Abstract
As an essential trace element, appropriate boron supplementation can promote immune function of animals. To illustrate the effects of boron in a rat model, RNA-Seq was conducted for the RNA from duodenum after treatment with different concentration of boron in which boron was given in the form of boric acid. More than 47 million reads were obtained in 0, 10, and 320 mg/L boron (0, 57.21, and 1830.66 mg/L boric acid) treatment groups that produced 58 965 402, 48 607 328, and 46 760 660 clean reads, respectively. More than 95% of the clean reads were successfully matched to the rat reference genome and assembled to generate 32 662 transcripts. A total of 624 and 391 differentially expressed candidate genes (DEGs) were found between 0 vs.10 and 0 vs. 320 mg/L boron comparison groups. We also identified transcription start site, transcription terminal site, and skipped exons as the main alternative splicing events. GO annotations revealed most of DEGs were involved in the regulation of immune activity. The DEGs were enriched in influenza A, herpes simplex infection, cytosolic DNA-sensing pathway, and antigen processing and presentation signaling pathways. The expression levels of genes enriched in these signaling pathways indicate that lower doses of boron could achieve better effects on promoting immune response in the duodenum. These effects on the immune system appear to be mediated via altering the expression patterns of genes involved in the related signaling pathways in a dose-dependent pattern. These data provide more insights into the molecular mechanisms of immune regulation in rats in response to dietary boron treatment.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Boron is a bioactive micronutrient and human bodies can obtain it primarily from fruits, vegetables, legumes, nuts and drinking water. It is widely used in healthcare, industry, agriculture and cosmetics [1,2,3]. Boron shows pleiotropic effects and plays various roles in bone development, osteoarthritis, osteoporosis, central nervous system, cell membrane integrity and function, metabolism of minerals, immune regulation, and endocrine regulation [4,5,6,7,8].
The biological effects of boron are dose-dependent but exhibit a U-shaped response pattern, rendering it necessary to confirm its active range [9]. It is shown that low doses of boron (3 and 10 mg/day) in humans facilitates kidney stone dissolution and prevents stone formation. However, higher doses of boron could not protect rats against nephrolithiasis [9]. In African ostrich chicks, low level of boron (up to 160 mg) inhibits cell apoptosis, and enhances the activity of antioxidant system to promote the development of the kidney whilst high levels of boron (320 or 640 mg) obviously affected the development of the kidney [10]. Higher concentration of boron activates the general amino acid control system and inhibits protein synthesis [11]. Borates inhibited the proliferation of cancer cells, in that they could bind to the site of the ryanodine receptor, which resulted in Ca2+ channel inactivation. In three hours after the physiological addition of boric acid, the lower concentration of Ca2+ induced the phosphorylation of eIF2α following the activation of ATF4 pathway which reconfigured gene expression to make cells manage stress conditions [12, 13].
More recently, there has been an increasing number of studies examining the effects of boron supplementation on immune functions in a range of farm animal and other species [14]. In growing beef steers, supplementation of the diet with boron has limited effects on humoral immune function [15]. Boron supplementation restores humoral immune response which was caused by low levels of Ca in the diet in lambs [16]. Xiao et al. (2019) verified boron modulates immune response via calcium and MAPK signaling pathways in the thymus of ostrich chicks [17].
The intestine represents the largest compartment of the immune system [18]. The immune system of intestine plays an important role in maintaining the balance between defense, tolerance, and tissue repair [18,19,20]. It is reported that normal mucosal T-cell function and cytokine production is necessary for physiologic growth and maturation of the intestine [21, 22]. It remains obscure whether boron could modulate immunity and other functions in the intestine.
In this study, to further elucidate the effects of boron in rats’ intestine, we examined genome-wide expression profile of the duodenum using transcriptome sequencing analysis after supplemented with low or high levels of boron. Our transcriptome analysis included the assembly of sequencing data, the identification of differentially expressed genes and their function, as well as the classification and enrichment of gene function.
Methods
Experimental Animals
The specific pathogen free (SPF) 23-day-old male Sprague Dawley (SD) rats (weight 53 ± 2 g, lot No.: SCXK Zhe 2014–001) were purchased from the Qinglongshan experimental animal breeding farm, Jiangning District, Nanjing, China. The use of all experimental animals was reviewed and approved by the Anhui Laboratory Animal Care Committee. All animal experimental procedures were performed in strict accordance with the Guide for Laboratory Animal Care and Use and complied with the Guide for National Laboratory Animal Healthcare and Use.
Experimental Design and Animal Diet
36 23-day-old weighting 53 ± 2 g male SD rats were randomly divided into a control group and two test groups (n = 12 each). The control group (NC) was given distilled water, while the experimental groups were given drinking water supplemented with different concentration of boron. The boric acid (Sinopharm Chemical Reagent Co. Ltd., Shanghai, China; purity > 99.5 g boron acid/g, boron content > 17.4 g boron/100 g boron acid) was used as an alternative to adding boron directly in the drinking water. The addition concentration of boron is 10 mg/L and 320 mg/L, which actually means adding 57.21 mg/L (10 mg/L boron/0.1748) and 1830.66 mg/L (320 mg/L boron/0.1748) boric acid in the drinking water, respectively. The period of the experiment was 60 days. Animals were housed in individual ventilated cages in the SPF Experimental Animal Center of Anhui Science and Technology University, at 24–28 °C, 50–65% humidity, a 12: 12 light/dark cycle, and had free access to food and water. During the experimental period, the water and food intake of rats were recorded. The rat feed was purchased from the Qinglongshan breeding farm. The main nutritional level of the feed was crude protein ≥ 18.15%, crude fat ≥ 4.03%, crude fiber ≥ 5.12%, crude ash ≥ 7.94%, 1.43% of calcium, 0.87% of phosphorus, and 1.96 mg/kg of boron.
Sample Collection and Preparation
Three rats from each group were anesthetized with the anesthetic breathing system (ABS), in which 4–5% isoflurane was used to induce anesthesia followed by 1–2% for maintenance post-60 days of experimental period. The rats were bled to death via cardiac puncture, and the duodenum was immediately harvested and stored in RNAfixer (BioTeke Co., Ltd, Beijing, China) at 4 °C overnight and then transferred to − 80 °C until RNA isolation.
RNA Sequencing
Total RNA was extracted from each sample using TRIzol reagent (Invitrogen, Carlsbad, CA, USA) following the manufacturer’s protocol. After isolation, the quality of total RNA was checked by examining its purity and integrity. The high-quality RNA samples were selected for library construction. The amount of total RNA for each sample should be up to 1 ug.
The sequencing libraries were prepared by the TruseqTM RNA sample prep Kit (Illumina, San Diego, CA) following the manufacturer’s guidelines. Briefly, the mRNA was purified from the total RNA using poly-T oligo-attached magnetic beads (Invitrogen, USA) to pull down the poly-A mRNA and then randomly cleaved into ~ 200 bp short fragments with fragmentation buffer. The fragmented mRNAs were used as templates to synthesize the first strand cDNA by reverse transcriptase and random hexamers, and then the second strand cDNA was subsequently synthesized to form stable double-stranded structure. The double-stranded DNA fragment was made into the blunt end with End Repair Mix and then adenylated which ligated to the adaptor. The PCR was performed to amplify the library by 15 cycles. The PCR product was purified using Certified Low Range Ultra Agarose (Bio-Rad). The library quality was measured by a Agilent 2100 Bioanalyzer and the library was quantified by TBS380 Picogreen (Invitrogen, USA). The clustering of the index-coded samples was performed on a cBot Cluster Generation System using HiSeq 4000 PE Cluster Kit (Illumia) according to the manufacturer’s instructions. Then the libraries were sequenced on the Illumina sequencing platform (Hiseq 4000) and 150 bp paired-end reads were generated.
Quality Control and Transcriptome Assembly
Raw sequence reads containing adapter sequences, reads below a threshold quality of 20, reads containing more than 10% of “N” base in one single end, reads shorter than 50 bp in length as well as the sequencing primer from the raw data were removed to produce clean reads [23]. All of the subsequent analyses and annotation were carried out using clean reads with high quality. Clean reads were aligned to the reference genome (Rnor_6.0) by Hisat2. The alternative splicing was analyzed by ASprofile [24]. Overall quality of the transcriptome was assessed by RSeQC-2.6.3 (http://rseqc.sourceforge.net/). All aligned reads were assembled for differentially expressed analysis. The gene abundance was estimated by fragments per kilobase of transcript per million mapped reads (FPKM) using RSEM (http://www.biomedsearch.com/nih/RSEM-accurate-transcript-quantification-from/21816040.html). Differential expression analysis between two groups was processed using edgeR (http://www.bioconductor.org/packages/2.12/bioc/html/edgeR.html) [25, 26]. Genes with FDR < 0.05 and fold-change ≥ 2 or ≤ 0.5 were considered differentially expressed [27, 28].
GO Annotation and KEGG Pathway Analysis
Gene ontology (GO) analysis was conducted to illustrate the biological implications of DEGs between the control group and boron-treated group by the Goatools software (https://github.com/tanghaibao/GOatools) [29]. DEGs at P-value < 0.05 and fold-change ≥ 2 or ≤ 0.5 were regarded as significantly enriched. Kyoto Encyclopedia of Genes and Genomes (KEGG) (http://www.genome.jp/kegg/) database was used to elucidate the potential biological pathways of DEGs by the KOBAS software (http://kobas.cbi.pku.edu.cn/home.do) (corrected P-value < 0.05) [30, 31].
Quantitative Real-Time PCR Confirmation of DEGs
The qRT-PCR was performed to evaluate the expression level of 8 randomly selected genes. Total RNA was reverse transcribed by EasyScript First-Strand cDNA Synthesis SuperMix (TransGen, Beijing, China) for gene detection. Specific primers of each gene were designed by the Primer Premier 5.0 and the sequences of the primers are listed in Supplementary Table S4. Real-time PCR reactions were performed by Power SYBR Green PCR Master Mix with the ABI 7500 system. The relative expression level of genes was calculated with reference to expression of GAPDH gene. All experiments were performed in triplicates for each biological repeat. The results are described as fold change determined by the 2−ΔΔCt method. The data are expressed as the mean ± standard error (SE).
Results
Global Overview of Transcriptome Sequencing
To globally decipher the effect of boron on the function of the duodenum in rats, we performed transcriptome sequencing after boron supplementation (at 0, 10, 320 mg/L) using the Illumina HiSeq 4000 platform. Three treatment groups with boron concentration of 0, 10, 320 mg/L were referred to negative control (NC), B10, and B320 groups. After removing sequencing adaptors and low-quality reads from raw data, approximately 59, 49 and 47 million clean reads were obtained for further analysis for the NC, B10, and B320 groups (Q20 > = ~ 97%). These clean reads were then used for aligning to the rat reference genome. According to the mapping results, approximately 95% of the clean reads were successfully mapped to the rat genome (Table 1). The CG content of each treatment group was all almost 50%. The details of sequencing data quality and mapping statistics are presented in Table 1. All of the mapped reads were further assembled into 32, 662 transcripts. Results suggested that the data of transcriptome sequencing was of sufficient quality for further biological analysis to analyze gene expression patterns.
Variation in Gene Expression Among Three Treatment Groups
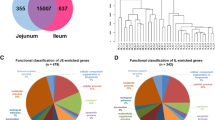
Through comparison of the transcriptome data from three different treatment groups, 624, 391 and 100 differentially expressed genes (DEGs) were screened out (false discovery rate < 0.05, and fold-change ≥ 2 or ≤ 0.5) (Table S1) for the comparison among the NC vs. B10, NC vs. B320 and B10 vs. B320 groups respectively. The fold changes induced by dietary supplementation with different concentration of boron ranged from − 8.13 to 9.14. Sample clustering analysis based on Pearson correlation coefficients showed there were high correlations (0.987–1.000) among samples in different treatment groups (Fig. S1) with greater similarity between the B10 and B320 groups. There were 455 upregulated and 169 downregulated genes in the 10 mg/L boron treatment group compared with control group, 316 upregulated and 75 downregulated genes in the 320 mg/L boron treatment group compared with control group as well as 53 upregulated and 47 downregulated genes in the 320 mg/L boron treatment group compared with 10 mg/L boron treatment group (Fig. 1A–C). There were 346, 125 and 10 specific DEGs in NC vs. B10, NC vs. B320 and B10 vs. B320 comparison groups, respectively (Fig. 1D). Samples between B10 and B320 groups showed some similar expression pattern (Fig. S1). The results of expression pattern showed that 777 DEGs in both NC vs. B10 and NC vs. B320 comparison groups were classified into 10 profiles, of which 702 genes were mostly clustered into two profiles using Short Time-series Expression Miner software (STEM) (Fig. 2A) [32]. Profiles 1 and 3 revealed significant clustering trends (Fig. 2B, C). Profiles 1 and 3 contained 177 and 525 DEGs respectively. The expression level of genes in the profile 1 firstly decreased and then increased slightly while the concentrations of boron increased. The expression level of genes in the profile 3 initially increased and then decreased slightly followed by the increasing concentration of boron.
Profiles of differentially expressed genes (DEGs) after treatment with different concentration of boron. Scatter plot of 624 DEGs between 0 and 10 mg/L boron treatment groups (A), 391 DEGs between 0 and 320 mg/L boron treatment groups (B) and 100 DEGs between 10 and 320 mg/L boron treatment groups (C). Red dots represent extremely significantly upregulated genes (FDR < 0.01, fold change ≥ 2); yellow dots represent significantly upregulated genes (FDR < 0.05, fold change ≥ 2); wathet dots represent significantly downregulated genes (FDR < 0.05, fold change ≤ 0.5); mazarine dots represent extremely significantly downregulated genes (FDR < 0.01, fold change ≤ 0.5); black dots represent insignificantly DEGs. (D) Venn plot of DEGs among 0, 10 and 320 mg/L boron treatment groups
Functional Annotation and Classification of DEGs
Gene Ontology (GO) could provide genes and their products with biological annotation in a wide variety of organisms [29]. The DEGs were used to conduct GO annotation analysis according to three categories including biological process, cellular component, and molecular function in different comparison groups (Fig. 3) [3]. A total of 624 genes between 0 and 10 mg/L of boron treatment groups were allocated to three primary GO categories, which were subsequently assigned to 60 functional terms (Fig. 3A, Table S2). The total upregulated genes were mainly related to cellular process (310), cell (309), cell part (309), single-organism process (302), and binding (300). The total downregulated genes were mostly related to single-organism process (122), cell (114), cell part (113), cellular process (112), and binding (107). A total of 391 genes between 0 and 320 mg/L of boron treatment groups were assigned to 59 functional terms (Fig. 3B, Table S2). The total upregulated genes were also related to cell (215), cell part (215), cellular process (214), single-organism process (210), and binding (203). The total downregulated genes were mainly related to metabolic process (54), binding (52), single-organism process (49), cellular process (48), cell part (44), and cell (44). A total of 100 genes between 10 and 320 mg/L of boron treatment groups were allocated to 52 functional terms (Fig. S3, Table S2).
Annotation of DEGs using Gene ontology after treatment with different concentration of boron. The number of genes for each GO annotation is exhibited in lower axis, and the proportion of genes for each GO annotation is listed in upper axis. A. GO annotation of DEGs between 0 and 10 mg/L boron treatment groups. B. GO annotation of DEGs between 0 and 320 mg/L boron treatment groups
GO Enrichment and KEGG Pathway Analysis of DEGs
We performed GO enrichment analysis to decipher the major functional categories represented in these DEGs. Of these categories, most DEGs were enriched in the “biological process” category (Fig. 4, Table S3), in which response to virus, defense response to virus, innate immune response, and cellular response to interferon-beta were the most primary subcategories. As for the “cellular component” category, most DEGs were assigned to extracellular space, external side of plasma membrane, brush border membrane, and cell surface. Within the “molecular function” category, the mainly enriched subcategories were double-stranded RNA binding, 2′-5′-oligoadenylate synthetase activity, zinc ion binding, and peptide antigen binding. The following genes participate in the above GO terms: Mx2, Rsad2, RT1-T24-3, Ifit3, Aim2, Oas1i, Car4, RT1-Bb, and RT1-Ba.
These DEGs were also used for the KEGG pathway analysis (Fig. 5, Table S3). The top 15 KEGG pathways with the highest representation of the DEGs were shown in Fig. 5. The DEGs were enriched in the main pathways including influenza A (ko05164), herpes simplex infection (ko05168), cytosolic DNA-sensing pathway (ko04623), and antigen processing and presentation (ko04612).
Confirmation of Gene Expression with Quantitative RT-PCR
To confirm the differentially expressed genes of duodenum between control and treatment groups obtained by RNA-Seq, eight differential expressed genes were randomly selected and their expression patterns were quantified by qRT-PCR. The results by qRT-PCR were consistent with that of RNA-Seq. The qRT-PCR results showed the similar down- or upregulated trend in the expression of these genes (Fig. 6). Overall, the validation of eight randomly selected genes by qRT-PCR confirmed the accuracy of the results of RNA-Seq.
Validation of eight randomly selected DEGs by qRT-PCR. A. Expression levels of 8 selected DEGs quantified by qRT-PCR. X-axis represents 8 selected DEGs for qRT-PCR. Left y-axis represents FPKM derived from RNA-Seq results, and right y-axis represents relative expression level derived from qRT-PCR results. Blue and yellow lines depict the RNA-Seq and qRT-PCR results, respectively
Alternative Splicing Analysis of Transcriptome Sequencing
Alternative splicing (AS) plays an important role in generating the protein diversity and complexity of gene expression regulation in higher organisms. In this study, we conducted AS analysis based on RNA-seq data. We identified there were 47 966, 45 151 and 44 555 AS events in NC, B10 and B320 groups respectively. There was no obvious change in AS after treatment with different concentration of boron. TSS, TTS, and SKIP accounted for the major types among the twelve known types of AS (Fig. S3).
Discussion
Our previous studies have found that addition of boron could promote the development and improve the immune capacity of main immune organs in rats and broilers [33, 34]. In the spleen, supplementation of 0.4 mmol/L boron accelerated the proliferation of spleen lymphocytes, while addition of 40 mmol/L boron had inhibitory effect [2]. Furthermore, addition of lower concentrations of boron (10, 20, or 40 mg/L) to drinking water increased IgG in the serum, lymphocyte proliferation and immune function, while high concentration above 320 mg/L had the inhibitory or toxic effect [35, 36]. In the thymus, lower doses of boron (10 and 20 mg/L) elevated the production of IL-2, IL-4, IFN-γ, and the number of proliferating cell nuclear antigen (PCNA) positive cells, while higher doses above 480 mg/L induced the production of pro-inflammatory cytokines to produce inhibitory effects [37]. Supplementation of boron (< 100 mg/L) could improve the growth of immune organs in Gushi chickens [34].
It is reported that lower doses of boron could improve the microstructure and function of duodenum, while higher doses of boron have an inhibitory or even toxic effect in rats. Lower doses of boron increased the number of intraepithelial lymphocytes (IELs) and goblet cells, the expression of secretory IgA, tight junction proteins Zonula occludens-1 and occludin, and PCNA in duodenum of rats [38]. These results indicated that appropriate amount of boron improves immune function in the duodenum and this provided the scientific rationale for this study to further analyze the underlying mechanism involved.
There were fewer DEGs enriched in NC vs. B320 and B10 vs. B320 comparison groups compared with NC vs. B10 comparison groups. Eight of the differentially expressed genes identified by RNA-Seq were randomly selected and validated by qRT-PCR. The expression trends of these genes were similar to the results obtained from the transcriptome data, and it implicated that the quality of sequencing data is accurate and credible.
The DEGs among three treatment groups were enriched in influenza A, herpes simplex infection, and cytosolic DNA-sensing pathway. Moreover, genes RIG-I and IRF7 were shared in these signal pathways, in which both genes were upregulated in the B10 group but decreased in the B320 group with the increasing boron concentrations. RIG-I is involved in viral double-stranded RNA recognition, positive regulation of cytokine production, and the regulation of immune response. RIG-I can maintain intestinal IELs via a type I interferon-independent, MAVS-IRF1-IL-15 axis-dependent manner [39]. RIG-I and MDA5 can sense viral dsRNA to activate the RIG-I-like receptor signaling pathway. During viral infection, RIG-I-like receptors recognize and combine with pathogen-associated molecular patterns (PAMPs), in which their interaction induces the expression of interferon regulatory transcription factor IRF3/7. The induction of IRFs benefits the secretion of type I IFN. Type I IFN secreted by infected cells acts in an autocrine and paracrine manner, leading to the expression of IFN stimulated genes to inhibit viral infection [40, 41]. Moreover, STAT1 and STAT2 proteins mediated type I and type II IFN (IFN-γ) signaling. The both proteins form heterotrimeric complex known as ISGF3 to associate with IFN regulatory factor 9 (IRF9) [42, 43], which results in the production of antiviral proteins MxA in the B10 group involved in the defense against viral infection. ISGF3 promotes the type I IFN-induced transcriptional activation, which acts as a primary defensive barrier for virus infection and promotes the activation of innate and adaptive antiviral immune responses [44].
TRIM-25 induces the ubiquitination of RIG-I and antiviral signaling adaptor MAVS to stabilize their interaction which promotes the downstream signaling [45, 46]. The expression levels of RIG-I, MDA5, and STAT1/2 were upregulated in the B10 vs. NC group, which benefits the activation of downstream RIG-I-like receptor signaling and Jak-STAT signaling pathways. There were fewer cytokines genes expressed in B320 vs. NC group compared with B10 vs. NC group. It indicates that supplement of 10 mg/L boron might be more effective to enhance intestinal immunity than that of 320 mg/L boron in rats.
The cytosolic DNA-sensing pathway provides immune defense against infection by a large variety of DNA-containing pathogens. Cytosolic DNA sensor cGAS was upregulated in B10 group and was involved in activation of innate immune responses through cGAS–STING pathway [47]. The pro-inflammatory cytokines CCL5 and CXCL10 were also upregulated in B10 group which resulted in anti-pathogens responses [48].
The DE genes between B10 and NC group were enriched in antigen processing and presentation. Compared with NC group, complement of 10 mg/L boron increased the gene expression levels of antigen binding and presentation, and induced the activation of MHC-I pathway, which mediated regulation of natural killer (NK) cell activity through NK cell mediated cytotoxicity. The upregulation of CD8a gene promoted the activation of T cell receptor signaling pathway and benefited the maintenance of the intestinal environment. TAPBP was upregulated in B10 compared with NC group. This gene mediated the interaction between newly assembled MHC-I molecules and the transporter associated with antigen processing (TAP), which is required for the transport of antigen peptides across the endoplasmic reticulum membrane [49, 50]. RT1 complex is the major histocompatibility complex (MHC) of the rat. It was predicted to have peptide antigen binding activity and signaling receptor binding activity. Moreover, it was predicted to be involved in antigen processing and presentation of endogenous and exogenous peptide antigen via MHC class I and class II respectively [51, 52]. RT1-T24-3, RT1-S3, RT1-N2, RT1-T24-1, RT1-CE7, RT1-N3, and RT1-T24-4 were upregulated in the B10 group compared with NC group. However, MHC class II molecules such as RT1-Ba and RT1-Bb were downregulated in B320 group compared with NC group [53]. It implicated that appropriate addition of boron might enhance the immune activity of the duodenum.
The differential genes can be enriched in some kinds of diseases such as type I diabetes mellitus, viral myocarditis, graft-versus-host disease, allograft rejection, and autoimmune thyroid disease pathways. The differential genes enriched in these pathways are mainly involved in the inflammatory response in these types of diseases and are used to produce autoimmune responses [54,55,56,57]. The addition of 10 mg/L boron activated more immune response genes than the addition of 320 mg/L boron. Some differential genes are involved in the process of S. aureus infection. The addition of 10 mg/L boron to this process facilitates the activation of complement and coagulation cascades and inhibits the formation of membrane attack complexes, thereby inhibiting the development of the infection process [58].
The DE genes were also enriched in the pancreatic secretion, protein digestion and absorption, fat digestion and absorption, and mineral absorption. As for pancreatic secretion pathway, the B10 vs. NC comparison group have four more DE genes than B320 vs. NC comparison group. The four genes were CLCA4, CAR4, ATP2B1, and ATP1A3, which promoted the secretion of pancreatic juice. As for protein/fat digestion and absorption, the B320 vs. NC comparison group have more DE genes than B10 vs. NC comparison group. But interestingly, the DE genes enriched in these signaling pathways between B10 and NC group were mostly downregulated. However, although the common DE genes between B10 vs. NC and B320 vs. NC comparison groups were downregulated, there were other genes upregulating between B320 and NC group. The addition of 320 mg/L elemental boron to drinking water is toxic to the small intestine of rats. We hypothesize that the organism compensates for the damage caused by high concentration of elemental boron by increasing the metabolic efficiency of the substance, in order to survive in an environment with high concentrations of elemental boron. It was found that high concentration of boron is toxic to cells which impaired protein synthesis by stimulating phosphorylation of eIF2α in the GCN2 kinase dependent manner [11]. Higher concentration of boron-containing compound inhibits the lipid deposition without no evident cytotoxicity on the cells [59]. It needs further study about the supplementary doses of boron to achieve the balance between capacity of digestion and absorption and response to immune reactions.
Conclusion
RNA-Seq analysis exhibited comprehensive gene expression of the duodenum treated with different concentration of boron in rats. Through transcriptome sequencing, we validated that lower dose of boron could achieve better effects on immune ability in the duodenum. Supplementation of 10 mg/L boron could improve the immune function, while boron concentration above 320 mg/L might inhibit the immune ability in the duodenum. These data provided more information for elucidating the underly mechanism of boron in the duodenum tissues.
Availability of Data and Material
The datasets generated and/or analyzed during the current study are available in its supplementary information files.
References
Cakir S, Eren M, Senturk M, Sarica ZS (2018) The effect of boron on some biochemical parameters in experimental diabetic rats. Biol Trace Elem Res 184(1):165–172
Wang C, Jin E, Deng J, Pei Y, Ren M, Hu Q, Gu Y, Li S (2020) GPR30 mediated effects of boron on rat spleen lymphocyte proliferation, apoptosis, and immune function. Food Chem Toxicol 146:111838
C Gene Ontology (2006) The Gene Ontology (GO) project in 2006. Nucleic Acids Res 34(Database issue):D322-326
Akram M, Matiullah IA, Husaini SN, Malik F (2011) Determination of boron contents in water samples collected from the Neelum valley, Azad Kashmir, Pakistan. Biol Trace Elem Res 139(3):287–295
Pizzorno L (2015) Nothing boring about boron. Integr Med (Encinitas) 14(4):35–48
Nielsen FH (2014) Update on human health effects of boron. J Trace Elem Med Biol 28(4):383–387
Hunt CD (2003) Dietary boron: an overview of the evidence for its role in immune function. J Trace Elem Exp Med 16(4):291–306
Gorustovich AA, Nielsen FH (2019) Effects of nutritional deficiency of boron on the bones of the appendicular skeleton of mice. Biol Trace Elem Res 188(1):221–229
Ergul AB, Kara M, Karakukcu C, Tasdemir A, Aslaner H, Ergul MA, Muhtaroglu S, Zararsiz GE, Torun YA (2018) High doses of boron have no protective effect against nephrolithiasis or oxidative stress in a rat model. Biol Trace Elem Res 186(1):218–225
Khaliq H, Jing W, Ke X, Ke-Li Y, Peng-Peng S, Cui L, Wei-Wei Q, Zhixin L, Hua-Zhen L, Hui S, Ju-Ming Z, Ke-Mei P (2018) Boron affects the development of the kidney through modulation of apoptosis, antioxidant capacity, and Nrf2 pathway in the African ostrich chicks. Biol Trace Elem Res 186(1):226–237
Uluisik I, Kaya A, Fomenko DE, Karakaya HC, Carlson BA, Gladyshev VN, Koc A (2011) Boron stress activates the general amino acid control mechanism and inhibits protein synthesis. PLoS One 6(11):e27772
Henderson KA, Kobylewski SE, Yamada KE, Eckhert CD (2015) Boric acid induces cytoplasmic stress granule formation, eIF2alpha phosphorylation, and ATF4 in prostate DU-145 cells. Biometals 28(1):133–141
Kobylewski SE, Henderson KA, Yamada KE, Eckhert CD (2017) Activation of the EIF2alpha/ATF4 and ATF6 pathways in DU-145 cells by boric acid at the concentration reported in men at the US mean boron intake. Biol Trace Elem Res 176(2):278–293
Romero-Aguilar KS, Arciniega-Martínez IM, Farfán-García ED, Campos-Rodríguez R, Reséndiz-Albor AA, Soriano-Ursúa MA (2019) Effects of boron-containing compounds on immune responses: review and patenting trends. Expert Opin Ther Pat 29(5):339–351
Fry RS, Lloyd KE, Jacobi SK, Siciliano PD, Robarge WP, Spears JW (2010) Effect of dietary boron on immune function in growing beef steers. J Anim Physiol Anim Nutr (Berl) 94(3):273–279
Bhasker TV, Gowda NKS, Pal DT, Bhat SK, Krishnamoorthy P, Mondal S, Pattanaik AK, Verma AK (2017) Influence of boron supplementation on performance, immunity and antioxidant status of lambs fed diets with or without adequate level of calcium. PLoS One 12(11):e0187203
Xiao K, Yang K, Wang J, Sun P, Huang H, Khaliq H, Naeem MA, Zhong J, Peng K (2019) Transcriptional study revealed that boron supplementation may alter the immune-related genes through MAPK signaling in ostrich chick thymus. Biol Trace Elem Res 189(1):209–223
Mowat AM, Agace WW (2014) Regional specialization within the intestinal immune system. Nat Rev Immunol 14(10):667–685
Agace WW, McCoy KD (2017) Regionalized development and maintenance of the intestinal adaptive immune landscape. Immunity 46(4):532–548
Mucida D, Esterhazy D (2018) SnapShot: gut immune niches. Cell 174(6):1600-1600 e1601
Thompson FM, Mayrhofer G, Cummins AG (1996) Dependence of epithelial growth of the small intestine on T-cell activation during weaning in the rat. Gastroenterology 111(1):37–44
Ruemmele FM, Seidman EG, Lentze MJ (2002) Regulation of intestinal epithelial cell apoptosis and the pathogenesis of inflammatory bowel disorders. J Pediatr Gastr Nutr 34(3):254–260
Cock PJ, Fields CJ, Goto N, Heuer ML, Rice PM (2010) The Sanger FASTQ file format for sequences with quality scores, and the Solexa/Illumina FASTQ variants. Nucleic Acids Res 38(6):1767–1771
Shen S, Park JW, Huang J, Dittmar KA, Lu ZX, Zhou Q, Carstens RP, Xing Y (2012) MATS: a Bayesian framework for flexible detection of differential alternative splicing from RNA-Seq data. Nucleic Acids Res 40(8):e61
Robinson MD, McCarthy DJ, Smyth GK (2010) edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26(1):139–140
Anders S, Huber W (2010) Differential expression analysis for sequence count data. Genome Biol 11(10):R106
Yoav B, Yosef H (1995) Controlling the false discovery rate: a practical and powerful approach to multiple testing. J R Stat Soc B (Methodol) 57(1):289–300
Young P (1989) p value adjustments for multiple tests in multivariate binomial models. J Am Stat Assoc 84(407):780
C Gene Ontology (2008) The Gene Ontology project in 2008. Nucleic Acids Res 36(Database issue):D440-444
Kanehisa M, Araki M, Goto S, Hattori M, Hirakawa M, Itoh M, Katayama T, Kawashima S, Okuda S, Tokimatsu T, Yamanishi Y (2008) KEGG for linking genomes to life and the environment. Nucleic Acids Res 36(Database issue):D480-484
Xie C, Mao X, Huang J, Ding Y, Wu J, Dong S, Kong L, Gao G, Li CY, Wei L (2011) KOBAS 2.0: a web server for annotation and identification of enriched pathways and diseases. Nucleic Acids Res 39(Web Server issue):W316-322
Ernst J, Bar-Joseph Z (2006) STEM: a tool for the analysis of short time series gene expression data. BMC Bioinformatics 7:191
Hu Q, Li S, Qiao E, Tang Z, Jin E, Jin G, Gu Y (2014) Effects of boron on structure and antioxidative activities of spleen in rats. Biol Trace Elem Res 158(1):73–80
Jin E, Gu Y, Wang J, Jin G, Li S (2016) Effect of supplementation of drinking water with different levels of boron on performance and immune organ parameters of broilers. Ita J Anim Sci 13(2):3152
Jin E, Li S, Ren M, Hu Q, Gu Y, Li K (2017) Boron affects immune function through modulation of splenic T lymphocyte subsets, cytokine secretion, and lymphocyte proliferation and apoptosis in rats. Biol Trace Elem Res 178(2):261–275
Jin E, Pei Y, Liu T, Ren M, Hu Q, Gu Y, Li S (2019) Effects of boron on the proliferation, apoptosis and immune function of splenic lymphocytes through ERα and ERβ. Food Agr Immunol 30(1):743–761
Jin E, Ren M, Liu W, Liang S, Hu Q, Gu Y, Li S (2017) Effect of boron on thymic cytokine expression, hormone secretion, antioxidant functions, cell proliferation, and apoptosis potential via the extracellular signal-regulated kinases 1 and 2 signaling pathway. J Agric Food Chem 65(51):11280–11291
Liu T, Wang C, Wu X, Ren M, Hu Q, Jin E, Gu Y (2021) Effect of boron on microstructure, immune function, expression of tight junction protein, cell proliferation and apoptosis of duodenum in rats. Biol Trace Elem Res 199(1):205–215
Liu L, Gong T, Tao W, Lin B, Li C, Zheng X, Zhu S, Jiang W, Zhou R (2019) Commensal viruses maintain intestinal intraepithelial lymphocytes via noncanonical RIG-I signaling. Nat Immunol 20(12):1681–1691
Broquet AH, Hirata Y, McAllister CS, Kagnoff MF (2011) RIG-I/MDA5/MAVS are required to signal a protective IFN response in rotavirus-infected intestinal epithelium. J Immunol 186(3):1618–1626
Loo YM, Gale MJ (2011) Immune signaling by RIG-I-like receptors. Immunity 34(5):680–692
Fu XY, Kessler DS, Veals SA, Levy DE, Darnell JE Jr (1990) ISGF3, the transcriptional activator induced by interferon alpha, consists of multiple interacting polypeptide chains. Proc Natl Acad Sci U S A 87(21):8555–8559
Stark GR, Darnell JE Jr (2012) The JAK-STAT pathway at twenty. Immunity 36(4):503–514
Au-Yeung N, Mandhana R, Horvath CM (2013) Transcriptional regulation by STAT1 and STAT2 in the interferon JAK-STAT pathway. JAK-STAT 2(3):e23931
Castanier C, Zemirli N, Portier A, Garcin D, Bidère N, Vazquez A, Arnoult D (2012) MAVS ubiquitination by the E3 ligase TRIM25 and degradation by the proteasome is involved in type I interferon production after activation of the antiviral RIG-I-like receptors. BMC Biol 10(1):44
Gack MU, Shin YC, Joo CH, Urano T, Liang C, Sun L, Takeuchi O, Akira S, Chen Z, Inoue S, Jung JU (2007) TRIM25 RING-finger E3 ubiquitin ligase is essential for RIG-I-mediated antiviral activity. Nature 446(7138):916–920
Chen Q, Sun L, Chen ZJ (2016) Regulation and function of the cGAS–STING pathway of cytosolic DNA sensing. Nat Immunol 17(10):1142–1149
Liu M, Guo S, Hibbert JM, Jain V, Singh N, Wilson NO, Stiles JK (2011) CXCL10/IP-10 in infectious diseases pathogenesis and potential therapeutic implications. Cytokine Growth F R 22(3):121–130
Cui D, Wang J, Zeng Y, Rao L, Chen H, Li W, Li Y, Li H, Cui C, Xiao L (2016) Generating hESCs with reduced immunogenicity by disrupting TAP1 or TAPBP. Biosci Biotechnol Biochem 80(8):1484–1491
Gonzalez OA, Novak MJ, Kirakodu S, Orraca L, Chen KC, Stromberg A, Gonzalez-Martinez J, Ebersole JL (2014) Comparative analysis of gingival tissue antigen presentation pathways in ageing and periodontitis. J Clin Periodontol 41(4):327–339
Gunther E, Walter L (2001) The major histocompatibility complex of the rat (Rattus norvegicus). Immunogenetics 53(7):520–542
Hurt P, Walter L, Sudbrak R, Klages S, Müller I, Shiina T, Inoko H, Lehrach H, Günther E, Reinhardt R (2004) The genomic sequence and comparative analysis of the rat major histocompatibility complex. Genome Res 14(4):631–639
Dressel R, Walter L, Günther E (2001) Genomic and funtional aspects of the rat MHC, the RT1 complex. Immunol Rev 184:82–95
Wang K, Li F, Cui Y, Cui C, Cao Z, Xu K, Han S, Zhu P, Sun Y (2019) The association between depression and type 1 diabetes mellitus: inflammatory cytokines as ferrymen in between? Mediat Inflamm 2019:1–11
Corsten MF, Schroen B, Heymans S (2012) Inflammation in viral myocarditis: friend or foe? Trends Mol Med 18(7):426–437
Kindermann I, Barth C, Mahfoud F, Ukena C, Lenski M, Yilmaz A, Klingel K, Kandolf R, Sechtem U, Cooper LT (2012) Update on myocarditis. J Am Coll Cardiol 59(9):779–792
Mikoś H, Mikoś M, Obara-Moszyńska M, Niedziela M (2014) The role of the immune system and cytokines involved in the pathogenesis of autoimmune thyroid disease (AITD). Endokrynol Pol 65(2):150–155
Killick J, Morisse G, Sieger D, Astier AL (2018) Complement as a regulator of adaptive immunity. Semin Immunopathol 40(1):37–48
Abdik EA, Abdik H, Tasli PN, Deniz AAH, Sahin F (2019) Suppressive role of boron on adipogenic differentiation and fat deposition in human mesenchymal stem cells. Biol Trace Elem Res 188(2):384–392
Funding
The work was supported in part by the National Natural Science Foundation of China (32002160, 32172816, and 31672502), Anhui Provincial Major Science and Technology Special Program (17030701004, 201903a06020002), Excellent Young Talents Fund Program of Higher Education Institutions of Anhui Province (gxyqZD2019061), Chuzhou Science and Technology Plan (2019ZN003), the University Research Project of Anhui Province (KJ2020A0081, KJ2019A0801), Anhui Provincial Natural Science Foundation (2108085MC117, 2008085QC140), Anhui Provincial Collaborative Innovation Program (GXXT-2019–035), Foundation of Anhui Science and Technology University (DKYJ201901), and the Innovation Funds for Undergraduate Students of Anhui Province (S202010879109, S202010879120, 202110879059, S202110879175).
Author information
Authors and Affiliations
Contributions
CZ, SL and EJ designed the study. CZ and YH performed the experiments. CZ, YH, and CW wrote the manuscript. CZ, MR, and QH analyzed the data. YG, PY, SL, and EJ helped revise the manuscript. All authors discussed the data. All authors reviewed the manuscript.
Corresponding authors
Ethics declarations
The use of all experimental animals was reviewed and approved by the Anhui Laboratory Animal Care Committee. All animal experimental procedures were performed in strict accordance with the Guide for Laboratory Animal Care and Use and complied with the Guide for National Laboratory Animal Healthcare and Use.
Conflict of Interest
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Zhao, C., Han, Y., Wang, C. et al. Transcriptome Profiling of Duodenum Reveals the Importance of Boron Supplementation in Modulating Immune Activities in Rats. Biol Trace Elem Res 200, 3762–3773 (2022). https://doi.org/10.1007/s12011-021-02983-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12011-021-02983-w