Abstract
American foulbrood is an important bacterial disease affecting the larvae of honey bees (Apis mellifera L.) caused by Paenibacillus larvae. Due to easy transmission of disease and the ability of bacteria to create spores, it is a bacterium resistant to both physical and chemical conditions. The study aims to isolate, perform microbiological analyses, and determine biochemical properties and genotypes P. larvae strains from AFB samples collected from Turkey's Central and Eastern Black Sea regions. An isolation study was conducted on adult bees, larvae, honey, and primary honeycomb samples from suspected colonies in the regions under study. After the purification of bacterial isolates from samples, P. larvae strains were identified using biochemical and molecular methods. The genetic diversity and ERIC types of P. larvae isolates were determined by rep-PCR DNA genotyping using BOX A1R and MBO REP1 primers and multiplex-PCR. A phylogenetic tree of P. larvae strains was constructed in the study. All P. larvae isolates were determined as ERIC I type. According to the rep-PCR results of P. larvae strains, 15 of the 28 isolates were Ab genotype (54%), 7 (25%) Aβ genotype, 4 (14%) AB genotype, 1 (3.5%) αB genotype, and 1 (3.5%) ab genotype. From an epidemiological viewpoint, it was determined that Ab and Aβ genotypes were widely distributed, while other genotypes (AB, αB, and ab) showed less spread. The results of the study will guide researchers in taking relevant measures to prevent and control American foulbrood.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
American foulbrood (AFB) is caused by the Gram-positive spore-forming bacterium Paenibacillus larvae. It is the most serious bacterial disease affecting honey bees at the larval stage. In larvae contaminated with the causative bacteria in the first period (12–48 h), the duration of death varies according to genotype. It was reported that it took about 12 days for genotype ERIC I to kill all infected larvae, while larvae infected with genotype ERIC II died within 7 days. In a study with other ERIC genotypes (LMG16252; ERIC III, LMG16247; ERIC IV, 138b; ERIC V), LT100 levels were reported to be reached after approximately three days (Genersch 2010; Beims et al. 2020). Paenibacillus larvae is a Gram-positive, facultative anaerobe, endospore-forming bacterium. Infected larvae have a brownish, semi-liquid, sticky, filamentous appendage and a foul, rotten odor. The spore resists chemical disinfectants and adverse physical conditions (heat and drying) (Hansen and Brødsgaard 1999). Its vegetative form is a chain or a single rod, while its spores are spindle-shaped, 1.3 × 0.6 µm in size, and swollen. When staining bacteria with Ziehl–Neelsen carbol-fuchsin, the bacterial walls are stained red–purple while the spore walls remain clear. In culture on MYPGP agar, colonies are 3–4 mm in diameter, small, regular, often rough, flat or raised, and whitish to beige or orange to red (Shimanuki and Knox 2000; OIE 2018). After killing the vegetative bacteria form by heat treatment, the Paenibacillus larvae bacteria were isolated (de Graaf et al. 2013). Determined bacterial phenotypes, such as the morphology of colonies in different culture media, biochemical testing, pathogenicity, and antibiotic resistance, can be used to differentiate between strains but need to be sufficiently variable between closely related strains (Li et al. 2009). Genotyping, which expresses the discrimination of bacterial strains according to their genetic content, has been widely used in identifying and subtyping strains due to their high solubility. Bacterial genotyping methods, include pulsed-field gel electrophoresis (PFGE) (Herschleb et al. 2007), ribotyping (Harvey and Minter 2005), restriction fragment length polymorphic DNA analysis (RFLP) (Todd et al. 2001), random amplified polymorphic DNA assay (RAPD) (Welsh and McClelland 1990), arbitrary primed PCR (AP-PCR) (Williams et al. 1990), amplified fragment length polymorphism (AFLP) (Vos et al. 1995), repetitive element based PCR (rep-PCR) (Koeuth et al. 1995) and multilocus sequence typing (MLST) method (Bertolotti et al. 2021), respectively. One of the methods used to genotype P. larvae strains is MLST(Morrissey et al. 2015). Rep-PCR, another analysis method, is a method for fingerprinting bacterial genomes by examining strain-specific patterns derived from PCR amplification of repetitive DNA elements in bacterial genomes. REP-PCR is a valuable method because it is more reproducible. After all, specific primers are used for amplification because they are easy to use, the cost is low, and they can get fast results (Li et al. 2009). Different genotypes of P. larvae have different reproductive abilities. For genotyping P. larvae isolates, three main sets of repetitive elements are used: the repetitive extragenic palindromic (REP) sequence, BOX elements, and the enterobacterial repetitive intergenic consensus sequence (ERIC) (Genersch et al. 2006; Beims et al. 2020). Genersch and Otten (2003) examined P. larvae isolates using the rep-PCR technique with BOX A1R and MBO REP1 primers and reported four genetic subgroups (AB, Ab, ab, and aB). Using the same combination (BOX A1R and MBO REP1 primer), Loncaric et al. (2009) described five different genotypes (ab, aB, Ab, AB, and αb).
AFB is a well-known and legally notifiable disease in Turkey. The prevalence of AFB in Turkey is monitored and diagnosed by the Samsun Veterinary Control Institute; however, the prevalence of specific P. larvae genotypes within the country is unknown. There needs to be more research on the identification and distribution of P. larvae genotypes in Turkey. This study aimed to isolate P. larvae from AFB suspected specimens collected from Turkey's Central and Eastern Black Sea regions to determine the isolates' microbiological and biochemical characteristics and their genetic diversity.
Materials and methods
Materials
The prevalence of AFB in Turkey is monitored and diagnosed by the Samsun Veterinary Control Institute Bee Diseases Laboratory. The samples from the Bee Diseases Laboratory consist of disease-suspicious honeycomb, larvae, and primary honeycomb samples from 10 Black Sea region (Central and Eastern) provinces. These P. larvae strains (P. larvae PB3.2a, PB3.2b2, PB3.1a, PB3.3a1, PB3.3b, PB3.3b1, PB4, PB5a, PB5b, PB6a, and PB6b) were isolated in a study by Baş and Alpay Karaoglu (2015).
Methods
Isolation of P. larvae from AFB suspect specimens
For P. larvae isolation, larvae, adult bees, and honey samples were collected from the blackened pores of clinically suspect combs. Five honey bee larvae from the combs were taken into 5 mL of sterile distilled water with the help of sterile forceps and suspended with a stirrer. Solid larval parts were fragmented with the use of a homogenizer. Honey samples scraped from the surface of the frames were used. Five grams of the honey sample taken with a sterile spoon was mixed homogeneously with 5 mL of sterile distilled water. Adult bees that completed the larval stage of the adult bee samples but could not get out of the comb cell were used. Five adult bees collected from the combs with sterile forceps for dissection were subjected to surface sterilization in 70% ethanol for 2 min. Then they were washed three times with 5 mL of sterile distilled water. Adult bees were crushed in a homogenizer by adding 5 mL of sterile distilled water. Homogenized samples were exposed to heat in a water bath at 80 °C for 10 min to eliminate vegetative bacterial forms (OIE 2008). The samples were left to cool at room temperature and precipitated by centrifugation at 5,000 rpm for 10 min. The supernatant was discarded, and the remaining pellet was homogenized by vortexing. Serial dilutions of 10–1 to 10–8 were prepared from the adult bee, honey, and honey bee larva homogenized samples. After being diluted for culture, the homogenate dilutions were spread on agar plates and incubated at 37 °C and 5% CO2 conditions for 3–4 days for growth (OIE 2008; de Graaf et al. 2013; Hamdi et al. 2013). At the end of the incubation, P. larvae-like colonies were checked for catalase activity with 3% (v/v) H2O2. Catalase-negative colonies were selected for morphological and biochemical tests. P. larvae isolated as a single colony was re-grown on MYPGP agar medium and stored at -80 °C for further studies (Hamdi et al. 2013).
Determination of the morphological and biochemical characteristics of isolates
Isolates of P. larvae were recognized on the basis of morphological and biochemical characteristics. Morphological characterization of bacterial isolates was performed by evaluating colony morphology, Gram staining, and motility of colonies. Biochemical properties tested include oxidase, catalase, urease, nitrate reduction, lecithinase, production of dihydroxyacetone, gelatinase, Voges-Proskauer (VP), citrate, indole, and sugar (glucose, maltose, fructose, lactose, trehalose, and xylose) fermentation (de Graaf et al. 2013).
DNA isolation and genetic diversity of P. larvae isolates
According to Sambrook's procedure, bacterial genomic DNA was extracted using the standard phenol: chloroform method (Sambrook et al. 1989). For DNA isolation, isolates were grown in MYPGP broth at 37 °C in a shaking incubator for 48 h. The isolates were transferred to a sterile Eppendorf tube and settled at 10,000 rpm for 10 min, and then the pellet was washed three times with sterile ultrapure water. Isolated DNA samples (5 µL) were visualized on 1% agarose gel electrophoresis. Primers were commercially obtained from Macrogen Inc.(Amsterdam, Netherlands). The 16S rRNA gene was amplified by PCR using primers 27F (5'-AGAGTTTGATCMTGGCTCAG-3') and 1492R (5'-TACGGYTAACCTTGTTACGACTT-3'). To determine the genetic diversity of P. larvae isolates, they were amplified by PCR using the repetitive element PCR (rep-PCR) technique using BOX A1R (5'-CTACGG CAAGGCGACGCTGACG-3') and MBO REP1 (5'-CCGCCGTTGCCGCCGTTGCCGCCG-3') primers. Reactions were prepared using a final volume of 1X Taq DNA polymerase buffer, 2.5 µM MgCl2, 250 µM dNTP mix, 0.3 pmol/µL primer, and 1 U/µL Taq DNA polymerase (Fermentas). The 16S rRNA gene PCR conditions were adapted according to the study by Sevim et al. (2017). The amplified PCR products were sequenced by sending 50 μL of each sample to the company of Macrogen (Amsterdam, Netherlands). The rep-PCR conditions for determining genetic diversity are as follows. The first denaturation step in the PCR cycle was performed at 94 °C for 15 min. PCR was performed with 35 cycles for BOX-PCR with an elongation step of 60 s at 94 °C, 60 s at 53 °C, and 2.5 min at 72 °C, respectively. The final elongation step was set at 72 °C for 7 min, and PCR products were preserved at 4 °C. BOX-PCR products were separated in a standard 1.0% agarose gel, visualized under UV light, and photographed with a gel Doc digital image capture system (Bio-Rad). The genetic diversity of the amplified PCR products was evaluated according to the nomenclature systems established by Genersch and Otten (2003) and Peters et al. (2006).
ERIC types of P. larvae isolates were determined according to the multiplex-PCR assay previously performed by Okamoto et al. (2022). The multiplex PCR assay was performed using the 2X PCR Master Mix (A140301, Ampliqon, Denmark) in a final reaction volume of 20 μl containing the appropriate primer concentrations previously described.
Alignment of 16S rRNA sequences and construction of a phylogenetic tree
After sequencing the 16S rRNA gene PCR products, the sequences were first edited with the BioEdit software (Hall 1999), and the nucleotide sequences were aligned using MEGA X software (Kumar et al. 2018). The resulting sequences were compared with sequences in the GenBank database using the NCBI-BLAST tool (Benson et al. 2012). A phylogenetic tree of the 16S rRNA gene region was constructed using neighbor-joining analysis. Finally, the sequences were subjected to neighbor-joining analysis with p-distance correction, gap omission, and 1.000 bootstrap pseudoreplicates using MEGA X (Kumar et al. 2018). NCBI GenBank accession numbers were recorded for the 16S rRNA gene sequences of P. larvae strains isolated in the study.
Results
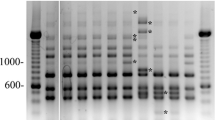
This study examined 64 suspect samples (honey bee larvae, adult bees, honey samples, and honeycombs) received by the institute in 2014 and 2015. Bacterial isolation from suspicious samples was performed at the Recep Tayyip Erdogan University Microbiology and Molecular Biology Research Laboratory. Because of culture, spore bacilli and cocci were isolated in 35 of 64 samples, while no bacterial isolation could be made from 29 samples. A total of 63 Gram-positive bacteria (58 Gram-positive bacilli and 5 Gram-positive cocci) were isolated from 35 samples with bacterial growth. The bacterial colonies were examined macroscopically (color, type, and size of colonies) and microscopically (Gram stain, bacterial morphology, and spore content). Eighteen isolates thought to be P. larvae in the morphological examination were then characterized and identified by biochemical and molecular methods. The other 10 P. larvae strains isolated in a study by Baş and Karaoglu (2015) were also included in the analyses. The isolation source and province of origin of the P. larvae strains in the study are given in Table 1. The P. larvae isolate was observed to have cream, transparent, and translucent-looking colonies on MYPGP agar medium, and all isolates were immobile and slow-growing (48–72 h). The results of the biochemical characteristics of P. larvae isolates are given in Table 2. In all the isolates, fructose fermentation was positive. Except for P. larvae SV21 and SV30b, they were able to metabolize glucose and at least one of the other tested sugars. Other than fructose, the SV21 and SV30b strains fermented only maltose. While catalase activity results were negative (weak positivity in 4 of them) in all strains, specificity could not be determined in the oxidase activity results. The strains analyzed by the morphological and biochemical methods were confirmed as P. larvae by 16S rRNA gene sequencing and the multiplex PCR developed by Okamoto et al. (2022) and typed by PCR assays with BOX A1R and MBO REP primers, methods commonly used for typing bacterial strains, including Bacillus species (Alippi and Aguilar 1998; Cherif et al. 2002). Although the 16S rRNA gene sequences of the strains showed relatively high similarities with those of P. larvae strains deposited in the GenBank database (Table 3), some strains (PB16.1b, PB24b, and PB31b) showed less than 98.65% sequence similarity, the value proposed as the threshold for differentiating two species (Kim et al. 2014), with other P. larvae strains. The phylogenetic tree generated according to the result of the analysis is shown in Fig. 1. However, the multiplex PCR (Okamoto et al. 2022) generated two products with sizes of 973-bp and 554-bp from all the strains analyzed and thus they were identified as P. larvae of genotype ERIC I (Fig. 2).
Phylogenetic tree based on 16S rRNA gene sequences of P. larvae isolates. Black circles indicate isolates used in this study. Accession numbers of the 16S rRNA gene sequences retrieved from the database and used for the analysis are shown in parentheses next to each strain name. The percentage of replicate trees in which the associated taxa clustered together in the bootstrap test are shown next to the branches
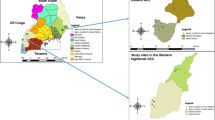
In the study, the genetic diversity of 28 Paenibacillus larvae isolates was determined by PCRs with BOX A1R and MBO REP primers. According to the presence or absence of BOX-PCR specific products around 750-bp, which was previously determined in the literature (Genersch and Otten 2003), it was determined that the isolates contained 3 different genotypes, "A", "α" and "a". With MBO-REP1 primers, 3 different genotypes named B, b and β were generated. Isolates with b patterns possess a band around 1000-bp moving slightly faster than B pattern strains having a more slowly running band at 1000-bp molecular marker fragments. However, isolates with the β pattern do not have a band around 1000-bp (Genersch and Otten 2003; Neuendorf et al. 2004). According to the BOX A1R and MBO REP1 PCR methods, 15 (54%) of the isolates were Ab genotype, 7 (25%) Aβ, 4 (14%) AB, 1 (3.5%) αB genotype, and 1 (3.5%) ab genotype. As a result of the study, it was determined that 28 Paenibacillus larvae strains had five different genotypic characteristics (Ab, Aβ, AB, αB, and ab) (Table 4). From an epidemiological viewpoint, Ab and Aβ genotypes showed a wide distribution, while other genotypes (AB, αB, and ab) showed a lower distribution (Figs. 3 and 4).
Discussion
In parallel with the importance of beekeeping in the world and Turkey, diseases (parasitic, bacterial, viral, and fungal) seen in honey bees are important for the sector. A lot of research is conducted worldwide to detect these diseases that cause economic and yield losses (Balkaya et al. 2016). Honey bee diseases and pests are some of the most important factors that slow down the development of beekeeping and affect production. The development stage of the bee creates suitable environments for many disease agents and pests (Çakmak et al. 2003). AFB, European foulbrood, Stonebrood, and Chalkbrood diseases are bacterial and fungal diseases that affect the honey bee in the larval stage (OIE 2008). Therefore, this study investigated the presence and genotype distribution of P. larvae causing AFB in honey bees from samples collected between 2014 and 2015 in Turkey's Central and Eastern Black Sea regions.
In a study conducted in Japan, seven P. larvae isolated in the western region of Aichi province between 2012 and 2014 were analyzed, and their genotypic and phenotypic characteristics were examined. Biochemical tests reported that all seven P. larvae isolates metabolized D-trehalose, D-glucose, and gelatin. The metabolic profiles of D-fructose, D-mannose, D-mannitol, salicin, glycerol, D-ribose, N-acetylglucosamine, and D-tagatose differed among the isolates (Hirai et al. 2016).
According to BOX and rep-PCR, the genotype distribution of these strains was found to be different. The biochemical characteristics of the strains in our study are given in Table 2, and their genotypes are given in Table 4. However, strains PB25b, PB34, and PB13.1b were positive for lactose fermentation, and strains PB25b, PB27b, PB32b, and SV28 were positive for xylose fermentation. Except for the PB13.1b strain, the strains positive for lactose and xylose fermentation were of the Ab genotype. P. larvae SV21 and SV30b strains, which could not ferment glucose and could only ferment maltose and fructose, two of the six sugars tested, were identified as having the Aβ genotype. Oxidase, citrate, gelatin hydrolysis, glucose, lactose, and trehalose fermentation of the P. larvae PB13.1b strain, whose genotype was determined as ab, were positive. The biochemical test results are likely related to the genotype, but more strains and biochemical tests should be performed to obtain definitive results. It was observed that P. larvae (PB3.3a, PB3.2b2, PB16.1b, and SV27b) strains with the AB genotype have the common feature of being able to ferment glucose and trehalose (Tables 2 and 4) (Sevim et al. 2017). Different genotypes of P. larvae have been identified in different regions. Rusenova and Parvanov (2014) determined the genotypes of P. larvae isolated in Bulgaria by rep-PCR. P. larvae were isolated from samples collected from fifteen apiaries in northern (3 apiaries) and southern (12 apiaries) Bulgaria between 2009 and 2014. Of the study's total 103 P. larvae isolates, 21 were reported as the AB genotype and 82 as the ab genotype.
It was reported that 70 P. larvae were isolated from samples from southern Bulgaria, while 37 P. larvae were isolated from apiary samples in northern Bulgaria. Genotyping of 107 P. larvae isolates was performed by the Rep-PCR method using BOX A1R and MBO REP1 primers. Based on the genotyping of the isolates, it was determined that there were ab and AB genotypes. The AB genotype, which was more lethal in southern Bulgaria, was reported to be more common (Rusenova and Parvanov 2016). According to the results of Rusenova and Parvanov (2016), the necessity of regular, mandatory screening of bee colonies for early disease diagnosis was stated. In the genotyping of 28 P. larvae strains analyzed in our study, the Ab (15) genotype was the most common. Then, (7) Aβ, (4) AB, (1) αB genotype, and (1) ab genotypes were determined, respectively (Fig. 4). It was observed that it is necessary to implement relevant measures for the prevention and control of AFB (Rusenova and Parvanov 2016). Germany reported four genotypes (AB, Ab, ab, and αβ) of P. larvae and that the metabolic relationship of Ab, ab, and aβ genotypes was similar to each other (Neuendorf et al. 2004). A study was conducted in Austria to examine the genotypic diversity of 214 Paenibacillus larvae strains. In the study, two genotypes could be distinguished using the ERIC-PCR method (ERIC I and II), while five different genotypes were detected by BOX- and REP-PCR (ab, aB, Ab, AB, and αb). The aB and αb genotypes were new and reported in this study for the first time among studies using the same techniques (Loncaric et al. 2009). According to the above literature results, four different genotypes were identified in a study conducted in Germany, while five were identified in a study conducted in Australia. While two genotypic diversities (AB and ab) were encountered in the Bulgarian study (Rusenova et al. 2013; Rusenova and Parvanov 2016), five genotypic diversities (Ab, Aβ, AB, αB, and ab genotypes) were identified in our study. It was found important that these genotypes were identified in our study, similar to the results in the literature.
Because of the established correlation between genotype, the virulence of strains, and the development of clinical symptoms, knowledge of genotypes is an important epidemiological tool for AFB risk assessment (Loncaric et al. 2009). Beekeeper activities such as itinerant beekeeping, unreported activities, queen purchases, and equipment exchange contribute to the rapid spread of bacterial strains and genotypes. Determining the relationship between rep-PCR genotypes AB and ab and biochemical phenotypes in studies is helpful in epidemiological studies to detect the source of infection and control the disease. The distribution of genotypes can be used to help locate the source of the disease. Genersch and Otten (2003) reported that only BOX primers would not be sufficient for the genetic subtyping of P. larvae and that the discrimination power of rep-PCR for typing could be increased by combining MBO REP1 and BOX A1R primers instead. In our study, the (using the BOX A1R and MBO REP1 primers) rep-PCR and multiple-PCR methods were used to ensure good genotyping and to better distinguish the isolated P. larvae strains.
In 2014, 22 P. larvae isolates from Turkey were genotyped using the rep-PCR method. The rep-PCR results show that no P. larvae were detected in the genotypes ERIC III and ERIC IV. All isolates genotyped by the molecular method belonged to genotypes ERIC I or ERIC II. It was found that 13 of the 22 P. larvae isolates belonged to genotype ERIC I and 9 isolates to genotype ERIC II (Schiesser 2014). According to the results of some investigations carried out in Europe, it was found that both genotypes, ERIC I and ERIC II, are widespread. The genotype ERIC II is common in Slovenia (Žugelj et al. 2021), the Czech Republic (Biová et al. 2021), and northern Italy (Bassi et al. 2015), while ERIC I is reported to be a more common genotype in the Republic of Kosovo (Hulaj et al. 2021). A total of 88 strains of P. larvae were isolated from samples collected in neighboring regions of the Czech Republic and Slovakia and analyzed using molecular methods that allow epidemiological inferences to be made. The majority of isolates (78 isolates) were isolated from samples from the Czech Republic, while 10 isolates were isolated from the western part of Slovakia on the border with the Czech Republic. According to the genotyping results, 78.9% of the isolates belonged to the ERIC II genotype and 21.1% belonged to the ERIC I genotype (Matiašovic et al. 2023). A total of 108 isolates of P. larvae collected in 2011 and 2021 in different geographical regions of Lithuania were analyzed by molecular methods. The results of molecular analyses of P. larvae isolates obtained by the rep-PCR method showed that 100% belonged to the ERIC I genotype (Amšiejute et al. 2022). In our study, both methods were combined to ensure good genotyping and differentiate the isolated P. larvae strains. The analysis of P. larvae isolated from the eastern Black Sea region by the multiplex PCR method revealed that all were genotype ERIC I. AFB disease, caused by P. larvae, is occasionally observed in many countries and regions. Global beekeeping activities, in general, and rarely atmospheric events, cause AFB-causing bacteria to spread between countries and/or regions. In the AFB disease detected in the Eastern Black Sea region, where beekeeping is an important sector geographically, it was found important to observe the ERIC I genotype, which is common worldwide and is pathogenic.
Conclusions
There are few studies on the genetic population structure of P. larvae in Turkey. Comprehensive studies of AFB cases and the phenotypic and genotypic characterization of isolated P. larvae will reveal this honeybee disease's the local and national epidemiology of this honeybee disease. Understanding the genotypic distribution and prevalence of P. larvae will help with disease prevention and control. These epidemiological data obtained in studies are critical in forming a strategy in the fight against notifiable diseases. Therefore, the scope can be expanded in future studies, and more detailed results can be revealed with samples isolated from all regions of Turkey.
Data Availability
Datasets used in this study will be made available by the corresponding author with no hesitation upon genuine request.
Abbreviations
- AFB:
-
American foulbrood
- PFGE:
-
Pulsed-field gel electrophoresis
- RFLP:
-
Restriction fragment length polymorphic DNA analysis
- RAPD:
-
Random amplified polymorphic DNA assay
- AP-PCR:
-
Arbitrary primed PCR
- AFLP:
-
Amplified fragment length polymorphism
- Rep-PCR:
-
Repetitive element based PCR
- ERIC:
-
Enterobacterial repetitive intergenic consensus sequence
- NCBI:
-
National Center for Biotechnology Information
- BLAST:
-
Basic Local Alignment Search Tool
References
Alippi AM, Aguilar OM (1998) Characterization of isolates of Paenibacillus larvae subsp. larvae from diverse geographical origin by the polymerase chain reaction and BOX primers. J Invertebr Pathol 72(1):21–27. https://doi.org/10.1006/jipa.1998.4748
Amšiejute P, Jurgelevičius V, Mačiulskis P, Butrimaite-Ambrozevičiene C, Pilevičiene S, Janeliunas Z, Kutyriova T, Jacevičiene I, Paulauskas A (2022) Molecular diversity of Paenibacillus larvae strains isolated from Lithuanian apiaries. Front Vet Sci 9:959636. https://doi.org/10.3389/fvets.2022.959636
Balkaya İ, Gülbaz H, Avcıoğlu H, Güven E (2016) Türkiye’de görülen bal arısı (Apis mellifera) hastalıkları. Atatürk Univ Vet Bil Derg 11(3): 339–347. https://doi.org/10.17094/ataunivbd.282993
Bassi S, Formato G, Milito M, Trevisiol K, Salogni C, Carra E (2015) Phenotypic characterization and ERIC-PCR based genotyping of Paenibacillus larvae isolates recovered from American foulbrood outbreaks in honey bees from Italy. Vet Q 35(1):27–32. https://doi.org/10.1080/01652176.2014.993095
Baş Y, Alpay Karaoglu Ş (2015) Characterization and antimicrobial susceptibility of spore forming bacilli isolated from honeycomb. Recep Tayyip Erdogan Univ Graduate School Natural Appl Sci Depart Biol, Master Thesis, pp123
Beims H, Bunk B, Erler S, Mohr KI, Spröer C, Pradella S, Günther G, Rohde M, der Ohe W, Steinert M (2020) Discovery of Paenibacillus larvae ERIC V: Phenotypic and genomic comparison to genotypes ERIC I-IV reveal different inventories of virulence factors which correlate with epidemiological prevalences of American Foulbrood. Int J Med Microbiol 310(2):151394. https://doi.org/10.1016/j.ijmm.2020.151394
Benson DA, Karsch-Mizrachi I, Clark K, Lipman, DJ, Ostell J, Sayers EW (2012) GenBank. Nucleic Acids Res 40(Database issue):D48–D53. https://doi.org/10.1093/nar/gkr1202
Bertolotti AC, Forsgren E, Schäfer MO, EuroPLarva Consortium, Sircoulomb, F, Gaïani N, Ribière-Chabert M, Paris L, Lucas P, de Boisséson C, Skarin J, Rivière MP (2021). Development and evaluation of a core genome multilocus sequence typing scheme for Paenibacillus larvae, the deadly American foulbrood pathogen of honeybees. Environ Microbiol 23(9):5042–5051. https://doi.org/10.1111/1462-2920.15442
Biová J, Bzdil J, Dostálková S, Petřivalský M, Brus J, Carra E, Danihlík J (2021) American Foulbrood in the Czech Republic: ERIC II genotype of Paenibacillus larvae is prevalent. Front Vet Sci 8:698976. https://doi.org/10.3389/fvets.2021.698976
Cherif A, Borin S, Rizzi A, Ouzari H, Boudabous A, Daffonchio D (2002) Characterization of a repetitive element polymorphism-polymerase chain reaction chromosomal marker that discriminates Bacillus anthracis from related species. J Appl Microbiol 93(3):456–462. https://doi.org/10.1046/j.1365-2672.2002.01712.x
Çakmak İ, Aydın L, Güleğen E, Korkut M (2003) Honeybee pests and diseases survey in Southern Marmara Region of Turkey. Uludag Bee J 3(2):33–35. https://dergipark.org.tr/en/pub/uluaricilik/issue/53709/162636
de Graaf DC, Alippi AM, Antúnez K, Aronstein KA, Budge G, Koker DDe, Smet LD, Dingman DW, Evans JD, Foster LJ, Fünfhaus A, Garcia-Gonzalez E, Gregore A, Human H, Murray KD, Nguyen BK, Poppinga L, Spivak M, Engelsdorp DV, Wilkins S, Genersch E (2013) Standard methods for American Foulbrood research. J Apicul Res 52(1):1–28. https://doi.org/10.3896/IBRA.1.52.1.11
Genersch E, Forsgren E, Pentikäinen J, Ashiralieva A, Rauch S, Kilwinski J, Fries I (2006) Reclassification of Paenibacillus larvae subsp. pulvifaciens and Paenibacillus larvae subsp. larvae as Paenibacillus larvae without subspecies differentiation. Int J Syst Evol Microbiol 56(Pt 3):501–511. https://doi.org/10.1099/ijs.0.63928-0
Genersch E, Otten C (2003) The use of repetitive element PCR fingerprinting (rep-PCR) for genetic subtyping of German field isolates of Paenibacillus larvae subsp. larvae. Apidologie 34(3):195–206. https://doi.org/10.1051/apido:2003025
Genersch E (2010) American Foulbrood in honeybees and its causative agent, Paenibacillus larvae. J Invertebr Pathol 103(Suppl 1):S10–S19. https://doi.org/10.1016/j.jip.2009.06.015
Hall TA (1999) BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp 41:95–98
Hamdi C, Essanaa J, Sansonno L, Crotti E, Abdi K, Barbouche N, Balloi A, Gonella E, Alma A, Daffonchio D, Boudabous A, Cherif A (2013) Genetic and biochemical diversity of Paenibacillus larvae isolated from Tunisian infected honey bee broods. BioMed Res Int 1-9. https://doi.org/10.1155/2013/479893
Hansen H, Brødsgaard CJ (1999) American foulbrood: a review of its biology, diagnosis and control. Bee World 80(1):5–23. https://doi.org/10.1080/0005772X.1999.11099415
Harvey SP, Minter JM (2005) Ribotyping of Burkholderia mallei isolates. FEMS Immunol Med Mic 44(1):91–97. https://doi.org/10.1016/j.femsim.2004.12.002
Herschleb J, Ananiev G, Schwartz DC (2007) Pulsed-field gel electrophoresis. Nat Prot 2(3):677–684. https://doi.org/10.1038/nprot.2007.94
Hirai Y, Suzuki T, Inaba N, Minoguchi N, Takamatsu D (2016) Existence of Paenibacillus larvae genotypes ERIC I-ST2, ERIC I-ST15 and ERIC II-ST10 in the western region of Aichi prefecture, Japan. J Vet Med Sci 78(7):1195–1199. https://doi.org/10.1292/jvms.16-0041
Hulaj B, Goga I, Cana A, Merovci X, Rossi F, Crudele S, Ricchiuti L, Mutinelli F (2021) Passive surveillance of American Foulbrood in the Republic of Kosovo: geographic distribution and genotype characterization. J Apicult Res 60:1–7. https://doi.org/10.1080/00218839.2021.1892400
Kim M, Oh HS, Park SC, Chun J (2014) Towards a taxonomic coherence between average nucleotide identity and 16S rRNA gene sequence similarity for species demarcation of prokaryotes. Int J Syst Evol Microbiol 64(2):346–351. https://doi.org/10.1099/ijs.0.059774-0
Koeuth T, Versalovic J, Lupski JR (1995) Differential subsequence conservation of interspersed repetitive Streptococcus pneumoniae BOX elements in diverse bacteria. Genome Res 5(4):408–418. https://doi.org/10.1101/gr.5.4.408
Kumar S, Stecher G, Li M, Knyaz C, Tamura K (2018) MEGA X: Molecular evolutionary genetics analysis across computing platforms. Mol Biol Evol 35(6):1547–1549. https://doi.org/10.1093/molbev/msy096
Li W, Raoult D, Fournier PE (2009) Bacterial strain typing in the genomic era. FEMS Microbiology Reviews 33(5):892–916. https://doi.org/10.1111/j.1574-6976.2009.00182.x
Loncaric I, Derakhshifar I, Oberlerchner JT, Köglberger H, Moosbeckhofer R (2009) Genetic diversity among isolates of Paenibacillus larvae from Austria. J Invertebr Pathol 100(1):44–46. https://doi.org/10.1016/j.jip.2008.09.003
Matiašovic J, Bzdil J, Papežíková I, Čejková D, Vasina E, Bizos J, Navrátil S, Šedivá M, Klaudiny J, Pikula J (2023) Genomic analysis of Paenibacillus larvae isolates from the Czech Republic and the neighbouring regions of Slovakia. Res Vet Sci 158:34–40. Advance online publication. https://doi.org/10.1016/j.rvsc.2023.03.007
Morrissey BJ, Helgason T, Poppinga L, Fünfhaus A, Genersch E, Budge GE (2015) Biogeography of Paenibacillus larvae, the causative agent of American foulbrood, using a new multilocus sequence typing scheme. Environ Microbiol 17(4):1414–1424. https://doi.org/10.1111/1462-2920.12625
Neuendorf S, Hedtke K, Tangen G, Genersch E (2004) Biochemical characterization of different genotypes of Paenibacillus larvae subsp. larvae, a honey bee bacterial pathogen. Microbiol 150(7):2381–2390. https://doi.org/10.1099/mic.0.27125-0
OIE (2008) In Manual of diagnostic tests and vaccines for terrestrial animals (mammals, birds and bees), Chapter. 2.2.2. American Foulbrood. vol. 1(Sixth Edition). pp. 395–404
OIE (2018) Manual of diagnostic tests and vaccines for terrestrial animals. Chapter 3.2.2. American foulbrood of honey bees (infection of honey bees with Paenibacillus larvae)(version adopted in May 2016). pp. 719–735
Okamoto M, Furuya H, Sugimoto I, Kusumoto M, Takamatsu D (2022) A novel multiplex PCR assay to detect and distinguish between different types of Paenibacillus larvae and Melissococcus plutonius, and a survey of foulbrood pathogen contamination in Japanese honey. J Vet Med Sci 84(3):390–399. https://doi.org/10.1292/jvms.21-0629
Peters M, Kilwinski J, Beringhoff A, Reckling D, Genersch E (2006) American foulbrood of the honey bee: occurrence and distribution of different genotypes of Paenibacillus larvae in the administrative district of Arnsberg (North Rhine-Westphalia). J Vet Med B Infect Dis Vet Public Health 53(2):100–104. https://doi.org/10.1111/j.1439-0450.2006.00920.x
Rusenova N, Parvanov P, Stanilova S (2013) Molecular typing of Paenibacillus larvae strains isolated from Bulgarian apiaries based on repetitive element polymerase chain reaction (rep-PCR). Curr Microbiol 66(6):573–577. https://doi.org/10.1007/s00284-013-0318-5
Rusenova N, Parvanov P (2014) Biochemical profile of Paenibacillus larvae repetitive element polymerase chain reaction (rep-PCR) genotypes in Bulgaria. Kafkas Univ Vet Fak Derg 20(2):313–316. https://doi.org/10.9775/kvfd.2013.9853
Rusenova N, Parvanov P (2016) Prevalence of American Foulbrood and Paenibacillus larvae Genotypes in Bulgaria. İstanbul Üniv Vet Fak Derg 42(1):98–102. https://doi.org/10.16988/iuvfd.2016.16660
Sambrook J, Fritsch EF, Maniatis T (1989) Molecular cloning: a laboratory manual, 2nd edn. Cold Spring Harbor Laboratory Press, Cold Spring Harbor
Schiesser A (2014) Determination of Paenibacillus larvae genotypes in Turkey and antimicrobial effect of different propolis against these genotypes. Hacettepe University Master Thesis. pp 125
Sevim E, Baş Y, Çelik G, Pınarbaş M, Bozdeveci A, Özdemir T, Akpınar R, Yaylı N, Alpay Karaoğlu Ş (2017) Antibacterial activity of bryophyte species against Paenibacillus larvae isolates. Turk J Vet Anim Sci 41(4):521–531. https://doi.org/10.3906/vet-1611-70
Shimanuki H, Knox DA (2000) Diagnosis of honey bee diseases. Agriculture Handbook No. AH690. United States Department of Agriculture, Beltsville
Todd R, Donoff RB, Kim Y, Wong DT (2001) From the chromosome to DNA: restriction fragment length polymorphism analysis and its clinical application. J Oral Maxil Surg 59(6):660–667. https://doi.org/10.1053/joms.2001.22707
Vos P, Hogers R, Bleeker M, Reijans M, Van De LT, Hornes M, Frijters A, Pot J, Peleman J, Kuiper M (1995) AFLP: a new technique for DNA fingerprinting. Nucleic Acids Res 23(21):4407–4414. https://doi.org/10.1093/nar/23.21.4407
Welsh J, McClelland M (1990) Fingerprinting genomes using PCR with arbitrary primers. Nucleic Acids Res 18(24):7213–7218. https://doi.org/10.1093/nar/18.24.7213
Williams JG, Kubelik AR, Livak KJ, Rafalski JA, Tingey SV (1990) DNA polymorphisms amplified by arbitrary primers are useful as genetic markers. Nucleic Acids Res 18(22):6531–6535. https://doi.org/10.1093/nar/18.22.6531
Žugelj A, Papić B, Zdovc I, Zajc U, Golob M, Avberšek J, Kušar D (2021) ERIC and WGS typing of Paenibacillus larvae in Slovenia: Investigation of ERIC I outbreaks. Insects 12(4):362. https://doi.org/10.3390/insects12040362
Funding
This study was financially funded by the Scientific Research Funding of the Recep Tayyip Erdogan University (Project Number: RTEU-BAP 2015.53001.102.03.04).
Author information
Authors and Affiliations
Contributions
Şengül Alpay Karaoglu: designed the experiments, analyzed the data, completed the final version manuscript, and provided funding acquisition. Arif Bozdeveci: performed the experiments, analyzed the data, and revised the final version of the manuscript. Müberra Pınarbaş Çetin: analyzed data and performed the experiments. Elif Sevim: designed the experiments, and performed the experiments. Şeyma Suyabatmaz: analyzed data and wrote the draft version of the manuscript. Rahşan Akpınar: collected sample materials and participated in the experiments. All authors have reviewed and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that there is no conflict of interest regarding the publication of this paper.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Alpay Karaoğlu, Ş., Bozdeveci, A., Pinarbaş Çetin, M. et al. Isolation, characterization, and genetic diversity of Paenibacillus larvae from AFB suspected specimens in the Central and Eastern Black Sea Regions. Biologia 78, 2919–2929 (2023). https://doi.org/10.1007/s11756-023-01448-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11756-023-01448-w