguyen-Tat, M. D., Giehl, K., Adler, G., Wedlich, D., & Menke, A. (2005). TGF beta-induced downregulation of E-cadherin-based cell-cell adhesion depends on PI3-kinase and PTEN. Journal of Cell Science, 118, 4901–4912.
Chang, W., Kim, R., Park, S. I., Jung, Y. J., Ham, O., Lee, J., Kim, J. H., Oh, S., Lee, M. Y., Kim, J., Park, M. S., Chung, Y. A., Hwang, K. C., & Maeng, L. S. (2015). Enhanced healing of rat calvarial bone defects with hypoxic conditioned medium from mesenchymal stem cells through increased endogenous stem cell migration via regulation of ICAM-1 targeted-microRNA-221. Molecules and Cells, 38, 643–650.
Etienne-Manneville, S. (2013). Microtubules in cell migration. Annual Review of Cell and Developmental Biology, 29, 471–499.
Watanabe, T., Noritake, J., & Kaibuchi, K. (2005). Regulation of microtubules in cell migration. Trends in Cell Biology, 15, 76–83.
Delaloy, C., Liu, L., Lee, J. A., Su, H., Shen, F., Yang, G. Y., Young, W. L., Ivey, K. N., & Gao, F. B. (2010). MicroRNA-9 coordinates proliferation and migration of human embryonic stem cell-derived neural progenitors. Cell Stem Cell, 6, 323–335.
Phinney, D. G., & Pittenger, M. F. (2017). Concise review: MSC-derived exosomes for cell-free therapy. Stem Cells, 35, 851–858.
Arslan, F., Lai, R. C., Smeets, M. B., Akeroyd, L., Choo, A., Aguor, E. N. E., Timmers, L., van Rijen, H. V., Doevendans, P. A., Pasterkamp, G., Lim, S. K., & de Kleijn, D. P. (2013). Mesenchymal stem cell-derived exosomes increase ATP levels, decrease oxidative stress and activate PI3K/Akt pathway to enhance myocardial viability and prevent adverse remodeling after myocardial ischemia/reperfusion injury. Stem Cell Research, 10, 301–312.
Lai, R. C., Arslan, F., Lee, M. M., Sze, N. S. K., Choo, A., Chen, T. S., Salto-Tellez, M., Timmers, L., Lee, C. N., el Oakley, R. M., Pasterkamp, G., de Kleijn, D. P. V., & Lim, S. K. (2010). Exosome secreted by MSC reduces myocardial ischemia/reperfusion injury. Stem Cell Research, 4, 214–222.
Vonk, L. A., van Dooremalen, S. F. J., Liv, N., Klumperman, J., Coffer, P. J., Saris, D. B. F., & Lorenowicz, M. J. (2018). Mesenchymal stromal/stem cell-derived extracellular vesicles promote human cartilage regeneration in vitro. Theranostics, 8, 906–920.
Bruno, S., Grange, C., Deregibus, M. C., Calogero, R. A., Saviozzi, S., Collino, F., Morando, L., Busca, A., Falda, M., Bussolati, B., Tetta, C., & Camussi, G. (2009). Mesenchymal stem cell-derived microvesicles protect against acute tubular injury. Journal of the American Society of Nephrology, 20, 1053–1067.
Xin, H. Q., Li, Y., Liu, Z. W., et al. (2013). MiR-133b promotes neural plasticity and functional recovery after treatment of stroke with multipotent mesenchymal stromal cells in rats via transfer of exosome-enriched extracellular particles. Stem Cells, 31, 2737–2746.
Li, T. F., Yan, Y. M., Wang, B. Y., et al. (2013). Exosomes derived from human umbilical cord mesenchymal stem cells alleviate liver fibrosis. Stem Cells and Development, 22, 845–854.
Zhang, B., Wang, M., Gong, A. H., et al. (2015). HucMSC-exosome mediated-Wnt4 signaling is required for cutaneous wound healing. Stem Cells, 33, 2158–2168.
Nakamura, Y., Miyaki, S., Ishitobi, H., Matsuyama, S., Nakasa, T., Kamei, N., Akimoto, T., Higashi, Y., & Ochi, M. (2015). Mesenchymal-stem-cell-derived exosomes accelerate skeletal muscle regeneration. FEBS Letters, 589, 1257–1265.
Tao, S. C., Yuan, T., Zhang, Y. L., Yin, W. J., Shang-Chun, G., & Zhang, C. Q. (2017). Exosomes derived from miR-140-5p-overexpressing human synovial mesenchymal stem cells enhance cartilage tissue regeneration and prevent osteoarthritis of the knee in a rat model. Theranostics, 7, 180–195.
Zhu, J., Lu, K., Zhang, N., et al. (2017). Myocardial reparative functions of exosomes from mesenchymal stem cells are enhanced by hypoxia treatment of the cells via transferring microRNA-210 in an nSMase2-dependent way. Artificial Cells, Nanomedicine, and Biotechnology, 1–12.
Acknowledgements
This work was supported by the National Natural Science Foundation of China (Grant no. 31371407, 30870642) and the Priority Academic Program Development of Jiangsu Higher Education Institutions.
Funding
This contribution is supported by the National Natural Science Foundation of China (Grant no. 31371407, 30870642) and the Priority Academic Program Development of Jiangsu Higher Education Institutions.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare no potential conflicts of interest.
Rights and permissions
About this article
Cite this article
He, L., Zhang, H. MicroRNAs in the Migration of Mesenchymal Stem Cells. Stem Cell Rev and Rep 15, 3–12 (2019). https://doi.org/10.1007/s12015-018-9852-7
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12015-018-9852-7