Abstract
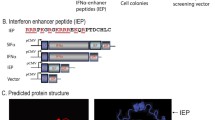
Interferon (IFN)-λ, also known as IL-28A, IL-28B, or IL-29, is a new type III IFN, which shares many functional characteristics with type I IFN (α/β). Currently, IFN-α is used in the treatment of certain forms of cancer with severe adverse effects. Some researches had stated that IFN-λs induced a similar but restricted growth inhibition of tumor cells relative to IFN-α; moreover, mutations of IFN-λs could strongly impact its biological properties. In this study, three hIL-29 mutants (K33R, R35K, and K33R/R35K) were generated by site-directed mutagenesis and efficiently expressed in Pichia pastoris GS115, which have considerable abilities to inhibit the growth of BEL-7402, HCT-8, and SGC-7901 tumor cells in vitro. The results showed that these mutants (K33R, R35K, and K33R/R35K) exhibited a significantly enhanced anti-proliferation activity against these tumor cells, compared with native hIL-29 in vitro. Further assay in vitro indicated that superior to K33R and R35K, K33R/R35K had a significant increase in anti-tumor activity compared with IFN-α2b, which suggested that the K33R/R35K could make improvement for the effectiveness of native hIL-29 in clinic and could be used as a potentially powerful candidate for cancer immunotherapy.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In 2003, three highly related cytokines were discovered: interleukin-29 (IL-29), IL-28A, and IL-28B. They are also designated as interferon (IFN)-λ1, λ2, and λ3, respectively, or together as type III IFNs, which use a distinct receptor complex consisting of two unique subunits, named IFN-λR1 and IL-10R2 [1, 2]. Both IFN-λs and type I IFNs are expressed in response to viral infection [3] and activate the JAK/STAT pathway [4], which imply that IFN-λs exhibit biological properties similar to type I IFNs. Like type I IFNs, IFN-λs could induce specific anti-tumor response [5]. However, compared with type I IFNs, only a few cells are responsive to IFN-λs, implying a less severe adverse effect profile [6].
Structurally, IFN-λs possess a characteristic topology consisting of six secondary structure elements named A–F. Helices A, C, D, and F form a four-helix bundle, which is the core of the structure. The helices A and F of IL-29 play important roles in receptor binding for exerting its biological effects [7, 8]. Compared with the crystal structure of IL-28B, 81 % identical to IL-29, only four amino acid residues of IL-29 were different from IL-28B at the positions in helices A and F: Lys-33, Arg-35, Lys-43, and Ala-143, which corresponded to Arg-34, Lys-36, Leu-44, and Pro-144 in IL-28B. Interestingly, mutagenesis at Arg-34 and Lys-36 of IL-28B revealed a profound impact on its biological activities [8]. Accordingly, it has been hypothesized that point mutations at positions Lys-33 and Arg-35 in IL-29 might influence its biological activities, such as the anti-tumor activity.
In this work, with the analysis of hydrophobicity (http://web.expasy.org/protscale) and binding energetics [9], three hIL-29 mutants were created by site-directed mutagenesis based on the corresponding residues of IL-28B. Positions at Lys-33, Arg-35, and Lys-33/Arg-35 in hIL-29 were mutated to Arg-33, Lys-35, and Arg-35/Lys-33, separately. In vitro, these mutants exhibited a significantly enhanced anti-proliferation activity against human BEL-7402, HCT-8, and SGC-7901 tumor cells compared with native hIL-29 and IFN-α2b, which suggested that the constructed hIL-29 mutants could be used as a potentially powerful candidate for cancer immunotherapy in the future.
Materials and Methods
Plasmids, Strains, and Cell Lines
Escherichia coli JM109 and plasmid pUCm-T (Sangon, Shanghai, China) were used for gene cloning and DNA sequencing. The recombinant cloning plasmid pUCm-T-hIL-29 was constructed in our laboratory, which contains full length hIL-29 complementary DNA (cDNA) sequence. The expression plasmid pPIC9KM (Invitrogen, USA) and Pichia pastoris GS115 were applied for the construction of recombinant expression plasmids and the expression of the target DNA fragments encoding hIL-29 mutants. Both the E. coli JM109 and P. pastoris GS115 were purchased from Invitrogen. E. coli JM109 was cultured at 37 °C in Luria broth (LB) medium containing the following antibiotics: ampicillin (100 μg/mL) or kanamycin (50 μg/mL), according to the requirement for maintaining the plasmids. P. pastoris GS115 was cultured at 30 °C in following media that were prepared according to the manual of Multi-Copy Pichia Expression Kit (Invitrogen): minimal dextrose (MD), yeast extract peptone dextrose (YPD), buffered glycerol-complex (BMGY), and buffered methanol-complex (BMMY).
Human hepatic carcinoma cell line BEL-7402, human colonic carcinoma cell line HCT-8, and human gastric carcinoma cell line SGC-7901 were obtained from the Cell Bank of the Chinese Academy of Sciences (Shanghai, China) and maintained in Roswell Park Memorial Institute (RPMI) 1640 supplemented with 10 % fetal bovine serum (FBS).
Chemicals and Enzymes
Dimethyl sulfoxide (DMSO), bicinchoninic acid (BCA) protein assay kit, and Cell Counting Kit-8 (CCK-8) were purchased from Beyotime Institute of Biotechnology (Haimen, China). RPMI 1640 medium and FBS were purchased from Hyclone (USA). Recombinant human IL-29 was purchased from ProSpec (Israel). Recombinant human IFN-α2b was purchased from Kaiyin Technology Co. (Beijing, China). Restriction enzymes and Pfu DNA polymerase were purchased from Takara Biotechnology Co. (Dalian, China). DNA gel extract kit and mini-preparation of plasmid kit were purchased from BBI (Canada).
Construction of hIL-29 Mutants by Site-Directed Mutagenesis
The plasmid pUCm-T-hIL-29 containing full length hIL-29 cDNA sequence was used as a template. Single amino acid substitutions of hIL-29 gene were introduced using megaprimer PCR method [10]. All primers used for sited-directed mutagenesis are listed in Table 1. Briefly, the desired mutations were introduced into hIL-29 gene by way of two rounds of PCR. Using the hIL-29 cDNA as the template, the first-round PCR was carried out using primers hIL-29-F and P1 (P2 or P3) with the following conditions: an initial denaturation at 94 °C for 2 min; 30 cycles of amplification at 94 °C for 30 s, 55 °C for 30 s, 72 °C for 20 s; and a final extension at 72 °C for 10 min. Then, the amplified fragments were purified using a DNA Gel Extract Kit to remove redundant primers. Afterward, each pair of these fragments was combined with primer hIL-29-R in the reaction mixture, and the second round of PCR was performed under the same conditions except with a different annealing condition (94 °C for 30 s, 57 °C for 30 s, 72 °C for 60 s). Finally, the resulting PCR products were purified again, inserted into pUCm-T separately, and identified by restriction enzyme analysis and DNA sequencing.
Construction and Transformation of the Recombinant Vector
The DNA fragments were excised from pUCm-T-hIL-29-K33R (R35K, and K33R/R35K) with restriction endonucleases Xho I and Not I, agarose gel-purified, and then inserted into the Xho I and Not I sites of pPIC9KM vector, followed by transforming it into E. coli JM109. The positive clones were identified by digestion with Xho I and Not I as well as DNA sequencing.
Furthermore, the resulting recombinant expression vectors, designated pPIC9KM-hIL-29-K33R (R35K, and K33R/R35K), were linearized with Sal I and transformed into P. pastoris GS115 by electroporation.
Screening and Expression of P. pastoris Transformants
All P. pastoris transformants were first spread on a MD plate and then successively inoculated on G418-containing YPD plates at increasing concentrations of 2.0 and 4.0 mg/mL for screening multiple copies. Expression of the mutant proteins in P. pastoris GS115 was performed according to the instruction of Multi-Copy Pichia Expression Kit with slight modification [11]. Each single colony of the transformants was inoculated into the BMGY medium and cultured at 30 °C with 220 rpm until the OD600 reached 2–4. The cells were then collected by centrifugation, resuspended in BMMY medium, and induced for expression of the recombinant proteins by adding methanol to a final concentration of 2.0 % (v/v) at 24-h intervals at 30 °C for 72 h. The transformants expressing the highest protein activity, named for P. pastoris GS115-K33R (R35K, and K33R/R35K), were selected and used for further studies.
Purification of the Expressed K33R (R35K, and K33R/R35K)
After fermentation process, the supernatants were harvested by centrifugation (8000 rpm, 10 min). The collected supernatants were filtered with a 0.45-μm cellulose membrane filter and further concentrated by 10-kDa cutoff membrane (Millipore, USA). The concentrated products were loaded onto SP Sepharose Fast Flow column pre-equilibrated with buffer A (20 mmol/L Na2HPO4, 10 mmol/L C6H8O7, pH 6.0). The column was washed with buffer A, and the bound proteins were eluted with 20, 40, 60, 80, and 100 % buffer B (20 mmol/L Na2HPO4, 10 mmol/L C6H8O7, 1 mol/L NaCl, pH 6.0). The eluted proteins were analyzed by 12 % sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE).
Sephadex G-75 column was used for the second-step purification. The column was pre-equilibrated with buffer A. The fractions containing mutant proteins were concentrated to 2.0 mL by ultrafiltration, loaded onto the column, and washed with buffer A. The eluents containing mutant proteins were pooled and analyzed by 12 % SDS-PAGE, HPLC, and BCA protein assay kit. Finally, K33R, R35K, and K33R/R35K were obtained by vacuum freeze-drying method. All purification procedures were performed at 4 °C.
Western Blot Analysis
The purity of the final products was assessed by HPLC method [12]. For western blot analysis, samples were subjected to 12 % SDS-PAGE and electroblotted onto a nitrocellulose (NC) membrane (Millipore) which was then stained with Ponseau S to confirm equivalent loads. Ponseau S was washed off, and the membrane was blocked by blocking solution containing 10 % rabbit serum overnight at 4 °C. Afterward, it was incubated with a 1:1000 dilution of goat anti-human IL-29 antibody (R&D systems, USA) at 37 °C for 2 h and then washed with TBS with 0.05 % Tween 20 (TBS-T) for three times (5 min each). Eventually, the membrane was incubated with a 1:2500 dilution of the rabbit anti-goat IgG (R&D systems) conjugated to HRP. Immunoreactivity was visualized with diaminobenzidine (DAB) as substrate.
Anti-Tumor Activity Assay
Tumor cells BEL-7402, HCT-8, and SGC-7901 were separately used as target cells. Target cells (1.0 × 104 cell/well) were added to a 96-well plate and incubated for 24 h in a humidified incubator at 37 °C in 5 % CO2. The medium was then removed and replaced with 100 μL of medium containing K33R (R35K, and K33R/R35K) and native hIL-29. Diluted proteins were separately added to the 96-well plate at 50, 500, or 1000 ng/mL in triplicate. Afterward, target cells were incubated for another 48 h. The blank wells (1640 medium only), unsettled BEL-7402 cells, HCT-8 cell or SGC-7901 cell control wells (tumor cells only), and IFN-α2b-treated wells (tumor cells and IFN-α2b) were used as control. The negative control was BSA.
Inhibitions of tumor cells growth acted by hIL-29 or hIL-29 mutants were determined by CCK-8 assay [13]. Absorbance value was measured with a microplate reader at a test wavelength of 450 nm against a reference wavelength of 630 nm. The final absorbance was the difference in absorbance read at 450 nm compared with that read at 630 nm. Tumor growth inhibition (%) was calculated with the equation: 100 − [(Abs value in protein-treated cells well − Abs value in blank control well) / (Abs value in unsettled tumor cells control well − Abs value in blank control well)] × 100.
Statistical Analysis
Results were presented as the mean ± SD of triplicates from at least three independently performed experiments. Statistical evaluation was performed using one-way ANOVA test. Differences between groups were considered statistically significant if P values <0.05.
Results
Cloning and Construction of the Expression Plasmid Vectors pPIC9KM-hIL-29-K33R (R35K, and K33R/R35K)
Site-directed mutagenesis was used to construct the hIL-29 mutants as described in “Materials and Methods” section. The mutant genes were generated by using the megaprimer PCR method and cloned into the pUCm-T vector. The positive colonies were verified by restriction endonuclease digestion and DNA sequencing. Sequence analysis of single clone showed that the desired mutations at positions 33 and 35 were successfully introduced.
For constructing pPIC9KM-hIL-29-K33R (R35K, and K33R/R35K), the digested mutant genes from pUCm-T-hIL-29-K33R (R35K, and K33R/R35K) were subcloned into the pPIC9KM expression vector based on its open reading frame (ORF), and the colonies were confirmed by DNA sequencing and double digestion of Xho I and Not I. The recombinant expression vectors were linearized with Sal I and transformed into P. pastoris GS115 cells via electroporation for expressing mutant proteins. Figure 1 shows the results of PCR amplification and restriction endonuclease digestion.
Verification of hIL-29 mutant PCR products and engineered hIL-29 mutant vectors. a Megaprimer PCR was performed by way of two rounds of PCR. Afterward, PCR fragments were separated by gel electrophoresis. Data from one out of three experiments are shown. b HIL-29 mutant vectors were digested by restriction endonucleases, and fragments were subsequently separated by gel electrophoresis. Data from one out of three experiments are shown. M: nucleic acid size marker
Screening and Expression of P. pastoris Transformants
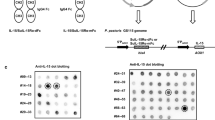
P. pastoris transformants that could resist higher concentrations of geneticin G418 might have integrated multiple copies of heterologous genes into its genome, which could potentially lead to higher expression levels of heterologous proteins [14]. Based on the manual of Multi-Copy Pichia Expression Kit, 20 transformants of each mutant resistant to 2.0 and 4.0 mg/mL of G418 were picked out respectively for flask expression tests. P. pastoris transformed with pPIC9KM empty plasmid, named for P. pastoris GS115-9K, was used as the negative control. After 72-h induction by adding 2.0 % (v/v) methanol at 24-h intervals, the cultured supernatants of these transformants were harvested for SDS-PAGE and western blot assays, respectively (data not shown). Among all tested transformants, three strains expressing the maximum recombinant mutant proteins, designated P. pastoris GS115-K33R (R35K, and K33R/R35K), were selected and used for subsequent studies (Fig. 2a). Figure 2 shows that the recombinant mutant proteins (22.6 kDa) were successfully secreted into the culture mediums. No band appeared for the supernate from the strain transfected with empty pPIC9KM vector (Fig. 2a, lane 1). Meanwhile, western blot assays indicated that the 22.6 kDa band corresponded to the mutant proteins (Fig. 3, lanes 1–3).
Identification of hIL-29 mutant fermentation products and purified hIL-29 mutants. a Subjected to the ethanol environment, hIL-29 mutant fermentation products were collected and centrifuged. Afterward, total proteins of hIL-29 mutants were separated by 12 % SDS-PAGE. Data from one out of four experiments are shown. b HIL-29 mutant fermentation products were successively purified by filtration, ultrafiltration, cation exchange chromatography, and gel filtration. Then, purified hIL-29 mutants were subsequently separated by 12 % SDS-PAGE. Data from one out of three experiments are shown. M: protein molecular weight marker
Western blot analysis of hIL-29 mutant proteins and purified hIL-29 mutants. After 12 % SDS-PAGE and electrotransfection, the identities of hIL-29 mutant fermentation proteins and corresponding purified hIL-29 mutants were confirmed with immunoreactivity with goat anti-human IL-29 antibody and rabbit anti-goat IgG in western blot analysis. Data from one out of six experiments are shown. M: protein molecular weight marker
Purification and Characterization of the Expressed K33R (R35K, and K33R/R35K)
After 72 h of induction with 2.0 % methanol, culture mediums were centrifuged and the supernatants were collected. The supernatants were then filtered (0.45 μm) to remove the particulates and further concentrated via 10-kDa cutoff ultrafiltration. Through cation exchange chromatography and gel filtration, the purity of recombinant mutant proteins K33R (R35K, and K33R/R35K) reached 95 % above with a yield of 3.6, 3.2, and 4.4 mg protein/L separately as revealed by SDS-PAGE (Fig. 2b), HPLC (data not shown), and BCA protein assay kit, which exhibited specific reaction with goat anti-human IL-29 antibody and rabbit anti-goat IgG. These purified recombinant mutant proteins displayed a single protein band with an apparent MW of 22.6 kDa (Figs. 2b and 3, lanes 4–6).
Anti-Tumor (Anti-Proliferation) Activity In Vitro
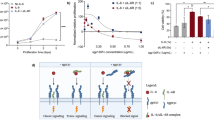
In order to study the anti-proliferation activity of hIL-29 mutants, the inhibitions of growth of BEL-7402, HCT-8, and SGC-7901 tumor cells induced by hIL-29 and mutant proteins were examined by CCK-8 assay in vitro. Compared with BSA, hIL-29, and K33R (R35K, and K33R/R35K), mutants could induce a significant anti-proliferation effect in vitro (P < 0.01) which were directly correlated with protein concentrations ranging from 50 to 1000 ng/mL (Fig. 4a–c). Among these constructed mutants, the mutant K33R/R35K displayed a strongly increased anti-proliferation effect on these tumor cells, whereas the other two induced a weaker response. In addition, at the same dosage level, especially at 1000 ng/mL, K33R/R35K exhibited a significantly enhanced growth inhibitory effect on these tumor cells whose inhibition rates were about 40 % in comparison with native hIL-29 (20 %, P < 0.01) and IFN-α2b (23–27 %, P < 0.05), which were consistent with the previous hypothesis (Fig. 4a–c).
Effects of native hIL-29 and hIL-29 mutants on inhibition of the growth of tumor cells in vitro. a Human hepatic carcinoma cells BEL-7402 were seeded in 96-well plate at a density of 1 × 104 cell/well, and proteins in different concentrations were added separately to triplicate wells to inhibit the growth of BEL-7402 cells by CCK-8 assay. b The inhibition of growth of human colonic carcinoma cells HCT-8 acted by hIL-29 and hIL-29 mutants were determined by CCK-8 assay. The cell density and protein concentration were consistent with the values used in a. c The inhibition of growth of human gastric carcinoma cells SGC-7901 acted by hIL-29 and hIL-29 mutants were determined by CCK-8 assay. The cell density and protein concentration were consistent with the values used in a. IFN-α2b and BSA were used as positive and negative controls, respectively. Data shown are representative of at least three separate experiments, and the values represent the mean ± standard error of the mean. Statistical evaluation was performed using one-way ANOVA test. Compared with native hIL-29, differences between the treatment groups were considered statistically significant if *P values <0.05 and **P values <0.01
Discussion
Type I IFNs are essential members in the family of anti-cancer drugs used as a treatment for several malignancies in clinic practice. IFN-λs, similar to type I IFNs, appear to share the analogous anti-viral and anti-tumor (anti-proliferation) activity. Type I IFNs are able to inhibit replication of a number of viruses in vitro, and the ability to interfere with viral replication in the absence of cells of the immune system has been defined as the anti-viral state [15]. The finding that IFN-λs also were able to inhibit viral replication in vitro was one of the first pieces of evidence suggesting an IFN-like function of these cytokines [1, 2]. However, it was noted that the IL-29 displayed significantly lower anti-viral activity than type I IFNs both in vitro and in vivo and that IL-29 was generally more potent than IL-28A/B [16], while another study reported that growth inhibitory responses induced by type I IFNs and IFN-λs are distinct and such effect correlated with differences in the duration of JAK/STAT pathway activation and prolonged ISG expression [4]. Although type I IFNs and IFN-λs activated the same components of the JAK/STAT pathway and shared the anti-proliferation activity, they differed at least in the magnitude of the anti-proliferative response. In this study, it was found that IFN-λs were more efficient than IFN-α in inducing an anti-proliferative effect that overlapped with the activation of apoptosis. The fact that IFN-λs could prolong the duration of STAT activation and promote apoptosis was of clinical interest, particularly since most patients receiving IFN-α must discontinue therapy due to severe side effects [17]. This comes as no surprise since IFN-α/β receptors are virtually expressed on all cell types. In addition, IFN-α signals are short-lived and tumors have the capacity to become desensitized to this cytokine [4]. However, the distribution of receptors of IFN-λs is much more restricted than that of type I IFNs, implying a less severe adverse effect profile [18]. In conclusion, these findings strongly suggested that prolonged duration of IFN-λs induced STAT activation and gene expression may account for the enhanced anti-growth and apoptotic effects that were not achievable even with high-dose IFN-α. Considering the adverse effects of IFN-α immunotherapy and that IFN-λR1 is not ubiquitously expressed as IFN-AR1/IFN-AR2, perhaps the combination of low-dose IFN-α and high-dose IFN-λs could be a more effective cancer therapy. Given that IFN-λs could induce a robust anti-proliferative effect in tumor cells that respond less optimally to IFN-α, this highlighted the potency of IFN-λs in suppressing the growth of tumors expressing both types of IFN receptors. Therefore, compared with type I IFNs, IFN-λs seem to become the alternative with similar functions but fewer adverse effects to satisfy current clinical needs.
Since the discovery of IFN-λs, IL-29 has been considered the most potent of the three members in clinical assays. In antecedent work, recombinant forms of each IFN-λ were produced and their biological activities compared. A cluster of six residues (Lys-33, Arg-34, Lys-36, and Asp-37 on helix A and Phe-155 and Phe-158 on helix F) was found at the center of the A and F helices in IL-28B. Mutation of any of these residues had a profound impact on its biological activities [7]. As compared with IL-29, four residues (Arg-34, Lys-36, Leu-44, and Pro-144) in helices A and F were probed in IL-28B (Fig. 5). It was demonstrated that Arg-34 and Lys-36 in IL-28B should be the remarkable residues to influence its biological activities, which corresponded to Lys-33 and Arg-35 in IL-29. Accordingly, it has been conjectured that these two residues in IL-29 could impact its biological activities such as the anti-tumor activity as well.
Comparison of crystal structure between hIL-29 and hIL-28B. hIL-29 is depicted in green while hIL-28B is depicted in red, with detected residues depicted in stick format. Four amino acid residues of IL-29 were probed at the positions in helices A and F which were different from that of IL-28B: Lys-33, Arg-35, Lys-43, and Ala-143, which corresponded to Arg-34, Lys-36, Leu-44, and Pro-144 in IL-28B
Meanwhile, previous studies had indicated that Lys-33 and Arg-35 in IL-29 resided in IL-10R2 and IFN-λR1 binding domains, and these residues played important role for IL-29 to interact with IL-10R2 or IFN-λR1 [8]; thus, it is possible that change of these two amino acids in IL-29 to the corresponding IL-28B residues may increase the receptor binding affinity of IL-29 and elicit an enhanced anti-tumor activity.
The spatial arrangement of atoms in a protein is called its conformation. The conformations existing under a given set of conditions are usually the ones that are thermodynamically the most stable, having the lowest binding free energy [19]. With the analysis of binding energetics (Table 2), we found that these mutants had lower binding free energy than wild-type hIL-29, among which K33R/R35K had the lowest binding free energy, that is, the most stable protein conformation. Simultaneously, the predictions of website displayed that the hydrophobicities of these designed mutants are basically unchanged.
For these reasons, in this work, we described the construction and expression of three mutant hIL-29 proteins by site-directed mutagenesis. The results indicated that the constructed mutant proteins K33R, R35K, and K33R/R35K exhibited significantly enhanced growth inhibitory effect on BEL-7402, HCT-8, and SGC-7901 tumor cells compared with native hIL-29, among which K33R/R35K displayed the dramatically obvious effect at a dosage of 1000 ng/mL. Consistently, compared with positive control IFN-α2b, K33R/R35K also exhibited the increased ability of inhibiting the growth of these tumor cells (P < 0.01), while the inhibition effects of K33R and R35K were similar with that of IFN-α2b (P < 0.05), which suggested that high concentration of these mutant proteins, especially K33R/R35K, could be used as potentially powerful substitute for IL-29 in cancer immunotherapy.
In conclusion, three hIL-29 mutants (K33R, R35K, and K33R/R35K) were constructed and characterized. The expression gave a yield of 3.6, 3.2, and 4.4 mg protein/L, respectively (with a purity of 95 % above). These mutants displayed a significantly enhanced anti-tumor (anti-proliferation) activity compared with native hIL-29 and IFN-α2b in vitro. Among them, K33R/R35K exhibited the superior growth inhibitory effect against BEL-7401, HCT-8, and SGC-7901 tumor cells, which might be a promising substitute for hIL-29 in cancer immunotherapy and provide a new strategy for the clinic application of IFN-λs. Meanwhile, further investigation of the role of K33R, R35K, and K33R/R35K in vivo is necessary in order to determine the perfect immunotherapeutic strategy.
References
Kotenko, S. V., Gallagher, G., Baurin, V. V., Lewis-Antes, A., Shen, M., Shah, N. K., & Donnelly, R. P. (2003). IFN-λs mediate antiviral protection through a distinct class II cytokine receptor complex. Nature Immunology, 4, 69–77.
Sheppard, P., Kindsvogel, W., Xu, W., Henderson, K., Schlutsmeyer, S., Whitmore, T. E., & Klucher, K. M. (2003). IL-28, IL-29 and their class II cytokine receptor IL-28R. Nature Immunology, 4, 63–68.
Onoguchi, K., Yoneyama, M., Takemura, A., Akira, S., Taniguchi, T., Namiki, H., & Fujita, T. (2007). Viral infections activate types I and III interferon genes through a common mechanism. Journal of Biological Chemistry, 282, 7576–7581.
Maher, S. G., Sheikh, F., Scarzello, A. J., Romero-Weaver, A. L., Baker, D. P., Donnelly, R. P., & Gamero, A. M. (2008). IFN-α and IFN-λ differ in their antiproliferative effects and duration of JAK/STAT signaling activity. Cancer Biology and Therapy, 7, 1109–1115.
Li, M., Liu, X., Zhou, Y., & Su, S. B. (2009). Interferon-λs: the modulators of antivirus, antitumor, and immune responses. Journal of Leukocyte Biology, 86, 23–32.
Kotenko, S. V. (2011). IFN-λs. Current Opinion in Immunology, 23, 583–590.
Gad, H. H., Dellgren, C., Hamming, O. J., Vends, S., Paludan, S. R., & Hartmann, R. (2009). Interferon-λ is functionally an interferon but structurally related to the interleukin-10 family. Journal of Biological Chemistry, 284, 20869–20875.
Miknis, Z. J., Magracheva, E., Li, W., Zdanov, A., Kotenko, S. V., & Wlodawer, A. (2010). Crystal structure of human interferon-λ1 in complex with its high-affinity receptor interferon-λR1. Journal of Molecular Biology, 404, 650–664.
Wang, T., Tomic, S., Gabdoulline, R. R., & Wade, R. C. (2004). How optimal are the binding energetics of barnase and barstar? Biophysical Journal, 87, 1618–1630.
Xie, Z. H., & Shi, X. J. (2009). Fast and almost 100% efficiency site-directed mutagenesis by the megaprimer PCR method. Progress in Biochemistry and Biophysics, 36, 1490–1494.
Li, J. F., Tang, C. D., Shi, H. L., & Wu, M. C. (2011). Cloning and optimized expression of a neutral endoglucanase gene (ncel5A) from Volvariella volvacea WX32 in Pichia pastoris. Journal of Bioscience and Bioengineering, 111, 537–540.
Huang, Y. S., Chen, Z., Yang, Z. Y., Wang, T. Y., Zhou, L., Wu, J. B., & Zhou, L. F. (2007). Preparation and characterization of a potent, long-lasting recombinant human serum albumin-interferon-α2b fusion protein expressed in Pichia pastoris. European Journal of Pharmaceutics and Biopharmaceutics, 67, 301–308.
Lou, J., Chu, G., Zhou, G., Jiang, J., Huang, F., Xu, J., & He, J. (2010). Comparison between two kinds of cigarette smoke condensates (CSCs) of the cytogenotoxicity and protein expression in a human B-cell lymphoblastoid cell line using CCK-8 assay, comet assay and protein microarray. Mutation Research, Genetic Toxicology and Environmental Mutagenesis, 697, 55–59.
Tan, Z. B., Li, J. F., Wu, M. C., Tang, C. D., Zhang, H. M., & Wang, J. Q. (2011). High-level heterologous expression of an alkaline lipase gene from Penicillium cyclopium PG37 in Pichia pastoris. World Journal of Microbiology and Biotechnology, 27, 2767–2774.
Pestka, S., Krause, C. D., & Walter, M. R. (2004). Interferons, interferon-like cytokines, and their receptors. Immunological Reviews, 202, 8–32.
Ank, N., West, H., & Paludan, S. R. (2006). IFN-λ: novel antiviral cytokines. Journal of Interferon & Cytokine Research, 26, 373–379.
Belardelli, F., Ferrantini, M., Proietti, E., & Kirkwood, J. M. (2002). Interferon-alpha in tumor immunity and immunotherapy. Cytokine & Growth Factor Reviews, 13, 119–134.
Ank, N., West, H., Bartholdy, C., Eriksson, K., Thomsen, A. R., & Paludan, S. R. (2006). Lambda interferon (IFN-λ), a type III IFN, is induced by viruses and IFNs and displays potent antiviral activity against select virus infections in vivo. Journal of Virology, 80, 4501–4509.
Kamiya, N., Yonezawa, Y., Nakamura, H., & Higo, J. (2008). Protein-inhibitor flexible docking by a multicanonical sampling: native complex structure with the lowest free energy and a free-energy barrier distinguishing the native complex from the others. Proteins, 70, 41–53.
Acknowledgments
We thank Junqing Wang, Haijun Zheng, Chunlei Ge, and Rong Zhu for previous research work.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Lu, Y., Li, L., Chen, W. et al. Enhanced Anti-Tumor (Anti-Proliferation) Activity of Recombinant Human Interleukin-29 (IL-29) Mutants Using Site-Directed Mutagenesis Method. Appl Biochem Biotechnol 177, 1164–1175 (2015). https://doi.org/10.1007/s12010-015-1804-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12010-015-1804-y