Abstract
Ethiopia is the center of origin and diversity for Arabica coffee with high morphological diversity between accessions as compared to commercial cultivars. Coffee germplasm collection and molecular characterization are crucial steps towards its conservation, breeding, and development of superior genotypes for various end uses. Hence, this study was initiated with the objective of studying the genetic diversity of Coffea arabica accessions collected from different regions of Ethiopia, using SSR markers. A total of 20 SSR markers were used to genotype 86 accessions and produced a total of 112 alleles, with an average of 5.6 alleles per locus. All the loci across the entire populations were found to be highly polymorphic with a mean of 0.6 PIC value. Average observed heterozygosity and allelic richness across all populations ranged from 0.22 to 0.27 and 3.52–4.26, with a mean of 2.43 and 3.97, respectively. AMOVA showed high variation within population based on geographical origin. The smaller Fst (0.037) observed indicates the presence of lower population genetic differentiation as a result of higher gene flow (Nm = 2.45) across populations and the lowest mean genetic distance (0.21) observed between populations. The UPGMA, PCoA, and structure analysis poorly grouped the individuals into distinct clusters indicating the presence of population admixture. The observed higher genetic variability in all populations indicates that the country has huge coffee genetic diversity which can be used for future coffee improvement. Our results revealed an unexploited highly diverse genetic resource particularly from Omo, Ilubabor, and Benchi Maji that should be considered in future coffee breeding program and germplasm conservation.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Coffee is one of the most valuable commodity crops in the world trade. More than 90% of its production comes from developing countries, providing an income for millions of smallholder farmers around the world that are dependent on coffee for their subsistence (Tran et al. 2016). It contributes largely to the economy of more than 50 countries in Asia, Latin America, and Africa (Davis et al. 2011). The share of coffee in total export earnings has a positive and significant reflection on economic growth for developing countries (Al-Abdulkader et al. 2018).
Since Ethiopia is the origin and the primary center of diversity for Coffea arabica, its production in Ethiopia has a long tradition which dates back to dozens of centuries. The country possesses a diverse genetic base for Arabica coffee and its wild relatives with considerable heterogeneity (Gebreselassie et al. 2018). The coffee grown in the country showed wide genetic variations within and between populations of different regions for yield, quality disease resistance and other traits. The availability of such genetic variations provides immense possibilities for improvement of the crop for any desirable traits of interest (Benti 2017; da Silva et al. 2019). Coffee is by far Ethiopia’s most important export crop which accounts the lion’s share of the foreign exchange earnings of the country. Furthermore, the livelihood of some 15 million people directly or indirectly depends on coffee. Ethiopia accounts for around 3% of the global coffee market and around 35% of foreign income comes from coffee (Benti 2017).
The quality standards of Ethiopian coffee are classified according to their origin of production. Among the best-known coffee varieties in Ethiopia are Yirgachefe, Harar, Wolegga, Limu, Jimma, Sidama, and others take the priority. Within Ethiopia, there are some distinctive varieties that are highly sought after. The majority of cultivated varieties in Ethiopia comes from Harar, where the Long berry variety is the most popular, having a wine-like flavor and tasting slightly acidic. Another Ethiopian coffee from Yirgachefe is the highest premium coffee in Ethiopia as well in the world (Gebreselassie et al. 2018). However, presently Ethiopian coffee genetic resource is under greatest threat, mostly due to deforestation, replacement of farmer’s variety by a few high yielding and disease-resistant varieties, establishment and expansion of modern plantation and illegal and legal settlements, drought, the spread, and escalating severity of devastating fungal pathogens (da Silva et al. 2019).
Coffee breeding programs invested intense efforts to release cultivars with high productivity, with climate change and simultaneous change in biotic and abiotic pressures stress tolerance and high biochemical quality of the beans (Tran et al. 2016). However, several factors are limiting the genetic gains in breeding programs (Vieira et al. 2010). Commercial coffee varieties are originated from a limited number of cultivars; consequently, only a narrow genetic base is available to support breeding programs. Susceptibility of commercial coffee cultivars to pests and diseases, climate change, and emerging pest and outbreak of diseases are important challenges to global coffee production (Bunn et al. 2015).
Many studies reported wide phenotypic diversity of Arabica coffee accessions collected from Ethiopia regarding leaf size, height, biotic and abiotic stress tolerance, and yield (Tran et al. 2016) and the presence of higher genetic variability compared with cultivars, demonstrating the potential of these accessions for breeding purposes (Aerts et al. 2013; Sant’Ana et al. 2018). The accessions also showed a great variability for chlorogenic acids, lipids, sucrose, and diterpenes contents of coffee beans and the discovery of decaffeinated coffee varieties (Silvarola et al. 2004) which underlines the great potential existing in Ethiopian coffee accessions for Arabica coffee future breeding programs.
A number of DNA-based techniques were used in different coffee genetic studies. These include RAPD (Diniz et al. 2005), AFLP (Steiger et al. 2002), ISSR (Tesfaye et al. 2014), and microsatellite (SSR) markers (Lashermes et al. 1995; Teressa et al. 2010; Geleta et al. 2012; Motta et al. 2014; Sousa et al. 2017; da Silva et al. 2019). Among the several types of molecular markers, microsatellites are most commonly used in genetic diversity study because of several advantages, including a high degree of polymorphism, repeatability, reproducibility, codominance, technical simplicity, speed, and multi allelism (Vieira et al. 2010).
Most of the studies, so far using these technologies, were done on commercial cultivars of Arabica coffee. Because of this, little is known about the genetic structure and a pattern of Arabica coffee accessions. However, the genetic diversity of a few number of coffee accession in Ethiopia has been studied (Teressa et al. 2010). The works have shown that there is sufficient diversity in Ethiopia, especially the South-Western part of the country. However, each coffee producing region has its own unique trait which should be studied independently and can be used for future breeding efforts. Therefore, analyzing the genetic diversity using different marker systems is vital to minimize the loss of coffee genetic resources and the best strategy for conservation and sustainable utilization for future breeding efforts. Hence, in this study, we analyze the genetic diversity among and within populations of Coffea arabica accessions collected from a range of eco-geographical zones of Ethiopia.
Materials and methods
Plant materials
In this study, a total of 86 Coffea arabica accessions were used. The accessions were obtained from Ethiopian Biodiversity Institute (EBI) and they were collected from different agro-ecologies varying in altitude, soil type, rainfall, temperature, and represent the major coffee growing regions of Ethiopia. The list of accessions used is given in Table 1.
DNA extraction
The entire experiment was conducted at the National Agricultural Biotechnology Research Center (NABRC), located at Holeta, 29 km west of Addis Ababa. Young leaf samples that are free from the disease were collected from the actively growing tips of the coffee branches and preserved and dried using silica gel (Blulux laboratory (p) Ltd.-121,005). The leaves were transported to NABRC for DNA extraction. Genomic DNA was extracted from dried leaf samples according to Khanuja et al. (1999) with some modifications. About 0.5 g of dried leaf tissue was placed in 2-ml Eppendorf tube and ground using a geno-grinder (Retsch MM200, Germany). Nearly 1 ml of freshly prepared extraction buffer (100 mM Tris–Cl (pH 8.0), 25 mM EDTA, 1.5 M NaCl, 2.5% CTAB, 0.2% β-mercaptoethanol (v/v), and 1% PVP (w/v) was added to the grinded material and mixed inversely. The mixed solution was incubated at 60 °C in a shaking water bath (100 rpm) for 2 h followed by addition of 1 ml chloroform: isoamyl alcohol (24:1). The solution was mixed by inverting the tubes for about 15 min and then centrifuged at 8000 rpm for 10 min at 25–30 °C. Carefully, the upper clear aqueous layer was transferred to another 2-ml Eppendorf tube and properly mixed with 0.75 ml of 5 M NaCl. About 0.6 volume of isopropanol was added and placed at room temperature for 1 h. After 1 h, the mixture was centrifuged at 10,000 rpm for 10 min at 25–30 °C and the supernatant was discarded and the pellet washed with 80% ethanol. The pellet was dried in a vacuum for 15 min and dissolved in 0.5 ml of high salt TE buffer (1 M NaCl, 10 mM Tris–Cl (pH 8.0) and 1 mM EDTA). After the pellet was dissolved in high salt TE buffer, 2 µl of RNase was added and incubated at 37 °C for 30 min to degrade RNA. An equal volume of chloroform: isoamyl alcohol (24:1) was added and the aqueous layer was transferred to fresh 1.5 ml microfuge tube and 2 volumes of cold ethanol was added to precipitate the DNA. The mixture was centrifuged at 10,000 rpm for 10 min at 25–30 °C and the pellet was washed with 80% ethanol. Finally, the pellet was dried in a vacuum and dissolved in 100 µl of nuclease free water. The genomic DNA concentration was estimated at 260 and 280 nm using the Nanodrop (Nano Drop® ND-800). Similarly, the quality of DNA was checked using 0.8% agarose gel by loading 5 µl genomic DNA after mixing with 2 µL 6X loading dye with gel red. The band of DNA was visualized using gel documentation (Bio-Doc-It™ imaging system) under UV light.
SSR markers used
Initially 24 SSR markers previously described as being polymorphic in C. arabica were selected from published literature (Rovelli et al. 2000; Combes et al. 2000; Moncada and McCouch 2004; Cubry et al. 2008). The annealing temperatures of the primers were optimized by performing gradient PCR. Finally, 20 polymorphic SSR markers were selected by excluding four markers that failed to amplify properly. Detail of the SSR markers used for this study is given in Table 2.
PCR amplification
PCR was conducted in 12.5 μl reaction volume containing: One Taq® 2X Master Mix with Standard Buffer, 10 µM of forward primers, 10 µM of reverse primers, Nuclease free water, and 20 ng/µl genomic DNA template. DNA free 10.5 µl reaction volume was used as a control. The PCR Conducted in Bio-Rad T100™ thermal cycler.
Gel electrophoresis
The amplified PCR products were separated using 3.5% (w/v) agarose gel by loading 10 μl of each of the PCR product mixed with 2 µl of loading dye and gel red. The gels were run in 1X TAE (40 mM Tris, 40 mM Acetate, 1 mM EDTA, pH 8) buffer and at 85 voltages, 100 mA for 3.5 h. Finally, the amplified bands were documented under UV light in gel documentation.
Data scoring and analysis
The amplified SSR fragment sizes on the gel were estimated using PyElph 1.4 software package. Locus based diversity indices including major allele frequency (MAF), number of alleles (Na), gene diversity; polymorphic information contents (PIC) were computed using Power Marker v3.25 software (Liu and Muse 2005). The effective number of alleles, Shannon's Information index (I), and Gene flow (Nm) were calculated using POPGENE version 1.31 (Yeh and Yang 1999). Allelic frequency, observed heterozygosity (Ho), expected heterozygosity (He), fixation index (F), and the estimate of the deviation from Hardy–Weinberg equilibrium (HWE) over the entire populations were computed with GenAlEx version 6.501 software (White and peakall 2015). The same software package was used to compute population differentiation test: Wrights fixation indices and pairwise population differentiation (Fst), analysis of molecular variance (AMOVA) and the estimate of the variance components. Moreover, rarified allelic richness (Ar) and private rarified allelic richness (Arp) were computed using HP-Rare 1.1 software (Kalinowski 2005). Genetic relationships among the different populations and genetic dissimilarity matrix were calculated using Jaccard’s coefficient and Unweighted Pair Group Method with Arithmetic Mean (UPGMA) clustering was generated using DARwin var. 6.0.14 (Perrier and Jacquemoud-Collet 2006). Principal coordinate analysis (PCoA) was performed using the GenALEX to produce Eigen values and Eigen vectors which reveal both total variance and the loci that are important to the variance. Biplots were generated to reveal associations among accessions. Population structure and admixture patterns were determined using STRUCTURE software ver. 2.3.4 based on Bayesian algorithm (Pritchard et al. 2000). To estimate the true number of population cluster (K), a burn-in period of 50,000 was used in each run, and data were collected over 500,000 Markov Chain Monte Carlo (MCMC) replications for K = 1 to K = 10 using 20 iterations for each K. The optimum K value was predicted following the simulation method of Evanno et al. (2005) using the web-based STRUCTURE HARVESTER ver. 0.6.92 (Earl and vonHoldt 2012). Bar plot for the optimum K was determined using Clumpak beta version (Kopelman et al. 2015).
Results and discussion
Polymorphism and allelic diversity among SSR markers
A total of 112 SSR alleles were produced across 20 loci, with the estimated product size range of 100 to 385 base pairs (Table 3). The mean number of alleles per locus was 5.6. The detected number of alleles ranged from 3 (AJ250253 and CFGA69) to 11 (CFGA465). This result is higher than that of a previous study by Anthony et al. (2002) who reported an average of 4.7 alleles per SSR using six SSR in Arabica coffee collections. Moncada and McCouch (2004) reported an average of 2.5 alleles per SSR in 11 wild Arabica coffee genotypes and 12 cultivated genotypes, respectively, with the number of alleles ranging from one to eight. Maluf et al. (2005) also reported an average number of 2.87 alleles in 28 cultivated Arabica lines using 23 SSR markers. Bigirimana et al. (2013) reported an average of 3.2 alleles in 14 genotypes using six SSR markers. In the present study, the higher average number of alleles (5.6) may be due to high diversity in Ethiopian C. arabica accessions. In another study, Teressa et al. (2010) reported a mean of 6.5 alleles per locus that ranged from 2 to 14 using 133 coffee genotypes. In another study, da Silva et al. (2019) reported 6.9 mean number of allele per locus in 36 Coffea genotypes using 30 SSR markers that is higher than the present study. Hussein et al. (2017) also reported higher mean number of allele per locus (10.7) using 15 SSR markers. Generally, the authors suggested the number of alleles depends on markers, the platform used for resolution of amplified products and the sources of genotypes used.
The polymorphic information content (PIC) value, ranged from 0.45 to 0.75 with a mean value of 0.6. About 16 (80%) of the markers were highly informative (i.e. PIC > 0.50). The rest four markers were moderately informative with PIC values less than 0.5. Al-Murish et al. (2013) reported 0.43 mean PIC value which is a lower than that of the present study. Similarly, Moncada and McCouch (2004) reported lower average PIC value of 0.35 among 11 Coffea arabica genotypes. In the present study, the higher mean PIC value (0.6) could be attributed to the diversity of genotypes used. In other studies, da Silva et al. (2019) reported 0.72 mean PIC values per locus in 36 Coffea arabica using 30 SSR markers that is higher than the present study. Such PIC value that observed in present and past studies provides an estimate of the discriminatory power of a locus (Marulanda et al. 2014).
About 17 (85%) of the markers showed significant (p < 0.05) deviation from Hardy–Weinberg equilibrium (HWE). Only three markers (CFGA465 p = 0.17), (AJ308774 p = 0.088) and (AJ308755 p = 0.062) showed non-significant deviation from HWE. The loci studied displayed differences between Ho and He in which some of them showed excess heterozygosity that led to a significant departure from HWE across populations. Such excess heterozygosity is expected in sexually reproducing organisms that can maintain their heterozygosity through sexual recombination, or other factors such as natural selection pressure and excess gene flow (Gadissa et al. 2018). In other ways, a high number of alleles and large gene diversity consequently give a high value of expected heterozygosity (Liu et al. 2005).
On the other hand, the lowest fixation index (− 0.42) was observed for AJ308774 marker (Table 3). The fixation index of the markers (F) ranged from 0 to 1 with a mean of 0.67 (Table 3). Five markers had negative fixation index values, indicating an excess of heterozygotes. For loci Aj308779 (72%), CFGA465 (77.4%), AJ250257 (73.2%), AJ308774 (68%), and AJ308755 (77.8%), genotypes were expected to be heterozygous under the specific locus under random mating conditions. However, 98.8%, 88.3%, 97.8%, 96.5%, and 91.9% of the genotypes were heterozygotes for Aj308779, CFGA465, AJ250257, AJ308774, and AJ308755, respectively. The F value revealed that five of the twenty markers showed an excess of heterozygotes (negative F value) which might be due to mutation at specific loci.
Genetic diversity within and among populations
Analysis of genetic diversity parameters
The comparative analysis showed that there is no much difference among the ten populations with regard to genetic diversity indices. Comparatively, the population from Omo had the higher genetic diversity indices. Next to population from Omo, the population from Ilubabor and Bench Maji scored the highest number of genetic diversity indices parameters. They are genetically more diverse than the other populations as estimated by genetic diversity indices parameters. Hence, the areas, where these populations were collected, could be considered as genetic diversity hot spots and potential in-situ conservation sites for C. arabica germplasm. Similarly, Tadele et al. (2014) reported higher genetic diversity in the population that was collected from South Omo. In addition to this, in terms of allelic richness the populations from Omo, Ilubabor, Benchi Maji and Kefficho are the top four in that order, and hence they are more interesting in terms of genetic and evolutionary studies (Leberg 1991). On other hand, the populations from Bale, Hararge and Harar scored the least number of genetic diversity indices parameters in order of their magnitude, which might suggest current rapid genetic erosion in the area.
The mean observed heterozygosity (0.24) observed in the present study is lower than the half mean of expected heterozygosity (0.65) observed (Ho < He) indicating that the population is inbreeding (Govindraj et al. 2015). The mean value of observed heterozygosity (Ho) observed in the present study is higher than that of Hussein et al. (2017) who reported 0.212 mean observed heterozygosity of Yemeni coffee (Coffea arabica L.). Combes et al. (2000) also reported 0.043 average Ho value in Coffea arabica using eleven SSR primers that is lower than the present study. In other studies, Al-murish et al. (2013) reported 0.32 mean observed heterozygosity (Ho) that higher than the present study. Overall the lower observed heterozygosity level in this study and other studies might be due to the autogamous nature of C. arabica which contributes to low heterozygosity levels.
The polymorphism detected in the present study was similar to the result reported by Hussein et al. (2017) who observed 100% polymorphism of markers in Yemeni coffee (Coffea arabica L.). This higher degree of polymorphism might be due to the difference in discrimination power of the genetic marker and the source of genotypes used in the present study. Hence, SSR markers are more powerful and very informative for genetic diversity analysis than any other markers (Vieira et al. 2010; Pestiana et al. 2015). In contrast to the present study, in several studies, a small number of polymorphic loci were observed for C. arabica (Combes et al. 2000; Vieira et al. 2010; Pestiana et al. 2015). These might be due to the low genetic variability of species and narrow genetic base of origin.
The mean diversity indices parameters obtained in the present study showed a high level of genetic variation within populations of Ethiopian coffee. This might be due to a relatively different genetic basis of the populations included in this study that resulted from different germplasm resources accessible to farmers, or due to addition in population size, both natural as well as human factors such as, sexual reproduction of the C. arabica and free movement of germplasm between different geographic areas (Table 4).
Population genetic differentiation and gene flow
Analysis of molecular variance (AMOVA) partitioned the total molecular variance withinand among the populations based on their area of collections. There were highly significant (P < 0.001) molecular variances among populations, among individuals within populations and within individuals. The highest proportion, 63% of the variation was attributed to genetic variability among individuals within populations, while 33% was due to variation within individuals in the population. In contrast, a smaller portion of the total variation (4%) was among populations suggesting that all populations are genetically similar (Table 5). In several studies, Aga et al. (2003), Tesfaye (2006), Balemi (2007), and Tadele et al. (2014) observed high variation within the population as compared to among population variation, which is in agreement with the results of this study.
The higher variation within populations’ genetic diversity might be accounted to contrary reasons. Coffea arabica is affected by multiple evolutionary forces which operate within historical and biological context of the plant species. This includes the mating types, gene flow, mode of reproduction and natural selection. In addition, the high genetic diversity observed within populations of the Coffea arabica might be due to preferential adaptive gene complexes adapted to environmental changes being evolved during the long evolutionary period in a given region (Loveless and Hamrik 1984; Tadele et al. 2014).
The overall observed gene flow (Nm) or gene migration value observed in this study on Ethiopian Arabica coffee was 2.45. Based on Slatkin (1985) and Waples (1987), Nm values grouped into three categories: Nm > 1.00 high, 0.25–0.99 intermediate, and 0.000–0.249 low. Therefore, the Nm value observed in this study indicates high gene flow between populations. Gene flow between population could occur through either seeds or seedlings exchange. Moreover, coffee farmers could also contribute to gene flow by exchanging seeds and seedlings of enhanced landraces among nearby districts. This leads to maximize the diversity of local genotypes and increase the distribution of alleles among different populations regardless of their geographical distance.
The magnitude between and within population differentiation was quantified using F-statistics (Fst, Fis, and Fit) also known as “Wright 1951”. According to Wright (1951), the threshold to determine the level of Fst value ranges from 0 to 0.05 considered as low, 0.05–0.15 moderate, 0.15–0.25 large and those greater than 0.25 mean very large genetic differentiation among populations. In the current study the overall F statistics used as a measure of population differentiation was low (Fst = 0.037 which is < 0.05), which indicates that C. arabica has very low genetic differentiation among populations, which accounted only for 3.7% of the total genetic variation (Table 5). This low genetic differentiation among population may be due to high gene flow that resulted from the movement of seeds from one location to another.
The pairwise genetic differentiation among populations within the source of origin ranged from 0.041 to 0.108 (Table 6). The lowest population differentiation was observed between Benchi Maji and Wolegga, Omo and Wolegga, and Jimma and Wolegga, whereas the highest population differentiation observed between Bale and Hararge, Kefficho and Hararge, and Jimma and Bale populations in order of their magnitude. The low value of Fst implies that there is high frequency of identical alleles among Coffea arabica accessions. The lowest Fst value observed between a pair of populations can be explained by high level of gene flow, which leads to genetic similarity of populations. The low population genetic differentiation is supported by high gene flow owing to step-wise pollen movement across C. arabica populations. This study showed the minimal effects of regions or geographic origins on genetic variation among Coffea arabica populations.
Genetic distance among populations
The magnitude of genetic distance between Coffea arabica populations was estimated based on Nei’s (1978). The pairwise Nei’s unbiased genetic distance value ranged from 0.048 to 0.426 whereas the mean Nei’s unbiased genetic distance of the populations ranged from 0.16 to 0.35 with an overall mean of 0.21 (Table 6). The Population from Hararge showed the highest (0.426) pairwise Nei’s unbiased genetic distance with Kefficho population. The population from Bale also, separated from Jimma and Hararge populations at higher pairwise Nei’s unbiased genetic distance of 0.406 and 0.4, respectively. The population from Bale separated from all populations at highest mean Nei’s unbiased genetic distance (0.35). This population was the most genetically distinct population with the highest mean Nei’s unbiased genetic distance. The population from Hararge also scored the higher mean Nei’s unbiased genetic distance (0.29) next to Bale population. Hence, these populations may serve as potential sources of new genetic variation of important traits that can be used in breeding programs.
Additionally, the smallest pairwise Nei’s unbiased genetic distance was observed between Jimma and Wolegga and between Benchi Maji and Wolegga populations in order of their magnitude. Thus, the population from Wolegga showed the smallest pairwise Nei’s unbiased genetic distance with Jimma and Benchi Maji populations with Nei’s unbiased genetic distance of 0.048 and 0.051, respectively (Table 6). The result of the present study indicates the availability of low pairwise genetic distance and low overall genetic distance among all populations of Coffea arabica (Table 7).
Genetic relationships among accessions
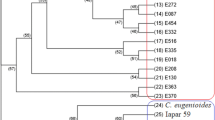
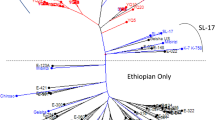
Cluster analysis of 86 Coffea arabica genotypes revealed three major clusters, consisting of 2.3%, 58%, and 39.5% of the total populations in clusters I, II, and III, respectively (Fig. 1). Similarly, Hussein et al. (2017) grouped Yemeni coffee into three major clusters that contain seventeen genotypes involving 16 commercial cultivars and one accession of Yemeni coffee (Coffea arabica L.) germplasm using 16 SSR markers. In another study, Tadele et al. (2014) reported two major clusters of 87 Coffea arabica that collected from southern Ethiopia using five ISSR markers. The first cluster (Cluster I) contains only two accessions (YC30 and WG40) from Yirgachefe and Wolegga populations. These two accessions grouped together in cluster I regardless of their geographical origin. Clusters II and III, classified in to different hierarchical sub-groups. The second cluster included most of the genotypes (50) from all populations. Cluster III included 34 genotypes from nine populations. Only the Accessions from Hararge were not included in cluster III.
Accessions from different populations were clustered together, which may imply the existence of gene flow between and within populations. Hararge accessions were grouped with geographically distant accessions from Omo in cluster II. Accessions from Bale were also grouped with Ilubabor accessions. In all of the clusters many accessions are grouped with geographically distant populations. This indicates accessions in one cluster might be evolved from different lines of ancestry. In addition, the independent events of evolutionary forces such as genetic drift, mutation, migration, natural and artificial selection and germplasm exchange might be separated them into related but different gene pools (Slatkin 1985).
Generally, the cluster revealed a weak clustering pattern confirming low genetic differentiation among the populations, suggesting the genetic background of Coffea arabica populations do not correlate with their geographical origin. Hence, the inter-mixed UPGMA dendrogram of 86 Coffea arabica individual plants indicate high level of genetic variation among individual genotypes investigated and the clustering pattern is weak to support the concept of “isolation by distance” (Fig. 1).
Principal coordinates analysis and population genetic structure
The PCoA analysis in the two dimensional plot displayed in Fig. 2 showed that accessions from different collection sites often grouped together. There was no separate group formed by a single population. This, in turn, agrees with the results of the UPGMA dendrogram in that there was no unique clustering among accessions from the same population. In some cases, accessions of the same population such as Hararge and Ilubabor formed sub cluster in the major groups. Even though, some of the accessions forming a sub cluster in their specific group, there was no separate group formed by a single population. The overall grouping pattern of PCoA corresponds with the clustering of UPGMA dendrogram (Fig. 1) which explains about conformity of the results obtained from the cluster analysis. The presence of seed exchange and high gene flow between and within populations or collection sites may be the probable explanation behind the mixed clustering of accessions from different populations.
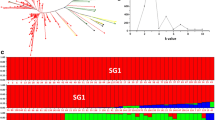
The Bayesian approach based assignment of the 86 individual plants to different populations and determination of their population structure, using STRUCTURE outputs, predicted K = 2 to be the most likely number of clusters (Fig. 3a). Based on this value, Clumpak result (bar plot) showed admixtures and hence there was no clear geographic origin based structuring of populations (Fig. 3b). Structure analysis revealed a close relationship (weak sub-division) among the samples from the ten collection zones and in general, two inferred groups (K = 2) with potential admixtures have been observed. It is interesting to indicate that all individual plants analyzed have alleles originated from the two clusters, which supports the presence of a strong gene flow that led to poor population genetic differentiation.
Delta K value estimated using Evano et al. (2005) method (a) and Bayesian model-based estimation of population structure (K = 2) (b) for the Coffea arabica L. individual plants in ten pre-determined populations
Availability of data and materials
Full passport data of the 86 coffee samples representing the 10 populations used in the present study is provided in Table 1. The other datasets generated and analyzed during the current study are available from the corresponding author on reasonable request.
References
Aerts R, Berecha G, Gijbels P, Hundera K, Van Glabeke S, Vandepitte K, Muys B, Roldan Ruiz I, Honnay O (2013) Genetic variation and risks of introgression in the wild Coffea arabica gene pool in south-western Ethiopian montane rain forests. Evol Appl 6(2):243–252. https://doi.org/10.1111/j.1752-4571.2012.00285.x
Aga E, Bryngelsson T, Bekele E, Balemi S (2003) Genetic diversity of forest arabica coffee (Coffea arabica L.) in Ethiopia as revealed by random amplified polymorphic DNA (RAPD) analysis. Hereditas 138:36–46. https://doi.org/10.1034/j.1601-5223.2003.01636.x
Al-Abdulkader AM, Al-Namazi AA, Al Turki TA, Al-Khuraish MM, Al-Dakhil AI (2018) Optimizing coffee cultivation and its impact on economic growth and export earnings of the producing countries: the case of Saudi Arabia. Saudi J Biol Sci 25(4):776–782. https://doi.org/10.1016/j.sjbs.2017.08.016
Al-Murish TM, Elshafei AA, Al-Doss AA, Barakat MN (2013) Genetic diversity of coffee (Coffea arabica L) in Yemen via SRAP, TRAP and SSR markers. J Food Agric Environ 11(2):411–416. https://doi.org/10.1234/4.2013.4315
Anthony F, Combes MC, Astorga C, Bertrand B, Graziosi G, Lashermes P (2002) the origin of cultivated Coffea arabica L. varieties revealed by AFLP and SSR markers. Theor Appl Genet 104:894–900. https://doi.org/10.1007/s00122-001-0798-8
Balemi S (2007) Genetic diversity analysis of the wild Coffea arabica L. populations from Harenna forest, Bale Mountains of Ethiopia, using Inter Simple Sequence Repeats (ISSR) marker. MSc thesis, Addis Ababa University, Addis Ababa, Ethiopia. P 89
Benti T (2017) Progress in Arabica coffee breeding in Ethiopia: achievements, challenges and prospects. Int J Sci Basic Appl Res 33(2):15–25
Bigirimana J, Njoroge K, Muthomi JW, Gahakwa D, Phiri NA, Gichuru EK, Walyaro DJ (2013) Genetic diversity among disease resistant coffee varieties and cultivars in Rwanda based on RAPD and SSR markers. J Renew Agric 1(6):106–112. https://doi.org/10.12966/jra.09.01.2013
Bunn C, Läderach P, Ovalle-Rivera O, Kirschke D (2015) a bitter cup: climate change profile of global production of Arabica and Robusta coffee. Clim Change 129:89–101. https://doi.org/10.1007/s10584-014-1306-x
Combes MC, Anderzejewski F, Anthony B, Bertrand P, Rovelli G, Graziosi and Lashermes, (2000) characterization of microsatellite loci in Coffea arabica and related coffee species. Mol Ecol 9:1178–1180. https://doi.org/10.1046/j.1365-294x.2000.00954-5.x
Cubry PP, Musali H, Legnate D, Pot FD, Bellis V, Poncet F, Anthony M, Dufour M, Leroy T (2008) diversity in Coffee assessed with SSR markers: structure of the genus Coffea and perspectives for breeding. Genome 51:50–63. https://doi.org/10.1139/G07-096
da Silva BSR, Sant’Ana GC, Chaves CL, Androcioli LG, Ferreira RV, Sera GH, Charmetant P, Leroy T, Pot D, Domingues DS, Pereira LFP (2019) population structure and genetic relationships between Ethiopian and Brazilian Coffea arabica genotypes revealed by SSR markers. Genetica 147(2):205–216. https://doi.org/10.1007/s10709-019-00064-4
Davis AP, Tosh J, Ruch N, Fay MF (2011) growing coffee: Psilanthus (Rubiaceae) subsumed on the basis of molecular and morphological data; implications for the size, morphology, distribution and evolutionary history of Coffea. Bot J Linn Soc 167(4):357–377. https://doi.org/10.1111/j.1095-8339.2011.01177.x
Diniz LEC, Ruas CDF, Carvalho VDP, Torres FM, Ruas EA, Santos MDO, Sera T, Ruas PM (2005) Genetic diversity among forty coffee varieties assessed by RAPD markers associated with restriction digestion. Braz Arch Biol Technol 48(4):511–521. https://doi.org/10.1590/S1516-89132005000500002
Earl DA, Von Holdt BM (2012) STRUCTURE HARVESTER: a website and program for visualizing STRUCTURE output and implementing the Evanno method. Conser Genet Resour 4:359–361. https://doi.org/10.1007/s12686-011-9548-7
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14:2611–2620. https://doi.org/10.1111/j.1365-294X.2005.02553.x
Gadissa F, Tesfaye K, Dagne K, Geleta M (2018) Genetic diversity and population structure analyses of Plectranthus edulis (Vatke) Agnew collections from diverse agro-ecologies in Ethiopia using newly developed EST-SSRs marker system. BMC Genet 19(1):1–15. https://doi.org/10.1186/s12863-018-0682-z
Gebreselassie H, Atinafu G, Degefa M, Ayano A (2018) Arabica Coffee (Coffea arabica L) Hybrid genotypes evaluation for growth characteristics and yield performance under Southern Ethiopian growing condition. Acad Res J Agric Sci Res 6(2):89–96. https://doi.org/10.14662/ARJASR2017.071
Geleta M, Herrera I, Monzón A, Bryngelsson T (2012) Genetic diversity of Arabica coffee (Coffea arabica L.) in Nicaragua as estimated by simple sequence repeat markers. Sci World J 2012:1–11. https://doi.org/10.1100/2012/939820
Govindaraj M, Vetriventhan M, Srinivasan M (2015) Importance of genetic diversity Assessment in crop plants and its recent advances: an overview of its analytical perspectives. A review article. Genet Res Int 2015:1–14. https://doi.org/10.1155/2015/431487
Hussein MAA, Al-Azab AAA, Habib SS, El Sherif FM, El-Garhy HA (2017) Genetic diversity, structure and dna fingerprint for developing molecular ids of Yemeni Coffee (Coffea arabica L) germplasm assessed by SSR markers. Egypt J Plant Breed 21(4):713–736
Kalinowski ST (2005) HP-RARE 1.0: a computer program for performing rarefaction on measures of allelic richness. Mol Ecol Notes 5:187–189. https://doi.org/10.1111/j.1471-8286.2004.00845.x
Khanuja SP, Shasany AK, Darokar MP, Kumar S (1999) Rapid isolation of DNA from dry and fresh samples of plants producing large amounts of secondary metabolites and essential oils. Plant Mol Biol Rep 17(1):74–74. https://doi.org/10.1023/A:1007528101452
Kopelman NM, Mayzel J, Jakobsson M, Rosenberg NA, Mayrose I (2015) CLUMPAK: a program for identifying clustering modes and packaging population structure inferences across K. Mol Ecol Resour 15:1179–1191. https://doi.org/10.1111/1755-0998.12387
Lashermes P, Combes MC, Cros J, Trouslot P, Anthony F, Charrier A (1995). Origin and genetic diversity of Coffea arabica L. based on DNA molecular markers. In: 16th Conference of ASIC, Kyoto (Japan), pp 528–535
Leberg PL (1991) Influence of fragmentation and bottlenecks on genetic divergence of wild turkey populations. Conserv Biol 5(4):522–530. https://doi.org/10.1111/j.1523-1739.1991.tb00359.x
Liu K, Muse SV (2005) Power marker: an integrated analysis environment for genetic marker analysis. Bioinformatics 21(9):2128–2129. https://doi.org/10.1093/bioinformatics/bti282
Loveless MD, Hamrick JL (1984) Ecological determinants of genetic structure in plant populations. Annu Rev Ecol Syst 15:65–95
Maluf MP, Silvestrini M, Ruggiero LM, Guerreiro Filho O, Colombo C (2005) Genetic diversity of cultivated Coffea arabica inbred lines assessed by RAPD, AFLP and SSR marker systems. Scientia Agricola 62(4):366–373. https://doi.org/10.1590/S010390162005000400010
Marulanda ML, Lopez AM, Isuzu L, Lopez P (2014) Microsatellite isolation and characterization for Colletotrichum spp, causal agent of anthracnose in Andean black berry. Genet Mol Res 13:7673–7685. https://doi.org/10.4238/2014.September.26.5
Moncada P, Couch M (2004) Simple sequence repeats diversity in diploid and tetraploid Coffea species. Genome 47:501–509. https://doi.org/10.1139/g03-129
Motta LB, Soares TCB, Ferrao MAG, Caixeta ET, Lorenzoni RM, Souza Neto JDD (2014) Molecular characterization of arabica and Conilon coffee plants genotypes by SSR and ISSR markers. Braz Arch Biol Technol 57(5):728–735. https://doi.org/10.1590/S1516-8913201402071
Nei M (1978) Estimation of average heterozygosity and genetic distance from a small number of individuals. Genetics 89:583–590
Perrier X, Jacquemoud-Collet JP (2006) DARwin software. http://darwin.cirad.fr/
Pestana KN, Capucho AS, Caixeta ET, de Almeida DP, Zambolim EM, Cruz CD, Zambolim L, Pereira AA, de Oliveira ACB, Sakiyama NS (2015) Inheritance study and linkage mapping of resistance loci to Hemileia vastatrix in Híbrido de Timor UFV 443–03. Tree Genet Genomes 11(4):72. https://doi.org/10.1007/s11295-015-0903-9
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155(2):945–959. https://doi.org/10.1093/genetics/155.2.945
Rovelli P, Mettulio R, Anthony F, Anzueto F, Lashermes P, Graziosi G (2000) Microsatellites in Coffea arabica L. In: Sera T, Soccol CR, Pandey A, Roussos S (eds) Coffee biotechnology and quality. Springer, Dordrecht, pp 123–133
Sant’Ana GC, Pereira LF, Pot D, Ivamoto ST, Domingues DS, Ferreira RV, Pagiatto NF, da Silva BSR, Nogueira LM, Kitzberger CS, Scholz MB, de Oliveira FF, Sera GH, Padilha LL, Labouisse JP, Guyot R, Charmetant P, Leroy T (2018) Genome-wide association study reveals candidate genes influencing lipids and diterpenes contents in Coffea arabica L. Sci Rep 8(1):465. https://doi.org/10.1038/s41598-017-18800-1
Silvarolla MB, Mazzafera P, Fazuoli LC (2004) Plant biochemistry: a naturally decaffeinated Arabica coffee. Nature 429(6994):826. https://doi.org/10.1038/429826a
Slatkin M (1985) Rare alleles as indicators of gene flow. Evolution 39(1):53–65. https://doi.org/10.1111/j.1558-5646.1985.tb04079.x
Sousa TV, Caixeta ET, Alkimim ER, de Oliveira ACB, Pereira AA, Zambolim L, Sakiyama NS (2017) Molecular markers useful to discriminate Coffea arabica L cultivars with high genetic similarity. Euphytica 213(3):75. https://doi.org/10.1007/s10681-017-1865-9
Steiger D, Nagai C, Moore P, Morden C, Osgood R, Ming R (2002) AFLP analysis of genetic diversity within and among Coffea arabica cultivars. Theor Appl Genet 105(2–3):209–215. https://doi.org/10.1007/s00122-002-0939-8
Tadele S, Mekbib F, Tesfaye K (2014) Genetic diversity of coffee (Coffea arabica L.) landraces from southern Ethiopia as revealed by inter simple sequence repeat marker. Glob Adv Res J Agric Sci 3(1):24–34
Teressa A, Crouzillat D, Petiard V, Brouhan P (2010) Genetic diversity of Arabica coffee (Coffea arabica L) collections. EJAST 1(1):63–79
Tesfaye K (2006) Genetic diversity of wild Coffea arabica populations in Ethiopia as a contribution to conservation and use planning. Ecology and development series. University of Bonn, Germany
Tesfaye K, Govers K, Bekele E, Borsch T (2014) ISSR fingerprinting of Coffea arabica throughout Ethiopia reveals high variability in wild populations and distinguishes them from landraces. Plant Syst Evol 300(5):881–897. https://doi.org/10.1007/s00606-013-0927-2
Tran HT, Lee LS, Furtado A, Smyth H, Henry RJ (2016) Advances in genomics for the improvement of quality in coffee. J Sci Food Agric 96:3300–3312. https://doi.org/10.1002/jsfa.7692
Vieira ESN, von Pinho EVR, Carvalho MG, Esselink GD, Vosman B (2010) Development of microsatellite markers for the identification of Brazilian Coffea arabica varieties. Genet Mol Biol 33(3):507–514. https://doi.org/10.1590/S1415-47572010005000055
Waples RS (1987) A multi species approach to the analysis of gene flow in marine shore fishes. Evolution 41(2):385–400. https://doi.org/10.1111/j.1558-5646.1987.tb05805.x
White Head and Peakall (2015) GENALEX 6.502: genetic analysis in Excel. Population genetic software for teaching and research. Mol Ecol Notes 6:288–295. https://doi.org/10.1111/j.1471-8286.2005.01155.x
Wright S (1951) The genetic structure of populations. Annu Eugen 15:323–354. https://doi.org/10.1111/j.1469-1809.1949.tb02451.x
Yeh FC, Yang R (1999) POPGENE VERSION 1.31: Microsoft Window-based Freeware for Population Genetic Analysis. Department of Renewable Resources University of Alberta, Canada. pp 1–29
Acknowledgements
The study was conducted by financial support provided by Ethiopian Institute of Agricultural Research Institute, National Agricultural Biotechnology Research Center. The authors are grateful to the Ethiopian Biodiversity Institute for providing the coffee germplasm.
Funding
This work was supported by grants from the Ethiopian Institute of Agricultural Research Institute, National Agricultural Biotechnology Research Center.
Author information
Authors and Affiliations
Contributions
Material preparation, data collection, and analysis were performed by GD and TD. The first draft of the manuscript was written by GD and all authors commented on previous versions of the manuscript. All authors participated in designing the experiment, interpreting the data, drafting and revising the manuscript and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors have no conflicts of interest to declare that are relevant to the content of this article.
Ethics approval and consent to participate
Not Applicable.
Consent for publication
Not Applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Dida, G., Bantte, K. & Disasa, T. Molecular characterization of Arabica Coffee (Coffea arabica L.) germplasms and their contribution to biodiversity in Ethiopia. Plant Biotechnol Rep 15, 791–804 (2021). https://doi.org/10.1007/s11816-021-00721-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11816-021-00721-1