Abstract
In the year 2019–2020, the whole world witnessed the spread of a disease called COVID-19 caused by SARS-CoV-2. A number of effective drugs and vaccine has been formulated to combat this outbreak. For the development of anti-COVID-19 drugs, the main protease (Mpro) is considered a key target as it has rare mutations and plays a crucial role in the replication of the SARS CoV-2. In this study, a library of selected lichen compounds was prepared and used for virtual screening against SARS-CoV-2 Mpro using molecular docking, and several hits as potential inhibitors were identified. Remdesivir was used as a standard inhibitor of Mpro for its comparison with the identified hits. Twenty-six compounds were identified as potential hits against Mpro, and these were subjected to in silico ADMET property prediction, and the compounds having favorable properties were selected for further analysis. After manual inspection of their interaction with the binding pocket of Mpro and binding affinity score, four compounds, namely, variolaric acid, cryptostictinolide, gyrophoric acid, and usnic acid, were selected for molecular dynamics study to evaluate the stability of complex. The molecular dynamics results indicated that except cryptostictinolide, all the three compounds made a stable complex with Mpro throughout a 100-ns simulation time period. Among all, usnic acid seems to be more stable and effective against SARS-CoV-2 Mpro. In summary, our findings suggest that usnic acid, variolaric acid, and gyrophoric acid have potential to inhibit SARS-Cov-2 Mpro and act as a lead compounds for the development of antiviral drug candidates against SARS-CoV-2.
Graphical Abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
A relatively new coronavirus, severe acute respiratory syndrome-related coronavirus (SARS-CoV-2), emerged in December2019, causing coronavirus disease 19 (COVID-19). As per the report, the first case of coronavirus is found in the city of Wuhan, Hubei province. With a high rate of spreading, COVID-19 became a global pandemic affecting the whole world [1, 2]. The 2019-CoV-2 epidemic was declared a “Public Health Emergency of International Concern (PHEIC)” by the World Health Organization (WHO) on January 30 [3]. The name of the coronavirus comes from its crown-like appearance, and these viruses are giant viruses having a single ss-RNA-positive sense genome enclosed in a membrane envelope. Generally, the membrane is covered with spike glycoprotein which is responsible for its appearance. Mainly coronaviruses are divided into four categories: alpha, beta, gamma, and delta. SARS-CoV, the middle east respiratory syndrome virus (MERS-CoV), and the novel SARS-CoV-2 are all members of the beta-coronavirus family (SARS-CoV-2) [4]. Even if all three viruses belong to the same classes, genes of SARS CoV-2 share less than 80% similarity (nucleotides) with other SARS-CoV and are more contagious than MERS-CoV and SARS-CoV [5, 6].
The genome of the SARS-CoV-2 virus is approximately 30,000 nucleotides in length. The viral genome encodes several structural and functional proteins such as RNA-dependent RNA polymerase (RdRp), papain-like protease and coronavirus main protease (Mpro), as well as glycosylated spike protein (S), an envelope protein (E), membrane protein (M), and nucleocapsid protein (N). After invading the host cell, the viral genome is released, and the genome is then translated into polypeptides by the host cell protein translation complex [1]. The viral proteases PLpro and Mpro ultimately cleave the polyproteins into effector proteins [7]. The crystal structure of Mpro (6lu7) was obtained by a co-crystallization along with a peptide-like inhibitor termed N3 (PRD_002214). The enzyme Mpro is dimeric in nature, where each monomer is composed of three domains domain I (8–101 residues), domain II (102–184), and domain III (201–301). The first two domains are antiparallel beta-barrel, and the third one is the alpha-helical which is required for enzymatic activity. It has a similar structure to cysteine protease, although it lacks the third functional catalytic residue in the active site. Hence, Histidine 41 (His 41) and Cysteine 145 (Cys145) are found in the pocket between domains I and II [8], whereas amino acids T24, L27, H41, F140, C145, H163, M165, P168, and H172 form a hydrophobic surrounding in the pocket” [9].
The concept of therapeutic repurposing is being frequently employed to find prospective COVID-19 disease treatments. The practice of repurposing pharmaceuticals can dramatically minimize the cost, time, and dangers associated with medication development [10]. Although several molecular docking investigations based on antiviral drugs are typically used to treat HIV, phytochemicals, antimalarial medicines, spices, or marine items have been established to uncover a possible inhibitor of Mpro activity [11]. Unfortunately, no particular medicines for COVID-19 disease are available to date, and research into the disease’s therapy is currently insufficient. As a result, it is critical to locate approved drugs that may limit SARS-CoV-2 viral proteins, as well as their appropriate association and concentration for the treatment of COVID-19 patients.
Consequently, in order to create novel therapeutics against COVID-19, we used computational screening to look for coronavirus-fighting compounds in lichen species, which might be a great natural source. Antiviral, antibacterial, and antifungal activities have been reported in some lichen species [12]. Many scientific studies have shown that lichen metabolites might be a potential source of antiviral drug candidates [13]. As a result, we conducted virtual screening to identify possible natural anti-SARS-CoV-2 drugs to see if lichen chemicals may also protect SARS-CoV-2. A very important role is played by Mpro or 3-chymotrypsin-like protease in the replication process of the virus. Polyproteins pp1a and pp1b are cleaved by Mpro, resulting in the formation of functional proteins such as endoribonuclease, exoribonuclease, and RNA polymerase. As Mpro is directly related to virus replication, targeting Mpro for new therapeutics drug development can be a new strategy to combat the spread of virus [14]. Hence, we have selected the viral Mpro enzyme as a drug target to find new SARS-CoV-2 inhibitors (Fig. 1). Targeting viral replication has shown to be a successful therapeutic development technique. SARS-CoV-2 Mpro is involved in the conversion of polyproteins into structural and nonstructural proteins. Natural products or their semi-synthetic derivatives are a major source of clinically used drugs, and lichens are an extremely important and unexplored source of natural products.
SARS-CoV-2 replication is inhibited by protease Mpro inhibitors. SARS-CoV-2 releases its genomic RNA after infecting the host cell. Polyproteins pp1a and pp1ab are produced during translation and are cleaved into the primary protease Mpro and nonstructural proteins (nsps). Nsps is required for the assembly of the viral replication transcription complex (RTC) in order for RNA synthesis to take place. When inhibitors like usnic acid, variolaric acid, gyrophoric acid, and remdesivir function in the cell, they bind to Mpro and inhibit its activity, resulting in virion assembly failure, which results into the host cell that was unable to release new complete virions
Two of the benefits of targeting Mpro are as follows: (a) while the rate of mutagenesis in viruses is significant, rare mutations occur in the major protease because any change here might be damaging to the virus [15]; (b) in humans, the lack of closely comparable homologs with identical specificity of cleavage [16]. In the following study, we evaluated the inhibitory capability of lichenic bioactive compounds against Mpro using molecular docking and a molecular dynamics approach. We have assessed the basic ADMET properties of all the lead compounds. Again, we have studied the toxicological properties of selected compounds, and the antiviral activity of selected compounds has been analyzed using PASS prediction.
Methods
Target Protein Retrieval from Databases and Target Preparation
The three-dimensional crystallographic (method: X-ray diffraction) structure of target protein Mpro was retrieved using the Protein Data Bank (PDB) (https://www.rcsb.org/) in PDB format [17]. The “Prepare protein” methodology in the BIOVIA Discovery studio was used for receptor preparation. At physiological pH 7.4, heteroatoms and water molecules from the crystal structure were eliminated and addition of polar hydrogens. The prepared protein structure was saved in pdbqt format using the PyRx tool for further docking analysis. Further, the prediction of binding sites of protein was made through the literature survey, additionally, with the help of online platform CASTp (http://sts.bioe.uic.edu) [18].
Determination and Preparation of Ligands
Through an extensive literature survey, bioactive compounds from lichens have been chosen, which show photoprotective nature, and are known to possess anti-inflammatory, antioxidant, antimicrobial properties [19]. The structures of bioactive compounds were obtained in SDF format from the PubChem database (https://pubchem.ncbi.nlm.nih.gov). The structures are then converted to PDB format and saved as pdbqt files utilizing the National Cancer Institute’s internet server online Structure File Generator and SMILES (Simplified Molecular Input Line Entry System) Translator (https://cactus.nci.nih.gov/translate/). These ligands were prepared using Autodock tools, charges of polar hydrogen were assigned, and nonpolar hydrogens were combined. Finally, for molecular docking, the compounds were converted in pdbqt format.
Molecular Docking Analysis
A molecular docking approach has been made in order to determine the molecular interaction between selected compounds and main protease (Mpro); docking was performed using the PyRx tool. PyRx is a virtual screening tool that employs Vina as well as Autodock 4.2 [20, 21]. Previously prepared target protein SARS-CoV-2 Mpro and prepared ligand molecule were loaded into the PyRx tool, and the former is remarked as a macromolecule, and later is commented as ligand. A grid box was set around the receptor having a dimension of (x, −10.729; y, 12.4176; z, 68.816122). All the ligands were docked, and the maximum exhaustiveness was computed for all of them. All other parameters in the software are in the default mode. For all the rigid-based docking studies, all the bonds in the ligand are inflexible and allowed to rotate freely. The docking results outcome was analyzed and ranked according to their dock score value.
Drug Likeness, Pharmacokinetic, and Oral Toxicity Evaluations
All selected compounds were submitted to ADMET (absorption, distribution, metabolism, excretion, and toxicity) analyzed with Swiss ADME to examine drug likeness and pharmacokinetics characteristics (http://www.swissadme.ch/) [22]. Those molecules which follow the Lipinski rule are further subjected to toxicity prediction. OSIRIS Property Explorer software was used to evaluate the toxicity risks (mutagenicity, tumorigenicity, irritation, and reproductive effect) as well as the physicochemical qualities (drug likeness and drug score) of the hit compounds [23].
Docked Structure Analysis and Visualization
BIOVIA discovery studio and PyMOL have been used to visualize protein and ligand models. The clusters from the previous stage were used to create a model. Amino acids engaging with the ligand, hydrogen bonds (H-bonds), and the individual atoms involved were examined in each ligand cluster. Each cluster’s interacting amino acids with the target were recorded. BIOVIA Discovery Studio was used to create 2-dimensional and 3-dimensional diagrams of the docked protein-ligand structure.
PASS Prediction for Antiviral Activity
The PASS online version (http://way2drug.com/PassOnline/) was used to obtain the biological activity spectra of selected bioactive compounds [24]. PASS software was used to assess the natural potential of a drug-like compound. Before chemical production and testing, PASS software can predict numerous biological activities of compounds based on the structure of those molecules. A substance’s estimated activity is estimated as probable activity (Pa) and probable inactivity (Pi). Substances with a Pa value greater than Pi are regarded as suitable for specific medical usage [25]. The compounds were assessed using the SMILES [26].
Molecular Dynamics Simulation
The ligand complexes of SARS-CoV-2 Mpro and screened compounds with the best binding conformations among the poses predicted by the molecular docking tool were submitted to 100 ns of MD simulation. All MD simulations were performed using GROMACS 2020.3 version [27]. The protein-ligand complex PDB obtained after the docking studies was used to build the system topology. The CHARMM-GUI [28] webserver was used to build the simulation system with the Charmm-36 [29] force field being used for describing the motions of the protein atoms. Each of the protein-ligand systems was solvated in a rectangular simulation box of dimensions 90 Å, 80 Å, and 65 Å and charges neutralized by adding Na+ and Cl− ions by Monte Carlo [28] ion placing method. Energy minimization was performed using a steepest descent algorithm [30] for 10,000 steps with the LINCS [31] algorithm being used for constraining bonds with hydrogen. Later the energy minimized system was equilibrated in a two-step process, first in a canonical ensemble for 100 ps with Nose-Hoover temperature coupling to bring the system temperature to desired temperature of 310.15 K, followed by an NPT equilibration step for 500 ps using Parrinello-Rahman pressure coupling [32], with the pressure maintained at 1 atm. The Newtonian equations of motion were integrated by the leapfrog algorithm [33], and production simulations were performed under NPT conditions for 100 ns, with the columbic interactions being calculated using the particle-mesh Ewald [34] method and Lennard-Jones interactions calculated with a 1.2 Å cutoff.
Results and Discussion
Estimating the physicochemical characteristics and bioavailability of effective lead compounds has been crucial in the outcome, and identification of new chemical compounds as unique and possible leads to the target. Computational chemistry and biology, molecular modelling, virtual screening, and MD simulations play a crucial role in understanding biological roles and structure-based drug discovery. These all play essential roles in the search for new and promising lead molecules for protein targets linked to human diseases [35]. The large-scale virtual screening’s efficiency, on the other hand, is mainly dependent on determining a precise target, selecting appropriate chemical compound databases, and critically assessing the pharmacokinetic profiles of lead compounds. The identification of various small chemical inhibitors of SARS-CoV Mpro was made possible by a thorough understanding of the strong similarities in amino acid sequence and structure between SARS-CoV and SARS-CoV-2 Mpro.
Target Protein Retrieval from Databases and Its Preparation
SARS CoV-2 Mpro is a multifunctional protein involved in viral RNA transcription and replication, which function as proteinases that cleave the polyprotein, Replicase polyprotein 1ab. It was chosen as a candidate protein for a purpose. Using the RCSB Protein Data Bank (PDB ID: 6LU7), the protein sequence was obtained in FASTA format [17].
Rao and colleagues developed N3, a Michael inhibitor that effectively reduced the proteolytic and catalytic activity of SARS CoV-1 Mpro and MERS CoV Mpro in 2005 [17, 36]. Using experimental experiments, they established that N3 reduces the enzymatic activity of SARS CoV-2 Mpro via a two-step irreversible inactivation process. The protein was prepared using BIOVIA Discovery studio’s “Prepare protein” methodology. At physiological pH 7.4, heteroatoms and water molecules were removed from the crystal structure, and polar hydrogens were added. Prediction of binding sites of protein was made by CASTp, based on current Computational Geometry theoretical and algorithmic discoveries. It has several advantages: (1) The bulk solvent-pocket demarcation is specifically defined, (2) all derived parameters were rotationally isotropic and do not require modelling or the use of dot surfaces or grid points, and (3) all derived parameters were circumferentially invariant yet do not require discretization or the use of dot substrate surface or grid points. The identification of a Mpro binding site in the chain, denoted by residue, was validated by a literature review and CASTp: “Thr24, Thr25, Leu27, His41, Cys44, Thr45, Ser46, Met49, Pro52, Ser139, Phe140, Leu141, Asn142, Gly143, Ser144, Cys145, His164, His163, Met165, Glu166, His172, Phe181, Gln189, Thr190, and Gln192” [37].
Docking Analysis
The molecular docking methodology was confirmed virtually by docking the inhibitor ligand N3 into a binding pocket predicted from the target protein Mpro’s crystal structure. The result displays various types of ligand-receptor interaction together with their respective binding energy (Kcal/mol). The lowest binding energy or highest docking score selected for further evaluation was the best and most stable binding mode. Table 1 shows all bioactive compounds, along with their PubChem compound ID, molecular formula, and receptor molecule docking score (Kcal/mol). The docking score of all the selected lead compounds ranges from −6.4 to −8.6 kcal/mol. The docking score is highest (lowest binding energy) of usnic acid (−8.6), followed by fumarprotrocetraric acid (−8.3) and norstictic acid (−8.3), and so on. The comparison of the docking score of all the selected lead compounds with that of remdesivir showed a docking score of −8.0. So, by taking the minimum threshold (−8.0 Kcal/mol) value into consideration for further studies, selection was made on the basis of docking scores of the compounds above −8.0 Kcal/mol. In comparison to the remdesivir, the reference molecule, many of the screened hits had reduced and significantly enhanced binding energy against Mpro, the target protein. According to the molecular docking findings, the screened compounds could work the same way as the reference molecule. These compounds may cause conformational changes in the receptor protein Mpro, inhibiting its function as proteases and limiting viral propagation into the cell protoplasm.
Drug Likeness, Pharmacokinetic, and Oral Toxicity Evaluations
Understanding a compound’s pharmacokinetic behavior and toxicity is critical for determining whether or not it is suitable for human administration. Drug-likeness studies also help to understand drug behavior [38]. Pharmacokinetic properties were obtained from the SwissADME online server, and Osiris software was used to retrieve various toxicological data for lead compounds. The drug likeness was determined using the Lipinski rule of five, which is an empirical thumb method for determining drug likeness. According to the rule of Lipinski’s RO5, most “drug-like” molecules contain a number of hydrogen bond donors: 5, number of hydrogen bond acceptors: 10, LogP 5, and molecular weight: 500.
Among 26 selected compounds, only eight compounds (cryptostictinolide, fumarprotrocetraric acid, gyrophoric acid, salazinic acid, stictic acid, norstictic acid, usnic acid, and variolaric acid) showed binding energy of more than −8.0 kcal/mol and were subjected to the ADMET analysis (Table 2). All the above-mentioned compounds follow Lipinski’s rule except fumarprotrocetraric, which has 12 hydrogen bond donors and violates Lipinski’s rule no 2. The molecular weight of each compound is less than 500. As a result, their internal mobility, diffusion, and absorption may be enhanced. All of the compounds had a TPSA value of less than 200 (220–182), suggesting high bioavailability. The topological polar surface area (TPSA) and atom molar refractivity (AMR) values were higher in all of the compounds. Among all selected compounds, the greatest AMR value (119.19) was found in gyrophoric acid, whereas the highest TPSA value was found in fumarprotrocetraric acid (193.96). The essential features of TPSA and AMR are associated with drug absorption, transport, and penetration mechanisms [39].
All compounds show a good bioavailability score with a value of 0.55 or 0.56, but fumarprotocetraric acid and gyrophoric acid show a bioavailability score of 0.11, respectively. The solubility of a molecule facilitates drug development activities such as designing and formulation [40]. Gastrointestinal absorption of cryptostictinolide, norstictic acid, usnic acid, and variolaric acid was high. In the case of fumarprotocetraric acid, gyrophoric acid, and salazinic acid, gastrointestinal absorption was low. All the screened compounds do not show any blood-brain barrier permeability, and there are no PAINS alerts for any of the selected compounds. The penetration of substances through the blood-brain barrier (BBB) reveals their ability to pass the BBB. Fumarprotrocetraric acid was found with one violation of Lipinski’s rule; we have eliminated that compound from further studies.
All the seven screened compounds which did not violate any Lipinski rule were subjected to their toxicity prediction through Osiris software. The toxicity risk predictor looks for fragments in a molecule that could indicate toxicity. These alerts suggest that the depicted structure may be hazardous to the given risk category. The results clearly reveal that all compounds except stictic acid, norstictic acid, and salazinic acid would be safe and expected to show no toxicity regarding mutagenicity, tumorigenicity, irritation, and effect on the reproductive system. According to the data shown in Table 3, the following are the toxic effects of the above-mentioned compounds: stictic acid (highly toxic-reproductive), norstictic acid (highly toxic-reproductive), and salazinic acid (highly toxic-reproductive). Simultaneously, most of the compounds performed well in screening, with the best drug score values (DS = 0.36–0.46), compared to the other compounds in Table 3 with lower values. In this step, we have eliminated those compounds which were showing carcinogenic behavior, such as norstictic acid, stictic acid, and salazinic acid. It reveals that the compounds chosen have the optimum pharmacokinetic qualities needed to develop promising and effective options for inhibiting SARS CoV-2 main protease.
Structural Analysis and Visualization
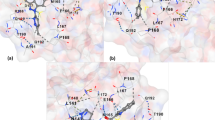
The analysis of docked protein-ligand complexes of screened hits with the Mpro, as well as the interacting residues accommodated in the binding pocket, is shown in Fig. 2. PyMOL and BIOVIA Discovery Studio 2019 were used to visualize the 3D interactions of the protein-ligand complex (docked structure). PyMOL was used to determine the amino acid residues of the Mpro proteins that bind ligands and the docked poses of the screened compounds with Mpro. Those compounds with the lowest binding energy, which follows all of the Lipinski rules and do not have any carcinogenic properties, were investigated further to determine interaction residues of the target protein. Lists of the four selected bioactive compounds along with their 2-dimensional structure, total no of hydrogen bonds, total number of polar contacts, the probable receptor protein-interacting amino acid residue along with their bond length, residue with hydrophobic interaction, and the information regarding the docking results are provided in Table 4. Usnic acid (dock score −8.6 kcal/mol) was the most stable ligand-protein interaction with the best and highest docking score (Fig. 3). It has six polar contacts with the target protein Mpro’s Gly143(2.07), Ser144(2.580), Met165(2.99), Glu168(2.8), Glu168(2.43), and Arg188(2.52) residues. All of the ligands’ binding residues were compared to the recognized drug remdesivir. Remdesivir interacts with the following amino acid residues: Gly143, Ser144, Cys145, Met49, Glu166, His41, and Met165. It was observed that screened bioactive compounds interact effectively with various amino acid residues in Mpro domains I and II. The 2-dimensional and 3-dimensional diagrams of docked protein-ligand structure were generated by using BIOVIA Discovery Studio (Fig. 4). The “display 2D diagram” revealed amino acid residues implicated in critical interactions and other key contacts that bind the ligand within the active crevice.
SARS-CoV-2 Mpro’s main binding cavity. A The superimposed structures of docked protein-ligand complexes of screened hits with the Mpro, as well as the interacting residues accommodated in the binding pocket are shown. B Amino acid residues that interact with the Ligands and the type of interaction; amino acids were represented by a three-letter code with the specified position. a Mpro-variolaric acid, b Mpro-gyrophoric acid, c Mpro-usnic acid, d Mpro-remdesivir
SARS-CoV-2 Mpro’s main binding cavity. A The structures of docked protein-ligand complex of usnic acid with the Mpro, as well as the interacting residue accommodated in the binding pocket, are shown. B Amino acid residues that interact with usnic acid and the type of interaction are highlighted along with amino acids represented by a three-letter code with the specified position
PASS Prediction for Antiviral Activity
Through the online PASS version, biological activity spectra of selected lead compounds were procured. All the predictions were analyzed, interpreted, and used in a flexible manner, as shown in Table 5. All four secondary metabolites show antiviral activity against the HIV virus and herpes virus. Gyrophoric acid and variolaric acid show antiviral activity against the picornavirus as well as rhinovirus. Including the above two compounds, usnic acid also showed antiviral activity against rhinovirus and influenza virus [41, 42]. Trachoma virus was inhibited by cryptostictinolide and gyrophoric acid. Cryptostictinolide and variolaric acid show antiviral activity against the hepatitis B virus. The probability of activity is more significant than the probability of inactivity against the virus in all compounds (Table 5).
Molecular Dynamics Simulation
The molecular dynamics (MD) simulation for a time period of 100 ns was performed using GROMACS 2020 with Charmm-36 force field to better understand the conformational changes and stability of the protein-ligand complexes of the four hits that were identified. The top scoring docking pose of usnic acid, gyrophoric acid, variolaric acid, as well as the conventional inhibitor remdesivir, in complex with Mpro, was used for MD system preparation. After completion of the simulation, the trajectory was analyzed for root mean square deviation (RMSD), root mean square fluctuation (RMSF), the radius of gyration (Rgyr), number of hydrogen bonds, and solvent accessible surface area (SASA). To analyze the structure’s flexibility during the simulation, we calculated the root mean square fluctuation (RMSF) of individual amino acid residues in the protein. The RMSD value denotes the average variance between the atoms of the trajectory frames; the lower the RMSD, the closer and more stable the two structures are. Because Mpro-Cryptostictinolide formed an unstable complex during MD, it was dropped out for further analysis. As indicated by the RMSD values, the ligand-protein complexes remained constant throughout the simulation time.
The results from the analysis of MD trajectory are shown in Fig. 5. The stability and quality of the system can be correlated to the structure’s deviation from its original coordinates over simulation time. Using the rmsd tool in GROMACS, the RMSD plot for the protein, Mpro-usnic acid, Mpro-gyrophoric acid, Mpro-variolaric acid, and the standard inhibitor Mpro-Remdesivir was obtained. As shown in Fig. 5A, the RMSD value for Mpro was 0.1 nm, in the beginning of MD simulation and later got stabilized (RMSD for protein and ligand was between 0.1 and 0.45 nm) during the simulation time interval, with little fluctuation. The RMSD values for usnic acid was stable in the range of 0.1 to 0.15 nm, for gyrophoric acid was in the range of 0.12 to 0.26 nm, and for variolaric acid was in the range of 0 to 0.1 nm indicating a stable binding of these ligands with the binding pocket of Mpro. Moreover, the RMSD value of remdesivir was in the range of 0.2 to 0.45 nm indicating a stable complex.
Further, root mean square fluctuation (RMSF) analysis was carried out in order to properly assess the dynamic behavior of the amino acid residues and the flexibility of the protein structure during the interaction phase of the simulation [43]. As demonstrated in Fig. 5C, the RMSF value of the protein complexes was within the range of 0.1 to 1 nm; the terminal part of the protein ligand complex showed higher RMSF value as compared to the rest of protein indicating the stability of the complexes.
Hydrogen bonding is a major protein ligand interaction involved in stabilizing a complex. We calculated average number of hydrogen bonds occurring throughout the simulation period using H-bond utility in GROMACS with a cutoff value of 0.35 nm. As shown in the Fig. 6A, the majority of the hydrogen bonds obtained in the crystal structure were retained throughout the 100-ns simulation. However, Mpro-gyrophoric acid complex showed major fluctuation in hydrogen bond number (6 to 4), and the minimum fluctuation in hydrogen bond number was observed in Mpro-usnic acid complex (0 to 2) throughout the 100-ns simulation.
Furthermore, we analyzed the solvent-accessible surface area (SASA), which determines the surface area of a biomolecule that is accessible by a solvent (biofluids) and aids in determining the stability of the protein’s hydrophobic core. These hydrophobic interactions are important indications of protein tertiary structure compactness [43]. The SASA profile of each protein complex was shown to be comparable with its radius of gyration, as a slight drop in the SASA profile represents a relative expansion of the protein surface area, resulting in a more variable radius of gyration. For protein complexes (Fig. 6B), the values ranged from 140 to 155 nm2, matching the complex system’s simulation time period (100 ns). The results were consistent with the SASA profiles being reproduced with relatively minimal variances.
GROMACS was also used to estimate the complex’s radius of gyration (RoG), as shown in Fig. 6C, in order to better comprehend the compactness and overall dimensions of the protein structure. For the first run, the RoG values for the protein-ligand complexes were consistent with the RMSD values, as well as steady and low, ranging between 2.2 and 2.35 nm. The results indicate that the RoG values were constant, with only minor variations within an acceptable range. Overall, it can be inferred that protein complexation with hit compounds enhances the Mpro structure’s compactness/rigidity, resulting in higher overall stability. The Mpro-usnic acid and Mpro-variolaric acid complexes seemed to have lower RMSD values, minimum fluctuation in RMSF values, a good number of hydrogen bonds, and low Rog (nm), indicating that they were forming a highly stable complex. Taking all of the findings into account, it is obvious that ligand interaction did not cause any conformational changes in Mpro’s active site. As a result, these predicted compounds may be able to limit Mpro’s catalytic activity.
Conclusions
Due to extensive outbreaks and a lack of therapy, COVID-19 has become a global problem. Certain investigations indicating antiviral treatment strategies are in the works to combat the life-threatening CoV infection. For the identification of new hit compounds as possible inhibitors for Mpro, a COVID-19 protein, several computational approaches were used, such as virtual screening, drug-likeness analysis, toxicity prediction, and molecular dynamics simulation (MDS). For this study, we screened an extensive library of lichen metabolites. A total of 26 selected compounds were chosen as lead drugs targeting Mpro based on molecular docking as well as binding affinity. The results indicated that all the top hits were binding to a similar cavity and interacted with residues like His41, Gly143, Ser144, Cys145, Met165, and Glu168 of the main protease. The “H” atom of the phenolic hydroxyl groups of the usnic acid has shown H-bond interaction with the Arg188 and Glu166 residues. The π electrons of the dimethyldibenzofuran-1,3-dione group made pi-pi interaction with Met165 residue and conventional hydrogen bond interaction with Gly143 and Ser144 residues of Mpro protein. Only seven compounds that obeyed Lipinski’s RO5 as well as PAINS filter using comprehensive pharmacokinetic drug-likeness analysis have been found . Four non-toxic compounds were predicted by the toxicity analysis because they showed no mutagenicity, carcinogenicity, or other consequences and had improved binding affinity toward Mpro. Finally, it has been found that usnic acid, along with three screened compounds, has the most stable Mpro binding affinities based on MD simulation. Usnic acid appears to be the most stable and efficient against SARS-CoV-2 Mpro among all screened compounds. These findings showed that these potent lichen bioactive compounds might be considered new lead compounds for the fast development of COVID-19 therapeutic drugs.
Data Availability
All the data related to the manuscript is included as figures and tables.
References
Gorbalenya, A. E., Baker, S. C., Baric, R. S., De Groot, R. J., Drosten, C., Gulyaeva, A. A., & Ziebuhr, J. (2020). The species severe acute respiratory syndrome-related coronavirus: Classifying 2019-nCoV and naming it SARS-CoV-2. Nature Microbiology, 5, 536–544. https://doi.org/10.1038/s41564-020-0695-z
Kupferschmidt, K., & Cohen, J. (2020). Will novel virus go pandemic or be contained? Science, 610–611. https://doi.org/10.1126/science.367.6478.610
Rodríguez-Morales, A. J., MacGregor, K., Kanagarajah, S., Patel, D., & Schlagenhauf, P. (2020). Going global–Travel and the 2019 novel coronavirus. Travel Medicine and Infectious Disease, 33, 101578. https://doi.org/10.1016/j.tmaid.2020.101578
Anthony, S. J., Johnson, C. K., Greig, D. J., Kramer, S., Che, X., Wells, H., & Goldstein, T. (2017). Global patterns in coronavirus diversity. Virus Evolution, 3(1), vex012. https://doi.org/10.1093/ve/vex012
Wu, F., Zhao, S., Yu, B., Chen, Y. M., Wang, W., Song, Z. G., & Zhang, Y. Z. (2020). A new coronavirus associated with human respiratory disease in China. Nature, 579(7798), 265–269. https://doi.org/10.1038/s41586-020-2008-3
Paraskevis, D., Kostaki, E. G., Magiorkinis, G., Panayiotakopoulos, G., Sourvinos, G., & Tsiodras, S. (2020). Full-genome evolutionary analysis of the novel corona virus (2019-nCoV) rejects the hypothesis of emergence as a result of a recent recombination event. Infection Genetics and Evolution, 79, 104212. https://doi.org/10.1016/j.meegid.2020.104212
Anand, K., Ziebuhr, J., Wadhwani, P., Mesters, J. R., & Hilgenfeld, R. (2003). Coronavirus main proteinase (3CLpro) structure: Basis for design of anti-SARS drugs. Science, 300(5626), 1763–1767. https://doi.org/10.1126/science.1085658
Boopathi, S., Poma, A. B., & Kolandaivel, P. (2021). Novel 2019 coronavirus structure, mechanism of action, antiviral drug promises and rule out against its treatment. Journal of Biomolecular Structure and Dynamics, 39(9), 3409–3418. https://doi.org/10.1080/07391102.2020.1758788
Yang, H., Yang, M., Ding, Y., Liu, Y., Lou, Z., Zhou, Z., & Rao, Z. (2003). The crystal structures of severe acute respiratory syndrome virus main protease and its complex with an inhibitor. Proceedings of the National Academy of Sciences, 100(23), 13190–13195. https://doi.org/10.1073/pnas.1835675100
Muralidharan, N., Sakthivel, R., Velmurugan, D., & Gromiha, M. M. (2021). Computational studies of drug repurposing and synergism of lopinavir, oseltamivir and ritonavir binding with SARS-CoV-2 protease against COVID-19. Journal of Biomolecular Structure and Dynamics, 39(7), 2673–2678. https://doi.org/10.1080/07391102.2020.1752802
Shah, B., Modi, P., & Sagar, S. R. (2020). In silico studies on therapeutic agents for COVID-19: Drug repurposing approach. Life Sciences, 252, 117652. https://doi.org/10.1016/j.lfs.2020.117652
Molnár, K., & Farkas, E. (2010). Current results on biological activities of lichen secondary metabolites: A review. Zeitschrift für Naturforschung C, 65(3–4), 157–173. https://doi.org/10.1515/znc-2010-3-401
Esimone, C. O., Grunwald, T., Nworu, C. S., Kuate, S., Proksch, P., & Überla, K. (2009). Broad spectrum antiviral fractions from the lichen Ramalinafarinacea (L.) Ach. Chemotherapy, 55(2), 119–126. https://doi.org/10.1159/000194974
Elmezayen, A. D., Al-Obaidi, A., Şahin, A. T., & Yelekçi, K. (2021). Drug repurposing for coronavirus (COVID-19): In silico screening of known drugs against coronavirus 3CL hydrolase and protease enzymes. Journal of Biomolecular Structure and Dynamics, 39(8), 2980–2992. https://doi.org/10.1080/07391102.2020.1758791
Ren, Z., Yan, L., Zhang, N., Guo, Y., Yang, C., Lou, Z., & Rao, Z. (2013). The newly emerged SARS-like coronavirus HCoV-EMC also has an “Achilles’ heel”: Current effective inhibitor targeting a 3 C-like protease. Protein and Cell, 4(4), 248. https://doi.org/10.1007/s13238-013-2841-3
Pillaiyar, T., Manickam, M., Namasivayam, V., Hayashi, Y., & Jung, S. H. (2016). An overview of severe acute respiratory syndrome–coronavirus (SARS-CoV) 3CL protease inhibitors: Peptidomimetics and small molecule chemotherapy. Journal of Medicinal Chemistry, 59(14), 6595–6628. https://doi.org/10.1021/acs.jmedchem.5b01461
Jin, Z., Du, X., Xu, Y., Deng, Y., Liu, M., Zhao, Y., & Yang, H. (2020). Structure of Mpro from SARS-CoV-2 and discovery of its inhibitors. Nature, 582(7811), 289–293. https://doi.org/10.1038/s41586-020-2223-y
Tian, W., Chen, C., Lei, X., Zhao, J., & Liang, J. (2018). CASTp 3.0: Computed atlas of surface topography of proteins. Nucleic Acids Research, 46(W1), W363–W367. https://doi.org/10.1093/nar/gky473
Manojlovic, N. T., Vasiljevic, P. J., Maskovic, P. Z., Juskovic, M., & Bogdanovic-Dusanovic, G. (2012). Chemical composition, antioxidant, and antimicrobial activities of lichen Umbilicaria cylindrica (L.) Delise (Umbilicariaceae). Evidence-Based Complementary and Alternative Medicine, 2012. https://doi.org/10.1155/2012/452431
Dallakyan, S., & Olson, A. J. (2015). Small-molecule library screening by docking with PyRx. Chemical Biology (pp. 243–250). Humana Press. https://doi.org/10.1007/978-1-4939-2269-7_19
Trott, O., & Olson, A. J. (2010). AutoDock Vina: Improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. Journal of Computational Chemistry, 31(2), 455–461. https://doi.org/10.1002/jcc.21334
Daina, A., Michielin, O., & Zoete, V. (2017). SwissADME: A free web tool to evaluate pharmacokinetics, drug-likeness and medicinal chemistry friendliness of small molecules. Scientific Reports, 7(1), 1–13. https://doi.org/10.1038/srep42717
Srivastava, R. (2021). Theoretical studies on the molecular properties, toxicity, and biological efficacy of 21 new chemical entities. ACS Omega, 6(38), 24891–24901. https://doi.org/10.1021/acsomega.1c03736
Filimonov, D. A., Lagunin, A. A., Gloriozova, T. A., Rudik, A. V., Druzhilovskii, D. S., Pogodin, P. V., & Poroikov, V. V. (2014). Prediction of the biological activity spectra of organic compounds using the PASS online web resource. Chemistry of Heterocyclic Compounds, 50(3), 444–457. https://doi.org/10.1007/s10593-014-1496-1
Lagunin, A., Stepanchikova, A., Filimonov, D., & Poroikov, V. (2000). PASS: Prediction of activity spectra for biologically active substances. Bioinformatics, 16(8), 747–748. https://doi.org/10.1093/bioinformatics/16.8.747
Weininger, D. (1988). SMILES, a chemical language and information system. 1. Introduction to methodology and encoding rules. Journal of Chemical Information and Computer Sciences, 28(1), 31–36. https://doi.org/10.1021/ci00057a005
Van Der Spoel, D., Lindahl, E., Hess, B., Groenhof, G., Mark, A. E., & Berendsen, H. J. (2005). GROMACS: Fast, flexible, and free. Journal of Computational Chemistry, 26(16), 1701–1718. https://doi.org/10.1002/jcc.20291
Jo, S., Kim, T., Iyer, V. G., & Im, W. (2008). CHARMM-GUI: A web‐based graphical user interface for CHARMM. Journal of Computational Chemistry, 29(11), 1859–1865. https://doi.org/10.1002/jcc.20945
Huang, J., & MacKerell, A. D., Jr. (2013). CHARMM36 all-atom additive protein force field: Validation based on comparison to NMR data. Journal of Computational Chemistry, 34(25), 2135–2145. https://doi.org/10.1002/jcc.23354
Averill, F. W., & Painter, G. S. (1992). Steepest-descent determination of occupation numbers and energy minimization in the local-density approximation. Physical Review B, 46(4), 2498. https://doi.org/10.1103/PhysRevB.46.2498
Hess, B., Bekker, H., Berendsen, H. J., & Fraaije, J. G. (1997). LINCS: A linear constraint solver for molecular simulations. Journal of Computational Chemistry, 18(12), 1463–1472. https://doi.org/10.1002/(SICI)1096-987X(199709)18:12%3C1463::AID-JCC4%3E3.0.CO;2-H
Rühle, V. (2008). Pressure coupling/barostats. Journal Club, 1–5. https://www2.mpipmainz.mpg.de/~andrienk/journal_club/barostats.pdf
Van Gunsteren, W. F., & Berendsen, H. J. (1988). A leap-frog algorithm for stochastic dynamics. Molecular Simulation, 1(3), 173–185. https://doi.org/10.1080/08927028808080941
Essmann, U., Perera, L., Berkowitz, M. L., Darden, T., Lee, H., & Pedersen, L. G. (1995). A smooth particle mesh Ewald method. The Journal of Chemical Physics, 103(19), 8577–8593. https://doi.org/10.1063/1.470117
Peterson, B., Weyers, M., Steenekamp, J. H., Steyn, J. D., Gouws, C., & Hamman, J. H. (2019). Drug bioavailability enhancing agents of natural origin (bioenhancers) that modulate drug membrane permeation and pre-systemic metabolism. Pharmaceutics, 11(1), 33.
Yang, H., Yang, M., Ding, Y., Liu, Y., Lou, Z., Zhou, Z., … Rao, Z. (2003). The crystal structures of severe acute respiratory syndrome virus main protease and its complex with an inhibitor. Proceedings of the National Academy of Sciences, 100(23), 13190–13195. https://doi.org/10.1073/pnas.1835675100
Chen, Y. W., Yiu, C. P. B., & Wong, K. Y. (2020). Prediction of the SARS-CoV-2 (2019-nCoV) 3C-like protease (3CL pro) structure: Virtual screening reveals velpatasvir, ledipasvir, and other drug repurposing candidates. F1000 Research, 9. https://doi.org/10.12688/f1000research.22457.2
Hoelder, S., Clarke, P. A., & Workman, P. (2012). Discovery of small molecule cancer drugs: Successes, challenges and opportunities. Molecular Oncology, 6(2), 155–176. https://doi.org/10.1016/j.molonc.2012.02.004
Ertl, P., Rohde, B., & Selzer, P. (2000). Fast calculation of molecular polar surface area as a sum of fragment-based contributions and its application to the prediction of drug transport properties. Journal of Medicinal Chemistry, 43(20), 3714–3717. https://doi.org/10.1021/jm000942e
Ritchie, T. J., Macdonald, S. J., Peace, S., Pickett, S. D., & Luscombe, C. N. (2013). Increasing small molecule drug developability in sub-optimal chemical space. Medicinal Chemistry Communications, 4(4), 673–680. https://doi.org/10.1039/C3MD00003F
Sokolov, D. N., Zarubaev, V. V., Shtro, A. A., Polovinka, M. P., Luzina, O. A., Komarova,N. I., … Kiselev, O. I. (2012). Anti-viral activity of (–)-and (+)-usnic acids and their derivatives against influenza virus A (H1N1) 2009. Bioorganic & Medicinal Chemistry Letters, 22(23), 7060–7064. https://doi.org/10.1016/j.bmcl.2012.09.084
Shtro, A. A., Zarubaev, V. V., Luzina, O. A., Sokolov, D. N., & Salakhutdinov, N. F. (2015). Derivatives of usnic acid inhibit broad range of influenza viruses and protect mice from lethal influenza infection. Antiviral Chemistry and Chemotherapy, 24(3–4), 92–98.
Baidya, A. T., Kumar, A., Kumar, R., & Darreh-Shori, T. (2022). Allosteric binding sites of Aβ peptides on the acetylcholine synthesizing enzyme ChAT as deduced by in silico molecular modeling. International Journal of Molecular Sciences, 23(11), 6073. https://doi.org/10.3390/ijms23116073
Acknowledgements
Mr. Amit Gupta (09/013(0912)/2019-EMR-I) and Ms. Niharika Sahu (09/013(0927)/2020-EMR-I) are thankful to the Council of Scientific & Industrial Research (CSIR), New Delhi, India, for the Senior Research Fellowship (SRF) and Junior Research Fellowship (JRF). Mr. Ashish P. Singh (NTA Ref. No. 191620014505) is thankful to the University Grants Commission (UGC), New Delhi, India, for the financial assistance in the form of fellowship as Junior Research Fellow. All authors would also like to take this opportunity to thank the Head of the Department of Botany and the Coordinator, School of Biotechnology, Banaras Hindu University, for providing the necessary facilities and encouragement to carry out this work. The support and the resources provided by “PARAM Shivay Facility” under the National Supercomputing Mission, Government of India at the Indian Institute of Technology (BHU), Varanasi, are gratefully acknowledged.
Author information
Authors and Affiliations
Contributions
AG, SCS, VJU, and RPS conceptualized the aims and methodology. AG performed formal data analysis, was involved in data curation, and produced the first draft of the manuscript under the supervision of SCS, VJU, and RPS. NS and APS performed formal data analysis and was involved in data curation. AT and RK performed the molecular dynamics study. VKS performed formal data analysis and validation. All the authors have read and approved the manuscript.
Corresponding author
Ethics declarations
Ethical Approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Consent to Participate
All authors participated in the study either performing experiments or writing the manuscript.
Consent to Publish
All authors agree to publish the manuscript.
Conflict of Interest
The authors declare no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Gupta, A., Sahu, N., Singh, A.P. et al. Exploration of Novel Lichen Compounds as Inhibitors of SARS-CoV-2 Mpro: Ligand-Based Design, Molecular Dynamics, and ADMET Analyses. Appl Biochem Biotechnol 194, 6386–6406 (2022). https://doi.org/10.1007/s12010-022-04103-3
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12010-022-04103-3