Abstract
Allelic variation is a valuable tool for displaying high levels of polymorphism within species and is closely correlated with crop productivity. In Marathawada, there is a significant amount of phenotypic heterogeneity among sorghum landraces. However, molecular variability needs to be reevaluated in order to identify any potential barriers that can interfere with current improvement initiatives. In the current work, we used 5 SSR markers to categorize 20 genotypes of elite (Sorghum bicolor L. Moench) accession from the Marathwada region, including one standard cultivar from various agro-economic zones. According to the results of this study, 14 alleles were found among the 20 genotypes, with a PIC value that ranged from 0.37 to 0.70 and a mean of 0.44 per locus. Each locus had anything from 1 (gpsb089) and 5 (mSbCIR223), with an average of 2.80 alleles per locus. A neighbor-joining tree was constructed and showed clustering of genotypes into two groups; this indicates that there is considerable diversity in genotypes compared with advanced cultivar for desired genotype (IS1042) by using SSR markers. Results show that most diverse cultivars were IS-4564, IS18357, and IS-18381, and significant variation was also reported in IS4566 and IS18379.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Sorghum (Sorghum bicolor L. Moench) is the third most important grain crop in India, after rice, wheat, maize, and millet [32]. Maharashtra is the leading producer of sorghum, ranking third after wheat and rice, regardless of the fact that prior to the agricultural revolution, sorghum was ranked second in India in terms of area and production [26]. Globally, it is the fifth most important cereal after rice, wheat, maize, and barley. In the production of alcohol, sorghum is used to produce ethanol, starch, and syrups [2].
Sorghum belongs to the Poaceae family, the genus Andropogoneae, and the species Sorghastrum. Sorghum originated from the Greek word “sorgo” that means to supersede, as in having grown taller than in other crops in the field. The Sorghum family has been arranged into two all-around emblazoned areas by specifically Eu-sorghum and Para-sorghum [8]. Divided the category Sorghum into six sub-genera, namely Eu-sorghum, Chaetero-sorghum, Hetero-sorghum, Para-sorghum, Sorghastrum, and Stipo-sorghum, and officially assigned the specific name Sorghum imlgare, to the detailed outline and its cross-viable wild and weed species members [3]. Despite, Price et al. [12] assumed the title Sorghum bicolor (L.) Moench was correct, and this idea has indeed been endorsed by Kong et al. [31]. Sorghum bicolor (L.) Moench includes yearly consist of varying with 10 sets of chromosomes (2n = 20), which includes all advanced grain sorghum and Sorghum-sudan grass crossovers, as well as their cross-viable wild and weedy relatives.
The primary focus is on genetic analyses of sorghum for its excellent quality. (Sorghum bicolor (L.), 2n = 2x = 20) is the nation’s fifth most important food crop after wheat, rice, maize, and barley, with over 80% of the crop grown in Africa and Asia [7]. The ICRISAT sorghum collection represents the major sorghum diversity centers. The National Plant Genetic Resources Centers house a wide range of sorghum landraces. The ICRISAT gene bank is a major source of diversity for subtropical agricultural crops [4].
SSR markers (simple sequence repeats) are suitable to study genetic variability. Typical landraces, optimized cultivars, and breeder’s cultivars are among the accessions. A further important feature of SSR markers is their convenience of its use [16]. DNA markers are typically nucleotide sequences that conform to a physical in a genome as well as their polymorphisms. SSR fingerprints are often extremely specific, and they are frequently used to differentiate various individuals and identities, as well as to expose polymorphism [14, 24].
Simple sequence repeats (SSR) markers—also called microsatellites—instantly became the DNA markers of choice for plant and animal genomes [5]. SSRs are tandem repeats of di-, tri-, tetra-, penta-, or hexanucleotide units in the DNA of plants [24]. A good number of microsatellite markers have been developed for sorghum that allows so much high rate of sorghum genotype assessment [20].
The analysis of genetic diversity in 20 sorghum accessions is from the ICRISAT gene bank, Hyderabad, India, both for agro-morphological character and SSR markers. Protein and mineral composition vary considerably within and between 20 sorghum accessions from the ICRISAT gene bank in Hyderabad, India [6]. The accessibility of such markers for evaluating genetic variation is a simple way for conventional breeding to offer integrated genetic traits for breeding programs. Predefined kinds of molecular markers used for sorghum diversity evaluations became relevant, as well as they differ in complexity, reliability, and information generating capacity [22]. When genetic diversity was assessed using microsatellite (SSR) markers, a p < 0.05 of genetic variation was evidenced among 20 sorghum accessions from different agro-ecological regions in Marathwada (70% between many accessions and 30% within accessions) [19].
Molecular markers and its value have indeed been demonstrated in studies of genetic variation (Morand et al., 2002; Zeid et al., 2003), breeding systems (Durand et al., 2000), and pollination biology (Durand et al., 2000), including pollination ecology (Durand et al., 2000) (White et al., 2002). Crop diversification has increased crop production such as wheat via domestication, yet this has diminished crop genetic diversity. As a consequence, a strategy to enhance the advancement of a crop is necessary. Advanced species is determined by genetic diversity. For the protection and use of germplasm of particular species diversity, genetic diversity is essential.
To broaden the genetic base for crop improvement, it is essential to reveal genetic variation. The employment of markers serves the purpose of revealing genetic diversity. Variety understanding also assists in the development of methods for integrating relevant variety into breeding plans. Use of markers to describe crops reveals both similarities (shared alleles) and diversity (typical alleles) across cultivars of the same crop. Also, it aids in determining the genetic pool or provenance of the selected crop. The genetic basis should be extended in order to develop elite cultivars. The species whose genetic diversity has indeed been discovered utilizing markers are utilized to generate superior hybrids via germ line exchanges. Markers are used to characterize and construct a genetic profile. Such crop DNA profiles are used to maintain a crop’s gene pool in gene bank.
Materials and Methods
The twenty seed material accessions for the current experiment were obtained from ICRISAT gene bank, originally obtained from various locations of Maharashtra, passport data for seeds listed in Table 1; these cultivars were selected because they had 90 to 100% germination rates and are cultivated in the semi-arid regions of Maharashtra, India. Selected seeds were allowed to germinate by blotter paper method, under controlled and optimum conditions on the Petri dishes lined with 75% humidity by Whatman no. 1 filter papers and incubated at 30 °C. After germination, 16 days old leaf tissues were harvested from 20 different sorghum accessions harvested and stored at − 20 °C for DNA extraction.
DNA Extraction
Genomic DNA extracted from 4 days old seedlings were performed by the CTAB extraction method modified by Khan et al. [10]. Three to four seeding from each sample was selected randomly and root and shoot both sampled and used for DNA isolation. Genomic DNA was purified by RNAse treatment. Concentration of purified DNA was measured both spectrophotometrically by using the UV visible spectrophotometer at 260 and 280 nm and was calculated, and purity was also confirmed by gel electrophoresis in electrophoresis system in 0.8% (w/v) agarose gel containing 0.5 µl/ml ethidium bromide at 6 V/Cm in 1X TBE buffer. About 20 to 100-ng total DNA were selected for the next experiment [18].
Polymerase Chain Reaction (PCR) Amplification of SSR
The PCR program was set in a thermal cycler amplification reaction mixture prepared in 0.2-ml thin-walled flat cap PCR tubes, containing the following components. The total volume of each reaction mixture was 20 µl; details of PCR master mixture are given in Table 2. Five pairs of SSR primers were chosen for the current experiment for analysis of diversity among selected accessions were part of commercial SSR primer, i.e., kit29 (http://sorghum.cirad.fr/SSR_kit), which provides reasonable coverage across the sorghum nuclear genome. Details of the primer sequences and type of microsatellite repeats are depicted.
In (Table 3) PCR, the PCR program consisted of an initial denaturation for 30 s at 94 °C and then 30 cycles of denaturation for 30 s at 94 °C, annealing at 57 °C (55 or 60 °C) for 60 s, depending on the annealing temperature for the primer, and extension at 72 °C for 60 s. The last PCR cycle was followed by a 5-min extension at 72 °C and then put on hold at 10 °C, at infinity (∞). The amplified products were stored at − 20 °C until they were needed to run gels [1, 15].
Gel Alectrophoresis of PCR Products
After amplification amplified product first separated by gel electrophoresis, for this purpose, gel was run using a 2.5% agarose gel. The gel contained 6.25 g of agarose, weighed using a Mettler Toledo electronic balance, dissolved in 250 ml TAE in a conical flask (volume of flask is required) and the mouth covered with cotton wool. The mixture was then heated in a microwave to dissolve finally and allowed to cool to about 60 °C, and 5 µl of ethidium bromide was added [28, 29]. This was swirled gently to avoid bubbling, and when cooled, the solution was poured into a mold with comb placed on a level surface and allowed to cool and solidify. The comb was gently removed, and the gel was transferred into an electrophoretic tank filled with a 1X TAE buffer. PCR products of 10 µl were loaded into each well and run at 90 V for 45 min. The gel was then observed under an UV trans illuminator and Gel Doc. Each set of PCR product data was obtained from each accession and was scored [27].
Scoring SSR Bands
Product from any of the eight primers with different dye labels were pooled into groups based on their respective agarose band length and resolution capacity of dye labels. One microliter from each co-loaded group was added to 8 µl of solution containing 0.108 µl of GSLIZ500 internal size standards to 8 µl master mix and vortexing at 900 RCF for solution mixing. Vortexted samples are denatured and immediately placed on ice for 3 min. Each fragment analyzes separately to estimate particular fragment size and allele sizing, and detection of homozygous and heterozygous was done with Gene Mapper software; the amplified fragments were visualized using the GEL DOCUMENTARY unit. Connected with Image Reader software version 1.01, size of DNA fragments was determined using Pe which could spontaneously recognize the lane and band and measure the fragment size. Genotypes of individual plants were indicated by the allele size at all SSR loci, and the pairwise different method was determined for each haplotype pair using the parameters. Polymorphic information content (PIC) values for each primer set were calculated using the algorithm described by Smith (1997), Varshney et al., and Davila et al. [11, 23].
where \({f}_{\mathrm{i}}^{2}\) is the frequency of ith allele. PIC is the estimated polymorphism of locus by taking into consideration that not only no of allele who expresses but also estimates frequency of alleles. PIC value ranges from 0 (monomorphic) to 1 (polymorphic). Statistical analysis of data was performed using Power Marker ver. 3.25 to estimates genetic distance, locus, hetrozygosity, PIC content, allele size, length, range, and abundance within chromosome. It is also reported for allele size of loci, and product of SSR was used for genotyping [21]. The genetic diversity of the landraces was measured at different levels of population structure. The genetic variation of each locus shows the utility and general information of each SSR tool [13].
Result and Discussion
Five SSR primers located variable regions of chromosomes that we selected for study of in vitro variability among sorghum accession were found highly polymorphic in nature and therefore provided a powerful data for evaluation of molecular diversity within sorghum landraces. These results also find close relation with Wang Min et al. [30] which concludes that wild sorghum and landraces possessed greater allelic diversity than cultivars and might contain alleles not present in the cultivars which can strengthen further conservation and utilization. It means that higher level of polymorphism is directly proportional with unique replication slippage and mutation responsible for generation of SSR allelic diversity [6, 14]. As results indicate that 5 markers involved in this study average 14 alleles were observed with 2.5 alleles per locus, this is a direct and indirect consequence of artificial selection for traits that improved agronomic qualities; the favorable alleles and variation allow to fixation of traits among population during introduction and acclimatization of species (Table 4). Similar results were found by Murray et al. and Ritter et al. [17, 25]. Number of microsatellite in SSR marker correlates with allele no and PIC (polymorphic information content) ratio which was found 0.37–0.70 with average mean of 0.44; SSR markers with di, tri, and tetra repeats show more no of PIC ratio than mono repeats; this also means no of repeats is associates with higher no of microsatellite and higher PIC number; similar results were found by Afolayan et al. [6], in which he also added that PIC cannot be more than 0.50, which is contrary to what is obtainable for multi-allelic markers, but according to Smita Shingane et al. [27], PIC value can give better and more information on variability, such as SSRs, in which the maximum no of repeats. This further confirms the fact that this detects more alleles than RAPD markers [8]. Similar PIC values found by Davila et al. and Afolayan et al. [6, 11] were significant with Akansha et al. [13] and El-Awady et al. [5] but significantly at par with Davila et al. [11]. Thus, the above shows that significant variation is observed in RAPD, SSR, and RFLP markers (Table 5). Findings confirm effective application of SSR markers over other in relation with evaluating allele frequency and PIC content. Average gene diversity is high, and according to Onyango and Ngugi [21], this strong allelic variation is due to di-nucleotide repeats containing SSRs and is the result of high mutation rates among such repeats. Earlier studies evaluate genetic relation between the cultivars and landraces of wild relatives were successfully identified by Ajeigbe et al. [7] and Maugham et al. (1995) on basis of his studies; 20 sorghum accessions can be divided into three different groups; hence, SSR marker analysis is effective tool for quantification of genetic diversity.
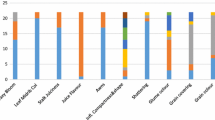
Out of five SSR markers, MsbCIR223 and XTSEP SSRs usually had more no of SSR loci in 20 accessions than the other; this may be due to the difference in allelic no and position in chromosome. XTSEP markers are reported to be extracted from a small tandem repeat library (Brown et al., 1996; Kong et al., 2000). These loci therefore were more likely to include non-coding regions than the other SSR primers. A neighbor-joining (NJ) tree was constructed based on genetic distance (Fig. 1). The group 1 in NJ tree consists of most of the traditional landraces, whereas the group 2 consists of mostly breeding material. Furthermore, in group 1 and 2, the accessions were classified into sub-sub groups based on their origin and types. Similar observations were arrived by many investigators in different crops.
Our results suggest that the sorghum accessions from different geographic origins or institutions could come from the same gene pool and may not be useful for continual sorghum improvements. Therefore, when planning for a new germplasm collection and conservation, it is imperative to characterize the available germplasm before commencing on a breeding program. Conclusively, our study revealed high genetic diversity within the sorghum germplasm. Hence, these sorghum accessions will be protected from genetic vulnerability to biotic and abiotic stress in environment, and breeder will guide how to utilize and conserve germplasm.
Data Availability
In this study, we conducted a comprehensive genetic analysis using SSR markers to investigate the diversity among various genotypes. Our findings reveal a clear segregation of genotypes into two distinct groups, highlighting the remarkable genetic diversity compared to the desired genotype, IS1042. Notably, we identified IS-4564, IS18357, and IS-18381 as the most diverse cultivars, underlining their potential significance for future breeding efforts. Additionally, we observed significant variability in IS4566 and IS18379, further enriching our understanding of genetic diversity within this crop. Among the five SSR markers utilized, MsbCIR223 and XTSEP SSRs consistently exhibited a higher number of SSR loci across 20 accessions. This variation is likely a result of differences in allelic numbers and chromosomal positions. Interestingly, XTSEP markers, derived from a small tandem repeat library, may encompass a greater proportion of non-coding regions compared to other SSR primers, shedding light on their unique genetic characteristics. In order to visually represent the genetic diversity, we constructed a neighbor-joining (NJ) tree based on genetic distance (Fig. 1). Group 1 in the NJ tree predominantly comprises traditional landraces, while Group 2 primarily consists of breeding materials. Moreover, within both groups, we classified accessions into sub-subgroups based on their origin and types, aligning with patterns observed in various crops studied by different researchers. These findings underscore the substantial genetic diversity present in the genotypes studied, with specific cultivars emerging as particularly diverse. The variability in SSR markers suggests potential implications for breeding strategies and underscores the importance of these markers in genetic studies. As part of our commitment to transparency and data sharing, we are pleased to provide a data availability statement in accordance with the journal's guidelines. Researchers interested in accessing the raw data used in this study are encouraged to contact us directly at [vikasshukre@gmail.com]. Sharing data promotes collaboration and enhances the reproducibility of our research, contributing to the scientific community's collective knowledge.
References
Murray, A. E., & J. T. (1996). Phylogenetic compositions of bacterioplankton from two California estuaries compared by denaturing gradient gel electrophoresis of 16S rDNA fragments. Applied and Enviromental Microbiology, pp. 2676–2680. https://doi.org/10.1128/aem.62.7.2676-2680.1996
Nayak, A. K., & Tripathi, R. T. (2022). Carbon and water footprint of rice, wheat & maize crop productions in India. Pedosphere-Elesvier. https://doi.org/10.1016/j.pedsph.2022.06.045
Chengcheng Tan, Y. W. (2010). Development of SSR markers for bermudamass from its expressed sequence tag sequence and prexisting sorghum SSR markers. Molecular Breeding, 29, 23–30.
Clarissa Kimber, J. D. (2012). The gene pool of sorghum bicolor and its improvement. Genomics of the Saccharinae, 11(2), 23–41. https://doi.org/10.1007/978-1-4419-5947-8_2
El-Awady, M., Sawsan, S. Y., Selim, E. E. M., & Marwa, M. G. (2008). Genetic diversity among sorghum bicolor genotypes using simple sequence repeats (SSRs) markers . Arab Journal of Biotechnology, pp. 181–192.
Afolayan, G., S. P.etal, (2019). Genetic diversity assessment of sorghum (Sorghum bicolor (L.) Moench) accessions using single nucleotide polymorphism markers. Plant Genetic Resources: Characterization and Utilization, pp. 1–9. https://doi.org/10.1017/S1479262119000212
Ajeigbe, H. A., & Angarawai, I. (2020). Improved agronomic practices of sorghum production in north east (pp. 1–13). ICRISAT.
Hariprasanna, K., Patil, J. V., & Rajendra Kumar P. (2015). Sorghum: Origin, classification, biology and improvement. In R. K. Madhusudhana R, Sorghum: Origin, Classification, Biology and Improvement (pp. 1–26). Elesivier. https://doi.org/10.1007/978-81-322-2422-8
Huo, Y., Zhao, Y., Xu, L., et al. (2021). An integrated strategy for target SSR genotyping with toleration of nucleotide variations in the SSRs and flanking regions. BMC Bioinformatics, 22, 429. https://doi.org/10.1186/s12859-021-04351-w
Khan, A., & Awan, F. S. (2004). A modified mini-prep method for economical and rapid extraction of genomic DNA in plants. Plant Molecular Biology Reporter, 22(1), 89.
Davila, J. A., & Loarce, Y. (2004). Diversity, comparison of RAMP and SSR markers for the study of wild barley genetic. Hereditas, 131(1), 5–13.
James, Price H., et al. (2005). Genome Evolution in the Genus Sorghum (Poaceae). Annals of Botany, 95(1), 219–227. https://doi.org/10.1093/aob/mci015
Joshi Akansha, R., & Kale Sonam, S. (2020). Genetic diversity among elite sorghum (Sorghum bicolor L.) accessions genotyped with SSR markers to enhance use of global genetic resources. International Journal of Chemical Studies, 8(2), 1691–1697.
Agarwal, M., Shrivastava, N., & Padh, H. (2008). Advances in molecular marker techniques and their applications in plant sciences. Plant Cell Rep, 27, 617–631. https://doi.org/10.1007/s00299-008-0507-z
Jain, R., Rivera, M. C., Moore, J. E., & Lake, J. A. (2003). Gene transfer accelerates genome innovation and evolution. Molecular Biology and Evolution,1598-602. https://doi.org/10.1093/molbev/msg154
Min, W., Run-zhi, L., Wan-ming, Y., & Wei-jun, D. (2010). Assessing the genetic diversity of cultivars and wild soybeans using SSR markers. African Journal of Biotechnology, 9(31), 4857–4866.
Murray, S. C., & Rooney, W., et al. (2009). Sweet sorghum genetic diversity and association mapping for brix and height. The Plant Genome, pp. 48–62. https://doi.org/10.3835/plantgenome2008.10.0011
Stein, N., & Herren, G. (2008). A new DNA extraction method for high-throughput marker analysis in a large-genome species such as Triticum aestivum. Plant Breeding, 120(4), 354–356.
NemeraGeleta, M. T. (2006). Genetic diversity analysis in sorghum germplasm as estimated by AFLP, SSR and morpho-agronomical markers. Biodiversity & Conservation, 15(6), 3251–3265.
Ng'uni, D., Geleta, M., & Hofvander, P., et al ,(2012). Comparative genetic diversity and nutritional quality variation among some important Southern African sorghum accessions ['Sorghum bicolor' (L.) Moench]. Australian Journal of Crop Science, pp. 56–6.
Onyango, K. N. (2012). Analysis of the molecular diversity of Kenyan sorghum germplasm using microsatellites. Journal of Crop Science and Biotechnology, 189–194.
Cavagnaro, P. F., & Chung, S.-M. (2011). Microsatellite isolation and marker development in carrot - genomic distribution, linkage mapping, genetic diversity analysis and marker transferability across Apiaceae. BMC Genomics, 12(1), 386. https://doi.org/10.1186/1471-2164-12-386
Varshney, R. K., & Marcel, T. C. et al. (2007). A high density barley microsatellite consensus map with 775 SSR loci. Theoretical and Applied Genetics, pp. 1091–1103. https://doi.org/10.1007/s00122-007-0503-7
Rajwant, K., & Kalia, M. K. et al. (2011). Microsatellite markers: An overview of the recent progress in plants. Euphytica, pp. 309–334. https://doi.org/10.1007/s10681-010-0286-9
Ritter, K. B., & McIntyre, C. L. (2007). An assessment of the genetic relationship between sweet and grain sorghums, within Sorghum bicolor ssp. bicolor (L.) Moench, using AFLP markers. Euphytica, pp. 161–167. https://doi.org/10.1007/s10681-007-9408-4
Saleem Shaik, A. J. (2004). Indian state-level rice productivity and its impact on poverty alleviation. New Delhi: IDEAS/RePEc.
SmitaShingane, J. V. (2018). Assessing genetic diversity among foxtail millet (Setaria italica (L.) P. Beauv.) Accessions Using. International Journal of Bio-resource and Stress Management, 9(1), 1–6.
Stirling, D. (1989). Multiplex amplification refractory mutation system for the detection of prothrombotic polymorphisms. In D. Stirling, PCR Protocols, pp. 323–325. https://doi.org/10.1385/1-59259-384-4:323
Dubey, S. C, & Tripathi, A., et al. (2014). Diversity of Rhizoctonia solani associated with pulse crops in different agro-ecological regions of India. World Journal of Microbiology and Biotechnology, pp. 1699–1715. https://doi.org/10.1007/s11274-013-1590-z
Min, W., L. R.-z.-m.-j., et al. (2010). Assessing the genetic diversity of cultivars and wild soybeans using SSR markers. African Journal of Biotechnology V, pp. 4857–4866.
WenQian Kong, C. K. et al.(2018). Genotyping by sequencing of 393 sorghum bicolor BTX623 X IS3620 recombinant inbreed lines improves sensitivity and resolution of QTL lines. Genes, Genomes and Genetics, pp. 2563–2572. https://doi.org/10.1534/g3.118.200173
Zabala, E., DeJesus, N. G., et al. (2015). Acceptability of food products from sweet sorghum Sorghum bicolor L. Moench grain developed at Pampanga Agricultural College, Philippines. International Journal on Advanced Science Engineering and Information Technology, p. 286. https://doi.org/10.18517/ijaseit.5.4.541
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Shukre, V.M., Tahakik, R., Kumar, K.G. et al. In Vitro Screening of Molecular Diversity Among Sorghums (Sorghum bicolor (L.) Landraces in Marathwada Region by Molecular Markers. Appl Biochem Biotechnol (2023). https://doi.org/10.1007/s12010-023-04724-2
Accepted:
Published:
DOI: https://doi.org/10.1007/s12010-023-04724-2