Abstract
The oomycetous phytopathogen Pythium myriotylum secretes cellulases for growth/nutrition of the necrotroph. Cellulases are multi-enzyme system classified into different glycoside hydrolase (GH) families. The present study deals with identification and characterization of GH gene sequence from P. myriotylum by a PCR strategy using consensus primers. Cloning of the full-length gene sequence using genome walker strategy resulted in identification of 1230-bp P. myriotylum GH gene sequence, designated as PmGH1. Analysis revealed that PmGH1 encodes a predicted cytoplasmic 421 amino acid protein with an apparent molecular weight of 46.77 kDa and a theoretical pI of 8.11. Tertiary structure of the deduced amino acid sequence showed typical (α/β)8 barrel folding of family 1 GHs. Sequence characterization of PmGH1 identified the conserved active site residues, viz., Glu 181 and Glu 399, that function as acid-base catalyst and catalytically active nucleophile, respectively. Binding sites for N-acetyl-D-glucosamine (NAG) were revealed in the PmGH1 3D structure with Glu181 and Glu399 positioned on either side to form a catalytic pair. Phylogenetic analysis indicated a closer affiliation of PmGH1 with sequences of GH1 family. Results presented are first attempts providing novel insights into the evolutionary and functional perspectives of the identified P. myriotylum GH.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Pythium myriotylum Drechsler, a broad-host-range necrotroph [1, 2] and distributed worldwide, is the causal agent of pre- or post-emergence damping-off of seedlings or soft-rot disease [1, 3–6]. Like fungal phytopathogens, Pythium spp. have been reported to secrete a suite of cell wall degrading enzymes (CWDEs) [7, 8] that facilitate penetration of the host plant to initiate infection [2]. The plant cell wall composed primarily of cellulosic polysaccahrides [9] is a major structural barrier that affords defense against the invading pathogens. Therefore, CWDEs that constitute hallmarks of filamentous pathogen secretomes [10, 11] facilitate host cell wall degradation and constitute a key factor to phytopathogenic infection/colonization [12–14] as revealed by studies involving targeted gene disruption [15]. Domain repertoire and EST analysis of Pythium spp. has revealed an abundance of protein domains that include glycoside hydrolase, involved in host tissue degradation [16, 17].
Our earlier studies on identification of the suite of secretory CWDEs by P. myriotylum have revealed cellulases to constitute one of the major enzymes that play an important role in colonization/nutrition [18]. Cellulase constitute a multi-enzyme system that hydrolyzes β-1,4 glycosidic bonds of cellulose to yield simple sugars and involves the synergistic action of endoglucanases [endo-β-1,4-glucanase or carboxyl-methyl cellulases (CMCase) EC 3.2.1.4], cellobiohydrolases (exo-β-1,4-glucanase, EC 3.2.1.91), and β-glucosidases (EC 3.2.1.21) [19–21]. Among these, β-glucosidases that belong to glycoside hydrolase (GH) families 1, 5, 6, 7, 9, 12, 44, and 45 as listed in the CAZY database [22–24] are a rate-limiting factor supplementing endoglucanase and exoglucanase activities to ensure final glucose release [25, 26] and hence are good candidates for many biotechnological processes. Earlier reports indicating an abundance of GH transcripts in Pythium spp. [17] combined with our own studies indicating cellulases to be one of the major extracellular enzymes produced by P. myriotylum [18] prompted us to undertake the present study that aims at (i) cloning of a β-glucosidase gene from P. myriotylum and (ii) diversity and relationship of the identified gene to members of the characterized GH families.
Materials and Methods
P. myriotylum Culture Conditions and DNA Isolation
P. myriotylum strain (RGCBN14) obtained from Rajiv Gandhi Centre for Biotechnology (RGCB), Kerala, India, was grown on potato dextrose agar (PDA; pH 6.4) and incubated at 25 °C. Periodic sub-culturing was done at an interval of 7 days to maintain the oomycete in actively growing phase.
Genomic DNA of P. myriotylum was isolated using cetyl trimethyammonium bromide (CTAB) method [27]. Briefly, P. myriotylum mycelia (1 g) from 7-day-old PDA cultures finely ground with liquid nitrogen were added to pre-warmed extraction buffer (100 mM Tris HCl, pH 7.0; 20 mM EDTA, pH 8.0; 1 % PVP; 2 M NaCl; 2 % CTAB; 50 μl β-mercaptoethanol), vortexed, and incubated at 65 °C for 1 h. Extraction of DNA was initiated by adding an equal volume of chloroform/isoamyl alcohol (24:1), centrifuged at 10,000g for 15 min at 4 °C. DNA was precipitated by adding 1/10 volume of sodium acetate (pH 5.2) and 1 volume of ice-cold isopropanol, centrifuged at 10,000g for 15 min at 4 °C, and washed with 70 % ethanol. The pellet was dried and suspended in TE buffer (pH 8.0). After RNase treatment, DNA was extracted followed by precipitation as mentioned above, dissolved in 1XTE (Tris-EDTA) (pH 8.0), and stored at −20 °C until further use.
PCR and Cloning of GH Homologs from P. myriotylum
PCR amplification of P. myriotylum GH homologs was carried out using degenerate primers designed based on Phythopthora GHs, GHF (5′-AARTAYATYGTCGTCYWSAAG-3′) and GHR (5′-TTAAGYRKHKCCRTTGWAG-3′), and synthesized by Integrated DNA Technologies (IDT, USA). PCR was performed in a total volume of 20 μl containing 1 U of Taq DNA polymerase (Genie, Bangalore), 1× PCR buffer with 1.5 mM MgCl2, 0.2 mM dNTP, 1 μM of each primer, and 20 ng template DNA. Amplification was performed in a mastercycler gradient (Eppendorf, Germany) programmed for an initial denaturation at 94 °C for 5 min, followed by 35 amplification cycles (94 °C for 1 min, 52 °C for 1 min, and 72 °C for 1 min) and a final extension step at 72 °C for 5 min. Amplicons were gel-purified using Wizard SV Gel and PCR Clean-up system (Promega, Madison, USA), cloned using pGEM-T easy vector system (Promega), and sequenced in an ABI Prism 3730 genetic analyzer (Applied Biosystems, USA).
Isolation of Flanking Sequences by Genome Walker PCR
Four genome walker libraries using P. myriotylum genomic DNA were constructed by restriction digestion with EcoRV, DraI, PvuII, or StuI using Universal Genome Walker kit (Clontech, USA) according to manufacturer’s instructions. The libraries were used as templates for primary and secondary genome walker PCR assays using P. myriotylum GH-specific primers PmGH53R1 (5′-ATCGACGAGCTTCGCGCGAACAAGATC-3′) and PmGH53R1N (5′-ATCTTGACGCTCTATCACTGGGAT-3′) in combination with the adaptor primers provided in the kit according to manufacturer’s instructions using Advantage2 polymerase mix (Clontech) in Mastercycler eps (Eppendorf, Germany). PCR cycling parameters for primary PCR consisted of an initial 7 cycles of 94 °C for 25 s and 72 °C for 3 min followed by 32 cycles of 94 °C for 25 s and 67 °C for 3 min and a final extension at 67 °C for 7 min. For secondary PCR, the same program was followed except that instead of 32 cycles, the second step of PCR comprised of 20 cycles. PCR products after electrophoresis on 1.5 % agarose gel were gel-purified, cloned, and sequenced.
Sequence Analysis
Sequence identities of the clones were confirmed by homology searches with BLAST and BLASTX algorithms [28]. Multiple sequence alignment using CLUSTALX [29] of deduced amino acid sequence of P. myriotylum GH, designated PmGH1, with homologous sequences identified following NCBI searches was used to identify sequence characteristics of PmGH1. Putative signal sequences were identified using SignalP 4.1 [30] while protein molecular masses and isoelectric points were calculated using ProtParam tool (http://web.expasy.org/protparam). Phylogenetic analysis of PmGH1 and 78 representatives of GHs belonging to GH families 1, 2, 5, 6, 7, 9, 12, 44, and 48 from the CAZy (Carbohydrate-Active Enzymes) database was performed using minimum evolution phylogeny test implemented in the software MEGA version 4 [31]. Robustness of clustering was checked by bootstrapping 1000 replicates.
Structure Prediction of PmGH1
Secondary structure of putative protein was predicted by SOPMA program (http://npsa-pbil.ibcp.fr/). The 3D model of PmGH1 was generated using homology modeling server, Protein Homology/analogy Recognition Engine V 2.0 (Phyre2) (http://www.sbg.bio.ic.ac.uk/phyre2/). PyMOL (http://www.pymol.org), the molecular graphics system, was used to render figures. The quality of 3D model was evaluated by generating Ramachandran plot using RAMPAGE (http://mordred.bioc.cam.ac.uk/). The ligand binding site was predicted using 3D Ligand site (http://www.sbg.bio.ic.ac.uk/3dligandsite/) [32].
Results
Amplification and Cloning of GH Homologs from P. myriotylum
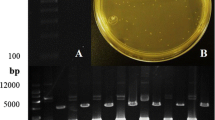
PCR amplification using degenerate primers designed based on Phythopthora GH family yielded an intense amplicon of 0.8 kb (Fig. 1a). The amplicon was cloned and randomly selected clones were sequenced. Sequences showed homology to β-glucosidase sequences cloned from other taxa. Based on this partial sequence, additional P. myriotylum GH-specific primers were designed for identification of flanking sequences (Fig. 1b). Downstream 3′ region of the partial P. myriotylum GH was identified by genome walker strategy by screening the P. myriotylum genomic DNA libraries using GH-specific primers and adaptor primers. Primary PCR of EcoRV library for identification of 3′ flanking sequences yielded an intense amplicon of size 0.9 kb besides additional products of 0.3-, 1.1-, and 2-kb sizes (Fig. 1c). Secondary PCR reactions using P. myriotylum GH-specific and nested primers GH53R1 and GH53R1N on the primary PCR products yielded single conspicuous amplicons of approximately 0.9-kb size (Fig. 1d). The resulting amplicon was eluted, cloned and sequenced. A minimum of five to six clones were randomly selected, sequenced, and subjected to homology searches. BLASTX searches revealed conserved domains of GH superfamily and significant homology to GHs cloned mainly from Phytophthora spp. with identity ranging from 44 % (e-value, 2e-70) from Oxytricha trifallax to 68 % (e-value, 1e-180) from Phytophthora infestans. The full-length P. myriotylum GH sequence designated as PmGH1 was identified by multiple sequence alignment of the 5′ and 3′ clones and amplified using specific primers.
PCR amplification of PmGH1. a Amplification products obtained by PCR with degenerate primers. b Schematic diagram showing the locations of primers used for genome walker PCR; identification of 3′ flanking region by c primary PCR using gene-specific primer, GH53R1, and d secondary PCR products obtained with nested primer, GH53R1N
Sequence and Phylogenetic Analysis of PmGH1
The isolated PmGH1 ORF encompassed 1230 bp, without any introns, and the conceptual PmGH1 protein comprising of 421 amino acids had an apparent molecular weight of 46.77 kDa and a theoretical pI of 8.11. Multiple alignment of deduced amino acid sequence of PmGH1 and homologous GHs from other taxa revealed the presence of conserved residues (Fig. 2). Sequence comparisons revealed that PmGH1 shared conserved amino acid residues with family 1 GHs and they all shared the conserved catalytic regions, namely, the NEP and ENG domains. Furthermore, Glu 181 and Glu 399, located in the conserved catalytic regions, were identified as residues that constitute the active site (Fig. 2; marked by arrowheads). Signal peptide prediction revealed lack of putative signal peptide sequences.
Multiple amino acid sequence alignment of PmGH1 with homologous glucanases (GH1). Dark-shaded regions represent highly conserved residues. The conserved regions for the catalytic centre in family 1 glycoside hydrolase are boxed, and the predicted GH1 active site residues (general acid/ base and nucleophile residue) are marked by an arrowhead. GH1 sequences from the following species were used for generating the alignment: Albugo laibachii (CCA20215), Aspergillus niger (CAK47813), Trichoderma reesei (AAP57758), and Fusarium fujikuroi (CCT66182)
Phylogenetic analysis was carried out to examine the relationships of PmGH1 to GHs listed in GH families 1, 2, 5, 6, 7, 9, 12, 44, and 48 from CAZy database (Fig. 3). The analysis clustered the GH families into distinct clades with PmGH1 clustering with sequences of GH family 1 clade (Fig. 3).
Phylogenetic analysis based on CLUSTALX analysis of PmGH1 and glucanases of GH families 5, 6, 7, 9, 12, 45, 44, and 48. Species of origin, accession number (NCBI), and GH family to which the sequences belong are given at the end of each node. Bootstrap values greater than 700/1000 and scale of genetic distance as computed from pairwise distance in ClustalX are indicated
Molecular Modeling of PmGH1
Phyre threading program revealed the X-ray crystallographic structure of wheat beta-glucosidase c2dgaA (pdb accession code: 2DGA) sharing 30 % sequence identity and coverage of 92 % to PmGH1, as the best template for homology modeling. Structural coordinates generated for the 3D model were visualized using PyMOL. Structure of PmGH1 showed the typical (α/β)8 barrel folding of family 1 GHs (Fig. 4a). Ramachandran plot generated to evaluate quality of 3D model revealed that the main chain conformations for 90.4 % of the amino acid residues were within the favored regions, 5.4 % in allowed regions, and 4.2 % in the outlier region (Fig. 4b). Furthermore PROSA analysis [33] revealed a favorable Z-score of −7.09, which is a good value for a protein of this size. Predicted PmGH1 3D structure revealed binding sites for NAG with the involvement of Glu181 and Glu399 residues in hydrogen bonding with the docked ligand (Fig. 4c). The calculated distance between Glu181 and Glu399 pair is 3 Å, which is well within the average distance of 5 Å that is required between catalytic residues for glucanase activity [34]. Calculation of electrostatic potential of the docked PmGH1 model revealed the active site to be acidic (Fig. 4d).
Predicted tertiary structure of PmGH1. a 3D structure of PmGH1 predicted using Phyre2; chains are colored in chainbows. The conserved Glu181 and Glu399 residues in the catalytic domain are represented as white sticks. b Ramachandran plot generated for the PmGH1 3D structure with red-shaded regions indicating the most favored regions; yellows indicate additional allowed regions and white areas are disallowed regions. c The 3DLigandSite server was used to identify predicted ligand-binding sites in the PmGH1 with the ligand NAG docked in PmGH1 catalytic groove and hydrogen bonded to the Glu181 and Glu399 residues. d The surface contour based on electrostatic potential with the substrate binding pocket shown in green patches
Discussion
Among the oomycetes, Pythium spp. with necrotrophic lifestyle are potent fungus-like organisms known to secrete GHs early during colonization and in many cases even precede it [18, 35]. GHs from P. myriotylum have not been cloned so far, and the present work is the first attempt toward cloning and sequence characterization of GHs from this understudied necrotrophic phytopathogen. PCR-based approaches have been previously reported to be selective and useful for identifying novel homologous fungal cellulases [36, 37]. The identified GH designated as PmGH1 that exhibited homology and shared conserved active site domains to the protein domains of family 1 GH was observed to lack putative signal peptide sequence suggesting it to be a cytosolic enzyme. As observed for various proteins with glucanase activity, conserved domains of the predicted PmGH1 protein contained glutamate residues that function as the proton donor and nucleophile pair in the catalytic center [38, 39]. From the tree topology observed following a detailed phylogenetic analysis with sequences of various GH families, it is clear that PmGH1 clustered closely with sequences of GH family 1.
SOPMA analysis indicated that the predicted secondary structure of predicted PmGH1 protein contained 34 % α-helices, 12 % β-strands, and 6 % random coils. To assess the function of PmGH1 protein, modeling tools and protein structure analysis were carried out. The predicted 3D structure closely resembled the template structure with the (α/β)8 barrel topology, characteristic of family 1 GHs. The distance of 3 Å between the catalytic Glu181-Glu399 pair observed for PmGH1 is well within the average distance of 5 Å that is a prerequisite for cleavage of glycosidic bond [34, 40]. NAG binding sites were predicted in the PmGH1 3D model. N-acetyl-β-D-glucosaminidase (NAGase) activity has been reported in members of GH families 3, 18, 20, 73, and 85 [41–43]. GH1 comprises enzymes with various activities that include β-glucosidase (EC 3.2.1.21), β-galactosidase (EC 3.2.1.23), β-mannosidase (EC 3.2.1.25), β-glucuronidase (EC 3.2.1.31), β-fucosidase (EC 3.2.1.38), 6-phospho-β-galactosidase (EC 3.2.1.85), and 6-phospho-β-glucosidase (EC3.2.1.86) (www.cazy.org/Glycoside-Hydrolases.html). Our results predicting NAG binding sites indicate NAGase activity for the predicted family 1 PmGH1 protein.
The present study that reports for the first time cloning of family 1 GH from P. myriotylum, a necrotrophic oomycete, is of importance considering the conservation of critical amino acid residues of PmGH1 with family 1 GHs of reported activity. Observations made in the present study support the likelihood that the predicted cytosolic PmGH1 enzyme is a typical β-glucosidase and, hence, from an economic perspective, is a potential enzyme source for hydrolyzing cellulosic biomass.
References
McCarter, S. M., & Littrell, R. H. (1970). Comparative pathogenicity of Pythium aphanidermatum and Pythium myriotylum to twelve plant species and intraspecific variation in virulence. Phytopathology, 60, 264–268.
Hardham, A. R. (2007). Cell biology of plant-oomycete interactions. Cellular Microbiology, 9(1), 31–39.
Drechsler, C. (1930). Some new species of Pythium. Journal of the Washington Academy of Sciences, 20(16), 398–418.
Levesque, C. A., & De Cock, A. (2004). Molecular phylogeny and taxonomy of the genus Pythium. Mycological Research, 108, 1363–1383.
Agrios, G. N. (2005). Plant pathology (5th ed., pp. 410–413). Burlington: Elsevier Academic Press.
Tomioka, K., Takehara, T., Osaki, H., Sekiguchi, H., Nomiyama, K., & Kageyama, K. (2013). Damping-off of soybean caused by Pythium myriotylum in Japan. Journal of General Plant Pathology, 79, 162–164.
Endo, R. M., & Colt, W. M. (1974). Anatomy, cytology and physiology of infection by Pythium. Proceedings of the American Phytopathological Society, 1, 215–223.
Campion, C., Massiot, P., & Rouxel, F. (1997). Aggressiveness and production of cell-wall degrading enzymes by Pythium violae, Pythium sulcatum and Pythium ultimum, responsible for cavity spot on carrots. European Journal of Plant Pathology, 103, 725–735.
Liepman, A. H., Wightman, R., Geshi, N., Turner, S. R., & Scheller, H. V. (2010). Arabidopsis—a powerful model system for plant cell wall research. Plant Journal, 61, 1107–1121.
Mueller, O., Kahmann, R., Aguilar, G., Trejo-Aguilar, B., Wu, A., & de Vries, R. P. (2008). The secretome of the maize pathogen Ustilago maydis. Fungal Genetics and Biology, 45(Suppl 1), S63–S70.
Tian, C., Beeson, W. T., Iavarone, A. T., Sun, J., Marletta, M. A., Cate, J. H., & Glass, N. L. (2009). Systems analysis of plant cell wall degradation by the model filamentous fungus Neurospora crassa. Proceedings of the National Academy of Sciences of the United States of America, 106, 22157–22162.
Pryce-Jones, E., Carver, T., & Gurr, S. J. (1999). The roles of cellulase enzymes and mechanical force in host penetration by Erisyphe graminis f.sp. hordei. Physiological and Molecular Plant Pathology, 55, 175–182.
Latijnhouwers, M., de Wit, P. J. G. M., & Govers, F. (2003). Oomycetes and fungi: similar weaponry to attack plants. Trends in Microbiology, 11(10), 462–469.
Levesque, C. A., Brouwer, H., Cano, L., Hamilton, J. P., Holt, C., Huitema, E., et al. (2010). Genome sequence of the necrotrophic plant pathogen Pythium ultimum reveals original pathogenicity mechanisms and effector repertoire. Genome Biology, 11(7), R73.
Ten Have, A., Tenberge, K.B., Benen, J.A.E., Tudzynski, P., Visser, J., & van Kan, J.A.L. (2002). The contribution of cell wall degrading enzymes to pathogenesis of fungal pathogens. In: The Mycota XI agricultural applications. Springer-Verlag, Heidelberg, pp 341–358.
Seidl, M. F., van den Ackerveken, G., Govers, F., & Snel, B. (2011). A domain-centric analysis of oomycete plant pathogen genomes reveals unique protein organization. Plant Physiology, 155(2), 628–644.
Adhikari, B. N., Hamilton, J. P., Zerillo, M. M., Tisserat, N., Lévesque, A., & Buell, C. R. (2013). Comparative genomics reveals insight into virulence strategies of plant pathogenic oomycetes. PLoS ONE, 8(10), e75072.
Geethu, C., Resna, A. K., & Aswati Nair, R. (2013). Characterization of major hydrolytic enzymes secreted by Pythium myriotylum, causative agent for soft rot disease. Antonie Van Leeuwenhoek, 104(5), 749–757.
Wood, T. M., & McCrae, S. I. (1977). Cellulase from Fusarium solani: purification and properties of the C1 component. Carbohydrate Research, 57, 117–133.
Ryu, D. D., & Mandels, M. (1980). Cellulases: biosynthesis and applications. Enzyme and Microbial Technology, 2, 91–102.
Bayer, E. A., Moraq, E., & Lamed, R. (1994). The cellulosome—a treasure trove for biotechnology. Trends in Biotechnology, 12, 379–386.
Cantarel, B. L., Coutinho, P. M., Rancurel, C., Bernard, T., Lombard, V., & Henrissat, B. (2009). The Carbohydrate-Active EnZymes database (CAZy): an expert resource for glycogenomics. Nucleic Acids Research, 37, D233–D238.
Henrissat, B. (1991). A classification of glycosyl hydrolases based on amino acid sequence similarity. Biochemical Journal, 280, 309–316.
Lombard, V., Golaconda Ramulu, H., Drula, E., Coutinho, P. M., & Henrissat, B. (2014). The carbohydrate-active enzymes database (CAZy) in 2013. Nucleic Acids Research, 42, D490–D495.
Zhang, Y.-P., & Lynd, L. R. (2004). Toward an aggregated understanding of enzymatic hydrolysis of cellulose: noncomplexed cellulase systems. Biotechnology and Bioengineering, 88, 797–824.
Zhang, Y.-P., Himmel, M. E., & Mielenz, J. R. (2006). Outlook for cellulase improvement: screening and selection strategies. Biotechnology Advances, 24, 452–481.
Moller, E. M., Bahnweg, G., Sandermann, H., & Geiger, H. H. (1992). A simple and efficient protocol for isolation of high molecular weight DNA from filamentous fungi, fruit bodies and infected plant tissues. Nucleic Acids Research, 20(22), 6115–6116.
Altschul, S. F., et al. (1997). Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Research, 25, 3389–3402.
Thompson, J., Gibson, T., Plewniak, F., Jeanmougin, F., & Higgins, D. (1997). The Clustalx windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Research, 25, 4876–4882.
Petersen, T. N., Brunak, S., von Heijne, G., & Nielsen, H. (2011). SignalP 4.0: discriminating signal peptides from transmembrane regions. Nature Methods, 8, 785–786.
Tamura, K., Dudley, J., Nei, M., & Kumar, S. (2007). MEGA4: molecular evolutionary genetics analysis (MEGA) software version 4.0. Molecular Biology and Evolution, 24(8), 1596–1599.
Wass, M. N., Kelley, L. A., & Sternberg, M. J. (2010). 3DLigandSite: predicting ligand-binding sites using similar structures. Nucleic Acids Research, 38, W469–W473.
Wiederstein, M., & Sippl, M. J. (2007). ProSA-web: interactive web service for the recognition of errors in three-dimensional structures of proteins. Nucleic Acids Research, 35, W407–W410.
Zechel, D. L., & Withers, S. G. (2001). Dissection of nucleophilic and acid-base catalysis in glycosidases. Current Opinion in Chemical Biology, 5(6), 643–649.
Marcus, L., Barash, I., Sneh, B., Koltin, Y., & Finkler, A. (1986). Purification and characterization of pectolytic enzymes produced by virulent and hypovirulent isolates of Rhizoctonia solani Kuhn. Physiological and Molecular Plant Pathology, 29, 325–336.
Jia, J., Dyer, P. S., Buswell, J. A., & Peberdy, J. F. (1999). Cloning the cbhI and cbhII genes involved in cellulose utilisation by the straw mushroom Volvariella volvacea. Molecular and General Genetics, 261, 985–993.
Sheppard, P. O., Grant, F. J., Oort, P. J., Sprecher, C. A., Foster, D. C., Hagen, F. S., Upshall, A., McKnight, G. L., & O’Hara, P. J. (1994). The use of conserved cellulase family-specific sequences to clone cellulose homologue cDNAs from Fusarium oxysporum. Gene, 150, 163–167.
Haiech, J., Chippaux, M., Barras, F., Py, B., & Bortoli-German, I. (1991). Cellulase EGZ of Erwinia chrysanthemi: structural organization and importance of His98 and Glu133 residues for catalysis. Protein Engineering, 4(3), 325–333.
Cui, C.-H., Kim, J.-K., Kim, S.-C., & Im, W.-T. (2014). Characterization of a ginsenoside-transforming β-glucosidase from Paenibacillus mucilaginosus and its application for enhanced production of minor ginsenoside F2. PLoS ONE, 9(1), e85727.
Ghosh, R., & Chakrabarti, C. (2008). Crystal structure analysis of NP24-I: a thaumatin-like protein. Planta, 228, 883–890.
Henrissat, B. (1998). Glycosidase families. Biochemical Society Transactions, 26, 153–156.
Stals, I., Karkehabadi, S., Kim, S., Ward, M., Van Landschoot, A., Devreese, B., & Sandgren, M. (2012). High resolution crystal structure of the endo-N-Acetyl-β-D-glucosaminidase responsible for the deglycosylation of Hypocrea jecorina cellulases. PLoS ONE, 7(7), e40854.
Tzelepis, G. D., Melin, P., Jensen, D. F., Stenlid, J., & Karlsson, M. (2012). Functional analysis of glycoside hydrolase family 18 and 20 genes in Neurospora crassa. Fungal Genetics and Biology, 49(9), 717–730.
Acknowledgments
The present research was supported by Faculty Research Grant (FRG) scheme (No. NITC/Dean(C&SR)/FRG10/0112) of NITC. PP thanks the Director of JNTBGRI, Kerala, India, for the research facilities extended.
Compliance with Ethical Standards
We hereby certify that the communicated manuscript is not submitted to any other journal for simultaneous consideration, nor been published previously (partly or in full). Furthermore, authors declare that they have no conflict of interest concerning this article. The investigations reported in the present manuscript do not involve any clinical studies engaging human participants or animals.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Nair, R.A., Geethu, C., Sangwan, A. et al. Sequence and Bioinformatic Analysis of Family 1 Glycoside Hydrolase (GH) 1 Gene from the Oomycete Pythium myriotylum Drechsler. Appl Biochem Biotechnol 176, 772–781 (2015). https://doi.org/10.1007/s12010-015-1610-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12010-015-1610-6