Abstract
Riemerella anatipestifer (RA) is one of the most harmful bacterial pathogens in waterfowl and causes enormous economic loss worldwide. Due to weak cross-immunity protection against different serotypes of RA, inactivated and attenuated vaccines are only effective for RA of specific serotypes. In this paper, outer membrane protein YaeT in RA was analyzed through bioinformatics, in vivo, and in vitro assays. Homology, physicochemical and structural properties, transmembrane domains, and B-cell binding epitopes were investigated. The recombinant outer membrane protein YaeT was then inoculated into Cherry Valley ducks to analyze its immune protection against RA. Results showed that the protein was conservative in different RA strains and had sufficient B-cell binding epitopes. The immunized duck serum contains high-affinity antibodies that could activate complement and promote the opsonophagocytosis of RA by phagocytes. After RA challenge, the survival rate of the YaeT protein-immunized ducks was 80%.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Riemerella anatipestifer (RA) is a Gram-negative member of the family Flavobacteriaceae that has no flagella or spores [18]. It produces a capsule and infects domestic birds such as ducks [6, 15], geese [9], and turkeys [27]. This widely distributed bacterium is currently a main pathogen affecting waterfowl breeding and is difficult to be eliminated from duck farms around the world. The RA infection is predominantly via respiratory system and skin contact (especially foot skin), causing acute or chronic septic bacterial infections [6, 26]. The prevalence of the disease can be as high as 90% or more. However, because the virulence of the various serotypes differs greatly, the mortality ranges from 5 to 75% [26, 34].
Long-term use of antibiotics has led to a significant increase in the resistance of pathogenic bacteria [16, 38]. The massive use of antibiotics enhances the resistance of pathogenic bacteria, which in turn further promotes the use of larger doses of antibiotics, resulting in a vicious cycle of stronger resistance of pathogenic bacteria. At the same time, the problem of antibiotic residues due to massive use is getting worse [19, 36]. These problems have become global and need to be addressed. In the long run, improving breeding conditions and vaccination are effective ways to reduce the abuse of antibiotics. Vaccines against RA mainly include attenuated vaccines [18], inactivated vaccines [17], and subunit vaccines [4]. Because of the weak cross-immunity protection among RA serotypes, inactivated vaccines and attenuated vaccines must be based on the local serotypes of RA. OmpA protein was used to prepare RA subunit vaccine, but its immune protection was inconsistent [32, 33, 35].
The YaeT protein, belonging to the Omp85 protein family [31], is an outer membrane protein that is ubiquitous among Gram-negative bacteria. The N-terminus, which is a periplasmic polypeptide-transport-associated (POTRA) domain, is located in the periplasmic space, and the C-terminus is a transmembrane region, comprising 16 discrete β-barrel transmembrane chains. As a part of the outer membrane assembly complex, YaeT is primarily involved in the proper assembly of outer membrane proteins [13]. Loss of the YaeT protein is fatal to Escherichia coli because it causes a series of outer membrane proteins (including TolC, OmpF, OmpC, and OmpA) to fail to assemble correctly on the outer membrane [3]. Reduced YaeT expression causes a reduction in the bacterial sensitivity to contact-dependent growth inhibitors and the downregulation of outer membrane protein synthesis [1]. The YaeT protein is also associated with cell adhesion. In previous studies, we showed that this protein is highly conserved in E. coli, Salmonella, and Shigella. When mice were immunized with a recombinant E. coli YaeT protein, the sera they produced had high affinity for E. coli, Salmonella, and Shigella and a strong immunoprotective effect on mice after E. coli challenge [8]. However, the immunoprotective effect of this protein against RA has not been studied.
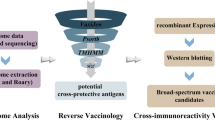
Bioinformatic analysis is used to predict and analyze unknown protein structure, physical and chemical properties, functions, etc., and provides certain guidance for subsequent protein study. Our previous results also proved that the bioinformatic analysis of vaccine candidate proteins is helpful for vaccine development. In this study, the outer membrane protein YaeT of RA was analyzed with bioinformatics and the recombinant protein was expressed and purified. Its immunoprotection against different RA infections was determined to develop an ideal vaccine candidate against RA.
Materials and Methods
Experimental Animals and RA Strains
One-day-old Cherry Valley ducklings were obtained from the Hainan Tianyuan Industrial Co., Ltd. (Haikou, China). The ducks were housed in cages, and provided water and food ad libitum during the study. The animal experiments were approved by Hainan University Institutional Animal Care and Use Committee.
RA strain ATCC 11,845 was purchased from Shanghai Yiyan Biological Technology Co. Ltd. (Shanghai, China). Serotype 1 RA strain CH-1 (RA-CH1) was isolated from sick ducks in Haikou, China, and grown in tryptic soy broth medium (TSB) at 37 °C. E. coli strain DH5α and BL21 (DE3) strains were purchased from TransGen Biotech, Inc. (Beijing, China) and grown on Luria–Bertani (LB) agar at 37 °C.
Protein Sequence
The sequence of YaeT (UniProt Accession no. E4TA62) of RA was retrieved from the Universal Protein Resource (UniProt) at http://www.uniprot.org saved in FASTA format. The status of the protein belonged to the computer-annotated TrEMBL section up to now (unreviewed). The protein was from RA strain ATCC 11,845 (DSM 15,868/JCM 9532/NCTC 11,014).
Homology Analysis
Sequence alignments of YaeT were carried out using the National Centre for Biotechnology Information (NCBI) at https://blast.ncbi.nlm.nih.gov/Blast.cgi. Representative sequences with identity higher than 50% were selected and phylogenetic tree was constructed by MEGA X with maximum likelihood method [14].
Physicochemical Property Analysis
Different physicochemical parameters of the YaeT protein, including number of amino acids, molecular weight, theoretical pI, amino acid (aa) composition, extinction coefficients, estimated half-life, instability index, aliphatic index, and hydrophobicity, were predicted using the Expasy ProtParam online server (https://web.expasy.org/protparam/).
Transmembrane Domain Prediction
PRED-TMBB tool (http://biophysics.biol.uoa.gr/PRED-TMBB/) was used to predict transmembrane domains of YaeT with Viterbi method, N-best method, and posterior decoding method. Subcellular location of the YaeT was predicted by QuickGO online services (https://www.ebi.ac.uk/QuickGO/).
Structure Prediction
Homology modeling of the protein was carried out by the SWISS-MODEL server at https://www.swissmodel.expasy.org/. The best homology models were selected according to Global Model Quality Estimation (GMQE) and QMEAN statistical parameters. The predicted model was then analyzed by Swiss-PdbViewer 4.1 software. The 3D structure of YaeT was also predicted in AlphaFold Protein Structure Database (https://alphafold.ebi.ac.uk/), by the mean of AlphaFold 2 [11, 29]. The 3D structure predictions of the two methods were then evaluated and superposed by in TM-align tool (https://zhanggroup.org/TM-align/), to get root-mean-square deviation (RMSD) and template modeling (TM) scores.
B-Cell Binding Epitope Prediction
The B-cell (http://tools.immuneepitope.org/bcell/) server was applied for linear B-cell epitope prediction based on physicochemical properties such as hydrophilicity, flexibility, polarity, and exposed surface on a non-redundant dataset. Discontinuous epitope was predicted by ElliPro (http://tools.iedb.org/ellipro/) server, with minimum score of 0.5, and maximum distance of 6 Å and number of residues of more than 7.
Expression of YaeT Protein
The construction of recombinant plasmids, recombinant protein expression in E. coli, and purification of the recombinant protein were performed as described previously [7]. The primers, F-BamHI-YaeT: 5′-GGATCCGGAGATGGGCAAGTATTATCATT-3′ and R-EcoRI-YaeT: 5′-GAATTCTCCTATTGTTCTATCAAATCCGTAAGCA-3′, were used for amplification of YaeT gene of ATCC 11,845. Briefly, the amplified YaeT gene and pET-28a vector were digested and purified by BamHI and EcoRI, and then ligated by T4 DNA ligase. The recombinant YaeT-28a plasmid was transformed into E. coli BL21 strain, and then transformed into self-induction medium after activation, cultured at 37 °C for 24 h. Purification and refolding of the recombinant protein were performed based on the previous protocol [7]. The bacterial cells were lysed by ultrasonic lysis on ice with lysis buffer and the lysed cells were centrifuged at 14,000 g for 20 min, and the supernatant was removed. Each gram of inclusion bodies was incubated in 50 ml Tris-HC1, pH 7.9 with 1.5% (v/v) lauryldimethylamine oxide (LDAO), and shaken at 20 °C for 1 h. The inclusion bodies were precipitated by centrifugation at 16,000 g for 30 min. Inclusion bodies were dissolved in 50 mM, Tris-HC1 solution, pH 7.9, and centrifuged at 16,000 g for 30 min, and the supernatant was removed. The sediment was added with denaturation buffer (10 mM Tris–HCl buffer, pH 7.5 containing 1 mM ethylenediamine tetraacetic acid (EDTA) and 8 M urea), and then centrifuged at 14,000 g for 20 min. The supernatant was gradually added with the refolding solution (20 mM Tris–HCl buffer, pH 7.9 containing 1 M NaCl and 5% (v/v) LDAO) to 1:1 with quick stirring for 1 h. Then, the mixture was dialyzed in 4 L permeabilization buffer (20 mM Tris–HCl buffer, pH 7.9 containing 0.5 M NaCl and 0.1% (v/v) LDAO) for 6–8 h. The mixture was then purified by HisTrap column and desalted by HPrepTM26/10 desalting column. The purity of the YaeT protein was detected by SDS-PAGE electrophoresis, and then freeze dried and stored at − 20 °C.
Animal Experiments
Thirty healthy 1-day-old Cherry Valley ducks were randomly divided into 3 groups of 10 ducks in each group. PBS group was immunized with PBS; PBS + adjuvant group was immunized with PBS and complete Freund’s adjuvant (catalogue number: F5881); YaeT group was immunized with YaeT protein and complete Freund’s adjuvant. The ducks were raised to 7-day-old to start the first immunization. The lyophilized YaeT protein was diluted with PBS and mixed thoroughly with complete Freund’s adjuvant in a 1:1 ratio. Each duck was injected subcutaneously with 0.2 mg YaeT protein. The second immunization started at 14 days of age, and the diluted YaeT protein was mixed with the incomplete Freund’s adjuvant (catalogue number: F5506) in a 1:1 ratio. Blood was collected at 6-day-old, 13-day-old, 20-day-old, 27-day-old, and 34-day-old, and the serum was stored at − 20 °C.
Indirect Enzyme-Linked Immunosorbent Assay (iELISA)
The iELISA was performed as described previously [8]. In short, 100 µl of 2 µg/ml YaeT protein and 108 CFU of RA strains were added to each well respectively, incubated overnight at 4 °C, and washed 4 times with 100 µl PBST buffer (1 × PBS [pH 7.4] and 0.5% Tween 20) washing solution, 5 min each time. The antigen was then blocked, and added with series dilution of duck serum and horseradish enzyme-labeled polyclonal rabbit anti-duck IgY (IgG). 1 × TMB was added as chromogenic reagent for 30 min and then added with 2 M H2SO4. The OD value of each well was measured at a wavelength of 450 nm. The ratio of the OD value of the test well to the negative control well (P/N) is greater than 2.1 as positive.
Serum Bactericidal Assay (SBA)
A single colony of RA-CH1 strain was inoculated into TSB liquid medium and cultivated to OD 600 = 1.5. After washing, the cells were diluted with PBS to 108 CFU/ml. Duck serum was diluted with PBS to a series of concentrations of 100%, 50%, 25%, 12.5%, and 5%. Heat-inactivated 100% duck serum and PBS were used as controls. Ten microliters of diluted bacterial solution was mixed with 190 µl of duck serum of different concentrations, and incubated at 37 °C for 30 min. The survival rate of bacteria was counted by the plate counting method.
Opsonophagocytosis Assay
A single colony of RA-CH1 strain was inoculated into TSB liquid medium and cultivated to OD 600 = 1.5. After washing, the cells were diluted with PBS to 5 × 108 CFU/ml. Twenty microliters of test serum and 20 µl of bacterial solution were added to a 96-well cell plate and incubated in a 5% carbon dioxide incubator at 37 °C for 15 min. Ten microliters of duck serum complement and 40 µl of whole duck blood were added to the cell plate, and incubated for 1 h at 37 °C in a 5% carbon dioxide incubator. The survival rate of bacteria was counted by the plate counting method.
Challenge Assay
The RA-CH1 strain was inoculated on a blood plate and cultured in a CO2 incubator at 37 °C for 12 h. Single colony was inoculated in 5-ml TSB medium and cultured with shaking at 37 °C for 18 h. The amount of bacteria was determined by a McDonald’s turbidity tube. The bacterial suspension is diluted 10 times with PBS buffer. Forty-eight healthy 31-day-old Cherry Valley ducks were randomly divided into 8 groups with 6 ducks in each group. The dilutions of bacterial suspension were injected into leg muscles of different groups of ducks. The mortality rate was recorded and calculated, and lethal dose of 50% (LD50) of RA-CH1 strain was calculated by the modified Karber method.
At 14 days after the second immunization of the duck (31 days old), 0.5 ml of 10 times LD50 dose of RA-CH1 bacterial suspension was injected intramuscularly in the leg. The mortality rate after the challenge was recorded.
Statistical Analysis
GraphPad Prism 8.0 was used to analyze the data of each experiment. ANOVA statistical method was used for the statistical analysis, where the symbol “*” indicates significant difference (0.01 < P < 0.05) and “**” indicates extremely significant difference (P < 0.01); P > 0.05 indicates no significant difference.
Results
Homology Analysis
The aa sequence of YaeT protein in RA was analyzed with a BLAST search in the NCBI database, which showed that it shares > 50% identity with homologues from R. anatipestifer (identity > 97%), Chryseobacterium spp. (identity 71–73%), Elizabethkingia sp. (identity 70%), and Ornithobacterium rhinotracheale (identity 71–73%). The aa sequences of the YaeT proteins were downloaded and a phylogenetic tree was constructed (Fig. 1). The tree showed that the YaeT protein is conserved across the different serotypes of RA.
Physicochemical Property Analysis
The physicochemical properties of the 850 aa of YaeT were analyzed with the Expasy ProtParam online server, which showed that the initial 19 s constituted the signal peptide, and the molecular weight of the protein was 93,518.73 Da. The total number of negatively charged residues (Asp + Glu) was 84, and the total number of positively charged residues (Arg + Lys) was 105. The theoretical isoelectric point (pI) was 9.35 and the estimated half-life in E. coli was 10 h. The extinction coefficient was 117,120. The instability index (II) was computed to be 27.95, which classifies the protein as stable. The aliphatic index of the protein was 76.68, and the grand average of hydropathicity (GRAVY) was − 0.476. The results of a hydrophobicity analysis performed with the Kyte–Doolittle method are shown in Fig. 2a.
Transmembrane Domains and Subcellular Localization Prediction
The transmembrane domains of YaeT were predicted with the Viterbi method, the N-best method, and the posterior decoding method, and are shown in Table 1. The Viterbi method and N-best method shared the same prediction in 1–828 aa domain, whereas the Viterbi method and the decoding method shared the same prediction in 829–847 aa domain. The domain predicted with the Viterbi method and visualized with the TMRPres2D tool is shown in Fig. 2b. The QuickGO program predicted that the subcellular location of YaeT is the outer membrane.
Structure Prediction
The 3D structure was predicted by SWISS-MODEL [30] server and AlphaFold Protein Structure Database, and superposed by TM-align tool, which was shown in Fig. 3. The global model quality estimation (GMQE) and qualitative model energy analysis (QMEAN) values with SWISS-MODEL were 0.56 and − 3.30, respectively. The Ramachandran favored value was 92.02%. The predicted three-dimensional (3D) structure was also predicted in AlphaFold Protein Structure Database based on AlphaFold method. Except for 1–41, 329–242 (pLDDT < 50), 131–138, 238–245, 342–347, 475–477, 498–503, 723–725, 847–850 (70 > pLDDT > 50), model confidence of remaining amino acid residues in AlphaFold Protein Structure Database were high (90 > pLDDT > 70) and very high (pLDDT > 90). RMSD and TM score of the two predictions were 7.11 and 0.64. Superposition of the two predicted protein structures showed that the protein comprised α-helices, random coils, and β-sheets that form the β-barrel protein, a representative structure of the outer membrane proteins of Gram-negative bacteria.
Predicted B-Cell Binding Epitopes
Twenty-eight, 16, and 33 linear B-cell epitopes were predicted in the YaeT protein with Bepipred-2.0, a linear epitope predictor, Emini’s surface accessibility prediction, and the Kolaskar and Tongaonkar [12] antigenicity method, respectively. When short sequences (less than 7 aa) were excluded, the three methods predicted 15 epitopes in the same sequence, 10 of which were located outside the cell membrane. The linear B-cell epitopes are listed in Table 2. Seven discontinuous epitopes were predicted, but only numbers 2–5 were outside the outer membrane. The residues and 3D structures of the epitopes are shown in Table 3 and Fig. 4, respectively.
Expression and Purification of YaeT Recombinant Protein
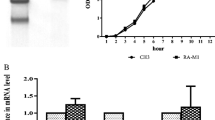
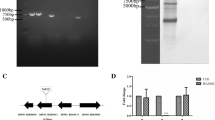
YaeT gene was cloned from ATCC 11,845 strain and sequenced. The sequence of this gene was with 100% identity to which in CH-1 serotype. The gene was then inserted into pET-28a. The recombinant protein was expressed in BL21 cells. After purification, the purity of the protein was about 85%, which was analyzed by gel electrophoresis and shown in Fig. 5.
YaeT Protein Induces Production of High-Affinity Antibodies
The results of iELISA are shown in Fig. 6. After immunization with YaeT protein, the antibody content in duck serum has been increasing, reaching the highest level at 14 days (27 days old) after the second immunization. The antibody titers to YaeT protein and RA strains reached 1:583,200 and 1:194,400, respectively. The antibody titers in the PBS group and the PBS + adjuvant group did not exceed 1:300. It showed that the YaeT protein has stimulated the production of high levels of antibodies in ducks.
Sera antibody titers of YaeT protein analyzed by iELISA. YaeT group-YaeT—the antibody titer of YaeT and adjuvant vaccination group against YaeT. PBS group-YaeT—the antibody titer of PBS vaccination group against YaeT. PBS + adjuvant group-YaeT—the antibody titer of PBS and adjuvant vaccination group against YaeT. YaeT group-CH1—the antibody titer of YaeT and adjuvant vaccination group against RA CH-1 strain. PBS group-CH1—the antibody titer of PBS vaccination group against RA CH-1 strain. PBS + adjuvant group-CH1—the antibody titer of PBS and adjuvant vaccination group against RA CH-1 strain
Serum Antibodies Could Promote Complement to Kill Pathogens
The RA strain was mixed with different concentrations of immune serum. The SBA results are shown in Fig. 7a. The number of colonies in the YaeT group decreased with increasing serum concentration. The number of colonies with 50% and 100% YaeT serum concentrations was lower than that of PBS group or PBS + adjuvant group, and the difference is extremely significant (P < 0.01). The number of colonies with 25% YaeT serum concentration was lower than that of PBS group or PBS + adjuvant group, and the difference was significant (P < 0.05). This result showed that serum antibodies could promote complement to kill pathogens in vitro.
Bactericidal activity and phagocytosis of the anti-YaeT sera in vitro. A Bactericidal activity of the anti-YaeT sera in vitro. RA CH-1 strain was incubated with serial dilution of YaeT group sera, PBS + adjuvant group sera and PBS group sera. B Phagocytosis of the anti-YaeT sera in vitro. RA CH-1 strain was incubated with complement, whole blood, and sera of different groups
Serum Antibodies Could Promote Phagocytosis of RA Strain
The results of phagocytosis are shown in Fig. 7b. After incubation, compared with the PBS group (100%) and the PBS + adjuvant group (58.3%), the percentage of colonies in the YaeT group was 2.08%, the difference was very significant (P < 0.01). This result shows that YaeT protein-immunized serum could promote phagocytosis of RA strain.
YaeT Protein Immunization Reduced the Mortality of RA-Infected Ducks
At 14 days (31 days old) after the second immunization, ducks were injected intramuscularly in the legs with 10 times of LD50 dose of the RA (2.5 × 106 CFU) each, and the survival rate was shown in Fig. 8. After challenge, the survival rate was 40% in the PBS group, 50% in the PBS + adjuvant group, and 90% in the YaeT group. This result showed that ducks vaccinated with YaeT protein could improve the survival rate of ducks after the RA infection.
Discussion
RA is harmful to waterfowl. Subunit vaccine has the advantages of good conservation and low harm to animals, so it is a good candidate for RA vaccine. OmpA protein is considered to have a certain immune protection [33]. However, there are few studies on RA outer membrane proteins, many of which were sequenced without expression and functional verification. This makes it difficult for RA protein to be used in a vaccine. Bioinformatics provides an analytical tool used to identify vaccine antigens in Staphylococcus aureus [28], E. coli [25], Burkholderia pseudomallei [22], and other pathogens [21, 23, 24]. Therefore, in this paper, bioinformatics tools were used to analyze the RA outer membrane protein YaeT.
An ideal vaccine should meet the following requirements. Firstly, this vaccine should confer strong immune protection. To evaluate the antigenicity of proteins, their B-cell epitopes are usually analyzed. There are many algorithms for predicting discontinuous epitopes, and the results obtained here with different programs were not identical. In this study, various models, including BepiPred-2.0 [10], Emini’s surface accessibility prediction [5], and the Kolaskar and Tongaonkar antigenicity methods, were used to ensure comprehensive prediction. The iELISA assay showed that Bama protein in RA contains high-affinity epitopes, which can stimulate humoral immune response in ducks. This is consistent with our bioinformatics analysis. Secondly, effective epitopes must be extracellular. So, the Viterbi method, N-best method, and posterior decoding methods [2] were used to analyze the topological structure of the YaeT protein at the outer membrane. These continuous and discontinuous epitopes require further screening, in combination with the topology of this outer membrane protein. The epitopes in the extracellular domain could be selected, whereas epitopes in the transmembrane and intracellular regions must be removed. The duck serum immunized with the recombinant YaeT protein had affinity for the protein and RA-CH1 strain, indicating that the predicted YaeT protein is actually located in the outer membrane.
In this paper, the conservation of the YaeT protein was also analyzed based on sequence similarities and it was found that the protein shares > 97% similarity among the RA serotypes, but < 75% with those of other genera. A phylogenetic tree showing the genetic distances between the YaeT proteins within RA and between RA and other bacteria confirmed that this protein is conserved in RA. If an antigen is strongly expressed in the pathogen and is well conserved throughout the bacterial serotypes, it may be a candidate vaccine antigen. In previous studies, we showed that some OmpA and BamA outer membrane proteins are conserved in E. coli, Salmonella, and Shigella, and exert good cross-immunoprotection against these bacteria. Antibodies from ducks immunized with the rOmpA of RA could detect RA serotypes 1 and 6 in western blot analysis [20]. The ATCC 11,845 strain shares the same YaeT gene as the CH-1 serotype strain. Therefore, we speculate that the recombinant YaeT protein also has an immunoprotection against the CH-1 serotype RA strain.
To understand the YaeT protein in more detail, its physical and chemical properties and structures were analyzed. The 3D structure was predicted by SWISS-MODEL and AlphaFold 2. SWISS-MODEL is an automatic online software for predicting the tertiary structure of proteins by homology modeling. The more consistent the target sequence is with the template sequence, the more accurate the homology modeling method is. However, this method relies on the similar spatial structure of proteins in the database. AlphaFold 2 provides a computational method that can predict protein structures with atomic accuracy even if no similar structure is known [11], and has good application potential. The structures of the two prediction methods were evaluated by RMSD value and TM score. RMSD value provides the average distance between alpha-carbon backbones of the two models, and higher RMSD values predict greater variant structure deviation. TM score provides the topological similarity between two proteins. If 0.5 < TM score < 1.00, the predictions can be thought about the same fold [37]. Superposition of the two proteins showed that the protein forms a β-barrel structure, which is a typical structure for an outer membrane protein of the Gram-negative bacteria. This further confirms that the protein is an outer membrane protein. Because of the hydrophobic region, transmembrane proteins cannot fold correctly in aqueous solution. So, after the inclusion body is complex, LDAO is added as a mild decontamination agent, which is conducive to the correct folding of Bama protein.
In vivo and in vitro experiments also further confirmed that the Bama protein immunization of RA had protective effect on ducks. After the second immunization, the affinity of duck serum antibody to purified YaeT protein was 1:583,200, indicating that YaeT protein can stimulate a strong immune response in ducks. At the same time, the affinity of the serum to the RA strain was 1:194,400, indicating that the YaeT protein was distributed in the outer membrane of RA, and part of the structure was an extramembrane structure. This is consistent with previous bioinformatics analyses. SBA and opsonophagocytosis assays were used to analyze the killing of RA strains by duck serum antibodies. Within a certain range (25 ~ 100% of YaeT serum), the higher the antibody concentration, the stronger the killing effect of complement on RA. Antibodies and complement also enhanced phagocytosis of RA by phagocytes. After the second immunization, ducks in different groups were injected with the same dose of RA. On the second day after immunization, the duck survival rate in the YaeT group remained at 90%, indicating that the YaeT group had a strong immune protection effect. The survival rates of the PBS + adjuvant group and the control group remained at 50% and 40% respectively after the 3rd day, which may be due to the stimulating effect of the adjuvant, which partially enhanced the immunity of ducks. These results indicate that YaeT protein has a great potential to be developed into a RA vaccine.
Data Availability
The data presented in the study are included in the article. The datasets generated during and/or analysed during the current study are available from the corresponding author on reasonable request. Further inquiries can be directed to the corresponding authors.
References
Aoki, S. K., Malinverni, J. C., Jacoby, K., Thomas, B., Pamma, R., Trinh, B. N., Remers, S., Webb, J., Braaten, B. A., Silhavy, T. J., & Low, D. A. (2008). Contact-dependent growth inhibition requires the essential outer membrane protein BamA (YaeT) as the receptor and the inner membrane transport protein AcrB. Molecular Microbiology, 70, 323–340.
Bagos, P. G., Liakopoulos, T. D., Spyropoulos, I. C., & Hamodrakas, S. J. (2004). PRED-TMBB: A web server for predicting the topology of beta-barrel outer membrane proteins. Nucleic Acids Research, 32, W400-404.
Bodelon, G., Marin, E., & Fernandez, L. A. (2009). Role of periplasmic chaperones and BamA (YaeT/Omp85) in folding and secretion of intimin from enteropathogenic Escherichia coli strains. Journal of Bacteriology, 191, 5169–5179.
Chu, C. Y., Liu, C. H., Liou, J. J., Lee, J. W., & Cheng, L. T. (2015). Development of a subunit vaccine containing recombinant Riemerella anatipestifer outer membrane protein A and CpG ODN adjuvant. Vaccine, 33, 92–99.
Emini, E. A., Hughes, J. V., Perlow, D. S., & Boger, J. (1985). Induction of hepatitis A virus-neutralizing antibody by a virus-specific synthetic peptide. Journal of Virology, 55, 836–839.
Flores, R. A., Cammayo, P. L. T., Nguyen, B. T., Fernandez-Colorado, C. P., Kim, S., Kim, W. H., & Min, W. (2021). Duck interleukin-22: Identification and expression analysis in Riemerella anatipestifer infection. Journal of Immunology Research, 2021, 3862492.
Guan, Q., Wang, X., Wang, X., Teng, D., Mao, R., Zhang, Y., & Wang, J. (2015). Recombinant outer membrane protein A induces a protective immune response against Escherichia coli infection in mice. Applied Microbiology and Biotechnology, 99, 5451–5460.
Guan, Q., Wang, X., Wang, X., Teng, D., & Wang, J. (2016). In silico analysis and recombinant expression of BamA protein as a universal vaccine against Escherichia coli in mice. Applied Microbiology and Biotechnology, 100, 5089–5098.
Gyuris, E., Wehmann, E., Czeibert, K., & Magyar, T. (2017). Antimicrobial susceptibility of Riemerella anatipestifer strains isolated from geese and ducks in Hungary. Acta Veterinaria Hungarica, 65, 153–165.
Jespersen, M. C., Peters, B., Nielsen, M., & Marcatili, P. (2017). BepiPred-2.0: Improving sequence-based B-cell epitope prediction using conformational epitopes. Nucleic Acids Research, 45, W24–W29.
Jumper, J., Evans, R., Pritzel, A., Green, T., Figurnov, M., Ronneberger, O., Tunyasuvunakool, K., Bates, R., Zidek, A., Potapenko, A., Bridgland, A., Meyer, C., Kohl, S. A. A., Ballard, A. J., Cowie, A., Romera-Paredes, B., Nikolov, S., Jain, R., Adler, J., … Hassabis, D. (2021). Highly accurate protein structure prediction with AlphaFold. Nature, 596, 583–589.
Karplus, P. A., & Schulz, G. E. (1985). Prediction of chain flexibility in proteins. Naturwissenschaften, 72, 212–213.
Kumar, G., Rathore, G., & El-Matbouli, M. (2014). Outer membrane protein assembly factor YaeT (omp85) and GroEL proteins of Edwardsiella tarda are immunogenic antigens for Labeo rohita (Hamilton). Journal of Fish Diseases, 37, 1055–1059.
Kumar, S., Stecher, G., Li, M., Knyaz, C., & Tamura, K. (2018). MEGA X: Molecular evolutionary genetics analysis across computing platforms. Molecular biology and evolution, 35, 1547–1549.
Li, D., Wang, X., Xu, X., Gu, J., Yang, Y., Liu, T., Wang, S., Chen, S., & Li, J. (2021). Duck complement factor H binds to outer membrane protein Omp24 of Riemerella anatipestifer. Avian Diseases, 65, 261–268.
Li, R., Jiang, Y., Peng, K., Wang, Y., Wang, M., Liu, Y., & Wang, Z. (2022). Phenotypic and genomic analysis reveals Riemerella anatipestifer as the potential reservoir of tet(X) variants. Journal of Antimicrobial Chemotherapy, 77, 374–380.
Liu, H., Wang, X., Ding, C., Han, X., Cheng, A., Wang, S., & Yu, S. (2013). Development and evaluation of a trivalent Riemerella anatipestifer-inactivated vaccine. Clinical and Vaccine Immunology, 20, 691–697.
Liu, M., Huang, M., Shui, Y., Biville, F., Zhu, D., Wang, M., Jia, R., Chen, S., Sun, K., Zhao, X., Yang, Q., Wu, Y., Chen, X., & Cheng, A. (2018). Roles of B739_1343 in iron acquisition and pathogenesis in Riemerella anatipestifer CH-1 and evaluation of the RA-CH-1DeltaB739_1343 mutant as an attenuated vaccine. PLoS One, 13, e0197310.
Ogunlaja, A., Ogunlaja, O. O., Olukanni, O. D., Taylor, K., Olorunnisola, C. G., Dougnon, V. T., Mousse, W., Fatta-Kassinos, D., Msagati, T. M., & Unuabonah, E. I. (2022). Antibiotic resistomes and their chemical residues in aquatic environments in Africa. Environmental Pollution, 312, 119783.
Phonvisay, M., Lee, J. W., Liou, J. J., Wang, H. Y., & Chu, C. Y. (2019). Evaluation of long-term antibody response and cross-serotype reaction in ducks immunised with recombinant Riemerella anatipestifer outer membrane protein A and CpG ODN. J Vet Res, 63, 543–548.
Saha, S., Vashishtha, S., Kundu, B., & Ghosh, M. (2022). In-silico design of an immunoinformatics based multi-epitope vaccine against Leishmania donovani. BMC Bioinformatics, 23, 319.
Shahab, M., Hayat, C., Sikandar, R., Zheng, G., & Akter, S. (2022). In silico designing of a multi-epitope vaccine against Burkholderia pseudomallei: Reverse vaccinology and immunoinformatics. Journal, Genetic Engineering & Biotechnology, 20, 100.
Shanthappa, P. M., Suravajhala, R., Suravajhala, P., Kumar, G., & Melethadathil, N. (2022). In silico based multi-epitope vaccine design against norovirus. Journal of Biomolecular Structure & Dynamics, 23, 1–11.
Shoushtari, M., Mafakher, L., Rahmati, S., Salehi-Vaziri, M., Arashkia, A., Roohvand, F., Teimoori-Toolabi, L., & Azadmanesh, K. (2022). Designing vaccine candidates against dengue virus by in silico studies on structural and nonstructural domains. Molecular and Cellular Probes, 63, 101818.
Soltan, M. A., Behairy, M. Y., Abdelkader, M. S., Albogami, S., Fayad, E., Eid, R. A., Darwish, K. M., Elhady, S. S., Lotfy, A. M., & AlaaEldeen, M. (2022). In silico designing of an epitope-based vaccine against common E coli pathotypes. Frontiers in Medicine (Lausanne), 9, 829467.
Tao, Z., Zhu, C., Xu, W., Shi, Z., Zhang, S., Song, W., Liu, H., & Li, H. (2020). Riemerella anatipestifer infection affects intestinal barrier structure and immune reactions in the duck caecum. Avian Pathology, 49, 572–580.
Tzora, A., Skoufos, S., Bonos, E., Fotou, K., Karamoutsios, A., Nelli, A., Giannenas, I., Tsinas, A., & Skoufos, I. (2021). Identification by MALDI-TOF MS and antibiotic resistance of Riemerella anatipestifer, isolated from a clinical case in commercial broiler chickens. Vet Sci, 8, 29.
Ullah, N., Anwer, F., Ishaq, Z., Siddique, A., Shah, M. A., Rahman, M., Rahman, A., Mao, X., Jiang, T., Lee, B. L., Bae, T., & Ali, A. (2022). In silico designed Staphylococcus aureus B-cell multi-epitope vaccine did not elicit antibodies against target antigens suggesting multi-domain approach. Journal of Immunological Methods, 504, 113264.
Varadi, M., Anyango, S., Deshpande, M., Nair, S., Natassia, C., Yordanova, G., Yuan, D., Stroe, O., Wood, G., Laydon, A., Zidek, A., Green, T., Tunyasuvunakool, K., Petersen, S., Jumper, J., Clancy, E., Green, R., Vora, A., Lutfi, M., … Velankar, S. (2022). AlphaFold Protein Structure Database: Massively expanding the structural coverage of protein-sequence space with high-accuracy models. Nucleic Acids Research, 50, D439–D444.
Waterhouse, A., Bertoni, M., Bienert, S., Studer, G., Tauriello, G., Gumienny, R., Heer, F. T., de Beer, T. A. P., Rempfer, C., Bordoli, L., Lepore, R., & Schwede, T. (2018). SWISS-MODEL: Homology modelling of protein structures and complexes. Nucleic Acids Research, 46, W296-w303.
Werner, J., & Misra, R. (2005). YaeT (Omp85) affects the assembly of lipid-dependent and lipid-independent outer membrane proteins of Escherichia coli. Molecular Microbiology, 57, 1450–1459.
Wu, H. C., Chang, W. C., Wu, M. C., Wang, H. Y., & Chu, C. Y. (2020). Assessment of immunization regimens of duck Riemerella anatipestifer vaccines. Journal of Applied Microbiology, 129, 1185–1192.
Xu, X., Xu, Y., Miao, S., Jiang, P., Cui, J., Gong, Y., Tan, P., Du, X., Islam, N., & Hu, Q. (2020). Evaluation of the protective immunity of Riemerella anatipestifer OmpA. Applied Microbiology and Biotechnology, 104, 1273–1281.
Yang, D., Mai, K., Zhou, Q., Zhu, Y., Xing, J., Luo, C., Liu, S., Zhou, Q., Huang, W., Luo, J., & Liu, J. (2020). The protective efficacy of specific egg yolk immunoglobulin Y(IgY) against Riemerella anatipestifer infections. Veterinary Microbiology, 243, 108642.
Yang, S., Dong, W., Li, G., Zhao, Z., Song, M., Huang, Z., Fu, J., Jia, F., & Lin, S. (2019). A recombinant vaccine of Riemerella anatipestifer OmpA fused with duck IgY Fc and Schisandra chinensis polysaccharide adjuvant enhance protective immune response. Microbial Pathogenesis, 136, 103707.
Zhang, Y., Cheng, D., Xie, J., Zhang, Y., Wan, Y., Zhang, Y., & Shi, X. (2022). Impacts of farmland application of antibiotic-contaminated manures on the occurrence of antibiotic residues and antibiotic resistance genes in soil: A meta-analysis study. Chemosphere, 300, 134529.
Zhang, Y., & Skolnick, J. (2005). TM-align: A protein structure alignment algorithm based on the TM-score. Nucleic Acids Research, 33, 2302–2309.
Zhang, Y., Zhang, R., Sy, S. K. B., Li, Z., Zhu, S., Zhou, M., Song, C., Zhang, J., Lv, Z., Liu, J., Qin, L., & Yu, M. (2022). Florfenicol/chlortetracycline effect on pharmacodynamic indices for mutant selection of Riemerella anatipestifer in ducks. Microbial Drug Resistance, 28, 832–840.
Funding
This study was funded by Hainan Provincial Natural Science Foundation of China (grant number 317030) and Hainan Science and Technology Project (grant numbers ZDYF2018085).
Author information
Authors and Affiliations
Contributions
HZ and QH conceived and designed research. QG and HZ conducted experiments. CL contributed new reagents or analytical tools. JZ and JW analyzed data. QG completed the writing and subsequent revisions of the manuscript. All authors read and approved the manuscript.
Corresponding authors
Ethics declarations
Ethics Approval
The study was approved by the Animal Welfare and Ethical committee of the Hainan University.
Consent for Publication
All the authors would like to submit the enclosed manuscript entitled “In silico analysis and immune response of YaeT protein against Riemerella anatipestifer in ducks,” which we wish to be considered for publication in Applied Biochemistry and Biotechnology.
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Qingfeng Guan and Hui Yang contributed equally to this work.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Guan, Q., Yang, H., Liao, C. et al. In Silico Analysis and Immune Response of YaeT Protein Against Riemerella anatipestifer in Ducks. Appl Biochem Biotechnol 195, 7483–7501 (2023). https://doi.org/10.1007/s12010-023-04445-6
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12010-023-04445-6