Abstract
Strawberry (Fragaria × ananassa Duch.) is one of the most widely cultivated fruit crop. Anthracnose caused by Colletotrichum spp. is a devastating disease of strawberry, causing large-scale strawberry losses worldwide. Chitinases act as defence proteins and are crucial for plant response to pathogens. Here, we isolated a class V Chitinase gene (designed as FnCHIT2, GenBank accession number MN709779) from Chinese wild diploid strawberry Fragaria nilgerrensis Schlecht (F. nilgerrensis), a species that exhibits high tolerance to anthracnose. Gene expression analysis showed that FnCHIT2 expression was highly induced after Colletotrichum gloeosporiodes inoculation and salicylic acid (SA) treatment. Subcellular localization analysis revealed the presence of FnCHIT2 in the plasma membrane. Recombinant FnCHIT2 protein was successfully expressed in E. coli Rosetta (DE3). Furthermore, we transformed FnCHIT2 into Col-0 wild type A. thaliana to perform functional analysis and evaluated the functions of Colletotrichum higginsianum and Pseudomonas syringae pv. tomato DC3000 (Pst DC3000). FnCHIT2 overexpression in A. thaliana showed enhanced resistance to C. higginsianum and Pst DC3000. Enhanced disease resistance of FnCHIT2 transgenic plants to C. higginsianum was correlated with pathogenesis-related gene 1 (AtPR1) and plant defensin 1.2 (AtPDF1.2) gene expression levels. These results provide evidence that FnCHIT2 may play an important role in response to fungal pathogens in strawberry. Our study provides an important theoretical reference for future strawberry resistance breeding.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Cultivated strawberry [Fragaria × ananassa Duch. (2n = 8x = 56)] is one of the most economically important horticultural crop throughout the world. However, fruit yield and quality are limited by a range of biotic and abiotic stresses. Anthracnose caused by Colletotrichum spp. is one of the most devastating fungal diseases in strawberry (Karimi et al. 2019). According to previous studies, C. acutatum, C. gloeosporioides, and C. fragariae are causative agents of strawberry anthracnose. However, some recent studies have reported that several new Colletotrichum spp. can cause anthracnose in strawberry (Chung et al. 2020). China has diverse germplasm resources of the genus Fragaria, including some diploid strawberries, such as F. nilgerrensis, F. hayatai, F. mandschurica, and F. chinensis. This diploid strawberry germplasm have been utilized for strawberry breeding programmes based on numerous characteristics such as distinct aroma and resistance to fungal diseases (Zhang et al. 2020a, b). F. nilgerrensis is a wild strawberry that is widely distributed in Southwest China this diploid strawberry has been exploited as a breeding material. For example, the special characteristic of F. nilgerrensis (aroma) was introduced into Fragaria × ananassa. This process added to the genetic diversity of cultivated strawberry (Noguchi et al. 2002). In addition to its special aroma, this wild strawberry also has high resistance levels against abiotic (drought and heat) and biotic (anthracnose and pests) stresses (Zhang et al. 2016). Understanding the mechanisms of disease resistance and characterizing genes involved in resistance in F. nilgerrensis can greatly accelerate strawberry disease resistance breeding.
Chitinases (EC3.2.1.14) are lytic enzymes that are present in a wide range of organisms from microorganisms to animals and higher plants (Dong et al. 2017). Chitinase catalyses the hydrolysis of b-1, 4 linkages of N-acetyl-D-glucosamine in chitin (Singh et al. 2015) and is the second most abundant biopolymer in nature after cellulose. Chitin is a major structural component of the cell walls of many pathogenic fungi and insect skeletons (Grover et al. 2012). Chitinases are strongly induced in plant responses to bacteria, fungi, and viruses (Grover et al. 2012). These proteins have an important role in plant defence against pathogen attack (Abeles et al. 1971; Punja and Zhang 1993). Hence, understanding exactly how chitinases are involved in plant immune systems and the roles they play is important for the study of plant resistance mechanisms (Dong et al. 2017). Previously, many scientists have reported the involvement of the chitinase gene in fungal disease resistance mechanisms. For example, overexpression of the chitinase gene EuCHIT2 from Eucommia ulmoides oliver (E. ulmoides) enhanced resistance to Erysiphe cichoracearum DC. In tobacco plants (Dong et al. 2017). Overexpression of a barley chitinase class-II gene in sugarcane enhanced resistance against red rot (Tariq et al. 2018). Furthermore, overexpression of a potato class I chitinase gene in transgenic tea enhanced resistance to blister blight (Singh et al. 2015). Transformation of barley antifungal genes chitinase and ribosome-inactivating protein-induced fungal resistance in black gram (Chopra and Saini 2014). Constitutive expression of a class II chitinase gene (Zjchi2) from zoysia grass significantly enhanced antifungal activity in transgenic zoysia grass compared with wild-type plants (Kang et al. 2017). Overexpression of an endo-chitinase gene from barley in potatoes decreased susceptibility after inoculation with Alternaria solani compared to the control (Khan et al. 2017). The pepper chitinase gene ChiIV3 acted as an antifungal protein and as a receptor for unidentified chitin in plants to trigger cell death and defence signalling against Phytophthora capsici infection (Liu et al. 2017). Studies have also demonstrated that genes encoding proteins that belong to chitinase families in plants are involved in abiotic-stress responses, such as osmotic, salt, cold, wounding, and heavy metal stresses (Brotmana et al. 2012). In melon, heat shock-induced resistance increased chitinase-1 gene expression (Widiastuti et al. 2012). In A. thaliana, Athot2 encodes an endo-chitinase-like protein that is essential for tolerance to heat, salt, and drought stresses (Kwon et al. 2007). According to some studies, abiotic and biotic stress conditions can elevate chitinase levels in P. ginseng (Brotmana et al. 2012; Pulla et al. 2011). In addition, thirty-two chitinase genes were identified in Ammopiptanthus nanus and 3 chitinase genes were strongly induced by low temperature and osmotic stress (Cao et al. 2019). The potential roles of chitinase genes in other plants species justify the need for functional analysis of chitinase genes in strawberry.
Two strawberry genes (class II chitinase) were partially characterized and were induced after inoculation with Colletotrichum fragariae or Colletotrichum acutatum (Akhan et al. 2004). The ch5B gene encodes a chitinase from Phaseolus vulgaris. This gene was transformed along with gln2 and ap24 into strawberry (cultivar Pájaro), and transgenic plants displayed high levels of resistance to grey mould disease (Botrytis cinerea) (Vellicce et al. 2006). A chitinase gene from Lycopersicon chilense was transformed into ‘Joliette’strawberry, and the transgenic strawberry exhibited increased resistance to Verticillium dahliae compared to non-transgenic plants (Chalavi and Tabaeizadeh 2003). However, characteristics and functional analysis of the chitinase gene from Chinese wild strawberry has not been reported.
In a previous study, we identified a Chitinase 2 gene through RNA-Seq analysis that was differently expressed after inoculation with C. gloeosporiodes in F. nilgerrensis. In the present study, we isolated this gene FnCHIT2 (GH-18 family) and expressed the recombinant FnCHIT2 protein in E. coli and overexpressed the gene in A. thaliana to evaluate its potential function in resistance to C. higginsianum and PstDC3000. Based on our results, we suggest that the FnCHIT2 gene may functions as a positive regulator in the strawberry defence response to anthracnose.
Materials and methods
Plant materials and pathogens
Chinese wild diploid strawberry (Fragaria nilgerrensis Schltdl, 2n = 2x = 14) was maintained at 23 ± 2℃ under a 14-h (h) day/10-h night photoperiod with 75% humidity. The plants were provided by Beijing Academy of Agricultural Sciences, Beijing, China. These plants were propagated as runner plants. After propagation, plants were maintained for an additional 2 weeks before inoculation with the pathogen C. gloeosporiodes (strain Lch-1911). A. thaliana wild-type (WT) and 35S::FnCHIT2 overexpression transgenic plants with a Col-0 ecotype background were used in this study. Plants were grown in a growth chamber at 21 °C, 70% relative humidity, and a 12-h light/12-h dark cycle for 5 weeks.
Cloning and bioinformatics analysis of FnCHIT2
Total RNA was extracted from leaves of F. nilgerrensis using the RNApre Pure Plant Kit (Tiangen, China). First-strand cDNA was synthesized from 1 μg of total RNA using the Primer Script™ 1st Strand cDNA Synthesis kit (TaKaRa Bio Inc., Dalian, China) according to the manufacturer’s guidelines. Polymerase chain reaction (PCR) master mix (Promega, Beijing, China) was used to amplify the ORF sequence of FnCHIT2. PCR was performed with an initial denaturation step of 94 ℃ for 90 s; 32 cycles of denaturation at 94 ℃ for 30 s, annealing at 58 ℃ for 30 s, and extension at 72 ℃ for 90 s; and a final extension at 72 ℃ for 10 min. PCR products were purified using the Universal DNA Purification Kit DNA (Tiangen Bio Inc., Beijing, China) and cloned into the pMD@ 18-T vector for sequencing (Shang Ya Bio Inc., Fuzhou, China). Molecular weight (MW) and isoelectric point (pI) predictions for the deduced protein were performed using the online ExPASy proteomics server database (http://www.expasy.org/tools/protparam.html). Sequence alignment analysis was undertaken with DNAMAN (version 7.0), and the phylogenetic tree was generated using the CLUSTALW2 program (http://ww w.ebi.ac.uk/Tools/clustalw2/index.html).
SA treatment of F. nilgerrensis
For SA treatment, strawberry plants were sprayed with 100 μM SA (Sigma), and 0.05% Tween-20 was used as a surfactant (Wang et al. 2006), plants were grown in a growth chamber at 21 ℃, 70% relative humidity. The leaves were harvested at 0, 6, 12, 24, and 48 h post spraying. The treatments were replicated thrice, and samples were stored at − 80 ℃ (for RNA analysis).
Subcellular localization of FnCHIT2 protein
ORF regions of FnCHIT2 were amplified using specific primers F-FnCHIT2-YFP and R-FnCHIT2-YFP (Table S1) that contained BamH I and Kpn I, and the product was subcloned into pYFPc vector. The resulting vectors were confirmed by sequencing, pm-rk CD3-1007 fusion protein as a plasma membrane-anchored marker (Nelson and Cai 2007) Four-week-old tobacco plants were used for transient expression (Wang et al. 2018) YFP fluorescence in tobacco leaves was examined by laser scanning confocal microscopy (Leica TCS SP8, Leica Microsystems, German).
Pathogen inoculation
The C. gloeosporiodes strain “Lch-1911” provided by Dr. Han Yongchao (Han et al. 2018) was cultured in liquid PDA medium (potato dextrose broth) at 25 ℃ for 7–10 days with shaking. Then, conidia were collected by centrifugation (3000 rpm/min) and subsequently filtrated with cheesecloth. Conidia were then re-suspended in distilled water and adjusted to 1 × 106 conidia/mL. Five mL of suspension was sprayed with a hand sprayer over each strawberry plant (F. nilgerrensis), and control plants were sprayed with distilled water. Twenty uniform, healthy, and micro propagated plantlets at the stage of ten compound leaves were used in the experiment. All inoculations were performed thrice. The inoculated plants were maintained in a moist environment at 28 ℃ (Suzuki et al. 2010), and the leaves were harvested at 0, 6, 12, 24, 48, and 72 h after inoculation.
Pseudomonas syringaepv. tomato DC3000 (Pst DC3000) was cultured in King's B medium containing rifampicin (60 µg/mL) and kanamycin (50 µg/mL) at 28 ℃ overnight. Then, pathogens were collected by centrifugation at 3000 g, then washed with 10 mmol MgCl2 and diluted with distilled water to concentrations of 1 × 105 and 107 cfu/mL for syringe and spray inoculations, respectively (Katagiri et al. 2002; Zhang et al. 2015). Control treatment was performed with distilled water. After inoculation, the plants were placed in a growth chamber under 16-h light (23 ℃)/8-h dark (21 ℃) cycles. Leaves were collected at 48 h post-inoculation. Furthermore, the bacterium growth assays in plants were performed, as previously described (Zhang et al. 2015).
C. higginsianum was used to inoculate transgenic Arabidopsis to elucidate the biological function of FnCHIT2 against anthracnose infection. C. higginsianum strain Ch-1 was provided by Dr. Zheng (Gu et al. 2019) and cultured on PDA at 25 ℃ in dark conditions for 7 days. Conidia were harvested from PDA culture plates by flooding the surface of culture with distilled water and filtering the suspension through cheese-cloth (Casado-Díaz et al. 2006). The spore suspension was examined with a haemocytometer and adjusted to 1 × 106 conidia/mL. Nine pots of four-week-old Col A. thaliana plants were sprayed with 5 mL spore suspension with a hand sprayer. The control plants were sprayed with distilled water. For qRT-PCR experiments, the leaves were collected at 0, 6, 12, 24, 48, 72, and 120 h after inoculation. For trypan blue staining and nitroblue tetrazolium (NBT) staining, fully developed rosette leaves were drop-inoculated with 8 μL of C. higginsianum spore suspension containing 1 × 106 spores /mL (Wen et al. 2015).
Recombinant FnCHIT2 protein expression in E. coli
pMD@-18-T-FnCHIT2 plasmid was used as a template to amplify a truncated gene encoding a signal-deleted gene FnCHIT2-1. Gene amplification using primer (F/R) resulted in the deletion of the first 84 nucleotides in the N-terminal of the FnCHIT2 gene, and PCR products were cloned into pMD@ 18-T vector and sequenced. The truncated gene was cloned into the multiple cloning site of the prokaryotic expression vector pET32A using BamH I and Xba I to generate recombinant vector pET32A-FnCHIT2-1 (Fig. 2S B). The recombinant plasmid was transformed to expression host E. coli strain Rosetta (DE3) (Yueyang Bio Inc., Beijing, China) competent cells. For induction of FnCHIT2-1 protein expression, isopropyl-β-D- thiogalactoside (IPTG) was added to a final concentration of 0.5 mmol/L. Bacterial cells were harvested by centrifugation, and cell pellets were further separated by sodium dodecyl sulphate polyacrylamide gel electrophoresis (SDS-PAGE) to analyse the expression of the recombinant proteins (Laemmli et al. 1970). Ni–NTA Agarose (Qiagen) was used to purify the recombinant protein. For Western blotting, proteins were electrophoresed on 12% SDS-PAGE, transferred to the polyvinylidene difluoride (PVDF) membrane, and then blocked with the membrane 5% skim milk powder in Tris-Buffered Saline Tween-20 (TBST) for 2 h. Then, blots were incubated with primary antibodies 1 h at room temperature followed incubation with the secondary antibody (goat-anti-rabbit IgG) (Thermo Pierce Fisher, Shanghai, China) at a dilution of 1:5000 for 45 min. The protein bands were visualized upon exposure to diaminobenzidine (DAB) substrate (Tiangen Biotechnology Co., Ltd., Beijing, China). Bacterial cells were broken using a supersonic technique, and the E. coli lysate was centrifuged. The resulting supernatant liquid was purified with Ni–NTA Agarose (Qiagen), and FnCHIT2 recombinant protein was detected with 12% SDS-PAGE.
Plasmid construction and plant transformation
To generate the 35S::FnCHIT2 construct, the ORF sequence FnCHIT2 was amplified, and the restriction sites BamH I and Kpn I were introduced on both ends of the sequence. Then, the FnCHIT2 coding sequence was introduced into the pCMBIA1300-HA construct under the control of the 35S promoter. The resulting construct was introduced into Agrobacterium tumeficiens strain GV3101 and then employed for A. thaliana (Col-0) transformation via the floral dip method (Clough et al. 1998). T0 seeds were collected and grown on Murashige and Skoog (MS) (1962) medium (pH 5.8, 10 g/L sucrose, 8 g/L agar) supplemented with 50 mg/L hygromycin. Transgenic plants were identified by PCR amplification using gene-specific primers. Three of twenty-seven T2 independent lines showed the highest resistance to C. higginsianum and Pst DC3000 inoculation and were selected for further experiments.
Expression analysis of FnCHIT2 in strawberry and defence-related genes in A. thaliana
Total RNA was extracted from C. gloeosporiodes-inoculated strawberry leaves and C. higginsianum-infected A. thaliana leaves. Briefly, 1 mg RNA was treated with amplification-grade DNase I (Invitrogen). Three biological replicates were used for each experiment. The first strand of cDNA was synthesized using the PrimeScript™ 1st Strand cDNA Synthesis Kit (TaKaRa Biotechnology, Dalian, China) following the manufacturer's instructions. Subsequently, the cDNA was diluted sixfold with the sterile water and stored at – 40 ℃ for future use. Quantitative real-time PCR analysis was performed using SYBR Green (TaKaRa Biotechnology, Dalian, China) with the Step One Plus Real-Time PCR System (Applied Biosystems, Foster, CA, USA). PCR conditions were as follows: a denaturing step of 95 ℃ for 30 s and 42 cycles of denaturing at 95 ℃ for 5 s and annealing at 60 ℃ for 30 s. The Actin gene (Acc. No. AB116565) and Actin1 gene (Acc. No. AT3G18780) were used as internal controls for strawberry and A. thaliana, respectively. The primer sequences used for qRT-PCR are listed in Table S2.
Statistical analysis
All the experiments were performed with three technical and biological replicates. Data analysis and plotting were performed using Microsoft Excel (Microsoft Corporation, Redmond, WA, USA) and SigmaPlot 12.0 (Systat, Inc., Point Richmond, CA, USA), respectively. Statistical data analysis (Student’s t test,*p < 0.05, **p < 0.01) was performed using SPSS 16.0 (IBM Corporation, Chicago, Illinois, USA).
Results
Characterization and expression analysis of FnCHIT2
A transcriptomic analysis of F. nilgerrensis was performed using RNA-seq and demonstrated FnCHIT2 (GenBank accession number MN709779) expression was highly induced after inoculation with C. gloeosporioides. Then, we isolated the FnCHIT2 cDNA sequence from the leaves of F. nilgerrensis. A 912-bp FnCHIT2 cDNA fragment located on chromosome 3 was cloned from F. nilgerrensis (Fig. 1S and Fig. 2S A). The predicted open reading frame encodes a polypeptide of 303 amino acid residues (Fig. 1S) with a predicted molecular mass of 33.96 Kda and a pI value of 7.09. The predicted protein contained a glycosyl hydrolase family 18 (GH18) domain from amino acid residues 95–156 with two carbohydrate-binding sites (CBD domain) in the N-terminal. However, the predicted protein lacks the chitin-binding domain and belongs to the class V plant chitinases. The deduced amino acid sequence shows relatively high homology with GH18-chitinase-like family members of Rosacea plants. For example, FnCHIT2 has 100%, 82%, and 78.11% similarity with FvCHIT2 (XP_004295963), RcCHIT2 (XP_024159678), and PmCHIT2 (XP_008223957), respectively (Fig. 1a). A phylogenetic tree was generated using putative amino acid sequences of FnCHIT2 and some counterpart GH18-chitinase-like family members from other species. FnCHIT2 clustered in the same clade as FvCHIT2 and RcCHIT2 (Fig. 1b). The responsiveness of FnCHIT2 to C. gloeosporioides was analysed. In F. nilgerrensis strawberry, FnCHIT2 expression was increased at 6–12 h post inoculation (hpi) (Fig. 1c) to levels greater than those of mock-inoculated plants. Upon SA treatment, FnCHIT2 mRNA levels were increased 3.4-fold to 32.9-fold at 6 to 48 hpi, reaching a maximum level at 6 hpi and then decreasing at 48 hpi (Fig. 1d).
Sequence analysis of the deduced amino acid sequence of FnCHIT2 from F. nilgrrensis and related proteins as well as qRT-PCR analysis of FnCHIT2 transcript levels in strawberry leaves after inoculation with C. gloeosporioides. a Multiple sequence alignment of FnCHIT2 and related proteins from Fragaria vesca (GenBank accession no.XP_004295963), Rosa chinensis (GenBank accession no. XP_024159678), and Prunus mume (GenBank accession no.XP_008223957). The FnCHIT2 sequence encodes a 303-amino acid protein containing one ChiA protein domain. The underlined amino acid residues (95 to 156) indicate the ChiA protein domain. b Evolution analysis of FnCHIT2. The accession numbers of protein sequences used to generate the phylogenetic tree are provided in Table S1. c Expression analysis of FnCHIT2 in F. nilgrrensis leaves after inoculation with C. gloeosporioides. d Expression analysis of FnCHIT2 in F. nilgrrensis leaves after exogenous SA treatment. Error bars represent the standard deviation from three independent replicates. Significance levels of *p < 0.05 and **p < 0.01 are indicated compared with control using Student’s t test
Subcellular location of FnCHIT2 protein
To investigate the subcellular location of FnCHIT2 protein, the ORF of FnCHIT2 was fused with a yellow fluorescent protein (YFP) under the control of the 35S promoter. Transient expression in tobacco leaves was examined by laser scanning confocal microscopy (Leica TCS SP8, German, Leica Microsystems, German) (Fig. 2a and Fig. 2S B). Using the pm-rk CD3-1007 fusion protein plasma membrane marker and free YFP vector fluorescence of mCHERRY was observed to distribute almost uniformly along the cell wall and the cell peripheral surface (Fig. 2b). Co-transient 35S::YFP with plasma membrane marker pm-rk CD3-1007, the YFP signals were merged with mCHERRY signals of the plasma membrane marker (Fig. 2c), suggesting that FnCHIT2 was localized to the plasma membrane.
Subcellular localization of FnCHIT2 protein. Transient expression of 35S::FnCHIT2-YFP and pm-rk CD3-1007 fusion protein in tobacco leaves, pm-rk CD3-1007 is a plasma membrane marker protein. Fluorescence signals were visualized using confocal laser scanning microscopy. a Structure of the CaMV35S promoter-FnCHIT2-YFP construct; LB left border, RB right border; 35S CaMV35S promoter, NOS terminator, Amp ampicillin; b indicate injection of pm-rk CD3-1007 and free YFP vector. c Indicate injection of 35S::FnCHIT2-YFP and pm-rk CD3-1007. The 35S::FnCHIT2-YFP signals can be merged with fluorescent signals of the plasma membrane
Construction of recombinant plasmid pET32A-FnCHIT2-1 and expression in E. coil
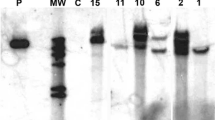
The truncated gene FnCHIT2-1 was inserted into the expression vector pET32A to generate the recombinant pET32A-FnCHIT2-1 construct (Fig. 3a and Fig. 2S B). The expression of recombinant protein products was detected using 12% Sodium dodecylsulphate polyacrylamide gel electrophoresis (SDS-PAGE) and the results showed that the recombinant protein with a predicted weight of approximately 32 Kda was successfully expressed in E. coli strain BL21 (DE3) with the induction of IPTG in the form of a fusion protein (Fig. 3b). According to Western blot analysis, FnCHIT2-1 recombinant protein has a specific reaction with anti-His-Tag rabbit monoclonal antibody (Fig. 3c). This finding suggests that the FnCHIT2-1 gene fusion protein was successfully expressed in E. coli BL21 (DE3). In addition, the samples contained recombinant pET32A-FnCHIT2-1 construct as demonstrated by a 32Kda band in the SDS-PAGE gel (Fig. 3c).
Generation of recombinant construct pET32A-FnCHIT2-1, expression of recombinant protein in E. coli (Rosetta, DE3), Western blot analysis of recombinant of pET32A-FnCHIT2-1 protein with His antibodies, and SDS-PAGE analysis of pET32A-FnCHIT2-1 protein expression induced with different concentrations of IPTG and temperature. a Diagram of FnCHIT2-1 prokaryotic expression vector; b Recombinant pET32A-FnCHIT2-1 vector in E. coli. M, protein molecular weight marker (Low); line 1, pET32A empty sample; line 2, recombinant protein expression of pET32A-FnCHIT2-1 un-induced with IPTG; lines 3–7, recombinant protein expression of pET32A-FnCHIT2-1 induced with IPTG (0.5 mmol). c Western blot of recombinant pET32A-FnCHIT2-1 protein. Protein molecular weight marker (Low); line 1, lysate of recombinant bacterial strain induced by IPTG for 4 h; line 2, positive control
Constitutive expression of FnCHIT2 in A. thaliana enhances resistance to C. higginsianum
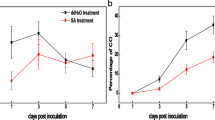
To investigate the biological role of FnCHIT2, the FnCHIT2 cDNA sequence was cloned under the control of the CaMV 35S promoter (Fig. 4a) to create 35S:: FnCHIT2 and overexpressed in A. thaliana plants. In total, 27 T2 transgenic plants were obtained. Three independent T2 transgenic lines (T2-6, T2-25 and T2-26) exhibiting the highest resistant against C. higginsianum and Pst DC3000 were selected for further studies. Colletotrichum species are hemibiotrophic fungi (Dubouzet et al. 2011). The response of Col-0 plants and three transgenic lines was noted against fungus inoculation. According to observations, all plants developed brown necrotic lesions surrounded by a yellow halo at three days post-inoculation (dpi), but more severe symptoms were observed in wild type plants (Fig. 4b). In addition, disease lesion diameters were significantly larger in wild type plants (Fig. 4c). To explore the molecular basis of the FnCHIT2 gene resistance mechanism against C. higginsianum inoculation, the transcription levels of genes related to SA- (AtPR1) and JA- (AtPDF1.2) dependent disease resistance pathways were determined (Fig. 4d and e). In general, the three transgenic lines showed increased AtPR1 expression after inoculation with C. higginsianum (Fig. 4d). AtPR1 expression was significantly enhanced after inoculation in three transgenic lines, and maximum levels were reached at 12 h in T2-6 and T2-26 lines. Initially, AtPDF1.2 expression was down regulated at 12 h in three transgenic lines and then up-regulated compared to wild type plants.
Overexpression of FnCHIT2 in A. thaliana conferred enhanced disease response to C. higginsianum inoculation. a Structure of the CaMV35S promoter-FnCHIT2 construct; LB, left border; RB, right border; 35S, CaMV35Spromoter; NOS, terminator; Hyg, hygromycin. b Disease symptoms in Col-0 and transgenic A. thaliana leaves three days post-inoculation. c The average lesion diameter on leaves at 72 hpi. d qRT-PCR analysis of AtPR1 transcripts after inoculation. e qRT-PCR analysis of AtPDF1.2 transcripts after inoculation. Four-week-old Col-0 and transgenic A. thaliana leaves were sprayed with a spore suspension of C. higginsianum (1 × 106 spores /mL) and harvested at 0, 12, 48, 72 and 120 hpi. Data are the mean ± SE of three replications. Asterisks indicate significant difference between wild type and transgenic lines (* p < 0.05, ** p < 0.01, Student's t test). Scale bars = 0.5 mm
Constitutive expression of FnCHIT2 in A. thaliana enhances resistance to Pst DC3000
To further elucidate the biological function of the FnCHIT2 gene against bacterial disease infection, transgenic and wild type Col-0 A. thaliana plants were inoculated with Pst DC3000. Plants and leaves were examined after 3 dpi. After inoculation, the leaves of Col-0 plants displayed symptoms of chlorosis and necrosis, while the three transgenic lines have no or less symptoms compared with Col-0 plants (Fig. 5a). After trypan blue staining, wild type plants showed more cell death compared to three transgenic lines (Fig. 5d). Regarding NBT staining, three transgenic lines exhibited more O2− accumulation compared with Col-0 plants (Fig. 5c). Bacterial populations were quantified, and the quantities of bacteria in three transgenic plants were significantly (p < 0.05) reduced compared with Col-0 plants (Fig. 5b). Compared with Col-0 plants, FnCHIT2 overexpression in three transgenic lines inhibited Pst DC3000 growth and activity in the leaves. Further, qRT-PCR analysis was performed to investigate the expression of AtPR1 and AtPDF1.2 in transgenic and wild plants. According to qRT-PCR results, the transgenic lines showed significantly increased AtPR1 gene expression after inoculation compared with wild type plants at 6, 12, 48, and 72 hpi (Fig. 5e). The AtPDF1.2 gene exhibited significantly increased expression at 48 or 72 hpi, and these levels subsequently decreased (Fig. 5f).
Effect of FnCHIT2 overexpression in A. thaliana on Pst DC3000 inoculation. Transgenic lines (T2-5, T2-25, and T2-26) and wild type A. thaliana were inoculated with Pst DC3000. a Images of leaves at 3 days post-inoculation. b Bacterial population assays counted at 3 dpi. c Nitro blue tetrazolium (NBT) staining of O2− at 3 dpi. d Trypan blue staining of leaves at 3 dpi. e Relative expression levels of AtPR1 examined using qRT-PCR in transgenic lines (T2-5, T2-25, and T2-26) and wild type A. thaliana. The leaves were collected at 0, 6, 12, 24, 72, and 120 hpi. (F) Relative AtPDF1.2 expression levels examined with qRT-PCR in transgenic lines (T2-5, T2-25, and T2-26) and wild type A. thaliana. The leaves were collected at 0, 6, 12, 24, 72 and 120 hpi. Data represent mean ± SD values from three independent experiments. Asterisks indicate significant differences between Col-0 and transgenic lines as determined by Student’s t-test (* p < 0.05, ** p < 0.01). Scale bars = 0.5 mm
Discussion
Colletotrichum is a genus of major plant pathogens causing anthracnose disease in many plants worldwide (Silva et al. 2017). This ascomycete genus is comprised of a highly diverse group of pathogenic fungi that can infect a wide range of commercially important crops (Gan et al. 2013). Several species of Colletotrichum spp. causes strawberry anthracnose in China. C. gloeosporioides has been defined as the major causal agent (Zhang et al. 2016). C. gloeosporioides is both intracellular hemibiotrophic and intramural necrotrophic (Gan et al. 2013; Xie et al. 2010; O’Connell et al. 2000; Kim et al. 2004; Moraes et al. 2013). C. higginsianum causes anthracnose disease in many wild and cultivated crucifers, including A. thaliana (Narusaka et al. 2004; O’Connell et al. 2000; Takahara et al. 2009). This fungus also has a hemibiotrophic lifestyle (Plaumann et al. 2018). The Arabidopsis-C. higginsianum interaction is a convenient model for system analysis of fungal pathogenicity and plant resistance (Takahara et al. 2009). In this study, we isolated the FnCHIT2 gene from F. nilgerrensis after C. gloeosporioides infection, overexpressed this gene in Arabidopsis and investigated its biological function after inoculation with C. higginsianum.
Chitinases are pathogenesis-related proteins that play important roles in host resistance to various pathogens and abiotic stress responses (Xu et al. 2016). Chitinase gene expression is strongly induced by infection with fungi, bacteria, and viruses. For example, 33 chitinase genes were identified in Brassica rapa, among them 14 genes were induced by P. brassicae infection (Chen et al. 2018). Eight chitinase genes were significantly induced by V. dahliae and quickly reached peak levels at different time points (Plaumann et al. 2018). In this study, we also found that FnCHIT2 from F. nilgerrensis was strongly induced after inoculation with C. gloeosporioides. According to previous studies, overexpression of chitinase genes in transgenic A. thaliana enhanced resistance to fungal infection (Hong et al. 2006) and inhibited fungal growth (Leah et al. 1991; Cao et al. 2009; Shah et al. 2010; Mercado et al. 2015; Durechova et al. 2019). According to our results, FnCHIT2 gene overexpression in A. thaliana enhanced resistance against C. higginsianum infection. Over-expression of the chitinase gene has been previously reported to impart enhanced disease resistance via two mechanisms: degradation of chitin in hyphae, which retards fungal growth, and release of pathogen-borne elicitors that induce defence reactions in plants (Prasad et al. 2013). Chitinases confer plant resistance against microbial attack by partially digesting isolated cell walls of several pathogenic fungi, such as ascomycetes, basidiomycetes, and deuteromycetes (Zhu et al. 1991; GonzáleZ et al. 2015). Furthermore, expression of the PR1 gene (SA-signalling pathway) in strawberry was up-regulated after C. gloeosporioides infection (Wang et al. 2017). In A. thaliana, the SA pathway was involved in resistance against C. higginsianum (Liu et al. 2017). According to our results, AtPR1 expression was strongly induced after inoculation with C. higginsianum in transgenic A. thaliana at 12 hpi. This finding suggests that SA-dependent pathways regulated defence response activation. Overexpression of chitinase genes in plants results in the release of pathogen-borne elicitors, which induce defence reactions in plants. In tobacco, overexpression of the chitinase gene conferred enhanced disease resistance to Trichoderma harzianum and enhanced levels of PR1 transcripts in transgenic lines compared with basal levels, suggesting the involvement of chitinase in the signal transduction of defence pathways (Voll et al. 2012). Therefore, we hypothesized that infection with C. higginsianum and overexpression of the FnCHIT2 gene in transgenic plants results in the release of pathogen-borne elicitors and induces defence responses via SA-dependent pathways. We also examined the expression pattern of the AtPDF1.2 gene (JA-signalling pathway marker gene) after infection with C. higginsianum. We found that AtPDF1.2 transcription levels in transgenic lines were reduced at 12 hpi and increased at 72 hpi compared with wild type plants. The JA-dependent signalling pathway was also activated after inoculation with C. higginsianum in transgenic plants. According to these results, SA and JA defence signalling pathways were activated in transgenic A. thaliana during C. higginsianum infection, suggesting the positive role of FnCHIT2 against Colletotrichum spp. infection. Our results are consistent with the previous findings that both SA and JA pathways are involved in plants resistance against C. higginsianum (Dana et al. 2006). Zhang et al. (2018) and Amil-Ruiz et al. (2016) reported that both SA and JA defence pathways were activated during Colletotrichum spp. infection in strawberry. We inferred that overexpression of FnCHIT2 in transgenic A. thaliana lead to increased chitinase expression and played an antifungal role by directly hydrolysing C. higginsianum chitin and subsequently activating SA and JA signalling pathways.
We also evaluated bacterial disease resistance in transgenic A. thaliana. We infected transgenic and wild type A. thaliana lines with Pst DC3000 and observed that transgenic plants exhibited enhanced resistance to Pst DC3000 and C. higginsianum. It was previously reported that CaChi2 overexpression in A. thaliana enhanced resistance to Pst DC3000 (Hong et al. 2006). In our study, transgenic lines showed reduced disease symptoms, more programmed cell death and increased O2− levels compared with wild type A. thaliana after inoculation. Reactive oxygen species (ROS) production is thought to be directly toxic to pathogens (Lambeth 2004), and restrict pathogens by triggering stomatal closure (Zhang et al. 2020a, b). In the SA-mediated signalling pathway, enhanced plant resistance against biotrophic and hemibiotrophic pathogens might be due to increased PR1 gene expression. We observed increased AtPR1 transcript levels in transgenic lines compared with wild type A. thaliana at 12–72 hpi. In transgenic lines, AtPDF1.2 exhibited increased expression after inoculation with Pst DC3000. These results suggest that overexpression of FnCHIT2 improved resistance to Pst DC3000 via activation of both SA and JA-mediated pathways and confirm that FnCHIT2 has a positive role in resistance against Pst DC3000 infection.
In the present study, we demonstrated that the FnCHIT2 gene from F. nilgerrensis was strongly induced following the inoculation of C. gloeosporioides and FnCHIT2 overexpression in A. thaliana conferred enhanced resistance to C. higginsianum and Pst DC3000. In the future, detailed studies are needed to evaluate disease resistance mechanisms against C. gloeosporiodes infection in transgenic strawberries.
Abbreviations
- F. nilgerrensis :
-
Fragaria nilgerrensis Schlecht
- C. gloeosporiodes :
-
Colletotrichum gloesporioides
- SA:
-
Salicylic acid
- JA:
-
Jasmonic acid
- A. thaliana :
-
Arabidopsis thaliana
- Pst DC3000:
-
Pseudomonas syringae Pv. tomato DC3000
- C. higginsianum :
-
Colletotrichum higginsianum
- PR1:
-
Pathogenesis-related gene 1
- PDF1.2:
-
Plant defensing 1.2
- PCR:
-
Polymerase chain reaction
- qRT-PCR:
-
Quantitative real-time PCR
- IPTG:
-
Isopropythio β-D- thiogalactoside
- PVDF:
-
Polyvinylidene difluoride
- TBST:
-
Tris-Buffered Saline Tween-20
- PEG:
-
Polyethylene glycol
- ROS:
-
Reactive oxygen species
- hpi:
-
Hours post inoculation
- dpi:
-
Days post inoculation
- NBT:
-
Nitro blue tetrazolium
References
Abeles FB, Bonhart RP, Forrence LE, Habig WH (1971) Preparation and purification of glucanase and chitinase from bean leaves. Plant Physiol 47:129–134
Akhan A, Shih DS (2004) Molecular cloning, characterization, and expression analysis of two class II chitinase genes from the strawberry plant. Plant Sci 166:753–762
Amil-Ruiz F, Garrido-Gala J, Gadea J, Blanco-Portales R, Muñoz-Mérida A, Trelles O, de Los SB, Arroyo FT, Aguado-Puig A, Romero F, Mercado JÁ, Pliego-Alfaro F, Muñoz-Blanco J, Caballero JL (2016) Partial activation of SA- and JA-defensive pathways in strawberry upon Colletotrichum acutatum interaction. Front Plant Sci 7:1036
Brotmana Y, Landau U, Pnini S, Lisec J, Balazadeh S, Mueller-Roeberd B, Zilberstein A, Willmitzera L, Chet I, Viterbo A (2012) The LysM Receptor-Like kinase LysM RLK1 is required to activate defense and abiotic-stress responses induced by overexpression of fungal chitinases in Arabidopsis Plants. Nat Plants 5:1113–1124
Cao RH, Liu XG, Gao KX, Mendgen K, Kang ZS, Gao JF, Dai Y, Wang X (2009) Mycoparasitism of endophytic fungi isolated from reed on soilborne phytopathogenic fungi and production of cell wall-degrading enzymes in vitro. Curr Microbiol 59:584–592
Cao SL, Wang Y, Li ZQ, Gao F, Zhou YJ, Zhang GF, Feng JC (2019) Genome-Wide identification and expression analyses of the chitinases under cold and osmotic stress in Ammopiptanthus nanus. Genes 10:472
Casado-Díaz A, Encinas-Villarejo S, Santos B, Schiliro E, Yuberoserrano E, Amilruiz F, Pocovi M, Pliegoalfaro F, Dorado G, Rey M, Romero F, Munozblanco J, Caballero JL (2006) Analysis of strawberry genes differentially expressed in response to Colletotrichum infection. Physiol Plantarum 128:633–650
Chalavi V, Tabaeizadeh Z (2003) Enhanced resistance to Verticillium dahliae in transgenic strawberry plants expressing a Lycopersicon chilense Chitinase gene. J Am Soc Hortic Sci 128:747–753
Chen JJ, Piao YL, Liu YM, Li XN, Piao ZY (2018) Genome-wide identification and expression analysis of chitinase gene family in Brassica rapa reveals its role in clubroot resistance. Plant Sci 270:257–276
Chopra R, Saini S (2014) Transformation of blackgram (Vigna mungo (L.) Hepper) by barley chitinase and ribosome-inactivating protein genes towards improving resistance to corynespora leaf spot fungal disease. Appl Bioch Biotech 174:2791–2800
Chung PC, Wu HY, Wang YW, Ariyawansa HA, Hu HP, Hung TH, Tzean SS, Chung CL (2020) Diversity and pathogenicity of Colletotrichum species causing strawberry anthracnose in Taiwan and description of a new species Colletotrichum miaoliense sp. nov. Sci Rep 10:14664
Clough SJ, Bent AF (1998) Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J 16:735–743
Dana MD, Pintor-Toro JA, Cubero B (2006) Transgenic tobacco plants overexpressing chitinases of fungal origin show enhanced resistance to biotic and abiotic stress agents. Plant Physiol 142:722–730
Dong X, Zhao YC, Ran X, Guo LX, Zhao DG (2017) Overexpression of a New chitinase gene EuCHIT2 enhances resistance to Erysiphe cichoracearum DC in tobacco plants. Int J Mol Sci 18:2361
Dubouzet JG, Maeda S, Sugano S, Ohtake M, Hayashi N, Ichikawa T, Kondou Y, Kuroda H, Horri Y, Matsui M, Oda K (2011) Screening for resistance against Pseudomonas syringae in rice-FOX Arabidopsis lines identified a putative receptor-like cytoplasmic kinase gene that confers resistance to major bacterial and fungal pathogens in Arabidopsis and rice. Plant Biotechnol J 9:466–485
Durechova D, Jopcik M, Rajninec M, Moravcikova J, Libantova J (2019) Expression of Drosera rotundifolia chitinase in transgenic tobacco plants enhanced their antifungal potential. Mol Biotechnol 61:916–928
Gan P, Ikeda K, Irieda H, Narusaka M, O’Connell RJ, Narusaka Y, Takano Y, Kubo Y, Shirasu K (2013) Comparative genomic and transcriptomic analyses reveal the hemibiotrophic stage shift of Colletotrichum fungi. New Phytol 197:1236–1249
GonzáleZ FF, Davey MR, Sanchez EC, Wilson ZA (2015) Conferred resistance to Botrytis cinerea in Lilium by overexpression of the RCH10 chitinase gene. Plant Cell Rep 34:1201–1209
Grover A (2012) Plant Chitinases: genetic diversity and physiological roles. Crit Rev Plant Sci 3:57–73
Gu QN, Yuan QF, Zhao D, Huang JB, Hsiang T, Wei YD, Zheng L (2019) Acetyl-coenzyme A synthetase gene ChAcs1 is essential for lipid metabolism, carbon utilization and virulence of the hemibiotrophic fungus Colletotrichum higginsianum. Mol Plant Pathol 20:107–123
Han YC, Zeng XG, Xiang FY, Zhang QH, Guo C, Chen FY, Gu YC (2018) Carbendazim sensitivity in populations of Colletotrichum gloeosporioides complex infecting strawberry and yams in Hubei Province of China. J Integr Agr 17:1391–1400
Hong JK, Hwang BK (2006) Promoter activation of pepper class II basic chitinase gene, CAChi2, and enhanced bacterial disease resistance and osmotic stress tolerance in the CAChi2-overexpressing Arabidopsis. Planta 223:433–448
Kang JN, Park MY, Kim WN, Kang HG, Jin SH, Yang DH, Min KS (2017) Resistance of transgenic zoysiagrass overexpressing the zoysiagrass class II chitinase gene Zjchi2 against Rhizoctonia solani AG2-2 (IV). Plant Biotechnol Rep 11:229–238
Karimi K, Arzanlou M, Pertot I (2019) Weeds as potential inoculum reservoir for Colletotrichum nymphaeae causing strawberry anthracnose in iran and rep-PCR fingerprinting as useful marker to differentiate C. acutatum complex on strawberry. Front Microbiol 10:129
Katagiri F, Thilmony R, He SY (2002) The Arabidopsis thaliana-Pseudomonas syringae interaction. Arabidopsis Book/Am Soc Plant Biol 1:1–35
Khan A, Nasir IA, Tabassum B, Aaliya K, Tariq M, Rao AQ (2017) Expression studies of chitinase gene in transgenic potato against Alternaria solani. Plant Cell Tiss Org 128:563–576
Kim KH, Jae-Bok Y, Hyo-Guen P, Eun Woo P, Young Ho K (2004) Structural modifications and programmed cell death of chili pepper fruit related to resistance responses to Colletotrichum gloeosporioides infection. Phytopathology 94:1295–1304
Kwon YR, Kim SH, Jung MS, Kim MS, Oh JE, Ju HW, Kim K, Vierling E, Lee H, Hong SW (2007) Arabidopsis hot2 encodes an endochitinase-like protein that is essential for tolerance to heat, salt and drought stresses. Plant J 49:184–193
Laemmli UK (1970) Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227:680–695
Lambeth JD (2004) NOX enzymes and the biology of reactive oxygen. Nat Rev Immunol 4:181
Leah R, Tommerup H, Svendsen I, Mundy J (1991) Biochemical and molecular characterization of three barley seed proteins with antifungal properties. J Biol Chem 266:1564–1573
Liu Z, Shi L, Yang S, Lin Y, Weng Y, Li X, Hussain A, Noman A, He S (2017) Functional and promoter analysis of ChiIV3, a Chitinase of pepper plant, in response to Phytophthora capsici infection. Int J Mol Sci 18:1661
Mercado JA, Barceló M, Pliego C, Rey JL, Muñoz-Blanco J, Ruano-Rosa D, López-Herrera C, Santos B, Romero-Muñoz F, Pliego-Alfaro F (2015) Expression of the β-1,3-glucanase gene bgn13.1 from trichoderma harzianum in strawberry increases tolerance to crown rot diseases but interferes with plant growth. Transgenic Res 24:979–989
Moraes SRG, Tanaka FAO, Massola NS (2013) Histopathology of Colletotrichum gloeosporioides on guava fruit (Psidium guajava L.). Rev Bras Frutic 35:657–664
Murashige T, Skoog F (1962) A revised medium for rapid growth and bioassays with tobacco tissue cultures. Physiol Plant 15:473–497
Narusaka Y, Narusaka M, Park P, Kubo Y, Hirayama T, Seki M, Shiraishi T, Ishida J, Nakashima M, Enju A, Sakurai T, Satou M, Kobayashi M, Shinozaki T (2004) RCH1, a Locus in Arabidopsis that confers resistance to the hemibiotrophic fungal pathogen Colletotrichum higginsianum. Mol Plant-Microbe In 17:749–762
Nelson BK, Cai X (2007) A multicolored set of in vivo organelle markers for co-localization studies in Arabidopsis and other plants. Plant J 51(6):1126–1136
Noguchi YJ, Mochizuki T, Sone (2002) Breeding of a new zromatic strawberry by interspecific hybridization Fragaria ananassa × F. nilgerrensis. Engei Gakkai zasshi 71:208–213
O’Connell RJ, Perfect S, Hughes B, Carzaniga R, Bailey JA, Green J (2000) Dissecting the cell biology of Colletotrichum. In: Infection processes. Colletotrichum, pp 57–76
Plaumann PL, Schmidpeter J, Dahl M, Taher LL, Koch C (2018) A dispensable chromosome is required for virulence in the hemibiotrophic plant pathogen Colletotrichum higginsianum. Front Microbiol 9:1–15
Prasad K, Bhatnagar-Mathur P, Waliyar F, Sharma KK (2013) Overexpression of a chitinase gene in transgenic peanut confers enhanced resistance to major soil borne and foliar fungal pathogens. J Plant Biochem Biot 22:222–233
Pulla RK, Lee OR, In JG, Paevin S, Kim YJ, Shim JS, Sun H, Kim YJ, Senthil K, Yang DC (2011) Identification and characterization of class I chitinase in Panax ginseng C. A Meyer Mol Biol Rep 38:95–102
Punja ZK, Zhang YY (1993) Plant Chitinases and their roles in resistance to fungal diseases. J Nematol 25:526–540
Shah MR, Mukherjee PK, Eapen S (2010) Expression of a fungal endochitinase gene in transgenic tomato and tobacco results in enhanced tolerance to fungal pathogens. Physiol Mol Biol Pla 16:39–51
Silva DE, Crous PW, Ades PK, Hyde KD, Taylor PW (2017) Life styles of Colletotrichum species and implications for plant biosecurity. Fungal Biol Rev 31(3):155–168
Singh HR, Deka M, Das S (2015) Enhanced resistance to blister blight in transgenic tea (Camellia sinensis [L.] O. Kuntze) by overexpression of class I chitinase gene from potato (Solanum tuberosum). Funct Integr Genomic 15:461–480
Suzuki T, Tanaka-Miwa Ebihara Y, Ito Y, Uematsu S (2010) Genetic polymorphism and virulence of Colletotrichum gloeosporioides isolated from strawberry (Fragaria × ananassa Duchesne). J Gen Plant Pathol 76:247–253
Takahara HD, Endl H (2009) Flow cytometric purification of Colletotrichum higginsianum biotrophic hyphae from Arabidopsis leaves for stage-specific transcriptome analysis. Plant J 59:672–683
Tariq M, Khan A, Tabassum B, Toufiq N, Bhatti MU, Riaz S, Nasir IA, Husnain T (2018) Antifungal activity of chitinase II against Colletotrichum falcatum Went.causing red rot disease in transgenic sugarcane. Turk J Biol 42:45–53
Vellicce GR, Ricci JCD, Hernández L, Castagnaro AP (2006) Enhanced resistance to Botrytis cinerea mediated by the transgenic expression of the chitinase gene ch5B in strawberry. Transgenic Res 15:57–68
Voll LM, Zell MB, Engelsdorf T, Saur A, Wheeler MG, Drincovich MF, Weber APM, Maurino VG (2012) Loss of cytosolic NADP-malic enzyme 2 in Arabidopsis thaliana is associated with enhanced susceptibility to Colletotrichum higginsianum. New Phytol 195:189–202
Wang LJ, Li SH (2006) Thermo tolerance and related antioxidant enzyme activities induced by heat acclimation and salicylic acid in grape (Vitis vinifera L.) leaves. Plant Growth Regul 49:137–144
Wang F, Zhang F, Chen MM, Liu ZH, Zhang ZH, Fu JF, Ma Y (2017) Comparative transcriptomics reveals differential gene expression related to Colletotrichum gloeosporioides resistance in the octoploid strawberry. Front Plant Sci 8:1–10
Wang M, Xu ZC, Ding Am, Kong YZ (2018) Genome-wide identification and expression profiling analysis of the xyloglucan Endotransglucosylase/Hydrolase gene family in tobacco (Nicotiana tabacum L.). Genes 9:273
Wen ZF, Yao LP, Wan R, Li Z, Liu CH, Wang XP (2015) Ectopic expression in Arabidopsis thaliana of an NB-ARC encoding putative disease resistance gene from wild Chinese Vitis pseudoreticulata enhances resistance to phytopathogenic fungi and bacteria. Front Plant Sci 6:1087
Widiastuti A, Yoshino M, Hasegawa M, Nitta Y, Sato T (2012) Heat shock-induced resistance increases chitinase-1 gene expression and stimulates salicylic acid production in melon (Cucumis melo L.). Physiol Mol Plant 82:51–55
Xie L, Zhang J, Wan Y, Hu D (2010) Identification of Colletotrichum spp. isolated from strawberry in Zhejiang Province and Shanghai City, China. J Zhejiang Univ Sci B 11:61–70
Xu J, Xu XY, Tian LL, Wang GL, Zhang XY, Wang XY, Guo WZ (2016) Discovery and identification of candidate genes from the chitinase gene family for Verticillium dahliae resistance in cotton. Sci Rep 6:29022
Zhang M, Kang HH, Zhang GQ, Chen YH, Kong XZ, Guo QF, Wang W (2015) Overexpression of TaUb2 enhances disease resistance to Pseudomonas syringae pv. tomato DC3000 in tobacco. Physiol Mol Plant Patho 90:98–104
Zhang QY, Zhang LQ, Song LL, Duan K, Li N, Wang YX, Gao QH (2016) The different interactions of Colletotrichum gloeosporioides with two strawberry varieties and the involvement of salicylic acid. Hortic Res 3:16007
Zhang LQ, Huang X, He CY, Zhang QY, Zou XH, Duan K, Gao QH (2018) Novel fungal pathogenicity and leaf defense strategies are revealed by simultaneous transcriptome analysis of Colletotrichum fructicola and strawberry infected by this fungus. Front Plant Sci 9:434
Zhang HY, Liu X, Zhang XY, Qin NN, Xu KF, Yin WH, Zheng YQ, Song YY, Zeng RS, Liu J (2020) Phosphoinositide 3-Kinase promotes oxidative burst, stomatal closure and plant immunity in bacterial invasion. Front Plant Sci. https://doi.org/10.3389/fpls.2019.01740
Zhang JX, Lei YY, Wang BT, Li S, Yu S, Wang Y, Li H, Liu YX, Ma Y, Dai HY, Wang JH, Zhang ZH (2020) The high quality of diploid strawberry (Fragaria nilgerrensis) provides new insights into anthocyanin accumulation. Plant Biotechnol J. https://doi.org/10.1111/pbi.13351
Zhu Q, Lamb CJ (1991) Isolation and characterization of a rice gene encoding a basic chitinase. Mol Gen Genet 226:289–296
Acknowledgements
This study was supported by the National Natural Science Foundation of China (31701907), the Natural Science Foundation of Fujian Province (2018J01703), the Educational and Scientific Research Program for Young and Middle-aged Instructor of Fujian Province (KLa17022A), and the Natural Science Funds for Distinguished Young Scholar of the Fujian Agriculture and Forestry University (xjq201723).
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
11816_2020_648_MOESM1_ESM.docx
Supplementary file1 (DOCX 141 KB) Supplementary Materials: Available online supporting material. Fig. S1. The nucleotide and deduced amino acid sequences of FnCHIT2 from F. nilgerrensis. The underlined sequences indicate the ChiA protein domain, and sequences in the square frame indicate initiation code and termination code, respectively. Fig. S2. Cloning of FnCHIT2 from F. nilgerrensis, generation of the 35S:FnCHIT2-YFP vector, and generation of the overexpression vector and prokaryotic expression vector. (A) PCR cloning of FnCHIT2 from wild F. nilgerrensis. a, DL 2000 DNA marker. (B). PCR product of FnCHIT2 amplification. Double digest of 35S:FnCHIT2-GFP vector with BamH I and Kpn I. c, 15Kb DNA marker; h, double digest of 35S: 35S:FnCHIT2-GFP vector with BamH I and Kpn I. (C) Double digest of 35S:FnCHIT2 over expression vector with BamH I and Kpn I. c, 15Kb DNA marker. d, Double digest of 35S:FnCHIT2 vector after FnCHIT2 under 35S promoter. D. Double digest of prokaryotic expression vector with BamH I and Kpn I. Double digest of prokaryotic expression vector after FnCHIT2 inserted into PET-32A. f, 12K DNA marker. Table S2primer sequences used in this study.
Rights and permissions
About this article
Cite this article
Wen, Z., Bai, J., Wang, L. et al. Over expression of a Chitinase 2 gene from Chinese Wild Strawberry improves resistance to anthracnose disease in transgenic Arabidopsis thaliana. Plant Biotechnol Rep 14, 725–736 (2020). https://doi.org/10.1007/s11816-020-00648-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11816-020-00648-z