Abstract
We report a fully automated DNA purification platform with a micropored membrane in the channel utilizing centrifugal microfluidics on a lab-on-a-disc (LOD). The microfluidic flow in the LOD, into which the reagents are injected for DNA purification, is controlled by a single motor and laser burst valve. The sample and reagents pass successively through the micropored membrane in the channel when each laser burst valve is opened. The Coriolis effect is used by rotating the LOD bi-directionally to increase the purity of the DNA, thereby preventing the mixing of the waste and elution solutions. The total process from the lysed sample injection into the LOD to obtaining the purified DNA was finished within 7 min with only one manual step. The experimental result for Salmonella shows that the proposed microfluidic platform is comparable to the existing devices in terms of the purity and yield of DNA.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
There have been significant advances in the development of microfluidics for biological analysis, such as micro total analysis systems (μTAS), during the last few decades [1–5]. The lab-on-a-chip (LOC) as a microfluidic system has the potential to play an important role because it represents a fully integrated analytic process, including reagents preparation, mixing, metering, reaction, and detection [6, 7]. Compared to the LOC, the lab-on-a-disc (LOD) as a centrifugal-microfluidic platform provides further advantages. For example, the LOD requires only a single motor to generate the force required for fluid propulsion, thereby eliminating the need for an external pump [5, 8].
Recently, molecular diagnostics has been attracting considerable attention and is recognized as an important process for biological and biomedical research [9–12]. The DNA purification process, as the initial step for molecular diagnostics, is mostly conducted with chemical reaction methods [13, 14]. Although novel DNA purification methods using adsorptive silica beads [15], magnetic beads [16–18], and dielectrophoretic forces [19] have been reported, they are expensive and the materials are hard to handle. The spin column using a centrifuge is widely used as a chemical reaction method for DNA purification with bench top protocol because it is easy and simple and shows good performance [20, 21]. However, the spin column method involves the laborious routine steps of pipetting, vortexing, and centrifugation when each reagent is injected into the column. Also, each reagent which passes the membrane in the spin column should be removed from it for the subsequent process. Most technologies used in a spin column are suitable for LODs, in that both technologies use the same centrifugal force. Therefore, we studied the possibility of developing an automated LOD based on the protocol of the spin column to improve on this inconvenient process.
According to the NDFP (National Database of Food Poisoning), the most common causes of food poisoning are Salmonella spp., Staphylococcus aureus (S. aureus), Escherichia coli (E. coli) O157:H7, Campylobacter jejuni (C. jejuni), Shigella, and Clostridium botulinum (C. botulinum) bacteria. Among these pathogens, Salmonella spp., S. aureus, and E. coli O157:H7 have been recognized as the leading causes of severe illness. In this paper, Salmonella spp. were used to show the performance of our automated LOD, which allows for the automation of the whole process, from sample injection to DNA purification, with one manual step. For the automated system, the LOD is employed to channel where we use the Coriolis effect [22, 23] to prevent the mixing of the regents. Also, a laser burst valve is utilized for controlling the flow order of the reagents. The designed LOD enables the integration of a complicated biological reaction that requires complete multi-step fluidic control.
The purified DNA results using cultured Salmonella are demonstrated by measuring the purity and yield of DNA obtained by absorbance measurements and the polymerase chain reaction (PCR) product obtained by purified DNA amplification. The results show that our novel automated LOD is comparable to a commercial device for DNA purification.
Materials and Methods
LOD Fabrication
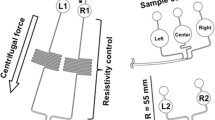
The conventional CD was modified to suit to our LOD system made from three polycarbonate (PC) layers through an injection molding machine (FA-DRA1F, Panasonic, Japan). These layers were combined using double-sided adhesive films (Tesa 4928, Germany; thickness, 0.125 mm), consisting of a transparent PET film backing and a modified acrylic adhesive, applied under pressure to finally construct channels and chambers for storing the DNA purification reagents. Figure 1 shows a schematic diagram of the proposed LOD system designed on a compact disc with a diameter of 120 mm, where the DNA purification reagents are contained in the chamber. The LOD was designed symmetrically for simultaneous experiments. The volume of the chambers was designed to be almost twice the volume of each reagent for the purpose of preventing the reagents from flowing backwards when they are injected. A micropored membrane (MM) was inserted between the PC structures by fixing it on the channel before the Coriolis hole, allowing all of the fluids to pass through the MM. The size of the MM (GF/B glass microfibre filters, Whatman, UK) was adjusted to vertical 4 mm × horizontal 28 mm × height 1 mm, having the same volume as the silica membrane of the spin column.
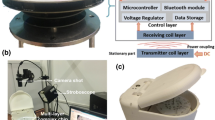
The disc-based system device for DNA purification. a Exploded view of the assembled lab-on-a-disc. b Schematics of the construction of the LOD. c Schematics of the whole LOD system which can be rotated bi-directionally as necessary. The ratio a:b of the channel widths; d photograph of the assembled prototype with LOD for the DNA purification system
Coriolis Hole
The Coriolis effect is generally more sensitive when the channel used for fluid separation is made longer according to Eq. (1) [22, 23]:
where ρ is the mass density of the fluid, \( \overset{\rightharpoonup }{\omega } \) is the angular velocity vector, and \( \overset{\rightharpoonup }{u} \) is the velocity of the fluid in the rotating LOD. The Coriolis effect is proportional to \( \overset{\rightharpoonup }{u} \), thus requiring a sufficient acceleration length of the channel to reach the velocity for generating the Coriolis effect reliably. However, there is a limitation for obtaining a sufficient length of Coriolis channel for sensitive operation, due to the integrated channels and chambers used on our LOD. To increase the sensitivity of the Coriolis effect, we designed the hole structure on the channel so as to allow for the separation of the reagents from the elution solution that passes through the MM. Figure 1c shows a diagram of the Coriolis hole. The Coriolis hole was made by drilling a hole with a diameter of 1 mm in the middle layer. The previous length of the Coriolis channel was 3 mm, but the Coriolis hole can make the channel longer to double the length 6 mm.
Laser Burst Valve
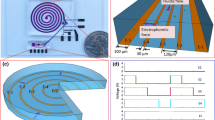
In this experiment, a laser burst valve [24] was employed to control the flow order of the reagents. The valve depends on the interplay of a laser beam with a polyethylene substrate. The valve is opened by optical energy from a laser diode (2 V, 1 W, 808 nm, QSI, Korea), melting the black color polyethylene substrate, whose melting point is typically 115∼135 °C. Since the transparent PC transmits the laser light in the near-infrared region of the spectrum, the laser beam can pass through the PC for the purpose of melting the black-colored polyethylene substrate, which absorbs the light energy. The distance between the laser diode and polyethylene is around 2 mm. By rotating the LOD slightly when the laser diode melts the substrate, the opening for spouting the fluid is made. The process of valve-opening is shown in Fig. 2 where the black polyethylene is torn while rotating the LOD with the diode on [24].
Schematic diagram showing the opening of the valve by the laser burst valve. Step (i): laser diode focused on polyethylene to optimize the thermal energy required to melt the material. Step (ii): widening of the valve opening by rotating the LOD in stepwise fashion with the diode on. Fluid can flow through the channel through the opened valve when the LOD is spinning
Reagents and Operation
We used a lysed cell which contained cultured Salmonella (OD600 = 0.8, 108 CFU/ml) as a sample for DNA purification. The lysis process was conducted on a clean bench. The sample and reagents (Genomic DNA extraction kit, Bioneer, Korea) were injected into the chambers in the LOD before the experimental setup for DNA purification. The volumes and mechanisms of the reagents followed the manual of the purchased kit. The reagents were controlled by the centrifugal force and laser burst valve, passing through the MM step by step. The process of DNA purification using the LOD is shown in Table 1. The DNA is negatively charged, because of the phosphate backbone and, thus, it binds to the MM, which is made from silica material by the cationic exchange method, when the sample passes through it. The W1 and W2 washed out the impurities on the MM except for DNA in order. Then, the DNA was eluted from the MM by the elution solution using the hydrolysis reaction. By spinning the LOD in different directions, the reagents are separated into the waste chamber and elution solution by the Coriolis effect to increase the purity of the DNA.
The mechanical system was composed of a motor (QSI, Korea) to generate a centrifugal force, laser module for the laser burst valve, and computer for motor control. From sample injection to injection into each chamber, all of the steps, ex. motor rpm acceleration, stop and valve open, were finished automatically by the program within 7 min. After the LOD has stopped spinning, the elution solution including the DNA is extracted from the DNA chamber using a syringe. The absorbance (A) of the DNA solution was measured at 260, 280, and 320 nm using a spectrometer (Multiskan GO, Thermo, Korea) to calculate the yield and purity of the DNA. The yield of DNA was calculated by Eqs. (2) and (3):
The total yield is obtained by multiplying the DNA concentration by the final total purified sample volume.
The purity of the DNA was calculated by Eq. (4):
The wavelength of maximum absorption for DNA is 260 nm, with each base (A, C, G, T) having its own individual characteristic value. Also, the amino acid sequence of the proteins absorbs light at 280 nm. The non-nucleic acids absorb at A 320 (nm). Therefore, the purity ratio can be calculated after subtracting the value of A 320 (nm) from the absorbances at 260 and 280 nm.
Experimental Results and Discussion
Coriolis Effect
In this experiment, the Coriolis effect in the LOD plays an important role in preventing the reagents from mixing with the purified DNA solution because the mixing of the purified DNA with the ethanol included in the reagents can affect the DNA amplification result. The LOD is turned counterclockwise (CCW) when the sample, W1 and W2 pass through the MM. On the other hand, the LOD is turned clockwise (CW) when the elution solution passes through the MM for the purpose of separating the reagents by means of the Coriolis effect. To reliably prevent the reagents from flowing toward the DNA chamber and only allow the elution solution to pass, we made use of the Coriolis effect by regulating the Coriolis channel-width ratio (a:b = toward waste chamber:toward DNA chamber). We measured the absorbance (at 635 nm) of the fluid in the chamber where it has injected with DIW for mixing with the leaked red-colored ink caused by the Coriolis effect according to the channel-width ratio. Based on the results in Fig. 3, we chose a channel-width ratio of 1.5:1. Even though there is some loss of volume caused by the narrowing of the width of the channel toward the DNA chamber, it was very small (≤7 μl on 1.5: 1), which is extra volume for requiring the volume (175 μl) of the measurement by the spectrometer.
Laser Burst Valve
The laser burst valve enabled the flow order of the fluids to be controlled in the LOD. The linked block polyethylene on the channel was melted by the optical energy from the laser diode to valve-open. The result of the melted valve is shown in Fig. 4. When the block polyethylene is used as a valve, no leaking occurs during the high rpm spin (10,000 rpm) for 15 min. Whenever the polyethylene is melted, the fluid escapes from the chamber. By using this method, the reagents pass through the MM, thus allowing the process to be automated.
DNA Purification
The combination of the MM and DNA was used as a chaotropic agent method for the purification of Salmonella DNA. To evaluate the performance of the LOD system, we measured the purity and yield of DNA. The experimental results of DNA purification are shown in Fig. 5. The calculated average yield of DNA from the LOD and spin column were 22.3 and 18.7 μg, respectively. Also, the DNA purity obtained using the automated LOD is 1.77 with an average coefficient of variation (CV) of 2.8 %, while that of the commercialized device was 1.88 with an average CV of 2.6 %. Even though the LOD system has a lower purity value than the spin column, it provides not only an automated system which can be accomplished with one manual step but also a comparable result value in terms of the yield and purity under laboratory conditions.
PCR Result
We amplified the purified DNA samples obtained using the LOD and spin column by PCR to confirm the performance of DNA purification. The PCR reagent was purchased from NobleZyme (E-220, Lot. No. E220S1402). The PCR process was done with a thermal cycling profile consisting of 30 cycles: initial denaturation at 95 °C for 2 min, 30 cycles at 95 °C for 45 s, at 52 °C for 45 s, 72 °C for 60 s, and a final extension at 72 °C for 5 min. The PCR products were loaded in a 2 % agarose gel (Bio solution, catalog no. BT002, Korea) prepared with 1× TAE buffer for gel electrophoresis at 100 V for 30 min. The experimental results are shown in Fig. 6. The PCR product results of the purified DNA obtained using the LOD demonstrate that the functions in the LOD performed normally for the purification of the DNA.
Gel electrophoresis image of the Salmonella DNA amplified by PCR. #1 and #2 are the amplified results from the diluted DNA concentration (by 10 times) with DIW to compare the limits of detection. Lane 1: PCR-positive control; lane 2: PCR-negative control; lanes 3–8: PCR products from Salmonella DNA purified by the LOD; lanes 9–15: PCR products from Salmonella DNA purified by spin column
Conclusion
The proposed LOD allows all of the steps for a DNA purification system to be automated using a laser burst valve, the Coriolis effect, and centrifugal force. The employed Coriolis effect system has the advantage of preventing the mixing of the purified DNA with the reagents, which might affect the DNA amplification result. The laser burst valve worked properly, controlling the fluid flow. The experimental results show that the purified DNA is obtained by the automated LOD system within 7 min with only one manual step. The proposed LOD is not only comparable to a commercialized device for obtaining purified DNA but also able to overcome the inconvenience of the spin column system with its laborious routine steps, by automating all of the steps of the DNA purification system.
References
Kwang-Seok, Y., & Euisik, Y. (2004). Microfluidic components and bio-reactors for miniaturized bio-chip applications. Biotechnology and Bioprocess Engineering, 9, 86–92.
Manz, A., Graber, N., & Widmer, H. M. (1990). Miniaturized total chemical analysis systems: a novel concept for chemical sensing. Sensors and Actuators B, 1, 244–248.
Lee, B. S., Lee, Y. U., Kim, H. S., Kim, T. H., Park, J., Lee, J. G., Kim, J. T., Kim, H. S., Lee, W. G., & Cho, Y. K. (2011). Fully integrated lab-on-a-disc for simultaneous analysis of biochemistry and immunoassay from whole blood. Lab on a Chip, 11, 70–78.
Duffy, D. C., Gillis, H. L., Lin, J., Sheppard, N. F., Jr., & Kellogg, G. J. (1999). Microfabricated centrifugal microfluidic systems: characterization and multiple enzymatic assays. Analytical Chemistry, 71, 4669–4678.
Nolte, D. D. (2009). Invited review article: review of centrifugal microfluidic and bio-optical disks. Review of Scientific Instruments, 80, 101101.
Srinivasan, V., Pamula, V. K., & Fair, R. B. (2004). An integrated digital microfluidic lab-on-a-chip for clinical diagnostics on human physiological fluids. Lab on a Chip, 4, 310–315.
Gaeberle, S., & Zengerle, R. (2007). Microfluidic platforms for lab-on-a-chip applications. Lab on a Chip, 7, 1094–1110.
Madou, M., Zoval, J., Jia, G., Kido, H., Kim, J., & Kim, N. (2006). Lab on a CD. Annual Review of Biomedical Engineering, 8, 601–628.
Liu, P., Li, X., Greenspoon, S. A., Scherer, J. R., & Mathies, R. A. (2011). Integrated DNA purification, PCR, sample cleanup, and capillary electrophoresis microchip for forensic human identification. Lab on a Chip, 11, 1041–1048.
Breadmore, M. C., Wolfe, K. A., Arcibal, I. G., Leung, W. K., Dickson, D., Giordano, B. C., Power, M. E., Ferrance, J. P., Feldman, S. H., Norris, P. M., & Landers, J. P. (2003). Microchip-based purification of DNA from biological samples. Analytical Chemistry, 75, 1880–1886.
Horn, N. A., Meek, J. A., Budahazi, G., & Marquet, M. (1995). Cancer gene therapy using plasmid DNA: purification of DNA for human clinical trials. Human Gene Therapy, 6, 565–573.
Stadler, J., Lemmens, R., & Nyhammar, T. (2004). Plasmid DNA purification. The Journal of Gene Medicine, 6, S54–S66.
Smrekar, V., Smrekar, F., Strancar, A., & Podgornik, A. (2013). Single step plasmid DNA purification using methacrylate monolith bearing combination of ion-exchange and hydrophobic groups. Journal of Chromatography A, 1276, 56–64.
Costioli, M. D., Fisch, I., Garret-Flaudy, F., Hilbrig, F., & Freitag, T. (2003). DNA purification by triple-helix affinity precipitation. Biotechnology and Bioengineering, 5, 535–545.
Cho, Y. K., Lee, J. G., Park, J. M., Lee, B. S., Lee, Y., & Ko, C. (2007). One-step pathogen specific DNA extraction from whole blood on a centrifugal microfluidic device. Lab on a Chip, 7, 565–573.
Yi-Lien, K., Liu, C. J., Lin, Y. C., Kuo, P. L., & Lee, G. B. (2009). Extraction of genomic DNA and detection of single nucleotide polymorphism genotyping utilizing an integrated magnetic bead-based microfluidic platform. Microfluidics and Nanofluidics, 6, 539–555.
Zhang, H. P., Bai, S., Liang, X., & Sun, Y. (2009). Fabrication of mono-sized magnetic anion exchange beads for plasmid DNA purification. Journal of Chromatography B, 877, 127–133.
Prinz, C., Tegenfeldt, J. O., Austin, R. H., Cox, E. C., & Sturm, J. C. (2002). Bacterial chromosome extraction and isolation. Lab on a Chip, 2, 207–212.
Sonnenberg, A., Marclnlak, J. Y., Krlshnan, R., & Heler, M. J. (2012). Dielectrophoretic isolation of DNA and nanoparticles from blood. Electrophoresis, 33, 2482–2490.
Yang, D. Y., Eng, B., Waye, J. S., Dudar, J. C., & Saunders, S. R. (1998). Technical note: improved DNA extraction from ancient bones using silica-based spin columns. American Journal of Physical Anthropology, 105, 539–543.
Tan, S.C., and Yiap, B.C. (2009). DNA, RNA, and protein extraction: the past and the present. Journal of Biomedicine and Biotechnology, 1–10.
Brenner, T., Glatzel, T., Zengerle, T., and Ducree J. (2003). A flow switch based on Coriolis force. 7th International Conference on Miniaturized Chemical and Biochemical Analysis System, Squaw valley, 903–906.
Brenner, T., Glatzel, T., Zengerle, T., & Ducree, J. (2005). Frequency-dependent transversal flow control in centrifugal microfluidics. Lab on a Chip, 5, 146–150.
Yoo J.C., and Na, W.H. (2014). A thin film valve apparatus using a hole closing membrane, Korean Patent Number: 10-1439483, Publication date: Sep. 2. 2014.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Choi, MS., Yoo, JC. Automated Centrifugal-Microfluidic Platform for DNA Purification Using Laser Burst Valve and Coriolis Effect. Appl Biochem Biotechnol 175, 3778–3787 (2015). https://doi.org/10.1007/s12010-015-1546-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12010-015-1546-x