Abstract
A tumor represents a highly intricate tissue entity, characterized by an exceptionally complex microenvironment that starkly contrasts with the typical physiological surroundings of healthy tissues. Within this tumor microenvironment (TME), every component and factor assume paramount importance in the progression of malignancy and exerts a pivotal influence on a patient’s clinical outcome. One of the remarkable aspects of the TME is its remarkable heterogeneity, not only across different types of cancers but even within the same histological category of tumors. In-depth research has illuminated the intricate interplay between specific immune cells and molecules and the dynamic characteristics of the TME. Recent investigations have yielded compelling evidence that several mutations harbored by tumor cells possess the capacity to instigate substantial alterations in the TME. These mutations, often acting as drivers of tumorigenesis, can orchestrate a cascade of events that remodel the TME, thereby influencing crucial aspects of cancer behavior, including its invasiveness, immune evasion, and response to therapies. It is within this nuanced context that the present study endeavors to provide a concise yet comprehensive summary of how specific mutations, within the genetic landscape of cancer cells, can instigate profound changes in TME features. By elucidating the intricate relationship between genetic mutations and the TME, this research aims to contribute to a deeper understanding of cancer biology. Ultimately, the knowledge gained from this study holds the potential to inform the development of more targeted and effective treatments, thereby offering new hope to patients grappling with the complexities of cancer.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
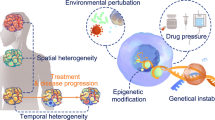
Cancer has a common characteristic of dysregulated cell growth composed of a heterogeneous group of ailments, complex interplay of genetic and environmental factors [1]. Despite a significant understanding of the molecular pathways involved, cancer remains the 2nd leading cause of death [2, 3]. In all their progression, tumors are considered a compound system including cancerous cells, extracellular matrix (ECM), and non-malignant cells known as the tumor microenvironment (TME), which comprises support cells and infiltrated inflammatory immune cells [4, 5]. TME is a concept that dramatically affects malignant cells. TME plays a significant role in tumor survival in every part of this complex ecosystem. However, infiltrated immune cells are the core operator of this organization [6]. The structure and cells of the TME related to the malignancy types, like features of hallmark like stromal cells, blood vessels, immune cells, and extracellular matrix (ECM). Throughout tumor development, relationship among malignant cells and immune cells and component of the TME led to tumor cell survival, metasets and malignant progression. Malignant cells infiltrated between adaptive and innate immune cells which can exhaust these cells [7]. According to the cancer cell microenvironment component, immune cells could stimulate or inhibit malignant cell progression [8]. Immune system cell arrangement in the TME is entirely different from healthy tissues [9]. In some cases, these cells cannot demonstrate anti-tumor roles and play a critical role in cancer growth. Some members of Tumor-infiltrating lymphocytes (TILs) and Tumor-associated macrophages (TAMs) families are the central cells with the tumor-supportive functions, and they play vital roles in immune system suppression which leads to tumor cells escaping from immune systems [10, 11]. In addition to TAMs and TILs, other immune cells like Natural killer (NK) cells and neutrophils have crucial effects on malignant cells’ fate. Every change in the resistant system population affects the prognosis of cancer dramatically. These days the presence or absence of these elements and the composition of their people in the TME are considered elements in a patient’s survival [12]. Current studies have approved that the immune cells population is related to the initiation of molecular pathways that encourage the anti-tumor effector cells apoptosis or immune cell functions suppression in the TME. Every malignant cell with particular mutations or gene expression profile invites a unique immune cell to TME and arranges their population differently. These facts absorb attention to single genes mutations and expression profiles affecting tumor infiltrated immune cells [13, 14]. In this study, we tried to summarize the functions of some critical components of the immune system in TME and explain how a single gene mutation or expression in malignant cells can change immune microenvironments of tumors (Fig. 1).
T-reg and monocytes effects on TME. In the context of tumors, T-reg cells can suppress the activity of other immune cells by producing several cytokines, allowing the tumor to grow and spread. Also these immune cells via producing cytokines lead to change M1 macrophage to M2 macrophage, inactivating NK-cells, inactivating CD8 + T cells, and inducing angiogenesis in tumor cells in TME. Overall, both T-reg cells and monocytes can have a negative impact on the immune response to tumors and contribute to the growth and spread of cancer in tumor microenvironment
Search Strategy
In this study, we conducted an exhaustive and systematic search for relevant scientific literature, leveraging prominent online databases such as PubMed, Embase, and Web of Science, up to the latest available data in August 2023. Our quest for pertinent articles was driven by a strategically chosen set of keywords that encompassed pivotal aspects of cancer research. These keywords included “Cancer,” “Tumor Microenvironment,” “CTLA-4,” “TP-53,” both individually and in combinations, reflecting the multifaceted nature of our investigation. Additionally, we complemented our database search with a thorough exploration of available resources on the EndNote software and the Google Scholar to ensure comprehensiveness in our data collection. Upon the compilation of the initial dataset, a meticulous curation process was employed to eliminate duplicate articles, thus ensuring the integrity and uniqueness of our final dataset. Subsequently, the selected articles were subjected to an in-depth analysis, where they were systematically summarized, critically evaluated, and rigorously analyzed to distill valuable insights and pertinent findings.
Immune Cells and Their Relation with TME
There are a number of different populations of T cells could be affect carcinogenesis. Cytotoxic CD8 + T cells sense unusual malignant cells antigens which expressed on tumor cells and destroyed malignant cells. CD4 + T cells divided into different subtypes and so affect to the immune cells responses within the microenvironment of the TME [15]. Regulatory T cells (Tregs) normally control autoimmunity and inflammatory responses during diverse settings. In the microenvironment of the TME, role of the Tregs is important as they might stimulate malignancy progression via exhausting immune cells [16]. Tumor infiltrating B-cells have significant role in the development of “tertiary lymphoid structures” that are designed within the TME. Tertiary lymphoid structures are a positive prognostic indicator in ovarian, melanoma and breast malignancy. The B-cells anti-carcinogenic role, contain secretion of cytokines, antigen-presentation to T cells, and production anti-cancer antibody that stimulate and regulate cytotoxic immune responses. Conversely, B-cells could promote tumor progression, and their existence in the TME can be sign of poor prognosis in renal cell carcinoma, prostate, and bladder tumor [7, 17]. Monocyte derived macrophages are divided into anti-inflammatory immune-suppressive M2 macrophages that take part in wound healing and M1 macrophages that phagocyte and kill cells. Both types of macrophages can be associated among a cancer cells, but TME supports the M2 phenotype via cytokines secretion like Interleukin (IL)-4 and hypoxia to support malignancy development. Usually, high macrophage existence is related with low prognosis in various kind of malignancy, like gastric, lung and breast tumor [18, 19]. When malignant cells start to progression, neutrophils which are in the TME, stimulate inflammation via releasing of cytokines and ROS to stimulate cancer cell apoptosis. Also, neutrophils stimulate malignancy progression via alteration of the extracellular matrix, releasing Matrix metalloproteinase (MMP)-9 and Vascular endothelial growth factor (VEGF) to stimulate angiogenesis and eventually cancer development [20, 21]. Antigen presenting cells like Dendritic cells (DCs) are and have a central role in the immune system. They sense, realized, trap and present antigens to T cells at secondary lymphoid organs. TME cytokines released from the triggered DCs promote tolerance to malignant cells and suppress the stimulation of an immune response [22]. Also, there are additional important non-immune which have significant role in tumor suppression, development and growth (adipocytes, cancer associated fibroblasts, endothelial cells, stromal cells).
The Tumor Immune Microenvironment is a Double-Edged Sword
Usually, the tumor immune microenvironment (TIME) must be a part of the battle against malignant cells. Still, since they can act as tumor-supportive or anti-tumor agents, the functions of every aspect of TIME in malignancy are sophisticated [23, 24]. Multiple factors contribute to the poor prognosis of solid tumors, one of them being the reduction or augmentation of particular immune cell kinds, which led to better or shrunk survival in some malignancies [25, 26]. It has been documented that an increased mass of tumor-infiltrating T cells (TILs) is responsible for a good prognosis and better results for chemotherapy in patients with colorectal tumor [9, 10]. However, all types of TILs do not correlate with a good prognosis; in this category of immune cells, CD8+ cytotoxic T cells, which act as straight killers of cancer cells, are involved with a better prediction in a large group of tumors [12]. On the other hand, CD8+ Tcells do not stand for the whole immune response, and other parts of this complex environment can influence and change the TIME roles from distractive to supportive. Collectively, the presence or absence of a special kind of cells are not responsible for good or bad prognosis, and they have been considered in connection with each other [27]. Other cells with a significant role in the TME are Tumor-associated macrophages (TAMs). Unlike cytotoxic T cells, increasing the amount of these cells is correlated with bad prognosis, increasing angiogenesis, tumor cell mobility, and tumor immune tolerance. The existence of TAMs is found to have a connection with the increase in the expression of VEGF, a protein directly related to metastasis [28]. Furthermore, TAMs are responsible for creating pre-metastatic niches, which can play a role in chemotherapy resistance [29, 30]. In TME, TAMs have been categorized in one of these two subgroups: M2 macrophages (CD206 + CD163 + HLA-DR+) or M1 macrophages (CD86 + CD68 + HLA-DR+). M2 macrophages stimulate from M1 by CD40 activation under the effects of interferon-gamma. These cells are responsible for lousy cancer progression and worse overall survival by affecting other TME immune cells [31, 32].
The role of signaling pathways in cancer cell growth, progression, and metastasis is pivotal in shaping the TME and subsequently influencing the tumor immune microenvironment. Signaling pathways encompass a complex network of molecular interactions that regulate various cellular processes, including proliferation, survival, angiogenesis, and invasion. Dysregulation of these pathways is a hallmark of cancer and plays a central role in tumor development and progression [33]. One of the critical ways in which signaling pathways impact the TME is through their influence on cancer cell behavior. Constitutive activation of specific pathways, often driven by genetic mutations or aberrant signaling molecule expression, can lead to uncontrolled cancer cell growth and survival. This unbridled proliferation results in the formation of hypoxic regions within the tumor due to inadequate blood supply, thereby altering the TME by inducing the secretion of proangiogenic factors such as VEGF. Consequently, the angiogenic switch in the TME promotes the recruitment of endothelial cells and the formation of new blood vessels, facilitating nutrient and oxygen delivery to the tumor and affecting its immune milieu [34]. Additionally, inflammation and high density of inflammatory cytokines are some of the main characteristics of the TME. The high concentration of inflammatory cytokines is responsible for changes in immune cell features. For example, secretion of IL-6 activates JAK/STAT3 pathway, leading to infiltrating tumor monocytes and differentiating to M2 macrophages, which express PD-L1 and consequently suppress cytotoxic T cell response [35, 36]. Furthermore, signaling pathways can modulate the TME by regulating the expression of immune checkpoint molecules. For example, the upregulation of the programmed cell death protein 1 (PD-1) and its ligand, PD-L1, is often orchestrated by oncogenic signaling pathways [37]. These immune checkpoint molecules play a pivotal role in dampening the anti-tumor immune response by inhibiting the activation of T cells [38]. Consequently, their overexpression can create an immunosuppressive TME, where immune cells are rendered ineffective in recognizing and attacking cancer cells. Metastasis, a critical hallmark of cancer progression, is also heavily influenced by signaling pathways. These pathways can induce epithelial-to-mesenchymal transition (EMT), a phenotypic shift in cancer cells that endows them with enhanced migratory and invasive properties. This process is orchestrated by various signaling pathways, including those involving transforming growth factor-beta (TGF-β) and Wnt [39]. The acquisition of EMT traits not only enables cancer cells to invade adjacent tissues and enter the bloodstream but also influences the pre-metastatic niche in distant organs, thus affecting the future metastatic site’s microenvironment.
NK cells are another type of cells whose presence in TME is related to the excellent prognosis; however, activation of TGF- β following M2 macrophages activation can reduce NK cells activation and reduce their attraction into tumors. In addition to M2 macrophage’s functions in inactivating CD8 + T and NK cells, Tregs promote tumor-supporting function in the immune system. Even though FOXP3 + CD25+ Treg acts as anti-inflammatory cells, they might release IL-10, TGF-β, and express cytotoxic T lymphocyte-associated protein 4 (CTLA-4) in TME and, thereby, suppress the immune cell responses against malignant cells. Some research has revealed that the existence and activation of Tregs in TME are associated with distant metastasis and poor prognosis [40, 41]. Despite the evidence that discloses the functions of every part of immune system activation in the TME, there are many questions about the main reasons for the differences of TIME population and their position between the tumors. Recently particular gene mutations or dysregulated expression in malignant cells are at the core of the attraction of scientists to explain this phenomenon.
Genomic Alterations of Malignant Cells and Features of Tumor Immune Microenvironment
Single-gene mutations can have intense effects on the TME, influencing patients’ treatment and survival rate in multiple cancers. For example, Kristen rat sarcoma viral oncogene homolog (KRAS) mutation in colorectal cancer is confirmed as a factor of change in immune cells population which is demonstrated as a predictive value and essential element to choose treatment career and outcome [42]. In addition to colorectal cancer, some studies have explained the correlation between gene mutation and TIME populations in glioma, non-small cell lung carcinoma, and melanoma, renal and ovarian cancers.
Glioma is categorized into two collections, CpG island methylator phenotype (CIMP) and non-CIMP [43]. Patients with CIMP gliomas show a high occurrence of mutations in the oncometabolite 2-hydroxygluratate (2-HG) and isocitrate dehydrogenase 1/2 (IDH1/2), that is associated with the hyper-DNA methylation phenotype in such patients [44]; these changes can explain the differences between the survival of two groups. The CIMP counterparts of gliomas are less invasiveness than their non-CIMP wild-type IDH1/2 (wtIDH1/2) [45]. While the brain is supposed to be an immune-isolated tissue, gliomas contain a unique element of immune cells; thus, macrophages can reach 20–30% of the entire cells in tumors with high aggression [46]. In gliomas, wtIDH1 can cause inflammation and immunologic response [47], which are conducted to increase the level of chemokines and particular types of immune cells penetration. In other words, the various genomic alterations in muIDH1 gliomas can change genes involved with extracellular matrix modification and formation. Following these variations, infiltrated immune cells are different in IDH1gliomas with varying mutations in compression to wtIDH1/2. For example, muIDH1 tumors attract fewer neutrophils [48].
High-grade serous tubo-ovarian cancer (HGSC) is another cancer in TME structures, and immune cells population differences are definable by particular types of mutation [49]. p53 encourages cell arrest in the presence of DNA damage or cellular stress so that the damage can be repaired, or self-mediated apoptosis can take place [50].
Commonly, HGSC malignancy starts with silencing, deletion, or modification of TP53 in the fallopian tube epithelium (FTE). PTEN, NF-1, Brca1, and MycOE, are other genes with a high level of mutation and direct effect in clinical features of patients and TME structures.
PTEN, P53, and NF1 mutations directly affect the CD45+ cell population in TME. It has been shown that Tp53-/-; Pten-/-; Nf1-/- cancers had a major number of macrophage (CD11b + F4/80 + ), monocytic myeloid-derived suppressor cells (m-MDSC, CD11b+Ly6GloLy6Chi), CD11b+Ly6CloLy6Ghi), smaller numbers of myeloid dendritic cells (mDC, CD11b + CD11C + ), granulocytic myeloid-derived suppressor cells (g-MDSC and sparse T lymphocytes (CD3+ cells). According to the lower fraction of total CD45+ cells, the total T cell number in Tp53-/-; Pten-/-; Nf1-/- cancers are even lesser than the other models. Moreover, the macrophages in Tp53-/-; Pten-/-; Nf1-/- malignancy had more “M2-like” character which can affect the survival rate of patients [51, 52]. Tp53-/-; Brca1-/-; MycOE cancers had large proportions of macrophages and lower parts of mDCs, g-MDSCs, and m-MDSCs. Unlike other models, CD8+ and CD4 + T cells in Tp53-/-; Brca1 -/-; MycOE tumors were predominantly (>60%), PD1 + , CTLA4 + , and CD44 + , proposing anti-tumor effect of them. Tp53-/-; Brca1 -/-; MycOE malignancy are more stable in their T helper (Th) 1 (Tbet + ), Th2 (GATA3 + ) cells and Th1/Th2: 0.7 population; the other models commonly have Th1 cells (Th1/Th2: 2.4 in Tp53-/-;Pten-/-;Nf1-/-; Th1/Th2: 3.4) [53]. In addition to HGSC, some studies have reported that lack of PTEN leads to reduced T cell penetration in prostate cancer and melanoma in mouse models [54]. Another highlighted illustration for TME affected by mutation is the Protein polybromo-1 (PBRM1) gene, which influences mast cells and T cells in renal cancer. PBRM1 is the common mutated gene in the Clear cell renal cell carcinoma (ccRCC), followed by VHL, SETD2, and BAP1 genes. According to the studies, the PBRM1 MUT ccRCC patients have lower immune scores. Furthermore, recruitment of resting mast cells to the TME and tumor purity is considerably higher in PBRM1 MUT ccRCC samples than PBRM1 WT, SETD2MUT, VHLMUT, and BAP1MUT ccRCC patients. In addition, PBRM1 mutations are involved in with Treg infiltration. [52]. ccRCC malignancy with PBRM1 mutations are also involved with up regulation of angiogenic genes [55]. In melanoma, a tumor with BRAF-mutation (usually BRAF V600E) displays low T cell infiltration and high level of pro-inflammatory cytokines like IL-10, IL-6 and VEGF induce an increase in the number of immunosuppressive cells, myeloid-derived suppressor cells, Tregs, leading to the inhibition of DC maturation in TME [56]. In one bio-informatics analysis on non-small-cell lung carcinoma (NSCLC), the prevalence of EGFR mutation was almost 14.30% (184/1287), and the co-mutation rate of EGFR and MAPK genes was 11.41% (21/184). Glycosaminoglycan-related pathways are importantly up-regulated in the EGFR mutant group. EGFR-mutated samples have low PDL1 protein levels than those in wild-type patients. Encouraging evidence has shown that augmented immature DCs infiltration and reduced NK CD56dim, Tγδ, cytotoxic, and Th2 cell infiltration are the primary immune variations in EGFR-mutated patients. On the other hand, PD-L1 protein expression levels in EGFR-MAPK co-mutations patients are high and parallel immune microenvironment with the wild-type group. Moreover, PD-L1 + /CD8+ tumor-infiltrating lymphocytes (TILs) ratio in in wild-type NSCLC was higher than EGFR-mutated NSCLC [57, 58]. In addition to their cytolytic functions and target cell killing, NK CD cells are the prime springs of pro-inflammatory chemokines [59]. Besides, Th2 cells improve B cell function and have detailed instruction of the immune environment [60]. Like NSCLC, other malignancies like gliomas, renal cell carcinoma, chronic myeloid leukemia, and pancreatic tumor demonstrate the same phenotype in co-mutated patients [61]. Tables 1 and 2 shows various gene mutations and their effect on the expression of different gene on the TIME cells population.
Single Gene Expressions Affect Tumor Fate
Cancer is a genetically complex and heterogeneous disorder. By considering the origin cell, the tumor genotype controls its susceptibility to intrinsic immunogenicity, conventional therapy, targeted therapy, and the range of produced chemokines and cytokines [23, 62, 63]. Cancer is principally a disease of copy number abnormalities (CNAs), as well as complex chromosomal rearrangements, deletions, and amplifications that regulate several genes and pathways [62, 63]. An extensive pan-cancer genomic analysis showed that high level of CCL5 protein and RNA involved with intra-tumor CD8 T cells in solid malignancy. On the other hand, high number of TAMs, especially MDSCs and M2-like TAMs, correlated with a poor result [64, 65]. Usually, a single gene mutation can initiate a gene expression pattern that affects the whole TME population. In Gliomas, wtIDH1 gliomas gene expression signatures are powerfully involved with immunological response including high levels of some interleukins, chemokines [46] and inflammation that may promote penetration of commune immune cells. The complex genomic losses and gains are realized in muIDH1 gliomas. For example, mutations in IDH1 are associated with the dysregulation of collagen and integrins gene expression, which is related to extracellular matrix modification and formation. As a result of these changes, immune cell infiltration differs between tumors with IDH1 mutations and wild-type IDH1. For example, muIDH1 tumors usually attract fewer neutrophils at basal levels [66, 67]. Human HGSC also has a complex TME, with various infiltrating tumor-associated chemokines/cytokines and immune cells which involved with the prognosis [68, 69]. As with many other tumors, intra-tumor CD8 + T cells and high CD8 + /Treg ratio involved with better survival, however high levels of Tregs are a negative prognostic sign [70, 71]. Intra-tumor T cells induced the expression of CCL22, CCL21, CCL5, CXCL10, and CXCL9 however high levels of VEGF inversely involved with T cell infiltration [72, 73]. The dual expression of chemokines CCL5 and CXCL9 is also involved with a valuable prognosis. Remarkably, ovarian malignancy with high low CCL5 RNA and intra-tumor CD8+ cells have shown up regulation of CXCL9 [74,75,76]. In the ccRCC, higher PDGFA, VCAM1, VEGFB, VEGFC, VEGFA, and PDGFB mRNA expression happens due to PBRM1 mutations. These outcomes have illustrated that PBRM1 mutation activate mast cell infiltration into the TME and triggers HIF-related signaling pathways, which initiate the progression of ccRCC. In this regard, the inflammatory response signaling pathway was curbed in PBRM1-overexpression. One of the significant consequences of PBRM1-overexpression is the downregulation of CCL5 which is one of the main genes in the inflammatory response pathway and is the core element of mast cells infiltration to TME. CCL5 mRNA and protein levels were meaningfully high express in PBRM1-mutated tumors. High CCL5 in ccRCC patients exhibited considerable enrichment in immune-related signaling pathways, as well as inflammatory signaling (TNF-α-NF-ΚB, IL-6-JAK-, STAT3, IFN-α, IFN-γ) [77]. Furthermore, in the tumor cells with mutated P53, miR-30 expression can dramatically increase hypoxia-responsive factor (HIF1) expression even in the condition of normoxia, signifying that mutp53 can arise HIF1-dependent responses in tumors independent of oxygen accessibility. Upon to the alteration of the miR-30d expression in melanoma, an increase in GalNAc transferase (GALNT7) occ, yours, regulating posttranslational protein O-glycosylation which led to high expression of the immunosuppressive cytokine like IL-10, promoting an immunosuppressive milieu in TME and subsequently fostering metastasis. In addition, miR-30d targets DGKZ, which related to the diacylglycerol (DAG) kinase family and regulates secretion and trafficking by multiple mechanisms. In Golgi membrane, DGKZ low expression increases the local concentration of DAG, helping vesicular secretion and transport through inducing protein kinase PKD signaling. VPS26B is second straight target of microRNA-30d, a core retromer complex factor that regulating the recycling of proteins through endosomal sorting. Defects in retromer recycling may perturb GA dynamics and result in abnormal secretion. The last impact of deregulation of VPS26B and DGKZ is the mis-glycosylation of ECM components. One of the critical factors for immune cell infiltration is the extracellular matrix condition [78]. In lower-grade glioma (LGGs), the analysis of immune checkpoints expression in both low and high-risk groups has revealed which patients in the low-risk group had low terms of LAG-3, PD-1, TIM-3 and CTLA-4, but higher expression of T cell immunoreceptor with Ig and ITIM domains (TIGIT). Besides immune checkpoints, the expression level of some other genes can regulate immune cell infiltration to TME. High immune cells (B cells, neutrophils, CD4 + T cells, dendritic cells, CD8 + T cells and macrophages) infiltration was confirmed to be connected with a poorer prognosis in LGGs. TNFRSF11B, LTF, GDF15, and BIRC5 expression levels were completely linked with infiltration levels of immune cells, while there was a negative relationship among PRLHR and CRLF1 expression levels [79].
Previous research has shown high expression of Prostaglandin-endoperoxide synthase (PTGS2) and SRY-box transcription factor 2 (SOX2) transcriptional factors in penile SCC and skin. SOX2 is a β-catenin transcriptional target, absent in normal epidermis and high expressed in malignant stem cells [80, 81]. At the same time, PTGS2 (COX2), is a pro-inflammatory gene that promote prostaglandins production from arachidonic acid, led to cytokine high expression and inflammatory reactions which related to the absorption of a particular type of immune cells to TME [82].
In NSCLC, higher Syndecan-2 (SDC2) expression levels happen due to EGFR mutation’s lack of MAPK mutation. SDC2 can support the clearance of the TCR/CD3 complex, which leads to T cell inactivation. Thus, SDC2 is correlated to the inhibitory microenvironment of patients with EGFR mutations, and down-regulation of SDC2 in EGFR-MAPK co-mutated patients can be a reason for up-regulation Immune-related pathways like FCγR-mediated phagocytosis, correlated with a good prognosis [83, 84].
Heparin sulfate (HS) is the primary glycosaminoglycan (GAG) essential extracellular matrix, several aggregating cytokines, and controlling immune activation. GAGs can act as chemokines protectors that preserve them from degradation and actively interfere with the action of chemokines. As a result, the degradation of HS can stimulate the immune system by reducing GAGs production [80]. Poor immune response in EGFR-mutated patients can result from GAG gathering, which causes an immune-suppressive microenvironment [85]. PD-L1 is one of the highlighted genes with a remarkable impact on TME, and its expression is regulated by a sophisticated communication between tumor cells and immune cells. Some studies have reported that the up-regulation of PD-L1 expression can be an outcome of the presence of IFN-γ produced by T cells [86]. However, there is some conflict on PDL-1 expression effects on cancer prognosis. Generally, PD-L1 expression in malignant cells reveals an antigen-induced antitumor immune response referred to TILs. As a result of this concept, patients with high number of T cell in TME and high expression of PD-L1, possibly demonstrate a better prognosis [81, 87]. Conversely, a high concentration of PD-L1 was linked to adverse results in several malignancies. However, some studies have found no linkage between PDL-1 expression and survival [88]. For example, in head and neck carcinoma, PDL-1 expression was related to a higher TIL count (≥30%), which was considerably involved with an expansive pattern of tumor invasion [83, 89]. LAG-3 is an inhibitory receptor, usually expressed on activated NK and T cells, and is the third inhibitory receptor after CTLA-4 and PD-1 which can be targeted in the clinic. Fibrinogen-like protein 1 (FGL1) is a new ligand of LAG-3, which is up-regulated in patients with lower PD-1 signals and less leucocyte infiltration in various types of cancer [90,91,92]. For example, in Small Cell carcinoma of the esophagus (SCCE), FGL1 is one of the genes involved in managing the TME immune cells population. In line with this demonstration, patients with significant up-regulation of FGL1 performed low leucocytes infiltration, representing that FGL1-LAG3 signaling can be an essential tool for immune inhibition in SCCE. In addition to FGL1-LAG3, B7-H3 (CD276), VEGFB, and sialic acid-binding Ig-like lectin (Siglec-15), which are known as immune inhibitors were considerably high expressed in malignant tissues which might be a practical reason to explain weak immune response in SCCE. [91, 93]. In renal cell carcinoma (RCC), a hypoxic TME initiates the up-regulation of CD73 (NT5E) expression on tumor cells which leads to CD39 and CD73 (ENTPD1) activation on cancer cells and extracellular adenosine generation by stromal cells. This issue applies an immunosuppressive influence on PD-1 [94, 95]. Adenosine causes the prevention of NK cell infiltration and cytotoxic T lymphocyte activity using intensifying Treg proliferation [96]. In preclinical models, increased adenosine signaling attenuates the antitumor immune response via the expansion of Tregs and myeloid-derived suppressor cells (MDSCs) as well as tumor-associated macrophages differentiation into the immunosuppressive M2 phenotype [84, 86, 97]. ADORA2A (A2AR) is another gene that have a substantial role in the Adenosine pathway, whose high expression was strongly correlated with the induction of angiogenesis. Mechanistically, A2AR stimulation can increase angiogenesis by reducing the production of thrombospondin-1 (TSP-1) and triggering the differentiation of macrophages to the M2 phenotype, which causes amplified expression of proangiogenic factors including nitric oxide synthase, IL-10, and VEGF [98,99,100]. Tumor cells and immune cells, like dendritic cells and macrophages are the primary resources for Lactate of TME [101]. The extraordinary lactate concentration is immune suppression in various types of cancers. Lactate, derived from cancer cells, inhibits the growth and cytokine production of cytotoxic T lymphocytes (CTLs) [102]. Monocarboxylate 1 (MCT1) participates in lactate transportation to simplify metabolic reprogramming during tumor progression. MCT1 is ubiquitously expressed in many kinds of malignancy, such as breast cancer, melanoma, and prostate cancer [103, 104]. In breast cancer, the overexpression of MCT1 is reported in HER2+ tumors are correlated with high CD163+ macrophages (M2) infiltration compared to other types of breast cancer. The main reason for this phenomenon is hidden, but some studies have suggested that high expression of MCT1 regulates the polarization of M2-like macrophages by influencing lactate uptake [104,105,106].
Limitations
The limitations of this study primarily stem from the relatively narrow scope of investigation, particularly with regard to the assessing of immune mutations within the TME and cancer cells. It is imperative to acknowledge that this constrained perspective may impose certain constraints on the depth and generalizability of our conclusions. First and foremost, the restricted focus on immune mutations within the TME and cancer cells implies that other critical factors within the broader cancer ecosystem may not have been comprehensively addressed. Cancer is a multifaceted disease with an intricate interplay of genetic, epigenetic, and microenvironmental elements. Consequently, an exclusive emphasis on immune mutations may inadvertently omit other crucial components, such as stromal cells, extracellular matrix remodeling, and metabolic alterations, all of which are integral to the TME’s dynamics. Thus, our conclusions may offer an incomplete picture of the complex interactions that govern cancer progression and immune responses. Furthermore, the limited scope may hinder our ability to capture the full spectrum of immune mutations and their nuanced effects. Immune-related genetic alterations can exhibit considerable heterogeneity across different cancer types and individual patients. By concentrating solely on a specific subset of these mutations, we may not fully appreciate the diverse array of genetic changes that can influence immune responses and immune evasion strategies employed by cancer cells. Consequently, our findings might not be fully representative of the broader landscape of immune-related genomic alterations and their implications in cancer biology. Another noteworthy limitation is that our study’s narrow focus might restrict its applicability to a specific subset of patients or cancer types, potentially limiting the generalizability of our conclusions to a more diverse population of cancer patients. The intricate relationship between immune mutations, the TME, and cancer cell behavior can exhibit substantial variations across different malignancies and patient cohorts. Therefore, extrapolating our findings to broader clinical contexts may require caution and consideration of these variations.
Conclusion and Future Perspectives
Tumors, far from being mere assemblages of malignant cells, are intricate structures comprised of various components. Within this disorder, non-cancerous cells intermingle with malignant cells, collectively creating a sophisticated and dynamic ecosystem. Remarkably, every element within this ecosystem assumes distinct roles and responsibilities. However, it is essential to underscore that at the heart of this complex milieu lie the cancer cells themselves. Any alteration in the characteristics or behavior of these malignant cells possesses the potential to exert profound repercussions on both the non-malignant cell populations and their respective functions. In the context of tumorigenesis, the cancer cells act as the central orchestrators, dictating the course of events within the TME. They secrete signaling molecules, engage in crosstalk with neighboring cells, and manipulate the surrounding ECM, thereby sculpting a microenvironment conducive to their survival and proliferation. However, it is not a unidirectional relationship; the non-malignant cells also exert influence on the cancer cells, creating a reciprocal interplay that can either promote or impede tumor progression. Crucially, the inherent complexity of cancer becomes even more pronounced when considering that within a single type of cancer, the introduction of a single genetic mutation or the dysregulation of a single gene’s expression can trigger a cascade of changes that dramatically transform the tumor environment. These alterations can encompass shifts in immune cell infiltration, modifications in the extracellular matrix composition, and adjustments in the nutrient and oxygen supply—each of which can profoundly impact the tumor’s behavior and therapeutic responsiveness. As we delve deeper into the intricate dynamics of the TME, it becomes evident that there is a wealth of uncharted territory awaiting exploration in future research. Unraveling the specific molecular mechanisms governing the interactions between malignant and non-malignant cells within the TME holds the promise of unveiling novel therapeutic targets. Additionally, identifying biomarkers indicative of these dynamic changes may facilitate early disease detection and the development of personalized treatment strategies.
Data availability
The datasets generated during and/or analyzed during the current study are available from the corresponding author on reasonable request.
Abbreviations
- TME:
-
Tumor microenvironment
- ECM:
-
Extracellular matrix
- TAMs:
-
Tumor-associated macrophages
- TILs:
-
Tumor-infiltrating lymphocytes
- MMP:
-
Matrix metalloproteinase
- CTLA-4:
-
Cytotoxic T lymphocyte-associated protein 4
- CIMP:
-
CpG island methylator phenotype
- 2-HG:
-
2-hydroxygluratate
- HGSC:
-
High-grade serous tubo-ovarian cancer
- FTE:
-
Fallopian tube epithelium
- g-MDSC:
-
Granulocytic myeloid-derived suppressor cells
- m-MDSC:
-
Monocytic myeloid-derived suppressor cells
- NSCLC:
-
Non-small-cell lung carcinoma
- CNAs:
-
Copy number abnormalities
- HIF1:
-
Hypoxia-responsive factor
- DAG:
-
Diacylglycerol
- LGGs:
-
Lower-grade glioma
- SDC2:
-
Syndecan-2
- GAG:
-
Glycosaminoglycan
- HS:
-
Heparin sulfate
- FGL1:
-
Fibrinogen-like protein 1
- SCCE:
-
Small Cell carcinoma of the esophagus
- VEGF:
-
Vascular endothelial growth factor
- RCC:
-
Renal cell carcinoma
- Tregs:
-
Regulatory T cells
- MDSCs:
-
Myeloid-derived suppressor cells
- MCT1:
-
Monocarboxylate transporter 1
References
Tabrez, S., Khan, A. U., Hoque, M., Suhail, M., Khan, M. I., & Zughaibi, T. A. (2022). Investigating the anticancer efficacy of biogenic synthesized MgONPs: An in vitro analysis. Frontiers in Chemistry, 10, 970193.
Alafaleq, N. O., Zughaibi, T. A., Jabir, N. R., Khan, A. U., Khan, M. S., & Tabrez, S. (2023). Biogenic Synthesis of Cu-Mn Bimetallic Nanoparticles Using Pumpkin Seeds Extract and Their Characterization and Anticancer Efficacy. Nanomaterials (Basel, Switzerland), 13, 7.
Ahmad, I., Hoque, M., Alam, S. S. M., Zughaibi, T. A., & Tabrez, S. (2023). Curcumin and Plumbagin Synergistically Target the PI3K/Akt/mTOR Pathway: A Prospective Role in Cancer Treatment. International Journal of Molecular Sciences, 24(7), 6651.
Hanahan, D., & Coussens, L. M. (2012). Accessories to the crime: functions of cells recruited to the tumor microenvironment. Cancer Cell, 21(3), 309–322.
Ebos, J. M., & Kerbel, R. S. (2011). Antiangiogenic therapy: impact on invasion, disease progression, and metastasis. Nature reviews Clinical oncology, 8(4), 210–221.
Segovia-Mendoza, M., & Morales-Montor, J. (2019). Immune Tumor Microenvironment in Breast Cancer and the Participation of Estrogen and Its Receptors in Cancer Physiopathology. Frontiers in immunology, 10, 348.
Anderson, N. M., & Simon, M. C. (2020). The tumor microenvironment. Current Biology, 30(16), R921–r5.
Peña-Romero, A. C., & Orenes-Piñero, E. (2022). Dual Effect of Immune Cells within Tumour Microenvironment: Pro- and Anti-Tumour Effects and Their Triggers. Cancers, 14, 7.
Ma, Y., Shurin, G. V., Peiyuan, Z., & Shurin, M. R. (2013). Dendritic cells in the cancer microenvironment. Journal of Cancer, 4(1), 36–44.
Balkwill, F. R., Capasso, M., & Hagemann, T. (2012). The tumor microenvironment at a glance. Journal of cell science, 125(Pt 23), 5591–5596.
Barzegari, A., Saeedi, N., Zarredar, H., Barar, J., & Omidi, Y. (2014). The search for a promising cell factory system for production of edible vaccine. Human Vaccines & Immunotherapeutics, 10(8), 2497–2502.
de Looff, M., de Jong, S., & Kruyt, F. A. E. (2019). Multiple Interactions Between Cancer Cells and the Tumor Microenvironment Modulate TRAIL Signaling: Implications for TRAIL Receptor Targeted Therapy. Frontiers in immunology, 10, 1530.
Ma, C., Luo, H., Cao, J., Zheng, X., Zhang, J., & Zhang, Y., et al. (2020). Identification of a Novel Tumor Microenvironment-Associated Eight-Gene Signature for Prognosis Prediction in Lung Adenocarcinoma. Frontiers in molecular biosciences, 7, 571641.
Xiao, B., Peng, J., Wang, Y., Deng, Y., Ou, Q., & Wu, X., et al. (2020). Prognostic value of tumor infiltrating lymphocytes combined with PD-L1 expression for patients with solitary colorectal cancer liver metastasis. Annals of translational medicine, 8(19), 1221.
Maimela, N. R., Liu, S., & Zhang, Y. (2019). Fates of CD8+ T cells in Tumor Microenvironment. Computational and structural biotechnology journal, 17, 1–13.
Ohue, Y., & Nishikawa, H. (2019). Regulatory T (Treg) cells in cancer: Can Treg cells be a new therapeutic target? Cancer science, 110(7), 2080–2089.
Fridman, W. H., Meylan, M., Petitprez, F., Sun, C. M., Italiano, A., & Sautès-Fridman, C. (2022). B cells and tertiary lymphoid structures as determinants of tumour immune contexture and clinical outcome. Nature reviews Clinical oncology, 19(7), 441–457.
Boutilier, A. J., & Elsawa, S. F. (2021). Macrophage Polarization States in the Tumor Microenvironment. International Journal of Molecular Science, 22, 13.
Poh, A. R., & Ernst, M. (2018). Targeting Macrophages in Cancer: From Bench to Bedside. Frontiers in Oncology, 8, 49.
Masucci, M. T., Minopoli, M., & Carriero, M. V. (2019). Tumor Associated Neutrophils. Their Role in Tumorigenesis, Metastasis, Prognosis and Therapy. Frontiers in Oncology, 9, 1146.
Tabrez, S., Khan, A. U., Mirza, A. A., Suhail, M., Jabir, N. R., & Zughaibi, T. A., et al. (2022). Biosynthesis of copper oxide nanoparticles and its therapeutic efficacy against colon cancer. Nanotechnology Reviews, 11(1), 1322–1331.
Prete, A. D., Salvi, V., Soriani, A., Laffranchi, M., Sozio, F. & Bosisio, D. et al. (2023). Dendritic cell subsets in cancer immunity and tumor antigen sensing. Cellular & molecular immunology, 20(5), 432–447.
Binnewies, M., Roberts, E. W., Kersten, K., Chan, V., Fearon, D.F. & Merad, M. (2018). Understanding the tumor immune microenvironment (TIME) for effective therapy. Nature Medicine, 24(5), 541–550.
Yu, Y. R., & Ho, P. C. (2019). Sculpting tumor microenvironment with immune system: from immunometabolism to immunoediting. Clinical and experimental immunology, 197(2), 153–160.
Quail, D. F., & Joyce, J. A. (2013). Microenvironmental regulation of tumor progression and metastasis. Nature medicine, 19(11), 1423–1437.
Kitamura, T., Qian, B. Z., & Pollard, J. W. (2015). Immune cell promotion of metastasis. Nature reviews Immunology, 15(2), 73–86.
Chaudhary, B., & Elkord, E. (2016). Regulatory T Cells in the Tumor Microenvironment and Cancer Progression: Role and Therapeutic Targeting. Vaccines., 4, 3.
Laviron, M., & Boissonnas, A. (2019). Ontogeny of Tumor-Associated Macrophages. Frontiers in immunology, 10, 1799.
Allavena, P., Sica, A., Solinas, G., Porta, C., & Mantovani, A. (2008). The inflammatory micro-environment in tumor progression: the role of tumor-associated macrophages. Critical reviews in oncology/hematology, 66(1), 1–9.
Gowd, V., Ahmad, A., Tarique, M., Suhail, M., Zughaibi, T. A., & Tabrez, S., et al. (2022). Advancement of cancer immunotherapy using nanoparticles-based nanomedicine. Seminars in cancer biology, 86(Pt 2), 624–644.
Capece, D., Fischietti, M., Verzella, D., Gaggiano, A., Cicciarelli, G., & Tessitore, A., et al. (2013). The inflammatory microenvironment in hepatocellular carcinoma: a pivotal role for tumor-associated macrophages. BioMed research international, 2013, 187204.
Wang, J., Li, D., Cang, H., & Guo, B. (2019). Crosstalk between cancer and immune cells: Role of tumor-associated macrophages in the tumor microenvironment. Cancer medicine, 8(10), 4709–4721.
Xu, L., Zou, C., Zhang, S., Chu, T. S. M., Zhang, Y., & Chen, W., et al. (2022). Reshaping the systemic tumor immune environment (STIE) and tumor immune microenvironment (TIME) to enhance immunotherapy efficacy in solid tumors. Journal of hematology & oncology, 15(1), 1–30.
Kim, H. J., Ji, Y. R., & Lee, Y. M. (2022). Crosstalk between angiogenesis and immune regulation in the tumor microenvironment. Archives of Pharmacal Research, 45(6), 401–416.
Groner, B., & von Manstein, V. (2017). Jak Stat signaling and cancer: Opportunities, benefits and side effects of targeted inhibition. Molecular and cellular endocrinology, 451, 1–14.
Mimura, K., Teh, J. L., Okayama, H., Shiraishi, K., Kua, L. F., & Koh, V., et al. (2018). PD-L1 expression is mainly regulated by interferon gamma associated with JAK-STAT pathway in gastric cancer. Cancer science, 109(1), 43–53.
Abaza, A., Idris, F. S., Shaikh, H. A., Vahora, I., Moparthi, K. P., & Al Rushaidi, M. T., et al. (2023). Programmed Cell Death Protein 1 (PD-1) and Programmed Cell Death Ligand 1 (PD-L1) Immunotherapy: A Promising Breakthrough in Cancer Therapeutics. Cureus., 15, 9.
Yenyuwadee, S., Aliazis, K., Wang, Q., Christofides, A., Shah, R. & Patsoukis, N, et al. (2022). Immune cellular components and signaling pathways in the tumor microenvironment. Seminars in cancer biology, 86(2), 187–201.
Han, Y.-L., Luo, D., Habaxi, K., Tayierjiang, J., Zhao, W., & Wang, W., et al. (2022). COL5A2 Inhibits the TGF-β and Wnt/β-Catenin signaling pathways to inhibit the invasion and metastasis of osteosarcoma. Frontiers in Oncology, 12, 813809.
Kim, H. R., Park, H. J., Son, J., Lee, J. G., Chung, K. Y., & Cho, N. H., et al. (2019). Tumor microenvironment dictates regulatory T cell phenotype: Upregulated immune checkpoints reinforce suppressive function. Journal for immunotherapy of cancer, 7(1), 339.
Leivonen, S. K., Pollari, M., Brück, O., Pellinen, T., Autio, M., & Karjalainen-Lindsberg, M. L., et al. (2019). T-cell inflamed tumor microenvironment predicts favorable prognosis in primary testicular lymphoma. Haematologica, 104(2), 338–346.
Shanehbandi, D., Zarredar, H., Asadi, M., Zafari, V., Esmaeili, S., & Seyedrezazadeh, E., et al. (2021). Anticancer impacts of Terminalia catappa extract on SW480 colorectal neoplasm cell line. Journal of Gastrointestinal Cancer, 52(1), 99–105.
Noushmehr, H., Weisenberger, D. J., Diefes, K., Phillips, H. S., Pujara, K., & Berman, B. P., et al. (2010). Identification of a CpG island methylator phenotype that defines a distinct subgroup of glioma. Cancer cell, 17(5), 510–522.
Dang, L., White, D. W., Gross, S., Bennett, B. D., Bittinger, M. A., & Driggers, E. M., et al. (2009). Cancer-associated IDH1 mutations produce 2-hydroxyglutarate. Nature, 462(7274), 739–744.
Turcan, S., Rohle, D., Goenka, A., Walsh, L. A., Fang, F., & Yilmaz, E., et al. (2012). IDH1 mutation is sufficient to establish the glioma hypermethylator phenotype. Nature, 483(7390), 479–483.
Kong, L. Y., Wu, A. S., Doucette, T., Wei, J., Priebe, W., & Fuller, G. N., et al. (2010). Intratumoral mediated immunosuppression is prognostic in genetically engineered murine models of glioma and correlates to immunotherapeutic responses. Clinical cancer research: an official journal of the American Association for Cancer Research, 16(23), 5722–5733.
Ceccarelli, M., Barthel, F. P., Malta, T. M., Sabedot, T. S., Salama, S. R., & Murray, B. A., et al. (2016). Molecular Profiling Reveals Biologically Discrete Subsets and Pathways of Progression in Diffuse Glioma. Cell, 164(3), 550–563.
Amankulor, N. M., Kim, Y., Arora, S., Kargl, J., Szulzewsky, F., & Hanke, M., et al. (2017). Mutant IDH1 regulates the tumor-associated immune system in gliomas. Genes & development, 31(8), 774–786.
Nazam, N., Jabir, N. R., Ahmad, I., Alharthy, S. A., Khan, M. S., & Ayub, R., et al. (2023). Phenolic Acids-Mediated Regulation of Molecular Targets in Ovarian Cancer: Current Understanding and Future Perspectives. Pharmaceuticals (Basel, Switzerland), 16, 2.
Alafaleq, N. O., Alomari, A., Khan, M. S., Shaik, G. M., Hussain, A., & Ahmed, F., et al. (2022). Anticancer potential of gold nanoparticles (AuNPs) using a battery of in vitro tests. Nanotechnology Reviews, 11(1), 3292–3304.
Peng, W., Chen, J. Q., Liu, C., Malu, S., Creasy, C., & Tetzlaff, M. T., et al. (2016). Loss of PTEN Promotes Resistance to T Cell-Mediated Immunotherapy. Cancer discovery, 6(2), 202–216.
Miao, D., Margolis, C. A., Gao, W., Voss, M. H., Li, W., & Martini, D. J, et al. (2018). Genomic correlates of response to immune checkpoint therapies in clear cell renal cell carcinoma. Science, 359(6377), 801–806.
Zhang, S., Iyer, S., Ran, H., Dolgalev, I., Gu, S., & Wei, W, et al. (2021). Genetically Defined, Syngeneic Organoid Platform for Developing Combination Therapies for Ovarian Cancer. Cancer discovery, 11(2), 362–383.
Wang, G., Lu, X., Dey, P., Deng, P., Wu, C. C., & Jiang, S., et al. (2016). Targeting YAP-Dependent MDSC Infiltration Impairs Tumor Progression. Cancer discovery, 6(1), 80–95.
Kapur, P., Peña-Llopis, S., Christie, A., Zhrebker, L., Pavía-Jiménez, A., & Rathmell, W. K., et al. (2013). Effects on survival of BAP1 and PBRM1 mutations in sporadic clear-cell renal-cell carcinoma: a retrospective analysis with independent validation. The Lancet Oncology, 14(2), 159–167.
Simiczyjew, A., Dratkiewicz, E., Mazurkiewicz, J., Ziętek, M., Matkowski, R. & Nowak, D. (2020). The Influence of Tumor Microenvironment on Immune Escape of Melanoma. International Journal of Molecular Sciences, 21(21), 8359.
Fauriat, C., Long, E. O., Ljunggren, H. G., & Bryceson, Y. T. (2010). Regulation of human NK-cell cytokine and chemokine production by target cell recognition. Blood., 115(11), 2167–2176.
Zarredar, H., Farajnia, S., Ansarin, K., Baradaran, B., Aria, M., & Asadi, M. (2019). Synergistic effect of novel EGFR inhibitor AZD8931 and p38α siRNA in lung adenocarcinoma cancer cells. Anti-Cancer Agents in Medicinal Chemistry (Formerly Current Medicinal Chemistry-Anti-Cancer Agents), 19(5), 638–644.
Thorsson, V., Gibbs, D. L., Brown, S. D., Wolf, D., Bortone, D. S., & Ou Yang, T. H., et al. (2018). The Immune Landscape of Cancer. Immunity., 48(4), 812–30.
Lambrecht, B. N., & Hammad, H. (2015). The immunology of asthma. Nature Immunology, 16(1), 45–56.
Izumi, H., Yamasaki, A., Takeda, K., Kodani, M., Touge, H., & Tanaka, N., et al. (2018). Acute-phase reaction induced by zoledronate and its effect on prognosis of patients with advanced non-small cell lung cancer. Lung Cancer, 122, 200–205.
Li, J., & Stanger, B. Z. (2019). The tumor as organizer model. Science, 363(6431), 1038–1039.
Wellenstein, M. D., & de Visser, K. E. (2018). Cancer-Cell-Intrinsic Mechanisms Shaping the Tumor Immune Landscape. Immunity, 48(3), 399–416.
Drakes, M. L., & Stiff, P. J. (2018). Regulation of Ovarian Cancer Prognosis by Immune Cells in the Tumor Microenvironment. Cancers, 10, 9.
Ruffell, B., & Coussens, L. M. (2015). Macrophages and therapeutic resistance in cancer. Cancer cell, 27(4), 462–472.
Suzuki, H., Aoki, K., Chiba, K., Sato, Y., Shiozawa, Y., & Shiraishi, Y., et al. (2015). Mutational landscape and clonal architecture in grade II and III gliomas. Nature genetics, 47(5), 458–468.
Eckel-Passow, J. E., Lachance, D. H., Molinaro, A. M., Walsh, K. M., Decker, P. A., & Sicotte, H., et al. (2015). Glioma Groups Based on 1p/19q, IDH, and TERT Promoter Mutations in Tumors. The. New England journal of medicine, 372(26), 2499–2508.
Rodriguez, G. M., Galpin, K. J. C., McCloskey, C. W. & Vanderhyden, B. C. (2018). The Tumor Microenvironment of Epithelial Ovarian Cancer and Its Influence on Response to Immunotherapy. Cancers, 10(8), 242.
Färkkilä, A., Gulhan, D. C., Casado, J., Jacobson, C. A., Nguyen, H. & Kochupurakkal, B. et al. (2020). Immunogenomic profiling determines responses to combined PARP and PD-1 inhibition in ovarian cancer. Nature Communications, 11(1), 1459
Curiel, T. J., Coukos, G., Zou, L., Alvarez, X., Cheng, P., & Mottram, P., et al. (2004). Specific recruitment of regulatory T cells in ovarian carcinoma fosters immune privilege and predicts reduced survival. Nature Medicine, 10(9), 942–949.
Zhang, L., Conejo-Garcia, J. R., Katsaros, D., Gimotty, P. A., Massobrio, M., & Regnani, G., et al. (2003). Intratumoral T cells, recurrence, and survival in epithelial ovarian cancer. The New England journal of medicine, 348(3), 203–213.
Bronger, H., Singer, J., Windmüller, C., Reuning, U., Zech, D., & Delbridge, C., et al. (2016). CXCL9 and CXCL10 predict survival and are regulated by cyclooxygenase inhibition in advanced serous ovarian cancer. British journal of cancer, 115(5), 553–563.
Nagarsheth, N., Wicha, M. S., & Zou, W. (2017). Chemokines in the cancer microenvironment and their relevance in cancer immunotherapy. Nature Reviews Immunology, 17(9), 559–572.
Dangaj, D., Bruand, M., Grimm, A. J., Ronet, C., Barras, D., & Duttagupta, P. A., et al. (2019). Cooperation between Constitutive and Inducible Chemokines Enables T Cell Engraftment and Immune Attack in Solid Tumors. Cancer cell, 35(6), 885–900.
Strickland, K. C., Howitt, B. E., Shukla, S. A., Rodig, S., Ritterhouse, L. L., & Liu, J. F., et al. (2016). Association and prognostic significance of BRCA1/2-mutation status with neoantigen load, number of tumor-infiltrating lymphocytes and expression of PD-1/PD-L1 in high grade serous ovarian cancer. Oncotarget, 7(12), 13587–13598.
Clarke, B., Tinker, A. V., Lee, C. H., Subramanian, S., van de Rijn, M., & Turbin, D., et al. (2009). Intraepithelial T cells and prognosis in ovarian carcinoma: novel associations with stage, tumor type, and BRCA1 loss. Modern pathology: an official journal of the United States and Canadian Academy of Pathology, Inc, 22(3), 393–402.
Liu, T., Xia, Q., Zhang, H., Wang, Z., Yang, W., & Gu, X., et al. (2020). CCL5-dependent mast cell infiltration into the tumor microenvironment in clear cell renal cell carcinoma patients. Aging, 12(21), 21809–21836.
Capaci, V., Mantovani, F., & Sal, G. D. (2020). A mutant p53/Hif1α/miR-30d axis reprograms the secretory pathway promoting the release of a prometastatic secretome. Cell stress, 4(11), 261–264.
Yin, W., Jiang, X., Tan, J., Xin, Z., Zhou, Q., & Zhan, C., et al. (2020). Development and Validation of a Tumor Mutation Burden-Related Immune Prognostic Model for Lower-Grade Glioma. Frontiers in oncology, 10, 1409.
Rovira-Clavé, X., Angulo-Ibáñez, M., Noguer, O., Espel, E., & Reina, M. (2012). Syndecan-2 can promote clearance of T-cell receptor/CD3 from the cell surface. Immunology, 137(3), 214–225.
Bastos, P., Gomes, T., & Ribeiro, L. (2017). Catechol-O-Methyltransferase (COMT): An Update on Its Role in Cancer, Neurological and Cardiovascular Diseases. Reviews of Physiology, Biochemistry and Pharmacology, 173, 1–39.
Ahmed, M. E., & Falasiri, S. (2020). The Immune Microenvironment in Penile Cancer and Rationale for. Immunotherapy, 9, 10.
Nechama, M., Kwon, J., Wei, S., Kyi, A. T., Welner, R. S., & Ben-Dov, I. Z., et al. (2018). The IL-33-PIN1-IRAK-M axis is critical for type 2 immunity in IL-33-induced allergic airway inflammation. Nature communications, 9(1), 1603.
Ohta, A., Kini, R., Ohta, A., Subramanian, M., Madasu, M., & Sitkovsky, M. (2012). The development and immunosuppressive functions of CD4(+) CD25(+) FoxP3(+) regulatory T cells are under influence of the adenosine-A2A adenosine receptor pathway. Frontiers in immunology, 3, 190.
Yang, W., Chen, N., Li, L., Chen, X., Liu, X., & Zhang, Y., et al. (2020). Favorable Immune Microenvironment in Patients with EGFR and MAPK Co-Mutations. Lung Cancer (Auckland, NZ), 11, 59–71.
Ryzhov, S., Novitskiy, S. V., Goldstein, A. E., Biktasova, A., Blackburn, M. R., & Biaggioni, I., et al. (2011). Adenosinergic regulation of the expansion and immunosuppressive activity of CD11b+Gr1+ cells. Journal of immunology (Baltimore, Md: 1950), 187(11), 6120–6129.
Ouyang, L., Zhang, K., Chen, J., Wang, J. & Huang, H. (2018). Roles of platelet-derived growth factor in vascular calcification. Journal of Cellular Physiology, 233(4), 2804–2814.
Wang, X., Teng, F., Kong, L., & Yu, J. (2016). PD-L1 expression in human cancers and its association with clinical outcomes. Onco Targets & Therapy, 9, 5023–5039.
Evrard, D., Hourseau, M., Couvelard, A., Paradis, V., Gauthier, H. & Raymond, E, et al. (2020). PD-L1 expression in the microenvironment and the response to checkpoint inhibitors in head and neck squamous cell carcinoma. Oncoimmunology, 9(1), 1844403
Zhao, Q., Chen, Y. X., Wu, Q. N., Zhang, C., Liu, M., & Wang, Y. N., et al. (2020). Systematic analysis of the transcriptome in small-cell carcinoma of the oesophagus reveals its immune microenvironment. Clinical & translational immunology, 9(10), e1173.
Shi, A. P., Tang, X. Y., Xiong, Y. L., Zheng, K. F., Liu, Y. J., & Shi, X. G., et al. (2021). Immune Checkpoint LAG3 and Its Ligand FGL1 in Cancer. Frontiers in immunology, 12, 785091.
Wang, J., Sanmamed, M. F., Datar, I., Su, T. T., Ji, L., & Sun, J., et al. (2019). Fibrinogen-like Protein 1 Is a Major Immune Inhibitory Ligand of LAG-3. Cell, 176(1-2), 334–347.
Guo, M., Yuan, F., Qi, F., Sun, J., Rao, Q., & Zhao, Z., et al. (2020). Expression and clinical significance of LAG-3, FGL1, PD-L1 and CD8(+)T cells in hepatocellular carcinoma using multiplex quantitative analysis. Journal of translational medicine, 18(1), 306.
Song, L., Ye, W., Cui, Y., Lu, J., Zhang, Y., & Ding, N., et al. (2017). Ecto-5’-nucleotidase (CD73) is a biomarker for clear cell renal carcinoma stem-like cells. Oncotarget., 8(19), 31977–31992.
Chambers, A. M., Wang, J., Lupo, K. B., Yu, H., Atallah Lanman, N. M., & Matosevic, S. (2018). Adenosinergic Signaling Alters Natural Killer Cell Functional Responses. Frontiers in immunology, 9, 2533.
Arab, S., & Hadjati, J. (2019). Adenosine Blockage in Tumor Microenvironment and Improvement of Cancer Immunotherapy. Immune network, 19(4), e23.
Takayama, H., Trenn, G., & Sitkovsky, M. V. (1988). Locus of inhibitory action of cAMP-dependent protein kinase in the antigen receptor-triggered cytotoxic T lymphocyte activation pathway. The. Journal of Biological Chemistry, 263(5), 2330–2336.
Tripathi, A., Lin, E., Xie, W., Flaifel, A., Steinharter, J. A., & Gatof, E. N. S., et al. (2020). Prognostic significance and immune correlates of CD73 expression in renal cell carcinoma. Journal for immunotherapy of cancer, 8(2), e001467.
Rafii, S., Ghouzlani, A., Naji, O., Ait Ssi, S., Kandoussi, S., & Lakhdar, A., et al. (2023). A(2A)R as a Prognostic Marker and a Potential Immunotherapy Target in Human Glioma. International Journal of Molecular Science, 24, 7.
Ernens, I., Bousquenaud, M., Lenoir, B., Devaux, Y., & Wagner, D. R. (2015). Adenosine stimulates angiogenesis by up-regulating production of thrombospondin-1 by macrophages. Journal of leukocyte biology, 97(1), 9–18.
Jin, M., Cao, W., Chen, B., Xiong, M., & Cao, G. (2022). Tumor-Derived Lactate Creates a Favorable Niche for Tumor via Supplying Energy Source for Tumor and Modulating the Tumor Microenvironment. Frontiers in cell and developmental biology, 10, 808859.
Romero-Garcia, S., Moreno-Altamirano, M. M., Prado-Garcia, H., & Sánchez-García, F. J. (2016). Lactate Contribution to the Tumor Microenvironment: Mechanisms, Effects on Immune Cells and Therapeutic Relevance. Frontiers in immunology, 7, 52.
Bola, B. M., Chadwick, A. L., Michopoulos, F., Blount, K. G., Telfer, B. A., & Williams, K. J., et al. (2014). Inhibition of monocarboxylate transporter-1 (MCT1) by AZD3965 enhances radiosensitivity by reducing lactate transport. Molecular cancer therapeutics, 13(12), 2805–2816.
Li, B., Yang, Q., Li, Z., Xu, Z., Sun, S., & Wu, Q., et al. (2020). Expression of Monocarboxylate Transporter 1 in Immunosuppressive Macrophages Is Associated With the Poor Prognosis in Breast Cancer. Frontiers in Oncology, 10, 574787.
Tiainen, S., Tumelius, R., Rilla, K., Hämäläinen, K., Tammi, M., & Tammi, R., et al. (2015). High numbers of macrophages, especially M2-like (CD163-positive), correlate with hyaluronan accumulation and poor outcome in breast cancer. Histopathology. 66(6), 873–883.
Kaleem, M., Dalhat, M. H., Azmi, L., Asar, T. O., Ahmad, W., & Alghanmi, M., et al. (2022). An Insight into Molecular Targets of Breast Cancer Brain Metastasis. International Journal of Molecular Sciences, 23(19), 11687.
Funding
This study was funded by Tuberculosis and Lung Disease Research Center, Tabriz University of Medical Science, Tabriz, Iran.
Author information
Authors and Affiliations
Contributions
Study Design: A.C., D.S.; Data Collection: M.A., H.Z.; Statistical Analysis: M.A., V.Z.; Data Interpretation: H.Z., Z.S.; Manuscript Preparation: Z.S., H.S.; Literature Search: A.C., D.S.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Asadi, M., Zarredar, H., Zafari, V. et al. Immune Features of Tumor Microenvironment: A Genetic Spotlight. Cell Biochem Biophys 82, 107–118 (2024). https://doi.org/10.1007/s12013-023-01192-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12013-023-01192-7