Abstract
Recent advances in gene/genome editing technologies, such as engineered meganucleases (EMNs), zinc finger nucleases (ZFNs), transcription activator-like effector nuclease (TALENs) and clustered regularly interspaced palindromic repeats (CRISPR/Cas9) allowed researchers to precisely modify or mutate genes. These genome editing tools make double-strand breaks (DSB) in DNA and then repair it by employing error-prone non-homologous end joining (NHEJ) or homology directed repair (HDR) mechanism which leads to mutation in specific location in genome. Since these editing techniques are simple to use, highly efficient and specific as compared to earlier mutation methods, their use in plant biology research is increasing rapidly to enhance biotic and abiotic stress tolerance, increased nutritional value and new trait development. Here, we review the applications of EMNs, ZFNs, TALENs and CRISPR/Cas9 in various plants (cereals, vegetable, oil crops and fruits), comparison of genome editing methods and their biosafety regulations.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The ability to modify genomes in a site-specific manner has the potential to transform plant biological research by addressing the basic questions relating to plant growth, development and responses to the environment. Therefore, continuous efforts were being taken for getting desired mutants and various methods have been evolved with the time. First generation mutagenesis methods were non-specific and cause random mutations in genome. The random mutagenesis was achieved with the help of mutation breeding, direct gene transfer, transposons and Agrobacterium mediated genetic modifications. Use of physical and chemical mutagenesis in breeding is characterised by large population size for screening, very low efficiency, time consumption and hazardous nature of mutagens. Further screening of mutation is also cumbersome process that requires sequencing, tilling, denaturing high pressure liquid chromatography, etc. (McCallum et al. 2000; Caldwell et al. 2004). For transposons and T-DNA insertion, target DNA sequence with high homology is required so that homologous recombination mechanism works well. T-DNA insertion lines of many crop plants thus produced help us to study and understand various mechanism and functions in plant biology. It is also accompanied by some limitations, such as multiple insertions, vector sequences insertion, and rearrangement in chromosome. (De Buck et al. 1999; Tax and Vernon 2001). Second generation mutagenesis methods were more specific and efficient. They are based on the recombinase (Cre-LoxP and Flp-FRT systems), rare cutting restriction nucleases such as homing/meganuclease based methods. Application of Cre-lox system for gene insertion and excision was reported way back in 1990s (Russell et al. 1992; Albert et al. 1995). They largely used in production of marker free transgenic plants, gene stacking and gene cassette replacement (Nanto et al. 2009; Petolino et al. 2010; D’Halluin et al. 2013). The popular reverse genetic techniques; micro-RNAs and RNA interference (RNAi) has been commonly used for gene functional studies in plants (Reddy et al. 2012; Khandagale et al. 2016). At the same time it has some limitations, such as variation in silencing level, not stable over several generations and the off-target effect.
Meanwhile, through next generation sequencing, sequence information of large number of plants became available. This DNA sequence information is used for the development of newer and more precise and efficient genome editing tools compared to the traditional methods. Third generation of mutagenesis was characterised by very rapid and precise genome editing tools which includes zinc finger nuclease (ZFN) (Porteus and Carroll 2005), transcription activator-like effector nucleases (TALEN) (Li et al. 2011) and clustered regularly interspaced palindromic repeats/CRISPR-associated system (CRISPR)/(Cas) (Barrangou 2012). These targeted genome editing is characterised by creation of a double-stranded breaks (DSBs) at specific position of target locus to be mutated (Carroll 2011). These DSBs are highly prone to the genetic mutations during repair process. Non-homologous end joining (NHEJ) and homology directed repair (HDR) pathways are involved in the repair of these DSBs in DNA. NHEJ led to random indels of varying lengths which finally cause frame shift mutations in coding as well as regulatory sequences of DNA (Cristea et al. 2013). In case of HDR, sequence homologous to the DSB must be available which can be exogenously supplied by donor DNA (Bortesi and Fischer 2015). Therefore, these repair pathways exploited for precise gene editing or targeted genome editing. Thus, creation of DSBs is perquisite and essential step in gene editing approach. With the discovery of TALEN and CRISPR/cas9 system, there was explosion of research papers describing the application of precise gene editing from last 2 to 3 years. Though precise gene editing is a new field, few review papers have been published describing the molecular mechanism and potential applications of various gene editing tools in plants such as ZFNs (Weinthal et al. 2010), TALEN (Wright et al. 2014), CRISPR/Cas9 (Bortesi and Fischer 2015). Various steps involved in the targeted genome engineering, viz. selection of a target nucleotide sequence in the genome; generation of a nuclease construct aimed at the selected target; delivery of this construct to the cell nucleus; and analysis of produced mutations were elaborated by Sander and Joung (2014). Various software and online resources are available for designing of these gene editing tools were depicted in Table 1.
As very little or no foreign DNA is inserted during gene editing process, scientists are claiming that it may not come under traditional definition of transgenic plant, and hence there is no need of stringent biosafety regulations for release of gene edited plants (Woo et al. 2015). Thus, biosafety regulation has become a matter of debate in case of genome edited plants. Recently, Swedish Board of Agriculture stated that CRISPR/Cas9 modified plants does not come under the EU definition of genetically modified organism (GMO). Hartung and Schiemann (2014) and Wolt et al. (2015) surveyed the status of biosafety regulation of plants modified using gene editing techniques and emphasized on development of regulatory policy for genome edited crops.
Present review will discuss the use of genome editing tools for highly precise and desired modifications in plants, comparison of various genome editing tools as well as biosafety concerns associated with the genome editing technology.
Tools used in precise genome editing
Homing/meganuclease
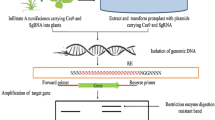
Meganucleases are kind of endonucleases which are characterised by their large recognition site of about 12–40 bp long. Such a long recognition site makes them most specific restriction enzyme. They are also called as homing endonucleases. Among the various families of meganuclease LAGLIDADG family is well understood and widely exploited for gene editing purpose. I-SceI (Saccharomyces cerevisiae), I-CreI (Chlamydomonas reinhardtii) and I-DmoI (Desulfurococcus mobilis) are a few well characterised meganucleases of LAGLIDADG family. They either form homodimers of two identical subunits (each 160–200 amino acid) or two tandem monomers joined by a linker to form single peptide (Stoddard 2011). α/β fold within structure of meganuclease could be modified to generate engineered meganuclease with desired specificity (Silva et al. 2011). The molecular structure of I–SceI is a complex thus makes it more challenging for re-engineering for customized applications, where as I–CreI is found to be more suitable for customized protein engineering (Arnould et al. 2007) (Fig. 1). Puchta et al. (1993), Salomon and Puchta (1998) pioneered the use of meganuclease in the study of DNA repair and gene integration. Further, Kirik et al. (2000) demonstrated that DSB repair by NHEJ led to knock out of genes in Arabidopsis and tobacco. To edit the gene of interest, we need to engineer the DNA recognition sites of the meganucleases. But DNA-binding domains and catalytic domain of meganucleases are not separated from each other, and therefore it becomes difficult to re-engineer the meganucleases compared to other genome targeting tools (Taylor et al. 2012). Continuous efforts of protein engineer resulted in a few successful attempts of gene editing in plants using engineered meganucleases such as Arabidopsis (Roth et al. 2012), Cotton (D’Halluin et al. 2013) and Maize (Djukanovic et al. 2013). Due to difficulty in manipulation of meganucleases, researchers were working on other simple, efficient and precise alternatives for the gene editing which gave rise to ZFN, TALEN and CRISPR.
Schematic representation of I-CreI meganuclease inducing break in double-stranded target DNA. I-CreI recombinase attaches to its target DNA at long recognition site and catalytic domain (shown in green colour) cleaves the DNA at specific target. Finally, mutations occur during the endogenous DNA repair process
ZFNs
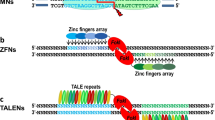
Engineered Zinc finger nuclease has become one of the powerful tools for targeted genome editing by making double-stranded break in genome. Zinc fingers are the first DNA-binding proteins which were engineered to target specific DNA sequence to make double-stranded breaks with the help of Fok1 nuclease (Porteus and Carroll 2005). NHEJ pathway is used to repair DSBs which is error prone leads in insertion and deletions (indels) during repair process. Structurally ZFNs monomer consists of two domains: DNA-binding domain, typically contain between three and six individual zinc finger repeats and each zinc finger repeat can recognize between nine and 18 base pairs, and DNA-cleavage domain, it is non-specific cleavage domain of type IIs restriction endonuclease Fok1, it is typically used as the cleavage domain in ZFNs. Two ZFN monomers attach their target sequences in reverse configuration flanking a 5–6 bp sequence of target DNA (Carroll et al. 2006). Dimerized Fok1 domains cut DNA within this flanking sequence (Fig. 2). DNA-binding domain of ZFNs generally recognizes sequence of 24–30 bp, and thus has unique or rare target sites in the genome. Therefore, specificity of ZFNs made their wide application in precision gene editing in variety of organisms. Modular assembly (Sander et al. 2007) and Oligomerized Pool ENgineering (OPEN) (Maeder et al. 2008) are two widely used methods for construction of engineered ZFNs. Use of ZFNs was initially demonstrated in model plants Arabidopsis (Lloyd et al. 2005) and tobacco (Wright et al. 2005) which ascertained scope of engineered nuclease in gene targeting and ignited the researchers for further application this technology in other crop plants. Few years later several reports had come describing the use ZFN in gene targeting in various model plants and crops plants: Arabidopsis (de Pater et al. 2013), tobacco (Townsend et al. 2009), maize (Shukla et al. 2009; Ainley et al. 2013), etc. Application of ZFN in gene targeting of different plants is described briefly in subsequent sections. ZFNs have limitation in binding to any DNA sequence due to context dependent nature of DNA as one zinc finger binds to 3 nucleotide in target DNA and off-target effect is also one limitation of ZFN.
Schematic representation of engineered zinc fingers fused with Fok1 restriction enzyme domain inducing DSBs in target DNA. Each zinc finger domain recognizes 3 nucleotides in target DNA, Fok1 nuclease is joined to C-terminal of three zinc finger domains to form a ZFN monomer. Dimerization of ZFN monomer essential for making DSBs by Fok1, thus two ZFN monomers bind to complementary strands of target DNA and cleave target DNA at the 6 bp spacer sequence in between recognition site of two monomers. Further, these DSBs are repaired through HDR or NHEJ DNA repair pathway
TALEN
TALEN (transcription activator-like effector nucleases) system for precise genome editing is named as method of year by nature methods in 2011 (Baker Becker 2012). TALENs are transcription activator-like effectors (TALEs) fused to the non-specific cleavage domain of the FokI endonuclease for binding target DNA. These TALEs are type III effector proteins secreted by phytopathogenic bacteria Xanthomonas spp. which attaches to DNA by the virtue of DNA-binding domain to alter the gene expression pattern in host plant (Boch et al. 2009). Here, DNA-binding domain has comprised conserved repeats of 34 amino acid and each repeat recognizes a single nucleotide in target DNA, whereas in case of ZFN, each repeat recognizes 3 nucleotides, and thus TALEN designing is comparatively easier and amenable to programming. Target recognition specificity of TALEs repeat is determined by the presence of repeat variable di-residue (RVD) at 12 and 13 positions of each repeat (Moscou and Bogdanove 2009). Various TALENs designing criteria and precautions for selection of target site, assembly of TALENs and their cloning were nicely elaborated by Bogdanove and Voytas (2011). With the available technology and kits one can assemble TALENs in less than 10 days. A dimerized FokI cleavage domain is responsible to make DSB, therefore two TALENs were used to target the DNA strands in an opposite orientation with proper spacing of 12–30 bp (Li et al. 2011) (Fig. 3). FokI domain self-dimerization is essential for that two separate TALE nucleases (TALEN) need to be co-expressed, it may cause off-target effect (Aouida et al. 2014). Monomeric TAL-based nucleases (mTALENs) are improvement over dimeric TALEN and it requires single domain for the activity, and thus has potential to decrease off-target cleavage by around 50%. Use of mTALENs have some benefits, such as less efforts required to design compared to dimeric TALEN, high specificity, low off-target cleavage and can be used for targeted mutagenesis of variety of organisms (Mahfouz et al. 2014). TALENs ability to target a specific genomic sequence allows highly precise targeted mutagenesis. Several research papers have been published reporting the TALEN mediated gene editing in various plants traits. Disease resistance rice (Wang et al. 2014), aroma in rice (Shan et al. 2015), oil quality in soybean (Haun et al. 2014) and browning of in potato (Clasen et al. 2016) and many more genes were targeted in model as well as other crop plants for functional studies as a proof of concept. TALENs are preferred over ZFN due to its comparative simplicity and cost effective design, and considerably less off-targeting. Though TALEN is convenient, efficient and highly specific tool for DNA editing, it is technically challenging due to need of protein engineering to alter the binding specificity of TALEs to suit our purpose or gene of interest and difficulty in multiplexing. TALEN is sensitive to methylation, and thus not able to edit methylated target (Bultmann et al. 2012). Continuous thrust of genomic craftsman for novel, easier and advanced tools resulted in the discovery of revolutionary gene editing tool called CRISPR/Cas9.
Schematic representation of engineered TALEs fused with Fok1 restriction enzyme domain that induces DSBs in target DNA. TALENs target sequence is detected by two monomers of TALE repeats. There is nuclear localisation signal at N-terminal and C-terminal gets fused with Fok1 nuclease, dimerization of monomer is necessary to make DSBs in target DNA. Further, ultimately mutations were induced at targeted site during repair of DSBs by HDR or NHEJ repair pathway
CRISPR/Cas9
Clustered regularly interspaced palindromic repeats (CRISPR)/CRISPR-associated system (Cas9) is recently emerged as a genome editing tool across eukaryotic species (Barrangou 2012). Unique CRISPR like repeats were initially discovered way back in 1987 in Eubacteria (Ishino et al. 1987) and in Archaea (Mojica et al. 1995) but later on in 2002 the term ‘CRISPR’ was coined for such repeats by Jansen et al. (2002). Earlier actual function of CRISPR was unknown, which was later experimentally proved as an adaptive immune system that protects bacteria and archaea from attack of invading bacteriophages (Barrangou et al. 2007; Wiedenheft et al. 2012). Initially it was assumed that CRISPR acts on RNA as in case RNA interference, but Marraffini and Sontheimer (2008) demonstrated that CRISPR targets DNA and it could be programmed. A few years later, Sapranauskas et al. (2011) demonstrated that CRSIPR system in one organism can be reconstituted in another distant organism. Further, Jinek et al. (2012) elucidated the molecular mechanism of CRISPR/Cas9 system in Streptococcus pyogenes. CRISPR/Cas9 system is composed of Cas9 endonuclease and two RNAs, viz. CRISPR RNA (crRNA) and trans-activating CRISPR RNA (tracrRNA), which guides the Cas9 endonuclease to its target DNA sequence flanked by a protospacer-adjacent motif (PAM) (Jinek et al. 2012) (Fig. 4). Cas9 has two cleavage domains which show the homology with RuvC and HNH nucleases. crRNA and tracrRNA are engineered to combine in one guide RNA (gRNA) which guides Cas9 nuclease to its target (Cong et al. 2013). From 2013, there is an outbreak of research on CRISPR technology from microbes, plants to human and is still continued. Many researchers demonstrated the use of CRISPR/Cas9 in plant genome editing (Shan et al. 2013; Li et al. 2013; Jiang et al. 2013; Feng et al. 2014; Ron et al. 2014). Further, Liang et al. (2016) demonstrated the rapid process for construction of CRISPR/Cas9 for multiplex editing of plant genes and parameters for most efficient gRNAs. Cas9 nuclease from Streptococcus pyogenes was most widely used in CRISPR mutagenesis but recently, Cas9 orthologues from Streptococcus thermophilus and Staphylococcus aureus were also found to be efficient in genome targeting (Steinert et al. 2015). Specificity of CRISPR is govern by small gRNA, and thus it is easy to design such small gRNA specific to particular target sequence, therefore within very short time CRISPR/Cas9 became method of choice for genome editing. Though it is simple and easy to develop and use, is accompanied limitations such as off-target effects and need of PAM sequence near the target. To minimise off-targeting, double-nicking strategy was developed, which uses mutant Cas9 enzyme (D10A) called DNA nickase. Mutations in RuvC domain converts Cas9 enzyme into DNA nickase (Cong et al. 2013). Cas9D10A which was used with a pair of equally efficient guide RNAs to reduce off-target effect by 50- to 1000-folds (Ran et al. 2013). Paired nickase mediated highly precise and efficient editing was performed in Arabidopsis (Fauser et al. 2014; Schiml et al. 2014). RNA guide Fok1 nuclease is another modification in CRISPR/Cas9 system, where fok1 nuclease was fused with catalytically inactive cas9 protein (Tsai et al. Tsai and Xue 2015). CRISPR technology can be used for gene expression modulation like RNAi and could be called as CRISPR interference (CRISPRi) (Qi et al. Qi et al. 2013a). Further length of sgRNA could be manipulated to overcome the problem of off-targeting (Cho et al. 2014). Thus, efficiency of CRISPR/Cas9 system is governed by the activity of Cas9 enzyme, designing of sgRNA, selection of target site and delivery methods used.
Schematic representation of CRISPR/Cas9 genome editing tool. sgRNA (20 nucleotide long complementary to target site) guides the Cas9 to the target DNA sequence. PAM (proto spacer adjacent motif) is needed to cut the DNA by cleavage domain of Cas9 (RuvC and HNH) to make DSBs. Thus, resulted DSBs are repaired by HDR or NHEJ to induce desired mutations (insertion, deletion, substitution) at targeted site
Applications of genome editing technologies in plants
With the advancement of precise genome editing technologies, their use in plants gene and genome modifications also increased in model plants as well as other crop plants. Initially, we discussed the genome editing in model plants, such as Arabidopsis, tobacco and rice, and then in general we described the application of genome editing in broadly in vegetable, oil crops, cereals and fruits.
Arabidopsis
It is model plant for genomic studies as lot of information available in public domain, so many researchers chose Arabidopsis for genome editing experiments. Efficiency of ZFN mutagenesis was assessed in Arabidopsis and analysis of mutants found that 78% were simple deletions of 1–52 bp, 13% were simple insertions of 1–4 bp, and 8% were deletions accompanied by insertions (Lloyd et al. 2005). ZFNs were designed using OPEN technique to target Alcohol dehydrogenase 1 (ADH1) and transparent testa 4 (TT4) genes under the oestrogen-inducible promoter resulted in 7 or 16% mutation frequencies, respectively (Zhang et al. 2010). Similarly, ABA-Insensitive4 (ABI4) (Osakabe et al. 2010) and protoporphyrinogen oxidase (de Pater et al. 2013) genes were targeted under heat shock promoter. Further, ZFNs were expressed under the egg apparatus-specific enhancer to introduce mutations in the germ line cells (Even-Faitelson et al. 2011). Use of engineered ZFNs in editing of tandemly arrayed genes was successfully demonstrated by editing the genes from the well-known RPP4 (Recognition of Peronospora Parasitica 4) resistance gene cluster, which contains eight genes. Mutation frequency was found to be 1% and deletions ranged from a few kb to 55 kb (Qi et al. 2013b).
Method and reagents for assembling of TALENs and guidelines for TAL effector and their binding sites were described by Cermak et al. (2011). Five Arabidopsis genes viz; ADH1, TT4, map kinase (MAPKKK1), ubiquitin receptor protein (DSK2B), and ATP-binding transport protein (NATA2) were targeted for TALEN induced mutagenesis. Mutation frequency was ranged from 41 to 73% of which 1.5–12% of mutations were inherited in next generation (Christian et al. 2013). Forner et al. (2015) expressed TALEN under the meristem specific promoter targeting the Clavata 3 (CLV3), few of the mutants showed loss of function of the CLV3 and slight off-target effect was seen upon re-sequencing.
Multiplex and homologous recombination-mediated editing of Arabidopsis Phytoene Desaturase (PDS3) gene with mutagenesis frequency was demonstrated by Li et al. (2013). Similarly, Zhang et al. (2015) designed a multiplex CRISPR/Cas9 platform for fast and efficient editing and demonstrated editing of six PYL families of ABA receptor genes in a single transformation experiment with 13–93% mutation frequency in T1 generation. Brassinosteroid Insensitive 1 (BRI1), Jasmonate-Zim-Domain Protein 1 (JAZ1) and Gibberellic Acid Insensitive (GAI) genes were precisely mutated using CRISPR/Cas9 system with 26–84% mutation frequency (Feng et al. 2013). Flowering Locus T (FT) and Squamosa Promoter Binding Protein-Like 4 were targeted using CRISPR/Cas9 and found that about 90% plants carrying mutation in T1 generation showed late flowering (Hyun et al. 2015). Similarly, CRISPR/Cas9 mediated gene targeting has been demonstrated at ADH1 locus (Schiml et al. 2014), GFP gene (Jiang et al. 2014), while Feng et al. (2014) analysed the patterns, efficiency, specificity and heritability of CRISPR/Cas-induced gene mutations in Arabidopsis without off-target effect. Ning et al. (2015) generated CRISPR-CAS9 mediated double mutant of NAC transcription factors (nac050/052), mutant plant flowered early which is similar to histone demethylase (jmj14) mutant; these results indicated the functional association between NAC050, NAC052 and JMJ14. CRISPR-CAS9 mediated gene targeting of auxin binding protein 1 (ABP1) revealed that ABP1 is not a key component in auxin signalling as mutants did not show any obvious developmental impairments (Gao et al. Gao et al. 2015a). Methylation of DNA makes TALEN and ZFN useless for genome targeting, but CRISPR/Cas9 is not affected by DNA modifications. AtFIS2 is imprinted in vegetative tissues by DNA methylation of promoter (Zemach et al. 2013); targeted editing of methylated CpG island in the AtFIS2 promoter resulted in significant activation of AtFIS2 transcription. Therefore, Cas9 system could act as transcriptional activator to activate silenced or imprinted genes in plants (Lowder et al. 2015).
Tobacco
Zinc finger nuclease mediated genome editing was performed in tobacco at the GFP:NPT (neomycin phosphotransferase) reporter transgene (Wright et al. 2005; Cai et al. 2009) and acetohydroxyacid synthase (SuRA and SuRB) genes (Maeder et al. 2008; Townsend et al. 2009) which showed the potential of ZFN as a genome editing tool. Mutation in SuRA and SuRB led to induce herbicidal resistance against the imidazolinone and sulfonylurea. A rapid, publicly available ‘OPEN’ strategy was used for construction of ZFN; found more efficient than modular assembly method. Petolino et al. (2010) deleted the transgene from the stably transformed plant using the ZFN. Transgene GFP-PAT construct was flanked by ZFN cleavage sites was transformed in tobacco, this transgenic plant was crossed with another tobacco plant expressing the corresponding ZFN gene and resultant progeny had a complete 4.3 kb deletion of transgene. Use of geminivirus based replicons for transient expression of zinc finger nucleases has been demonstrated by Baltes et al. (2014).
Precise genome modification has been successfully attempted in tobacco using TALEN (Mahfouz et al. 2011; Cermak et al. 2011). Further, Zhang et al. (2015) targeted acetolactate synthase (ALS) gene in tobacco protoplasts, 30% of transformed cells were mutated at ALS locus with 14% approximated gene insertion frequencies.
Targeted mutagenesis of GFP in Nicotiana benthamiana using Cas9 RNA-guided endonuclease has been demonstrated (Nekrasov et al. 2013). Similarly, Li et al. (2013) reported multiplex and homologous recombination-mediated genome editing in tobacco. Further, two genes of tobacco: NtPDS and NtPDR6 (pleiotropic drug resistance) were targeted with CRISPR/Cas9 and observed indel frequencies of 16.2–20.3% in protoplasts. Transgenic plants had 81.8 and 87.5% mutation in (NtPDS) and (NtPDR6), respectively, with no significant off-target effect (Gao et al. 2015b). Comparative study of different designed nucleases showed that cleavage of luciferase gene by CRISPR-Cas9 was more efficient than by TALENs, and also emphasized on optimization of Cas9 nuclease to make sure maximum cleavage efficiency (Johnson et al. 2015). Ali et al. (2015a) demonstrated tobacco rattle virus (TRV) mediated delivery of CRISPR/Cas9 without any off-target effect thus opened avenues for discovery of alternate methods for delivery of nucleases for genome editing. Moreover, Cas9 was delivered via TRV to manipulate in planta gene regulation by editing of transcriptional factors (Piatek et al. 2015). Marta et al. (2016) improvised the CRISPR/Cas9 by adapting the RNA-guided Cas9 system with a modular DNA construction framework (GoldenBraid). They developed a new software assembly for CRISPR/Cas9 with GoldenBraid. Feasibility and efficiency of this new approach was proved in N. benthamiana, therefore it can be extrapolated to other species for gene editing.
Rice
Sites for gene integration in chromosome are called as safe harbour loci. DNA damage and repair property of ZFN was exploited to locate these safe harbour loci in rice genome using different ZFN constructs with β-glucuronidase (GUS) as a reporter gene. 28 regions were identified for safe integration after DNA sequencing of which single is located in a non-coding region (Cantos et al. 2014).
Protocol for TALEN mediated mutagenesis was described by Chen et al. (2014) which consisted of constructing TALEN vectors, nuclease activity assay, and transformation of TALENs in callus or protoplast and identification and analysis of mutants. Using their protocol, one can generate TALEN mutant in T0 rice plants within 4–5 months. Similarly, Li et al. (2014) reported the procedure for TALEN mutagenesis in rice which included construction of modularly assembled TALENs, transformation, and screening for mutants. TALEN mediated gene editing led to generate blast resistance in rice. OsSWEET14 gene which codes for sucrose-efflux transporter family was taken over by X. oryzae pv. oryzae, using its endogenous TAL effectors thus re-route sugars from the plant cell to pathogen. Due to TALEN-based disruption of OsSWEET14, pathogens TAL effector unable to bind OsSWEET14 and ultimately resulted in disease resistance (Li et al. 2012). Similarly, Zhou et al. (Zhou et al. 2015a) edited OsSWEET13 and resultant mutants were resistant to bacterial blight. TALEN with chimeric RNA/DNA oligonucleotides was used to substitute a single base in the rice OsEPSPS (5-enolpyruvylshikimate-3-phosphate (EPSP) synthase) gene. Mutant plants were more glyphosate sensitive and seed setting was also less as compared to wild-type plants (Wang et al. 2015). 2-acetyl-1-pyrroline (2AP) is a major aroma compound in the aromatic rice and presence of nonfunctional betaine aldehyde dehydrogenase2 (BADH2) leads to synthesize 2AP and ultimately aroma in rice (Hinge et al. 2015). Disruption of BADH2 with TALEN increased grain 2AP content from 0 to 0.35–0.75 mg/kg, which was similar to the positive control aromatic variety (Shan et al. 2015). Yokoi et al. (2015) investigated classical non-homologous end joining and alternative-NHEJ repair mechanism in TALEN mediated targeted gene editing. For this they used DNA ligase4 (Lig4) mutants of rice and targeted waxy gene in rice. It was found that mutations frequency in lig4 null mutant was higher than lig4 heterozygous mutant or wild-type. Similar results were reported by Endo et al. (2015) with Cas9 mediated targeting of acetolactate synthase (ALS) gene in lig4 mutant rice calli.
Genome editing with CRISPR/Cas9 has been demonstrated in rice (Miao et al. 2013; Jiang et al. 2013; Xie and Yang 2013; Xu et al. 2014). The CRISPR/Cas9 system generated precise and homozygous gene editing in single generation in rice (Zhang et al. 2014). Zhou et al. (2014) reported large chromosomal deletions in gene cluster and heritable small genetic changes during CRISPR/Cas9 editing in rice and found 87–100% genome editing in T0 plants. Endo et al. (2016) exploited the off-target effect of CRISPR/Cas9 system to edit the multigene clusters in rice. A single guide RNA (sgRNA) was designed to recognize 20 bp sequences of cyclin-dependent kinase B2 (CDKB2). Sequence analysis of mutant revealed that single, double and triple mutants of CDKA2, CDKB1 and CDKB2 produced by a single sgRNA. Ma et al. (2015) reported a robust, convenient, efficient and multiplex CRISPR/Cas9 system for targeted genome modification in monocot and dicot plants using golden gate ligation or Gibson assembly. With this system, 46 target sites in genome were mutated with 85.4% average mutations. CRISPR/Cas9 mediated mutagenesis was demonstrated in alternative oxidase genes (OsAox1a, OsAox1b, OsAox1c) and OsBEL genes of rice, and also reported the transgene clean genome modification that inherited stably in next generation (Xu et al. 2015). Similarly, Young Seedling Albino (OsYSA) and OsROC5 genes were precisely and simultaneously targeted and sequencing of T0 plants revealed 33.3–53.3% mutation frequencies and most mutations were bi-allelic in nature (Lowder et al. 2015). OsSWEET13 is the bacterial blight susceptibility gene as it is a target for TAL effector of pathogen Xanthomonas oryzae pv. oryzae. Modification in OsSWEET13 with CRISPR/Cas9 resulted in blight resistance rice (Zhou et al. 2015a). Similarly, Wang et al. (2016) edited OsERF922 gene which encodes for ERF transcription factor to develop resistant to rice blast disease. Li et al. (2016) targeted carbon starved anther (csa) gene in rice and the resultant mutant line showed photosensitive genic male sterility. Parameters affecting frequency of CRISPR/Cas9 targeted mutagenesis in rice was investigated. A positive correlation was found between mutagenesis frequencies and level of Cas9 expression and extended culture period of rice calli (Mikami et al. 2015a). Further, Mikami et al. (2015b) also studied the CRISPR/Cas9 mutagenesis with different Cas9 and gRNA expression cassettes. It was found that mutagenesis frequencies varied with the Cas9 expression cassette used, in addition to that superiority of OsU6 promoter to OsU3 promoter in gRNA expression was also reported.
Oil seeds
A ZFN mediated genome editing was performed in soybean targeting DICER-Like (DCL) genes. The mutation showed efficient heritable transmission of the ZFN-induced mutation. A context-dependent assembly platform, a rapid and open-source method was used for generating ZFN array (Curtin et al. 2011). Further, Curtin et al. (2013) described a method for targeted mutagenesis in the paleopolyploid soybean genome using ZFN that was capable for efficiently targeting either single or multicopy gene families. Zinc finger protein has been used for transcription activation of b-ketoacyl-ACP Synthase II in Brasicca napus. Mutant plants showed significant reduction in palmitic acid, increased total C18 and reduced total saturated fatty acid contents. Therefore, designed ZFP-TFs can play an important role in modification of endogenous genes to specifically modify agronomic trait (Gupta et al. 2012).
Soybean oil is often partially hydrogenated to increase its shelf life and improve oxidative stability due to polyunsaturated fats. Fatty acid desaturase 2 genes (FAD2-1A and FAD2-1B), which converts the monounsaturated oleic acid into polyunsaturated linoleic acid. TALENs were employed to cleave conserved DNA sequences in both of these genes. In homozygous mutant plants, oleic acid increased from 20 to 80% and linoleic acid decreased from 50 to below 4% (Haun et al. 2014). Soybean genes GmPDS11 and GmPDS18 were targeted for modification using TALENs with efficiency of 17.5–21.1% under the AtU6-26 promoter (Du et al. 2015).
Jacobs et al. (2015) demonstrated the CRISPR/Cas9 genome editing in soybean. GFP transgene and other nine endogenous loci were targeted using hairy roots and somatic embryos. A novel In-Fusion® cloning strategy was developed for easier design of CRISPR/Cas9 vectors, which should be applicable for targeting any gene in any organism. A transgene (bar) and different sites of two endogenous soybean genes (GmFEI2 and GmSHR) were successfully targeted using CRISPR/Cas9 system (Cai et al. 2015). Cas9-guide RNA (gRNA) was applied for targeted mutagenesis, gene integration and gene editing of two genomic sites DD20 and DD43 on chromosome 4 of soybean with mutation frequency of 59 and 76%, respectively (Li et al. 2015). Du et al. (2015) compared the TALEN and CRISPR/Cas9 system in soybean GmPDS11 and GmPDS18 gene editing under AtU6-26 and GmU6-16 g-1 promoter. The mutation efficiency of CRISPR/Cas9 (11.7–18.1%) was slightly lower than the TALENs (17.5–21.1%) using the AtU6-26 promoter but much higher (43.4–48.1%) under GmU6-16 g-1 promoter. This suggested that Cas9/sgRNA system is more efficient for simultaneous editing of multiple homoeo alleles of PDS gene.
Cereals
Use of homing endonuclease derived from the I-CreI for specific genome editing was demonstrated in maize liguleless locus. 3% T0 plants showed mutation at desired locus (Gao et al. 2010). Similarly, Djukanovi et al. (Djukanovic et al. 2013) re-designed the I–CreI homing endonuclease from Chlamydomonas reinhardtii to recognize a 22 bp target sequence of a maize fertility gene (MS26). The homozygous mutants were male sterile due to abnormal development of tapetum cells. Martin-Ortigosa et al. (2014) developed a mesoporous silicon nanoparticle (MSN) for the intracellular delivery of Cre protein for maize genome editing. ZFN mediated genome editing in maize endogenous loci was demonstrated by Shukla et al. (2009). Further, ZFN were used for PAT (phosphinothricin acetyl transferase) and AAD1 (aryloxyalkanoate dioxygenase) herbicide resistance gene stacking at single locus in maize (Ainley et al. 2013).
Liang et al. (2014) targeted four genes viz; ZmPDS, ZmIPK1A (Isopentenyl phosphate kinase), ZmIPK, ZmMRP4 (multidrug resistance-associated proteins 4) using TALEN in protoplast system. 23.1% of protoplasts were mutagenized whereas, about 13.3–39.1% of the transgenic plants were found to be mutated. Further, TALENs were used to generate stable, heritable mutations at the maize glossy2 (gl2) locus of maize with mutation frequency of about 10% (Char et al. 2015). TALENs were also employed in genome editing of barley (Wendt et al. 2013; Gurushidze et al. 2014).
CRISPR/Cas9 mediated targeted mutagenesis was employed to target the ZmIPK gene in maize protoplasts with mutagenesis efficiency of 13.1% (Liang et al. 2014). Similarly, Svitashev et al. (2015) attempted site-specific gene insertion in liguleless1 (LIG1) gene, male fertility genes (Ms26 and Ms45), and acetolactate synthase genes (ALS1 and ALS2) of maize. Feng et al. (2016) demonstrated that gene present in heterochromatic region of the chromosome could be efficiently targeted using CRISPR/Cas9 technology. They targeted Zmzb7 as a marker and mutant plant had an albino phenotype. Similarly, phytoene synthase gene (PSY1) in maize was targeted and the mutations thus obtained showed stable inheritance in next generation (Zhu et al. 2016). Moreover, GFP transgene in Sorghum was also edited using CRISPR/Cas9 (Jiang et al. 2013). TALEN and CRISPR-Cas9 technologies were used in hexaploid bread wheat to precisely mutate three homoeoalleles of Mildew-Resistance Locus (MLO) to achieve broad spectrum resistance against the powdery mildew disease (Wang et al. 2014). Similarly, CRISPR/Cas9 mutations were successfully targeted in the inositol oxygenase (inox) and phytoene desaturase (pds) genes of wheat in suspension culture (Upadhyay et al. 2013). Barley, HvPM19 gene codes for ABA-inducible membrane protein which is involved in regulation of grain dormancy. Cas9-induced mutation in two copies HvPM19 was attempted and found 23 and 10% precise mutant frequency. They also reported off-target effects, and thus showed limitations in targeting multicopy genes in crops (Lawrenson et al. 2015).
Vegetables
DELLA gene in tomato is called as PROCERA (PRO) which negatively regulates the signalling of gibberellic acid. Lor et al. (2014) mutated PRO gene in tomato using TALEN under the control of an oestrogen-inducible promoter and resultant mutant showed phenotypes consistent with increased GA response. A pipeline for tetraploid potato genome editing based on transient expression of TALEN in protoplasts has been developed. Transfection efficiency of protoplasts was 38–39%, whereas mutation frequency in calli was 11–13% (Nicolia et al. 2015). Vacuolar invertase gene (VInv) is responsible for accumulation of reducing sugars in potato tubers during cold storage. High temperature processing results in brown, bitter-tasting products and elevated levels of acrylamide. TALENs were used to knockout VInv gene, VInv-knockout tubers had untraceable levels of reducing sugars, and reduced levels of acrylamide in chips and were lightly coloured (Clasen et al. 2016). RNAi approach was also used for the development of acrylamide free potatoes (Chawla et al. 2012).
Brooks et al. (2014) demonstrated the CRISPR/Cas9 mediated disruption of ARGONAUTE7 (SlAGO7) gene which results in needle like or wiry leaves. Functional conservation of SHOR-TROOT and SCARECROW genes between Arabidopsis and tomato was showed using CRISPR mutagenesis (Ron et al. 2014). MADS-box transcription factor encoding RIN gene regulates fruit ripening in tomato. CRISPR/Cas9 system was designed to target three regions within the gene, homozygous RIN mutant tomato plants exhibited incomplete ripening with reduction in red pigmentation than wild-type confirming the important role of RIN in ripening (Ito et al. 2015). Cermak et al. (2015) used geminivirus replicons as vector for delivery of CRISPR/Cas9 system. They reported overexpression and ectopic accumulation of anthocyanin in tissues upon insertion of strong promoter, and also found that more than 2/3 insertions were specific with no off-target effect. Geminivirus replicons were found to be more efficient than Agrobacterium for genome targeting. In Arabidopsis, GA4 knockouts exhibit dwarf phenotype due to inhibition of gibberellin biosynthesis pathway and reduced fruit dehiscence. GA4 orthologues in B. oleracea, BolC.GA4.a was targeted and reported 10% Cas9 induced mutations and also showed dwarf phenotype associated GA4 knockouts (Lawrenson et al. 2015).
Fruits and woody tree
ZFN mediated genome editing was demonstrated first time in perennial fruits, viz. Fig and Apple by targeted repairing of uidA gene under heat shock promoter. GUS staining and PCR product sequencing was employed to confirm the repaired uidA gene (Peer et al. 2015). Jia and Wang (2014) developed a novel Xanthomonas citri facilitated agro-infiltration technique to deliver CRISPR/Cas9 targeting the CsPDS gene, into sweet orange leaves. The mutation rate was approximately 3.2–3.9% without any off-target effect. Similarly, targeted editing of phytoene desaturase (PDS) gene in Apple was also demonstrated (Nishitani et al. 2016). Recently, Ren et al. (2016) demonstrated CRISPR mediated editing in Grapes cultivar ‘Chardonnay’, where they targeted L-idonate dehydrogenase (IdnDH) gene with 100% mutation frequency. There is a huge potential for gene editing in fruit crops to elucidate the mechanism behind flowering, ripening and shelf life, etc. This highly specific genome editing does not involve the introduction of foreign DNA so it will help in consumer acceptance of genome edited fruits. Kanchiswamy et al. (2015) and Xiong et al. (2015) nicely elaborated the potential application of genome editing in breeding of horticultural crops.
CRISPR/Cas9 mediated genome editing was also demonstrated in woody species like poplar (Populus tomentosa Carr). Phytoene desaturase 8 (PtoPDS) was targeted using four gRNAs. Transgenic poplar plants were albino in phenotype with mutation frequency of 51% (Fan et al. 2015). 4-coumarate:CoA ligase genes (4CL1 and 4CL2) are involved in lignin and flavonoid synthesis in poplar. 4CL1 and 4CL2 were mutated using CRISPR/Cas9 system under U6.6 promoter of Medicago with 100% efficiency having bi-allelic mutation. But another member of 4CL family, 4CL5 was not mutated despite of 89% similarity with 4CL1 due to the presence of SNP in target site near PAM sequence (Zhou et al. 2015b). CRISPR/Cas9 system is highly specific and a single nucleotide change can affect the efficacy of genome editing. These findings suggested that we need to design gRNA carefully by considering the occurrence of SNP in out crossing species. Similarly Tsai et al. (2015) published commentary entitled ‘CRISPRing into the woods’ regarding the potential of genome editing in poplar and other woody trees. Breeding of fruits and woody perennial plants is difficult due to their long generation time. With the help of this, fast and specific genome editing tools we can generate homozygous mutation in very short time and ultimately can accelerate the breeding program in perennial plants.
Plant immunity against virus
Recently, CRISPR/Cas9 system has been used to increase the resistance of plants to geminiviruses (Ali et al. 2015b; Ji et al. 2015; Baltes et al. 2015). Ali et al. (2015b) engineered sgRNAs targeting open reading frames encoding viral genes such as Rep, coat proteins and conserved non-coding intergenic region (IR) were targeted. It was reported that conserved intergenic region was most effective target for reducing the virus titre of tomato yellow leaf curl virus (TYLCV). Further, simultaneous resistance to TYLCV, beet curly top virus (BCTV) and Merremia mosaic virus (MeMV) could be achieved with the use of sgRNA targeting conserved sequence from IR region. Similarly, Ji et al. (2015) and Baltes et al. (2015) targeted different region of geminivirus genome to develop virus resistance in plants. Cotton leaf curl disease is caused by begomoviruses and is one of the major diseases of cotton. A multiplex CRISPR editing technique was developed as a broad spectrum method for control of leaf curl diseases of cotton (Iqbal et al. 2016). Chandrasekaran et al. (2016) targeted eIF4E (eukaryotic translation initiation factor 4E) gene in cucumber and the resultant homozygous mutant lines were immune to three different viruses, (Cucumber vein yellowing virus, Zucchini yellow mosaic virus and Papaya ring spot mosaic virus-W).
Chaparro-Garcia et al. (2015) postulated three mechanisms to explain reduction in virus titre and infection symptoms with the aid of CRISPR/Cas9 mediated genome editing: (1) blocking of replication due to binding of Cas9/sgRNA at the origin of replication, (2) replication may be affected due to fragmentation of viral genome by Cas9/sgRNA, and (3) non-homologous end joining (NHEJ) may incorporate error during DNA repair process.
Comparison of EMN, ZFN, TALEN and CRISPR/Cas9
Protein engineering in case of meganucleases is difficult for researchers as DNA-binding and cleavage domains are not separated (Silva et al. 2011). Engineering of DNA-binding domain is essential in case of TALEN and ZFN, and it is cost and labour intensive to have engineered protein for each gene of interest in different organism. TALENs are easier to design and engineer than ZFN, but have off-target effects whereas CRISPRs are easiest of all to design and to use and more efficient. ZFN and TALEN binding depends on protein-DNA interactions that might have repeat context dependence and methylation sensitive (Zemach et al. 2013), while CRISPR/Cas9 system depends on Watson–Crick base pairing, which is highly predictable and specific, therefore is a method of choice for genome editing (Ran et al. 2013). Similarly, dimerization of monomer is needed to make DSBs at target site in case of EMN, ZFN, TALEN, whereas single sgRNA is enough to edit multiple genes. Multiplex gene editing could be easily achieved using CRISPR to edit several genes at a time while studying gene families. With rapid progress and innovations in improvement of efficiency and preciseness of CRISPR/Cas9 editing system, it has clear edge over the other editing techniques (Khatodia et al. 2016). Nevertheless, there is no perfect answer to proceed with which of the genome editing systems and logically one needs to critically analyse different systems and choose the most appropriate for their scientific exploration (Table 2).
Biosafety regulations
ZFNs, TALENs and CRISPR/Cas9 systems are the robust genome editing techniques used for precise modification in many plant genome without or little foreign DNA insertion (Weeks et al. 2015). Woo et al. (2015) demonstrated the genome editing without insertion of foreign DNA with the help of preassembled CRISPR/Cas9 proteins and are similar to naturally occurring mutations. Kanchiswamy (2016) elaborated potential and importance of DNA-free genome editing of crops. Therefore, there is a debate in the scientific community and law makers whether genome edited plants come under the regulation of GMO or not. Scientists and policy makers have extensively discussed the issues regarding the regulation of genome edited plants. They considered various parameters related to genome editing; pathway of DNA repair (NHEJ/HDR), phenotype modified/developed, off-target effect and the available regulatory framework in various countries (European Food Safety Authority Panel on Genetically Modified Organisms 2012; Araki et al. 2014; Hartung and Schiemann 2014; Wolt et al. 2015; Araki and Ishii 2015; Swedish Board of Agriculture 2015). Several countries have already formed regulatory framework for genome editing technologies; product based and process based framework. Process based framework take into account the procedure and techniques used in genome editing as per the FAO guidelines and Cartagena protocol (Hartung and Schiemann 2014). Product based regulation gives less importance to process used, but emphasizes on public and environmental risk analysis of final product developed.
USDA in 2012 stated that plant edited using ZFNs without transgene insertions not come under regulation as in case of GMOs. Similarly in EU also, ZFN mediated genome edited crops were assessed under European Community regulations. Further, the New Zealand Environmental Protection Authority (EPA) committee announced that ZFN-1 and TALENs mediated editing of plants are not GMOs (The McGuinness Institute 2013). Recently, Swedish Board of Agriculture also stated that CRISPR-Cas9 does not fall under EU definition of GMO. Similarly, USDA also allowed cultivation and sell of CRISPR edited mushroom without passing through any regulation (Waltz 2016). Araki et al. (2014) emphasized on careful handling of genome edited crops to avoid misleading of society. Genome or transcriptome sequencing and other novel methods need to be employed for investigation of genome edited crops. Though some decisions were made in regulation of gene edited crops, still there is a need of international consensus for careful regulation of these technologies and risk management associated with them for better relationship between science/technology and the society.
Conclusion and future perspectives
Emergence of ZFNs, TALENs and CRISPR as a gene editing techniques has transformed the plant biology research as they have ability to generate highly specific and efficient mutations in short time span. These genome editing techniques are highly specific, rapid and cost effective, so can be used as support for the labour and time intensive classical plant breeding. This precise gene editing was successfully applied for functional genomics study, transcriptional regulation, disease and pest resistance and new trait development in model plants as well as cereals, vegetable, fruit crops. All the gene editing techniques have their own pros and cons, but it was seen that due to its simple, versatile nature and affordability, CRISPR/Cas9 has become a method of choice among the plant molecular biologist. It is proved by seeing the comparative number of publication churning out in plant genome editing (Fig. 5).
Though these techniques are highly specific, some degree of off-target effect has been reported but careful designing these tools and selection of target will reduce them completely. The degree of off-targeting can be overcome by designing and discovering highly specific nucleases as the development in these technologies is advancing rapidly. Zhang’s group at Broad institute recently discovered Cpf1, an another nuclease for CRISPR editing which is smaller compare to earlier reported nucleases and require single RNA for activity (Zetsche et al. 2015). Such advances and discoveries of different new nucleases have widened our choices for editing trait of our interest. Similarly, various modes of delivery of engineered nuclease into the cell were used and each one has its own properties, such as reduced off-targeting, efficiency and DNA-free editing. It was reported that ribonucleoprotein mediated delivery of nucleases reduced off-target effect and also resulted in DNA-free editing in plants (Woo et al. 2015). Still it is a challenge to completely reduce the off-targeting as well as accurate detection of off-targeting in whole genome. Further, genome editing tools enable researchers to study gene expression at various spatiotemporal stages of plant development by modulating the cis-acting element and transcription factors involved expression of trait of interest (Piatek et al. 2015; Chandrasekaran et al. 2016). It is also possible to edit or modify multiple genes at a time. Therefore, it can be a powerful tool for functional genomic analysis of plants. There is need of sgRNA libraries for genome wide detection of multiple mutations in a single experiment like animal system (Shalem et al. 2014). Such large sgRNA libraries for plant system will further improve the speed and accuracy of detection of mutation in coming days.
The increase in use of genome editing tools raised the regulatory issues as there is a debate on whether genome edited plant considered as GMO or not. There are differences among the regulatory bodies of the world on product based or process based approach to be used for regulation. So, there is a need of harmony at international level for regulation of genome editing crops to access the potential risk associated with them. Proactive discussion between researchers, public and regulatory institutions will be useful for regulation and as well as social acceptance of genome edited crops. Overall, these genome editing tools enhanced our ability to edit genome for better understanding of biological processes and development of desired traits in plant species.
References
Ainley WM, Sastry-Dent L, Welter ME, Murray MG, Zeitler B, Amora R et al (2013) Trait stacking via targeted genome editing. Plant Biotech J 11:1126–1134
Albert H, Dale EC, Lee E, Ow DW (1995) Site-specific integration of DNA into wild-type and mutant lox sites placed in the plant genome. Plant J 7:649–659
Ali Z, Abul-faraj A, Piatek M, Mahfouz M (2015a) Activity and specificity of TRV-mediated gene editing in plants. Plant Signal Behav. doi:10.1080/15592324.2015.1044191
Ali Z, Abulfaraj A, Idris A, Ali S, Tashkandi M, Mahfouz MM (2015b) CRISPR/Cas9- mediated viral interference in plants. Genome Biol 16:238. doi:10.1186/s13059-015-0799-6
Aouida M, Piatek MJ, Bangarusamy DK, Mahfouz MM (2014) Activities and specificities of homodimeric TALENs in Saccharomyces cerevisiae. Curr Genet 60:61–74
Araki M, Ishii T (2015) Towards social acceptance of plant breeding by genome editing. Trends Plant Sci 20(3):145–149
Araki M, Nojima K, Ishii T (2014) Caution required for handling genome editing technology. Trends Biotechnol 32(5):234–237
Arnould S, Perez C, Cabaniols JP, Smith J, Gouble A, Grizot S, Epinat JC, Duclert A, Duchateau P, Paques F (2007) Engineered I-CreI derivatives cleaving sequences from the human XPC gene can induce highly efficient gene correction in mammalian cells. J Mol Biol 371:49–65
Baltes NJ, Gil-Humanes J, Cermak T, Atkins PA, Voytas DF (2014) DNA replicons for plant genome engineering. Plant Cell 26:151–163
Baltes NJ, Hummel AW, Konecna E, Cegan R, Bruns AN, Bisaro DM et al (2015) Conferring resistance to geminiviruses with the CRISPR-Cas prokaryotic immune system. Nat Plants. doi:10.1038/nplants.2015.145
Barrangou R (2012) RNA-mediated programmable DNA cleavage. Nat Biotechnol 30:836–838
Barrangou R, Fremaux C, Deveau H, Richards M, Boyaval P, Moineau S, Romero DA, Horvath P (2007) CRISPR provides acquired resistance against viruses in prokaryotes. Science 315:1709–1712
Becker M (2012) Method of the year 2011. Nat Methods. doi:10.1038/nmeth.1852
Boch J, Scholze H, Schornack S, Landgraf A, Hahn S, Kay S, Lahaye T, Nickstadt A, Bonas U (2009) Breaking the code of DNA-binding specificity of TAL-type III effectors. Science 326:1509–1512
Bogdanove AJ, Voytas DF (2011) TAL effectors: customizable proteins for DNA targeting. Science 333:1843–1846
Bortesi L, Fischer R (2015) The CRISPR/Cas9 system for plant genome editing and beyond. Biotech Adv 33:41–52
Brooks C, Nekrasov V, Lippman ZB, Van Eck J (2014) Efficient gene editing in tomato in the first generation using the clustered regularly interspaced short palindromic repeats/CRISPR-associated9 system. Plant Physiol 166:1292–1297
Bultmann S, Morbitzer R, Schmidt CS, Thanisch K, Spada F, Elsaesser J, Lahaye T, Leonhardt H (2012) Targeted transcriptional activation of silent oct4 pluripotency gene by combining designer TALEs and inhibition of epigenetic modifiers. Nucleic Acids Res 40:5368–5377
Cai CQ, Doyon Y, Ainley WM, Miller JC, Dekelver RC, Moehle EA, Rock JM et al (2009) Targeted transgene integration in plant cells using designed zinc finger nucleases. Plant Mol Biol 69:699–709
Cai Y, Chen L, Liu X, Sun S, Wu C, Jiang B et al (2015) CRISPR/Cas9-mediated genome editing in soybean hairy roots. PLoS One. doi:10.1371/journal.pone.0136064
Caldwell D, McCallum N, Shaw P, Muehlbauer GJ, Marshall DF, Waugh R (2004) A structured mutant population for forward and reverse genetics in Barley (Hordeum vulgare L.). Plant J 40:143–150
Cantos C, Francisco P, Trijatmiko KR, Slamet-Loedin I, Chadha-Mohanty PK (2014) Identification of “safe harbor” loci in indica rice genome by harnessing the property of zinc-finger nucleases to induce DNA damage and repair. Front Plant Sci. doi:10.3389/fpls.2014.00302
Carroll D (2011) Genome engineering with zinc-finger nucleases. Genetics 188:773–782
Carroll D, Morton JJ, Beumer KJ, Segal DJ (2006) Design, construction and in vitro testing of zinc finger nucleases. Nat Protoc 1(3):1329–1341
Cermak T, Doyle EL, Christian M, Wang L, Zhang Y, Schmidt C, Baller JA, Somia NV, Bogdanove AJ, Voytas DF (2011) Efficient design and assembly of custom TALEN and other TAL effector-based constructs for DNA targeting. Nucleic Acids Res. doi:10.1093/nar/gkr218
Cermak T, Baltes NJ, Čegan R, Zhang Y, Voytas DF (2015) High-frequency, precise modification of the tomato genome. Genome Biol. doi:10.1186/s13059-015-0796-9
Chandrasekaran J, Brumin M, Wolf D, Leibman D, Klap C, Pearlsman M, Sherman A, Arazi T, Gal-On A (2016) Development of broad virus resistance in non-transgenic cucumber using CRISPR/Cas9 technology. Mol Plant Pathol. doi:10.1111/mpp.12375
Chaparro-Garcia A, Kamoun S, Nekrasov V (2015) Boosting plant immunity with CRISPR/Cas. Genome Biol. doi:10.1186/s13059-015-0829-4
Char SN, Unger-Wallace E, Frame B, Briggs SA, Main M, Spalding MH, Vollbrecht E, Wang K, Yang B (2015) Heritable site-specific mutagenesis using TALENs in maize. Plant Biotech J. doi:10.1111/pbi.12344
Chawla R, Shakya R, Rommens CM (2012) Tuber-specific silencing of asparagine synthetase-1 reduces the acrylamide-forming potential of potatoes grown in the field without affecting tuber shape and yield. Plant Biotech J 10:913–924
Chen K, Shan Q, Gao C (2014) An efficient TALEN mutagenesis system in rice. Methods 69:2–8
Cho SW, Kim S, Kim Y, Kweon J, Kim HS, Bae S, Kim JS (2014) Analysis of off-target effects of CRISPR/Cas-derived RNA-guided endonucleases and nickases. Genome Res 24:132–141
Christian M, Qi Y, Zhang Y, Voytas DF (2013) Targeted mutagenesis of Arabidopsis thaliana using engineered TAL effector nucleases. G3 (Bethesda) 3:1697–1705
Clasen BM, Stoddard TJ, Luo S, Demorest ZL, Li J, Cedrone F, Tibebu R et al (2016) Improving cold storage and processing traits in potato through targeted gene knockout. Plant Biotech J. doi:10.1111/pbi.12370
Cong L, Ran FA, Cox D, Lin S, Barretto R, Habib N, Hsu PD, Wu X, Jiang W, Marraffini LA, Zhang F (2013) Multiplex genome engineering using CRISPR/Cas systems. Science 339:819–823
Cristea S, Freyvert Y, Santiago Y, Holmes MC, Urnov FD, Gregory PD et al (2013) In vivo cleavage of transgene donors promotes nuclease-mediated targeted integration. Biotechnol Bioeng 110:871–880
Curtin SJ, Zhang F, Sander JD, Haun WJ, Starker C, Baltes NJ et al (2011) Targeted mutagenesis of duplicated genes in soybean with zinc-finger nucleases. Plant Physiol 156:466–473
Curtin SJ, Anderson JE, Starker CG, Baltes NJ, Mani D, Voytas DF, Stupar RM (2013) Targeted mutagenesis for functional analysis of gene duplication in legumes. Methods Mol Biol 1069:25–42
D’Halluin K, Vanderstraeten C, Van Hulle J, Rosolowska J, Van Den Brande I, Pennewaert A, D’Hont K, Bossut M, Jantz D, Ruiter R, Broadhvest J (2013) Targeted molecular trait stacking in cotton through targeted double-strand break induction. Plant Biotech J 11:933–941
De Buck S, Jacobs A, Van Montagu M, Depicker A (1999) The DNA sequences of T-DNA junctions suggest that complex T-DNA loci are formed by a recombination process resembling T-DNA integration. Plant J 20:295–304
de Pater S, Pinas JE, Hooykaas PJ, Van der Zaal BJ (2013) ZFN mediated gene targeting of the Arabidopsis protoporphyrinogen oxidase gene through Agrobacterium-mediated floral dip transformation. Plant Biotech J 11:510–515
Djukanovic V, Smith J, Lowe K, Yang M, Gao H, Jones S, Nicholson MG, West A, Lape J, Bidney D, Carl Falco S, Jantz D, Alexander L (2013) Male-sterile maize plants produced by targeted mutagenesis of the cytochrome P450-like gene (MS26) using a re-designed I-CreI homing endonuclease. Plant J 76:888–899
Du H, Zeng X, Zhao M, Cui X, Wang Q, Yang H, Cheng H, Yu D (2015) Efficient targeted mutagenesis in soybean by TALENs and CRISPR/Cas9. J Biotech. doi:10.1016/j.jbiotec.2015.11.005
EFSA Panel on Genetically Modified Organisms (2012) Scientific opinion addressing the safety assessment of plants developed using zinc finger nuclease 3 and other site-directed nucleases with similar function. EFSA J 10:2943
Endo M, Mikami M, Toki S (2015) Multi gene knockout utilizing off-target mutations of the CRISPR/Cas9 system in rice. Plant Cell Physiol 56:41–47
Endo M, Mikami M, Toki S (2016) Bi-allelic gene targeting in rice. Plant Physiol. doi:10.1104/pp.15.01663
Even-Faitelson L, Samach A, Melamed-Bessudo C, Avivi-Ragolsky N, Levy AA (2011) Localized egg-cell expression of effector proteins for targeted modification of the Arabidopsis genome. Plant J 68:929–937
Fan D, Liu T, Li C, Jiao B, Li S, Hou Y, Luo K (2015) Efficient CRISPR/Cas9-mediated targeted mutagenesis in Populus in the first generation. Sci Rep. doi:10.1038/srep12217
Fauser F, Schiml S, Puchta H (2014) Both CRISPR/Cas-based nucleases and nickases can be used efficiently for genome engineering in Arabidopsis thaliana. Plant J 79:348–359. doi:10.1111/tpj.12554
Feng Z, Zhang B, Ding W, Liu X, Yang DL, Wei P, Cao F, Zhu S, Zhang F, Mao Y, Zhu JK (2013) Efficient genome editing in plants using a CRISPR/Cas system. Cell Res 23:1229–1232
Feng Z, Mao Y, Xu N, Zhang B, Wei P, Yang DL, Wang Z, Zhang Z, Zheng R, Yang L, Zeng L, Liu X, Zhu JK (2014) Multi-generation analysis reveals the inheritance, specificity, and patterns of CRISPR/Cas induced gene modifications in Arabidopsis. PNAS 111:4632–4637
Feng C, Yuan J, Wang R, Liu Y, Birchler JA, Han F (2016) Efficient targeted genome modification in maize using CRISPR/Cas9 system. J Genet Genomics 43:37–43
Forner J, Pfeiffer A, Langenecker T, Manavella P, Lohmann JU (2015) Germline- transmitted genome editing in Arabidopsis thaliana using TAL-Effector-Nucleases. PLoS One. doi:10.1371/journal.pone.0121056
Gao H, Smith J, Yang M, Jones S, Djukanovic V, Nicholson MG, West A, Bidney D, Falco SC, Jantz D, Lyznik LA (2010) Heritable targeted mutagenesis in maize using a designed endonuclease. Plant J 61:176–187
Gao J, Wang G, Ma S, Xie X, Wu X, Zhang X, Wu Y, Zhao P, Xia Q (2015a) CRISPR/Cas9-mediated targeted mutagenesis in Nicotiana tabacum. Plant Mol Biol 87:99–110
Gao Y, Zhang Y, Zhang D, Dai X, Estelle M, Zhao Y (2015b) Auxin binding protein 1 (ABP1) is not required for either auxin signalling or Arabidopsis development. PNAS 112:2275–2280
Gupta M, DeKelver RC, Palta A, Clifford C, Gopalan S, Miller JC, Novak S, Desloover D et al (2012) Transcriptional activation of Brassica napus beta-ketoacyl-ACP synthase II with an engineered zinc finger protein transcription factor. Plant Biotech J 10:783–791
Gurushidze M, Hensel G, Hiekel S, Schedel S, Valkov V, Kumlehn J (2014) True-breeding targeted gene knock-out in barley using designer TALE nuclease in haploid cells. PLoS One. doi:10.1371/journal.pone.0092046
Hartung F, Schiemann J (2014) Precise plant breeding using new genome editing techniques: opportunities, safety and regulation in the EU. Plant J 78:742–752
Haun W, Coffman A, Clasen BM, Demorest ZL, Lowy A, Ray E, Retterath A, Stoddard T, Juillerat A, Cedrone F, Mathis L, Voytas DF, Zhang F (2014) Improved soybean oil quality by targeted mutagenesis of the fatty acid desaturase 2 gene family. Plant Biotech J 12:934–940
Hinge V, Patil H, Nadaf AB (2015) Comparative characterization of aroma volatiles and related gene expression analysis at vegetative and mature stages in basmati and non-basmati rice (Oryza sativa L.) cultivars. Appl. Biochem. doi:10.1007/s12010-015-1898-2
Hyun Y, Kim J, Cho SW, Choi Y, Kim JS, Coupland G (2015) Site-directed mutagenesis in Arabidopsis thaliana using dividing tissue-targeted RGEN of the CRISPR/Cas system to generate heritable null alleles. Planta 241:271–284. doi:10.1007/s00425-014-2180-5
Iqbal Z, Sattar MN, Shafiq M (2016) CRISPR/Cas9: A tool to circumscribe cotton leaf curl disease. Front Plant Sci. doi:10.3389/fpls.2016.00475
Ishino Y, Shinagawa H, Makino K, Amemura M, Nakata A (1987) Nucleotide sequence of the iap gene, responsible for alkaline phosphatase isozyme conversion in E coli and identification of the gene product. J Bacteriol 169:5429–5433
Ito Y, Nishizawa-Yokoi A, Endo M, Mikami M, Toki S (2015) CRISPR/Cas9-mediated mutagenesis of the RIN locus that regulates tomato fruit ripening. Biochem Biophys Res Commun. doi:10.1016/j.bbrc.2015.09.117
Jacobs TB, LaFayette PR, Schmitz RJ, Parrott WA (2015) Targeted genome modifications in soybean with CRISPR/Cas9. BMC Biotechnol. doi:10.1186/s12896-015-0131-2
Jansen R, Embden JDAV, Gaastra W, Schouls LM (2002) Identification of genes that are associated with DNA repeats in prokaryotes. Mol Microbiol 43:1565–1575
Ji X, Zhang H, Zhang Y, Wang Y, Gao C (2015) Establishing a CRISPR-Cas-like immune system conferring DNA virus resistance in plants. Nat Plants. doi:10.1038/nplants.2015.144
Jia H, Wang N (2014) Targeted genome editing of sweet orange using Cas9/sgRNA. PLoS One. doi:10.1371/journal.pone.0093806
Jiang W, Zhou H, Bi H, Fromm M, Yang B, Weeks DP (2013) Demonstration of CRISPR/Cas9/sgRNA-mediated targeted gene modification in Arabidopsis, tobacco, sorghum and rice. Nucleic Acids Res. doi:10.1093/nar/gkt780
Jiang W, Yang B, Weeks DP (2014) Efficient CRISPR/Cas9-mediated gene editing in Arabidopsis thaliana and inheritance of modified genes in the T2 and T3 generations. PLoS One. doi:10.1371/journal.pone.0099225
Jinek M, Chylinski K, Fonfara I, Hauer M, Doudna JA, Charpentier E (2012) A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337:816–821
Johnson RA, Gurevich V, Filler S, Samach A, Levy AA (2015) Comparative assessments of CRISPR-Cas nucleases’ cleavage efficiency in planta. Plant Mol Biol 87:143–156
Kanchiswamy CN (2016) DNA-free genome editing methods for targeted crop improvement. Plant Cell Rep. doi:10.1007/s00299-016-1982-2
Kanchiswamy CN, Sargent DJ, Velasco R, Maffei ME, Malnoy M (2015) Looking forward to genetically edited fruit crops. Trends Biotechnol 33(2):62–63
Khandagale KS, Zanan RL, Nadaf AB (2016) RNA interference and targeted genome editing for improvement of rice (Oryza sativa L.). Israel J Plant Sci (communicated)
Khatodia S, Bhatotia K, Passricha N, Khurana SMP, Tuteja N (2016) The CRISPR/Cas genome-editing tool: application in improvement of crops. Front Plant Sci. doi:10.3389/fpls.2016.00506
Kirik A, Salomon S, Puchta H (2000) Species-specific double-strand break repair and genome evolution in plants. EMBO J 19:5562–5566
Lawrenson T, Shorinola O, Stacey N, Li C, Østergaard L, Patron N, Uauy C, Harwood W (2015) Induction of targeted, heritable mutations in barley and Brassica oleracea using RNA-guided Cas9 nuclease. Genome Biol. doi:10.1186/s13059-015-0826-7
Li T, Huang S, Zhao X, Wright DA, Carpenter S, Spalding MH, Yang B (2011) Modularly assembled designer TAL effector nucleases for targeted gene knockout and gene replacement in eukaryotes. Nucleic Acids Res 39(14):6315–6325
Li T, Liu B, Spalding MH, Weeks DP, Yang B (2012) High-efficiency TALEN-based gene editing produces disease-resistant rice. Nat Biotechnol 30:390–392
Li JF, Norville JE, Aach J, McCormack M, Zhang D, Bush J, Church GM, Sheen J (2013) Multiplex and homologous recombination mediated genome editing in Arabidopsis and Nicotiana benthamiana using guide RNA and Cas9. Nat Biotechnol 31:688–691
Li T, Liu B, Chen CY, Yang B (2014) TALEN utilization in rice genome modifications. Methods 69:9–16
Li Z, Liu ZB, Xing A, Moon BP, Koellhoffer JP, Huang L, Ward RT, Clifton E, Falco SC, Cigan AM (2015) Cas9-guide RNA directed genome editing in Soybean. Plant Physiol. doi:10.1104/pp.15.00783
Li Q, Zhang D, Chen M, Liang W, Wei J, Qi Y, Yuan Z (2016) Development of japonica photo-sensitive genic male sterile rice lines by editing carbon starved anther using CRISPR/Cas9. J Genet Genomics. doi:10.1016/j.jgg.2016.04.011
Liang Z, Zhang K, Chen K, Gao C (2014) Targeted mutagenesis in Zea mays using TALENs and the CRISPR/Cas system. J Genet Genomics 41:63–68
Liang G, Zhang H, Lou D, Yu D (2016) Selection of highly efficient sgRNAs for CRISPR/Cas9 based plant genome editing. Sci Rep. doi:10.1038/srep21451
Lloyd A, Plaisier CL, Carroll D, Drews GN (2005) Targeted mutagenesis using zinc-finger nucleases in Arabidopsis. PNAS 102:2232–2237
Lor VS, Starker CG, Voytas DF, Weiss D, Olszewski NE (2014) Targeted mutagenesis of the tomato PROCERA gene using TALENs. Plant Physiol 166:1288–1291
Lowder LG, Zhang D, Baltes NJ, Paul JW, Tang X, Zheng X, Voytas DF, Hsieh TF, Zhang Y, Qi Y (2015) A CRISPR/Cas9 toolbox for multiplexed plant genome editing and transcriptional regulation. Plant Physiol. doi:10.1104/pp.15.00636
Ma XL, Zhang Q, Zhu Q, Liu W, Chen Y, Qiu R et al (2015) A robust CRISPR/Cas9 system for convenient, high-efficiency multiplex genome editing in monocot and dicot plants. Mol Plant. doi:10.1016/j.molp.2015.04.007
Maeder ML, Thibodeau-Beganny S, Osiak A, Wright DA, Anthony RM et al (2008) Rapid “open-source” engineering of customized zinc-finger nucleases for highly efficient gene modification. Mol Cell 31:294–301
Mahfouz MM, Li L, Shamimuzzaman M, Wibowo A, Fang X, Zhu JK (2011) De novo-engineered transcription activator-like effector (TALE) hybrid nuclease with novel DNA-binding specificity creates double-strand breaks. PNAS 108:2623–2628
Mahfouz MM, Piatek A, Stewart CN Jr (2014) Genome engineering via TALENs and CRISPR/Cas9 systems: challenges and perspectives. Plant Biotech J 12:1006–1014
Marraffini LA, Sontheimer EJ (2008) CRISPR interference limits horizontal gene transfer in staphylococci by targeting DNA. Science 322:1843–1845
Marta V, Joan MB, Asun F, Pello Z, Jose B, Antonio G, Diego O (2016) A modular toolbox for gRNA–Cas9 genome engineering in plants based on the GoldenBraid standard. Plant Methods. doi:10.1186/s13007-016-0101-2
Martin-Ortigosa S, Peterson DJ, Valenstein JS, Lin VS, Trewyn BG, Lyznik LA, Wang K (2014) Mesoporous silica nanoparticle-mediated intracellular cre protein delivery for maize genome editing via lox-P site excision. Plant Physiol 164:537–547
McCallum CM, Comai L, Greene EA, Henikoff S (2000) Targeted screening for induced mutations. Nat Biotechnol 18:455–457
Miao J, Guo D, Zhang J, Huang Q, Qin G, Zhang X, Wan J, Gu H, Qu LJ (2013) Targeted mutagenesis in rice using CRISPR-Cas system. Cell Res 23:1233–1236
Mikami M, Toki S, Endo M (2015a) Comparison of CRISPR/Cas9 expression constructs for efficient targeted mutagenesis in rice. Plant Mol Biol. doi:10.1007/s11103-015-0342-x
Mikami M, Toki S, Endo M (2015b) Parameters affecting frequency of CRISPR/Cas9 mediated targeted mutagenesis in rice. Plant Cell Rep. doi:10.1007/s00299-015-1826-5
Mojica FJM, Ferrer C, Juez G, Rodrı´guez-Valera F (1995) Long stretches of short tandem repeats are present in the largest replicons of the Archaea Haloferax mediterranei and Haloferax volcanii and could be involved in replicon partitioning. Mol Microbiol 17:85–93
Moscou MJ, Bogdanove AJ (2009) A simple cipher governs DNA recognition by TAL effectors. Science 326:1501
Nanto K, Sato K, Katayama Y, Ebinuma H (2009) Expression of a transgene exchanged by the recombinase-mediated cassette exchange (RMCE) method in plants. Plant Cell Rep 28:777–785
Nekrasov V, Staskawicz B, Weigel D, Jones JD, Kamoun S (2013) Targeted mutagenesis in the model plant Nicotiana benthamiana using Cas9 RNA-guided endonuclease. Nat Biotechnol 31:691–693
Nicolia A, Proux-Wera E, Ahman I, Onkokesung N, Andersson M, Andreasson E, Zhu LH (2015) Targeted gene mutation in tetraploid potato through transient TALEN expression in protoplasts. J Biotechnol 204:17–24
Ning YQ, Ma ZY, Huang HW, Mo H, Zhao TT, Li L, Cai T, Chen S, Ma L, He XJ (2015) Two novel NAC transcription factors regulate gene expression and flowering time by associating with the histone demethylase JMJ14. Nucleic Acids Res 43:1469–1484
Nishitani C, Hirai N, Komori S, Wada M, Okada K, Osakabe K, Yamamoto T, Osakabe Y (2016) Efficient Genome Editing in Apple Using a CRISPR/Cas9 system. Sci Rep 6:31481. doi:10.1038/srep31481
Osakabe K, Osakabe Y, Toki S (2010) Site-directed mutagenesis in Arabidopsis using custom-designed zinc finger nucleases. PNAS 107:12034–12039
Peer R, Rivlin G, Golobovitch S, Lapidot M, Gal-On A, Vainstein A, Tzfira T, Flaishman MA (2015) Targeted mutagenesis using zinc-finger nucleases in perennial fruit trees. Planta 241:941–951
Petolino JF, Worden A, Curlee K, Connell J, Moynahan TL, Larsen C, Russell S (2010) Zinc finger nuclease-mediated transgene deletion. Plant Mol Biol 73:617–628
Piatek A, Ali Z, Baazim H, Li L, Abulfaraj A, Al-Shareef S, Aouida M, Mahfouz MM (2015) RNA-guided transcriptional regulation in planta via synthetic dCas9-based transcription factors. Plant Biotechnol J 13:578–589
Porteus MH, Carroll D (2005) Gene targeting using zinc finger nucleases. Nat Biotechnol 23:967–973
Puchta H, Dujon B, Hohn B (1993) Homologous recombination in plant-cells is enhanced by in vivo induction of double-strand breaks into DNA by a site-specific endonuclease. Nucleic Acids Res 21:5034–5040
Qi LS, Larson MH, Gilbert LA, Doudna JA, Weissman JS, Arkin AP, Lim WA (2013a) Repurposing CRISPR as an RNA-guided platform for sequence-specific control of gene expression. Cell 152:1173–1183. doi:10.1016/j.cell.2013.02.022
Qi Y, Li X, Zhang Y, Starker CG, Baltes NJ, Zhang F, Sander JD, Reyon D, Joung JK, Voytas DF (2013b) Targeted deletion and inversion of tandomly arrayed genes in Arabidopsis thaliana using zinc finger nucleases. G3 (Bethesda) 3:1707–1715
Ran FA, Hsu PD, Wright J, Agarwala V, Scott DA, Zhang F (2013) Genome engineering using the CRISPR-Cas9 system. Nat Protoc 8:2281–2308
Reddy DCL, Radhika V, Bharadwaj A, Khandagale KS, Aswath C (2012) miRNAs in brinjal (Solanum melongena) mined through an in silico approach. J Hort Sci Biotechnol 87(2):186–192
Ren C, Liu X, Zhang Z, Wang Y, Duan W, Li S, Liang Z (2016) CRISPR/Cas9-mediated efficient targeted mutagenesis in Chardonnay (Vitis vinifera L.). Sci Rep 6:32289. doi:10.1038/srep32289
Ron M, Kajala K, Pauluzzi G, Wang D, Reynoso MA, Zumstein K, Garcha J, Winte S, Masson H, Inagaki S, Federici F, Sinha N, Deal RB, Bailey-Serres J, Brady SM (2014) Hairy root transformation using Agrobacterium rhizogenes as a tool for exploring cell type-specific gene expression and function using tomato as a model. Plant Physiol 166:455–469
Roth N, Klimesch J, Dukowic-Schulze S, Pacher M, Mannuss A, Puchta H (2012) The requirement for recombination factors differs considerably between different pathways of homologous double-strand break repair in somatic plant cells. Plant J 72:781–790
Russell SH, Hoopes JL, Odell JT (1992) Directed excision of a transgene from the plant genome. Mol Gen Genet 234:49–59
Salomon S, Puchta H (1998) Capture of genomic and T-DNA sequences during double-strand break repair in somatic plant cells. EMBO J 17:6086–6095
Sander SD, Joung JK (2014) CRISPR-Cas systems for genome editing, regulation and targeting. Nat Biotechnol 32(4):347–355
Sander JD, Zaback P, Joung JK, Voytas DF, Dobbs D (2007) Zinc Finger Targeter (ZiFiT): an engineered zinc finger/target site design tool. Nucleic Acids Res 35:599–605
Sapranauskas R, Gasiunas G, Fremaux C, Barrangou R, Horvath P, Siksnys V (2011) The Streptococcus thermophilus CRISPR/Cas system provides immunity in Escherichia coli. Nucleic Acids Res 39:9275–9282
Schiml S, Fauser F, Puchta H (2014) The CRISPR/Cas system can be used as nuclease for in planta gene targeting and as paired nickases for directed mutagenesis in Arabidopsis resulting in heritable progeny. Plant J 80:1139–1150
Shalem O, Sanjana NE, Hartenian E, Shi X, Scott DA, Mikkelsen TS, Heckl D, Ebert BL, Root DE, Doench JG et al (2014) Genome-scale CRISPR-Cas9 knockout screening in human cells. Science 343:84–87
Shan Q, Wang Y, Li J, Zhang Y, Chen K, Liang Z et al (2013) Targeted genome modification of crop plants using a CRISPR-Cas system. Nat Biotechnol 31:686–688
Shan Q, Zhang Y, Chen K, Zhang K, Gao C (2015) Creation of fragrant rice by targeted knockout of the OsBADH2 gene using TALEN technology. Plant Biotech J 13:791–800
Shukla VK, Doyon Y, Miller JC, DeKelver RC, Moehle EA, Worden SE et al (2009) Precise genome modification in the crop species Zea mays using zinc-finger nucleases. Nature 459:437–441
Silva G, Poirot L, Galetto R, Smith J, Montoya G, Duchateau P, Pâques F (2011) Meganucleases and other tools for targeted genome engineering: perspectives and challenges for gene therapy. Curr Gene Ther 11:11–27
Steinert J, Schiml S, Fauser F, Puchta H (2015) Highly efficient heritable plant genome engineering using Cas9 orthologues from Streptococcus thermophiles and Staphylococcus aureus. Plant J 84(6):1295–1305
Stoddard BL (2011) Homing endonucleases: From microbial genetic invaders to reagents for targeted DNA modification. Structure 19:7–15. doi:10.1016/j.str.2010.12.003
Svitashev S, Young JK, Schwartz C, Gao H, Falco SC, Cigan AM (2015) Targeted mutagenesis, precise gene editing and site-specific gene insertion in maize using Cas9 and guide RNA. Plant Physiol 2:931–945
Swedish Board of Agriculture (2015) Green light in the tunnel”! Swedish Board of Agriculture: a CRISPR-Cas9-mutant but not a GMO. http://www.upsc.se/about-upsc/news/4815-green-light-in-the-tunnel-swedish-board-of-agriculture-a-crispr-cas9-mutant-but-not-a-gmo.html. Accessed 10 Sept 2016
Tax FE, Vernon DM (2001) T-DNA-associated duplication/translocations in Arabidopsis implications for mutant analysis and functional genomics. Plant Physiol 126:1527–1538
Taylor G, Petrucci L, Lambert A, Baxter S, Jarjour J, Stoddard B (2012) LAHEDES: the LAGLIDADG homing endonuclease database and engineering server. Nucleic Acids Res 40:110–116
The McGuinness Institute (2013) An Overview of Genetic Modification in New Zealand 1973–2013: The First Forty Years, Auckland Council. http://www.aucklandcouncil.govt.nz/EN/planspoliciesprojects/plansstrategies/unitaryplan/Documents/Section32report/Appendices/Appendix%203.49.16.pdf. Accessed 8 Sept 2016
Townsend JA, Wright DA, Winfrey RJ, Fu F, Maeder ML, Joung JK, Voytas DF (2009) High-frequency modification of plant genes using engineered zinc-finger nucleases. Nature 459:442–445
Tsai C, Xue L (2015) CRISPRing into the woods. GM Crops Food. doi:10.1080/21645698.2015.1091553
Upadhyay SK, Kumar J, Alok A, Tuli R (2013) RNA-guided genome editing for target gene mutations in wheat. G3 (Bethesda) 3:2233–2238
USDA (2012) http://www.aphis.usda.gov/biotechnology/downloads/reg_loi/APHIS_response_DO W_ZFN_PK1_030812.pdf]
Waltz E (2016) Gene-edited CRISPR mushroom escapes US regulation. Nature 532:293. doi:10.1038/nature.2016.19754
Wang Y, Cheng X, Shan Q, Zhang Y, Liu J, Gao C, Qiu JL (2014) Simultaneous editing of three homoeoalleles in hexaploid bread wheat confers heritable resistance to powdery mildew. Nat Biotechnol 32:947–951
Wang M, Liu Y, Zhang C, Liu J, Liu X, Wang L et al (2015) Gene editing by co-transformation of TALEN and chimeric RNA/DNA oligonucleotides on the rice OsEPSPS gene and the inheritance of mutations. PLoS One. doi:10.1371/journal.pone.0122755
Wang F, Wang C, Liu P, Lei C, Hao W, Gao Y, Liu YG, Zhao K (2016) Enhanced rice blast resistance by CRISPR/Cas9-targeted mutagenesis of the ERF transcription factor gene OsERF922. PLoS One. doi:10.1371/journal.pone.0154027
Weeks DP, Spalding MH, Yang B (2015) Use of designer nucleases for targeted gene and genome editing in plants. Plant Biotech J. doi:10.1111/pbi.12448
Weinthal D, Tovkach A, Zeevi V, Tzfira T (2010) Genome editing in plant cells by zinc finger nucleases. Trends Plant Sci 15(6):308–321
Wendt T, Holm PB, Starker CG, Christian M, Voytas DF, Brinch- Pedersen H, Holme IB (2013) TAL effector nucleases induce mutations at a pre-selected location in the genome of primary barley transformants. Plant Mol Biol 83:279–285
Wiedenheft B, Sternberg SH, Doudna JA (2012) RNA-guided genetic silencing systems in bacteria and archaea. Nature 482:331–338. doi:10.1038/nature10886
Wolt JD, Wang K, Yang B (2015) The Regulatory Status of Genome-edited Crops. Plant Biotech J. doi:10.1111/pbi.12444
Woo JW, Kim J, Kwon S, Corvalán C, Cho SW, Kim H, Kim S, Kim S, Choe S, Kim J (2015) DNA-free genome editing in plants with preassembled CRISPR-Cas9 ribonucleoproteins. Nat Biotechnol 33:1162–1164. doi:10.1038/nbt.3389
Wright DA, Townsend JA, Winfrey RJ, Irwin PA, Rajagopal J, Lonosky PM, Hall BD, Jondle MD, Voytas DF (2005) High frequency homologous recombination in plants mediated by zinc-finger nucleases. Plant J 44:693–705
Wright DA, Li T, Yang B, Spalding MH (2014) TALEN-mediated genome editing: prospects and perspectives. Biochem J 462:15–24
Xie K, Yang Y (2013) RNA-guided genome editing in plants using a CRISPR-Cas system. Mol Plant 6:1975–1983
Xiong JS, Ding J, Li Y (2015) Genome-editing technologies and their potential application in horticultural crop breeding. Hort Res. doi:10.1038/hortres.2015.19
Xu R, Li H, Qin R, Wang L, Li L, Wei P, Yang J (2014) Gene targeting using the Agrobacterium tumefaciens-mediated CRISPR-Cas system in rice. Rice (N Y) 7:5. doi:10.1186/s12284-014-0005-6
Xu RF, Li H, Qin RY, Li J, Qiu CH, Yang YC, Ma H, Li L, Wei PC, Yang JB (2015) Generation of inheritable and “transgene clean” targeted genome-modified rice in later generations using the CRISPR/Cas9 system. Sci Rep. doi:10.1038/srep11491
Yokoi AN, Cermak T, Hoshino T, Sugimoto K, Saika H, Mori A, Osakabe K, Hamada M, Katayose Y, Starker C, Voytas DF, Toki S (2015) A defect in DNA ligase 4 enhances the frequency of TALEN-mediated targeted mutagenesis in rice. Plant Physiol. doi:10.1104/pp.15.01542
Zemach A, Kim MY, Hsieh PH, Coleman-Derr D, Eshed-Williams L, Thao K, Harmer SL, Zilberman D (2013) The Arabidopsis nucleosome remodeler DDM1 allows DNA methyl transferases to access H1-containing heterochromatin. Cell 153:193–205
Zetsche B, Gootenberg JS, Abudayyeh OO, Slaymaker IM, Makarova KS, Essletzbichler P, Volz SE, Joung J, van der Oost J, Regev A, Koonin EV, Zhang F (2015) Cpf1 is a single RNA-guided endonuclease of a class 2 CRISPR-Cas system. Cell 163:759–771
Zhang F, Maeder ML, Unger-Wallace E, Hoshaw JP, Reyon D, Christian M, Li X, Pierick CJ, Dobbs D, Peterson T, Joung JK, Voytas DF (2010) High frequency targeted mutagenesis in Arabidopsis thaliana using zinc finger nucleases. PNAS 107:12028–12033
Zhang H, Zhang J, Wei P, Zhang B, Gou F, Feng Z, Mao Y, Yang L, Zhang H, Xu N, Zhu JK (2014) The CRISPR/Cas9 system produces specific and homozygous targeted gene editing in rice in one generation. Plant Biotech J 12:797–807
Zhang Z, Mao Y, Ha S, Liu W, Botella J, Zhu JK (2015) A multiplex CRISPR/Cas9 platform for fast and efficient editing of multiple genes in Arabidopsis. Plant Cell Rep. doi:10.1007/s00299-015-1900-z
Zhou H, Liu B, Weeks DP, Spalding MH, Yang B (2014) Large chromosomal deletions and heritable small genetic changes induced by CRISPR/Cas9 in rice. Nucleic Acids Res 42:10903–10914
Zhou J, Peng Z, Long J, Sosso D, Liu B, Eom JS, Huang S, Liu S, Cruz CV, Frommer WB, White FF, Yang B (2015a) Gene targeting by the TAL effector PthXo2 reveals cryptic resistance gene for bacterial blight of rice. Plant J 82:632–643
Zhou X, Jacobs TB, Xue L, Harding SA, Tsai C (2015b) Exploiting SNPs for biallelic CRISPR mutations in the outcrossing woody perennial Populus reveals 4-coumarate: CoA ligase specificity and redundancy. New Phytol 208:298–301
Zhu J, Song N, Sun S, Yang W, Zhao H, Song W, Lai J (2016) Efficiency and inheritance of targeted mutagenesis in maize using CRISPR–Cas9. J Genet Genomics 43:25–36
Acknowledgements
KK acknowledges Council of Scientific and Industrial Research (CSIR), New Delhi, India (Sanction No. 09/137/(0541)/2012-EMR-1) for the award of senior research fellowship.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Khandagale, K., Nadaf, A. Genome editing for targeted improvement of plants. Plant Biotechnol Rep 10, 327–343 (2016). https://doi.org/10.1007/s11816-016-0417-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11816-016-0417-4