Abstract
Plumbago auriculata Lam. is an ornamental plant native to South Africa and widely cultivated in China, but the cultivated plants are dominated by a single variety. The development of new varieties is of great commercial interest, and genetic diversity is the foundation of breeding programs. In this study, 85 progenies were obtained by crosses between Plumbago auriculata and Plumbago auriculata f. alba. The genetic diversity of these hybrids was evaluated using horticultural traits and ISSR and SRAP markers. Of the 25 horticultural traits evaluated, the largest variation was found in the beginning of the blooming period, and sepal length was the least variable trait. Correlation analysis showed that the wider the plant, the greater the number of inflorescences and the earlier the flowering. Seven factors explained 65.171% of the total variance; the first factor was leaf morphology, and the second factor was flower morphology. The genetic diversity of the 85 progenies was analyzed using seven pairs of SRAP primers and eight ISSR primers. The average number of effective alleles for 85 hybrids was 1.638, and the average Shannon index value was 0.507. The Nei genetic similarity coefficient indicated that the similarity between WLBS and WSBL was the highest, while that between BLWS and BSWL was the lowest. Analysis of molecular variance (AMOVA) indicated that the main variation was within populations. Cluster analysis based on horticultural traits and molecular markers divided all progenies into seven and five groups, respectively, and there were obvious differences between the two clusters. In this study, we created intermediate materials for future breeding, taking the first step in the cross-breeding of P. auriculata.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The development and utilization of plant germplasm resources are at the core of the commercial ornamental plant industry, especially for plants that are naturally both robust and beautiful (De Souza et al. 2017). Plumbago auriculata Lam., a perennial shrub originating from South Africa, belongs to the family Plumbaginaceae (Jaradat et al. 2016). P. auriculata is the most popular ornamental plant of the genus Plumbago because of its blue flowers and blooming period from early summer to late autumn. P. auriculata f. alba is a form of P. auriculata whose flowers are white. Both of these species are heterostylous and self-incompatible (Ferrero et al. 2009; Lakshmanan et al. 2016). Plumbagin is a secondary metabolite existing in P. auriculata with high biological activity, so previous research focused almost exclusively on the medicinal value of this species (Van de Vijver and Lötter 1971; Deshpande et al. 2014). However, as an excellent ornamental plant, nearly no commercial cultivars exist in P. auriculata, so the breeding of new varieties of this plant should not be neglected.
Although genetic engineering is a current hot topic in ornamental plant breeding, cross-breeding is a traditional breeding method that has long been useful for the development of new varieties (Afkhami-Sarvestani et al. 2012; Kishi-Kaboshi et al. 2018). Genetic diversity is the foundation of breeding programs and is essential for selecting parents and justifying the potential value of new varieties. In research, genetic diversity has generally been evaluated using phenotypic characteristics and molecular markers for efficiency. The combined analysis of morphological traits and molecular markers has been successfully applied to the study of genetic diversity in plants such as bitter gourd (Dey et al. 2006), pepper (Rivera et al. 2016) and chrysanthemum (Baliyan et al. 2014).
Morphological examination is a powerful method for the estimation of genetic diversity, and it is essential for selecting, evaluating and certifying the cultivars obtained through breeding programs (Govindaraj et al. 2015; Yazdanpour et al. 2018). Mutation and correlation analysis can reveal the internal relationships between characteristics, so we can choose those traits that have a large degree of variation and achieve early selection for some traits (Pluta et al. 2012; Rakonjac et al. 2010).
Molecular markers are considered important tools for providing data on genetic polymorphisms between different individuals in a population (Barbosa et al. 2018). This compensates for the fact that morphological characteristics are vulnerable to environmental influence. Among the various types of molecular markers, analysis of inter simple sequence repeats (ISSRs) and sequence-related amplified polymorphisms (SRAPs) does not require prior knowledge of the genome of a species (Li and Quiros 2001; Reddy et al. 2002; Zietkiewicz et al. 1994). In addition, these methods have simple technical requirements, and only a small amount of DNA can be analyzed, so these methods are widely used in genetic analysis (Wang et al. 2009).
This study aimed to evaluate the diversity of progenies between P. auriculata and P. auriculata f. alba as well as to evaluate the potential of these hybrids in the cross-breeding of P. auriculata.
Materials and methods
Plant materials
Eighty-five seeds were produced by crosses between P. auriculata and P. auriculata f. alba and sown in 21-cell trays on February 26, 2019 (Table 1). The plants were transplanted into 15-cm-diameter pots when 5–6 true leaves had developed. The four genitors used in this work were seed-propagated seedlings planted in 35-cm-diameter pots in a plastic greenhouse on campus. The experiment was conducted at Sichuan Agricultural University in Chengdu, Sichuan Province, China (30°42′19″N, 103°51′28″E, 608 m elevation).
Because both of the parents are heterostylous and only compatible with different style types, the parents were distinguished for artificial hybridization by style type (Fig. 1).
Phenotypic evaluation
A total of 25 horticultural traits were evaluated in the 85 hybrids. The 25 traits are defined in Table 2. The traits include flower, leaf blade and plant characteristics. The plant height and width were evaluated 3 months after seeds were sown on May 27, 2019. All the other traits were evaluated in the blooming period in July 2019 except for the time of the first flower bloom.
Measurements of inflorescence length, inflorescence diameter, flower diameter, flower length, pedicel length, sepal length, leaf length, leaf width, stipule width, stipule length, stem thickness and internode length were taken using a Vernier caliper. Plant height and width were measured by a flexible rule. The other qualitative characteristics were measured by visualization. To reduce the error, all experiments are carried out by one person. The beginning of the blooming period was graded in the range of 1–4 based on cluster analysis. Mutation analysis and Pearson correlation coefficients were calculated to analyze the traits with large variation and the relationship between pairs of the 25 horticultural traits. Factor analysis was applied to extract information from the multivariate dataset. Standard deviation transformation was used to standardize the morphological traits, and cluster analysis of 85 hybrids based on morphological traits was performed using the intergroup connectivity Euclidean distance method. All analyses were performed using SPSS 22.0.
DNA extraction
Total genomic DNA of 85 progenies was extracted from a 0.1 g mixture of young leaves for each individual following the CTAB procedure in May 2019 (Doyle 1991). DNA quality was checked on a 1% agarose gel, and DNA quantity was measured by a Nanodrop 2000 (Thermo Scientific, USA) and diluted to 50 ng/μL.
ISSR analysis
We used 10 progenies to screen the 100 ISSR primers (Table S1) published by the University of British Columbia collection (developed by the Biotechnology Laboratory at the University of British Columbia, collection number 9). The screening criteria were that the amplified bands were both clear and polymorphic in the 10 progenies.
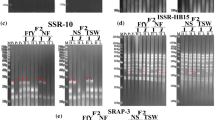
The PCR mixture contained the following: DNA (100 ng), Taq DNA polymerase 1 U, dNTPs 0.15 mmol/L, primer 0.4 μmol/L, Mg2+ 0.5 mmol/L, 10 × PCR buffer 2 μL, and ultrapure water for a final volume of 25 μL. DNA amplification was conducted using a T100 thermocycler (BIO-RAD, Cambridge, MA, USA). PCR conditions included the following: 1 cycle at 94 °C for 5 min, followed by 35 cycles at 94 °C for 30 s, 55 °C for 50 s, and 72 °C for 90 s and 1 cycle at 72 °C for a final extension of 10 min. The amplified fragments were then separated using electrophoresis on a 1% (w/v) agarose gel at 150 V for 30 min in a horizontal tank, and subsequent imaging in a gel scanner (Universal HoodII, BIO-RAD, USA) under ultraviolet light revealed the DNA fragments for analysis.
SRAP analysis
According to the SRAP primer design standard published by Li and Quiros (Li and Quiros 2001), we designed 9 forward primers and 10 reverse primers (Table S1), with a total of 90 pairs of SRAP primers. Similarly, 10 progenies were used to screen the 90 pairs of primers for polymorphism.
The PCR mixture contained the following: DNA (50 ng), Taq DNA polymerase 1.25 U, dNTPs 0.25 mmol/L, forward primer 0.6 μmol/L, reverse primer 0.6 μmol/L Mg2+ 1.0 mmol/L, 10 × PCR buffer 2 μL, and ultrapure water for a final volume of 25 μL. DNA amplification was conducted using a T100 thermocycler (BIO-RAD, Cambridge, MA, USA). PCR conditions are as follows: one cycle at 94 °C for 5 min, followed by five cycles at 94 °C for 1 min, 35 °C for 1 min, and 72 °C for 1 min. This was followed by 35 cycles of 94 °C for 1 min, 50 °C for 1 min, and 72 °C for 1 min and one cycle at 72 °C for a final extension of 10 min. The amplified fragments were then separated using electrophoresis on a 2% (w/v) agarose gel at 150 V for 30 min in a horizontal tank, subsequent imaging in a gel scanner (Universal HoodII, BIO-RAD, USA) under ultraviolet light revealed the DNA fragments for analysis.
Molecular marker data analysis
ISSR and SRAP bands in all of the gel profiles were scored visually as present (1) or absent (0) at least twice for each individual. Only reproducible and unambiguous bands were used for scoring. Data were compiled in a binary data matrix using Excel 2016. The genetic diversity, Nei genetic similarity coefficient, genetic distance and AMOVA of four populations were computed using GenALEx 6.502 (Peakall and Smouse 2010). According to the obtained molecular marker data, cluster analysis for the hybrids was performed. The SimQual program in the NTSYS 2.10e software was used to calculate the Nei-Li similarity coefficient between the offspring, then the UPGMA (Unweighted Pair Group Method Analysis) method in the SHAN program was used for cluster analysis, and finally, the tree plot was used to generate the clustering diagram.
Results
Phenotypic variation
Descriptive statistical analysis of 25 horticultural traits for the 85 hybrids included the minimum, maximum, mean, range, standard deviation and coefficient of variation for the traits recorded (Table 2). The specific morphological data of 85 hybrids are shown in Table S2. The results showed that the coefficient of variation of these traits ranged from 5.95% to 61.22%. The variation range of the beginning of the blooming period was the largest, with the earliest and the latest flowering occurring 44 d apart. The sepal length showed the lowest variation coefficient at just 5.95%. The inflorescence number, pedicel length, petal apex shape, style type, leaf apex shape, stipule shape, branch number, and degree of divergence of plants also had high variation coefficients of more than 30%. Most of these traits are related to ornamental value.
The correlation coefficient between pairs of 25 horticultural traits showed that the inflorescence number was positively correlated with plant width but negatively correlated with the beginning of the blooming period (p < 0.01) (Table S3). The beginning of the blooming period was extremely significantly negatively correlated with style type, plant width and internode length. Flower length and flower diameter were positively correlated with inflorescence length, inflorescence diameter, and sepal length, but flower diameter was negatively correlated with flower number per inflorescence. The leaf traits evaluated included leaf length, leaf width, stipule length, and stipule width, which are positively correlated with each other and correlated with the thickness of the stem and the length of the internode. The degree of divergence of the plant, which influenced the whole plant type, was negatively correlated with plant height and branch number.
In factor analysis, seven factors explained 65.171% of the total variance among the 85 hybrids, and each factor explained more than 5% of the total variance (Table 3). Factor 1 represented leaf traits such as stipule length (r = 0.926), stipule width (r = 0.916), internode length (r = 0.799) and leaf length (r = 0.546), which accounted for 15.049% of the total variance. Factors 2 and 3 represented the flowers, such as inflorescence length, inflorescence diameter, flower length and flower diameter, which accounted for 12.908% and 10.279% of the total variance, respectively. Factor 4 represented the whole plant type, such as the degree of divergence of the plant and the number of branches, which explained 8.158% of the total variance. Factors 5, 6 and 7 represented the qualitative characteristics of flowers and leaves, which accounted for 6.848%, 6.397% and 5.531% of the total variance, respectively.
Cluster analysis based on morphological traits divides all offspring into seven main groups at a Euclidean distance of 20 (Fig. 2). Group I was the largest group, and most of the hybrids flowered in the first blooming period, i.e., bloomed between May 6 and June 4. The hybrids in group II also flowered in the first blooming period, but the inflorescence length and inflorescence diameter were generally smaller than those of the hybrids in group I. All the plants that flowered in the fourth blooming period (i.e., bloomed between July 6 and July 11) were concentrated in group III. Consistent with group I and group II, the hybrids in group IV also flowered in the first blooming period, but the plants were higher than those plants in group I and group II, with obvious growth. Group V had only one individual (No. 85), and all the horticultural traits of this plant were smaller than those of the other offspring. Similar to group V, group VI also had only one individual (No. 4), but the difference was that the floral and leaf organs of this plant were significantly larger than those of other offspring, especially in inflorescence diameter, flower length, leaf length, leaf width and stipules. Group VII had two individuals (No. 7 and No. 8), and the leaves of these two plants were wider than those of the other offspring.
Polymorphism analysis of primers
According to the primers’ ability to yield clear, polymorphic, and reproducible patterns of amplification, eight ISSR primers and seven pairs of SRAP primers were selected for subsequent genetic diversity analysis (Table 4). The number of effective alleles generated by the SRAP primers ranged from 1.323 (ME5-EM3) to 1.741 (ME9-EM10), with an average of 1.615 (Table 4). The number of effective alleles of the ISSR primers ranged from 1.400 (UBC828) to 1.779 (UBC808), with an average of 1.629. The Shannon information index ranged from 0.266 (ME5-EM3) to 0.598 (ME3-EM9), with an average of 0.487 for SRAP primers. This index was higher for the ISSR primers than for the SRAP primers, with an average of 0.503. The uHe values were greater than the He values, but their variance trends were the same.
Genetic diversity analysis
A summary of the mean genetic variation statistics of all four populations as well as the mean of all loci are presented in Table 5. Under the assumption that the population is in Hardy–Weinberg equilibrium, the effective number of alleles ranged from 1.530 to 1.711 with a mean value of 1.638. For the Shannon information index, the lowest value was 0.431, and the largest value was 0.565. The population WLBS had the highest genetic diversity, with 91.80% polymorphic loci. The mean values for the number of alleles, number of effective alleles, Shannon information index, expected heterozygosity and the percentage of polymorphic loci of the 85 hybrids were 1.787, 1.638, 0.507, 0.352, and 82.38%, respectively.
The pairwise population matrix of Nei genetic distance and Nei genetic identity further elucidated the gene differentiation between populations (Table 6), which showed that the populations WSBL and WLBS had the highest Nei genetic identity (0.963), and the lowest values were found for BSWL and BLWS (0.932).
The results obtained by AMOVA illustrated that most of the genetic variations existed within populations (Table 7). It was observed that 93% of the total variation corresponded to intra-population variation, and 7% occurred between populations.
Based on molecular marker data, 85 hybrids were divided into five groups at a similarity coefficient of approximately 0.66 (Fig. 3). Combined with the horticultural traits, we found that clustering based on molecular markers was not significantly associated with the phenotype. Group I contained the vast majority of hybrids. Group II had three offspring from BSWL combinations (No. 13, 18 and 20) as well as one offspring from WLBS (No. 54) and WSBL (No. 66). Group III consisted mainly of offspring of BSWL and WLBS (No. 17, 41, 42 and 44). Group IV consisted mainly of offspring, with P. auriculata f. alba as the female parent (No. 45 and 55). Similar to group IV, the plants in group V almost came from the offspring with P. auriculata f. alba as the female parent, and only one came from BLWS (No. 25). No. 25 was also the only individual of BLWS not divided into group I.
Discussion
Phenotypic variation
Morphological traits are used in the protection of intellectual property associated with new varieties based on the regulations of the international Union for the Protection of New Varieties of Plants (UPOV) (Hong et al. 2015). These traits are widely used to estimate phenotypic variation and select parental lines for crossings because of the convenience of this method (Lee and Park 2017). The method employs simple tools, such as flexible rules and Vernier calipers, to obtain a large amount of data in a short time. In this study, we estimated 25 horticultural traits in 85 hybrids. Generally, if the traits showed coefficient variance values of more than 30%, we can consider the given character to have potential in breeding (Luo and Dai 2010). The beginning of the blooming period had the highest coefficient variance (61.22%); however, the data were collected from the first year and might be affected by the plant growth rate, so the selection potential of these traits should be judged after another one or two years of observation. In addition, among the nine traits whose coefficient variance was more than 30%, the number of inflorescences, number of branches and degree of divergence of plants directly influenced the ornamental value of plants, so we can strengthen the selection of these traits in the breeding program. A surprising finding was that although we found that the flower diameter showed high variance over the course of measurement, the final results indicated that this trait was stable, with a coefficient variance of only 8.41%.
Correlation analysis can reveal the potential relationships between characters, so we can make early selection for objective traits through these relationships (Hui et al. 2016; Litrico and Violle 2015). In this study, we found that those traits related to flowers had significant correlations, such as flower diameter, flower length, inflorescence length, inflorescence width and sepal length. In addition, we also found some useful information for breeding programs. The plant width is significantly positively correlated with inflorescence numbers and negatively correlated with the beginning of the blooming period, which means that the larger the plant width, the more inflorescences and the earlier the blooming period. Therefore, we can measure the plant width to predict the plants that are early flowering and will have higher inflorescence numbers. In addition, leaf length and leaf width were significantly positively correlated with flower length, and a longer flower usually means larger inflorescences through their correlations. Therefore, we aim to breed large-flowered varieties and can achieve this target by examining leaves in the vegetative growth period to shorten the breeding time. Another interesting finding was that plant height was significantly negatively correlated with flower width, possibly because vegetative growth competes with reproductive growth for nutrition.
Factor analysis is a method of data reduction to decrease original multitrait variation to a limited number of uncorrelated new variables (Yanai and Ichikawa 2006). In factor analysis, seven factors explained approximately 65% of the total variance. Factor 1 and factor 2 explained 27.957% of the total variance; factor 1 mainly represented leaf blade traits, and factor 2 represented flower traits. This means that the leaves and flowers have an important effect on the variation of morphological traits.
Genetic diversity analysis revealed by ISSR and SRAP markers
In recent decades, molecular markers based on PCR techniques have been demonstrated to be effective in studying the relationship or diversity between different species (Ibarra-Torres et al. 2015). Seven pairs of SRAP primers and 8 ISSR primers were used in the present study. All of them showed high polymorphism, which indicated that those primers had high identification abilities. In other species, such as Salvia (Peng et al. 2014) and apricot (Li et al. 2014), it was also found that ISSR and SRAP markers were efficient. The most efficient primer was UBC808, with Na, Ne, I, He and uHe values of 2.0, 1.779, 0.618, 0.429, and 0.441, respectively. In this study, we found that the number of alleles (1.772), number of effective alleles (1.629), Shannon index (0.503), expected heterozygosity (0.349), and unbiased heterozygosity (0.352) generated by ISSR primers were higher than those generated with SRAP primers (Na = 1.743, Ne = 1.615, I = 0.487, He = 0.338, uHe = 0.347). The same results were also found in research on Salvia miltiorrhiza (Song et al. 2010). The possible reason for these findings might be that the ISSR markers revealed the diversity of the whole genome, while the SRAP markers only amplified the target region of the open reading frame (ORF). However, the disparity between the two markers was small, and we think that both are effective.
The Shannon diversity index is an important genetic parameter that can estimate phenotypic or genotypic diversity within a population. This parameter ranged from 0.431 to 0.565 for the four populations, indicating high variability within the populations. Obviously, genetic recombination was found in the four populations after crossing. The results of the Nei genetic identity pairwise population matrix showed that populations WLBS and WSBL had the highest genetic similarity, but when P. auriculata was the female parent, the offspring had the lowest Nei genetic identity. This might be because the long and short styles of P. auriculata undergo genetic differentiation over the course of evolution. In addition, populations WLBS and WSBL had a higher percentage of polymorphic loci than did BLWS and BSWL. Thus, genetically speaking, with P. auriculata f. alba as the female parent, we can obtain more plant material with genetic differences. However, the Nei genetic identity between the four populations was higher than 0.9, which shows that the genetic relationship of the parents was very close and further proves the plant classification. AMOVA can divide the total variance into its components between and within populations. In the present work, it was shown that the highest proportion of variation was found within populations (93%), while the variation between populations was just 7%. Barbosa et al. (2018) also reported that the highest proportion of variation was found within full-sib families (73.49%) in research on Jatropha curcas. Therefore, we should consider the internal variance to avoid genetic narrowing, which may impact selection and recombination in future breeding programs (Bhering et al. 2015).
Cluster analysis
Cluster analysis based on phenotypic traits divided all offspring into seven groups. Group I contained most of the hybrids, indicating that the hybrids were consistent in phenotype. Additionally, we found that the clusters were divided by floral quantitative traits and leaf quantitative traits, which also confirmed that flowers and leaves are the main factors that caused the difference in hybrids in factor analysis. Interestingly, group V and group VI both have only one hybrid. No. 4 in group VI was significantly larger than the other offspring in florescence diameter, flower length and leaf size. Therefore, we could use No. 4 as a potential material with large flower breeding value.
Cluster analysis based on molecular markers divides all offspring into 5 groups. This clustering is quite different from phenotypic traits, probably because the markers we selected are universal markers and cannot cover the genes that control horticultural traits. However, from the results of the clustering, we also found that when P. auriculata f. alba was the female parent, the offspring were distributed in each group, confirming the results of the Shannon index, that is, when P. auriculata f. alba was the female parent, the genetic diversity of offspring was higher. As a result, the development of specific molecular markers, such as EST-SSR, should be carried out in a later breeding program, which also shows that it is very effective and necessary to combine morphological traits with molecular markers to analyze the genetic diversity of hybrids.
In conclusion, 85 progenies with high genetic diversity were obtained by intraspecific crossing. The relationship between the traits can guide us to select plants that have a large amount of inflorescence through the examination of plant width. The genetic diversity analyzed by ISSR and SRAP markers showed that population WLBS presented the greatest genetic divergence, and we will continue to observe this population and judge its value in the follow-up breeding program. These results are valuable for the innovation of P. auriculata germplasm used in cross-breeding.
Author contribution statement
XC and SP-G conceived the manuscript and its components. All authors researched the literature. TL, WJ-L, YR-L, XF-Y, LJ-Y and JN-L conceived and designed the Figures. PS, YQ-L and LS-S wrote the manuscript. All authors read and approved the final document
References
Afkhami-Sarvestani R, Serek M, Winkelmann T (2012) Interspecific crosses within the Streptocarpus subgenus Streptocarpella and intergeneric crosses between Streptocarpella and Saintpaulia ionantha genotypes. Sci Hortic 148:215–222. https://doi.org/10.1016/j.scienta.2012.10.006
Baliyan D, Sirohi A, Kumar M, Kumar V, Malik S, Sharma S (2014) Comparative genetic diversity analysis in chrysanthemum: a pilot study based on morpho-agronomic traits and ISSR markers. Sci Hortic 167:164–168. https://doi.org/10.1016/j.scienta.2013.12.029
Barbosa DA, Silva GW, Silvia N, Toledo PMC, Alves RBR, Borges FT, Medeiros PPT (2018) Genetic diversity between and within full-sib families of Jatropha using ISSR markers. Ind Crop Prod 124:899–905. https://doi.org/10.1016/j.indcrop.2018.08.066
Bhering LL, Peixoto LdA, Ferreira-Leite NLS, Laviola BG (2015) Molecular analysis reveals new strategy for data collection in order to explore variability in Jatropha. Ind Crop Prod 74:898–902. https://doi.org/10.1016/j.indcrop.2015.06.004
De Souza EH, Versieux LM, Duarte SFV, Lanzoni RM, de Carvalho CMAP, Pinheiro MA (2017) Interspecific and intergeneric hybridization in Bromeliaceae and their relationships to breeding systems. Sci Hortic 223:53–61. https://doi.org/10.1016/j.scienta.2017.04.027
Deshpande J, Labade D, Shankar K, Kata N, Chaudhari M, Wani M, Khetmalas M (2014) In vitro callus induction and estimation of plumbagin content from Plumbago auriculata Lam. Indian J Exp Biol 52:1122–1127
Dey SS, Singh AK, Chandel D, Behera TK (2006) Genetic diversity of bitter gourd (Momordica charantia L.) genotypes revealed by RAPD markers and agronomic traits. Sci Hortic 109:21–28. https://doi.org/10.1016/j.scienta.2006.03.006
Doyle J (1991) DNA protocols for plants. Mol Tech in Tax 57:283–293. https://doi.org/10.1007/978-3-642-83962-7_18
Ferrero V, de Vega C, Stafford GI, Van Staden J, Johnson SD (2009) Heterostyly and pollinators in Plumbago auriculata (Plumbaginaceae). S Afr J Bot 75:778–784. https://doi.org/10.1016/j.sajb.2009.06.014
Govindaraj M, Vetriventhan M, Srinivasan M (2015) Importance of genetic diversity assessment in crop plants and its recent advances: an overview of its analytical perspectives. Genet Res Int 2015:431487. https://doi.org/10.1155/2015/431487
Hong JH, Kwon YS, Mishra RK, Kim DH (2015) Construction of EST-SSR databases for effective cultivar identification and their applicability to complement for lettuce (Lactuca sativa L.) Distinctness. Test Am J Plant Sci 06:113–125. https://doi.org/10.4236/ajps.2015.61013
Ibarra-Torres P, Valadez-Moctezuma E, Pérez-Grajales M, Rodríguez-Campos J, Jaramillo-Flores ME (2015) Inter- and intraspecific differentiation of Capsicum annuum and Capsicum pubescens using ISSR and SSR markers. Sci Hortic 181:137–146. https://doi.org/10.1016/j.scienta.2014.10.054
Jaradat NA, Zaid AN, Hussein F (2016) Investigation of the antiobesity and antioxidant properties of wild Plumbago europaea and Plumbago auriculata from North Palestine. Chem Biol Technol Agric 3:31. https://doi.org/10.1186/s40538-016-0082-4
Kishi-Kaboshi M, Aida R, Sasaki K (2018) Genome engineering in ornamental plants: current status and future prospects. Plant Physiol Biochem 131:47–52. https://doi.org/10.1016/j.plaphy.2018.03.015
Lakshmanan G, Bupesh G, Vignesh A, Sathiyaseelan A, Murugesan K (2016) Micropropagation and anticancer activity of methanolic extract of Plumbago auriculata Lam. Int J Adv Biotech Res 7:2001–2011. https://www.researchgate.net/publication/312197033
Lee ON, Park HY (2017) Assessment of genetic diversity in cultivated radishes (Raphanus sativus ) by agronomic traits and SSR markers. Sci Hortic 223:19–30. https://doi.org/10.1016/j.scienta.2017.05.025
Li G, Quiros CF (2001) Sequence-related amplified polymorphism (SRAP), a new marker system based on a simple PCR reaction: its application to mapping and gene tagging in Brassica. Theor Appl Genet 103:455–461. https://doi.org/10.1007/s001220100570
Li M, Zhao Z, Miao XJ (2014) Genetic diversity and relationships of apricot cultivars in North China revealed by ISSR and SRAP markers. Sci Hortic 173:20–28. https://doi.org/10.1016/j.scienta.2014.04.030
Litrico I, Violle C (2015) Diversity in plant breeding: a new conceptual framework. Trends Plant Sci 20:604–613. https://doi.org/10.1016/j.tplants.2015.07.007
Luo XY, Dai SL (2010) Taxonomic analysis of morphological characters of large-flowered chrysanthemum cultivars. J Beijing Forestry Uni 32:135–140. https://doi.org/10.13332/j.1000-1522.2010.03.013
Peakall R, Smouse PE (2010) GENALEX 6: genetic analysis in Excel. Population genetic software for teaching and research. Mol Ecol Resour 6:288–295. https://doi.org/10.1111/j.1471-8286.2005.01155
Peng L, Ru M, Wang BQ, Wang Y, Li B, Yu J, Liang ZS (2014) Genetic diversity assessment of a germplasm collection of Salvia miltiorrhiza Bunge. based on morphology, ISSR and SRAP markers. Biochem Syst Ecol 55:84–92. https://doi.org/10.1016/j.bse.2014.01.020
Pluta S, Mądry W, Sieczko L (2012) Phenotypic diversity for agronomic traits in a collection of blackcurrant (Ribes nigrum L.) cultivars evaluated in Poland. Sci Hortic 145:136–144. https://doi.org/10.1016/j.scienta.2012.07.036
Rakonjac V, Akšić MF, Nikolić D, Milatović D, Čolić S (2010) Morphological characterization of ‘Oblačinska’ sour cherry by multivariate analysis. Sci Hortic 125:679–684. https://doi.org/10.1016/j.scienta.2010.05.029
Reddy MP, Sarla N, Siddiq EA (2002) Inter simple sequence repeat (ISSR) polymorphism and its application in plant breeding. Euphytica 128:9–17. https://doi.org/10.1023/A:1020691618797
Rivera A, Monteagudo AB, Igartua E, Taboada A, García-Ulloa A, Pomar F, Riveiro-Leira M, Silvar C (2016) Assessing genetic and phenotypic diversity in pepper (Capsicum annuum L.) landraces from North-West Spain. Sci Hortic 203:1–11. https://doi.org/10.1016/j.scienta.2016.03.006
Song ZQ, Li XF, Wang HG, Wang JH (2010) Genetic diversity and population structure of Salvia miltiorrhiza Bge in China revealed by ISSR and SRAP. Genetica 138:241–249. https://doi.org/10.1007/s10709-009-9416-5
Van de Vijver LM, Lötter AP (1971) The constituents in the roots of Plumbago auriculata Lam. and Plumbago zeylanica L. responsible for antibacterial activity. Planta Med 20:8–13. https://doi.org/10.1055/s-0028-1099658
Wang HZ, Feng SG, Lu JJ, Shi NN, Liu JJ (2009) Phylogenetic study and molecular identification of 31 Dendrobium species using inter-simple sequence repeat (ISSR) markers. Sci Hortic 122:440–447. https://doi.org/10.1016/j.scienta.2009.06.005
Xia H, Zhao GH, Zhang LS, Sun XY, Yin SP, Liang DY, Li Y, Zheng M, Zhao XY (2016) Genetic and variation analyses of growth traits of half-sib Larix olgensis families in northeastern China. Euphytica 212:1–11. https://doi.org/10.1007/s10681-016-1765-4
Yanai H, Ichikawa M (2006) Factor Analysis. . Handbook Statistics 26:257–296. https://doi.org/10.1016/S0169-7161(06)26009-7
Yazdanpour F, Khadivi A, Etemadi-Khah A (2018) Phenotypic characterization of black raspberry to select the promising genotypes. Sci Hortic 235:95. https://doi.org/10.1016/j.scienta.2018.02.071
Zietkiewicz E, Rafalski A, Labuda D (1994) Genome fingerprinting by simple sequence repeat (SSR)-anchored polymerase chain reaction amplification. Genom 20:176–183. https://doi.org/10.1006/geno.1994.1151
Acknowledgements
Special thanks to Sichuan Gaonong Ecological Science and Technology Co., Ltd and SICHUAN TIANYI ECOLOGICAL GARDEN GROUP CO., LTD and American journal experts-AJE Company for editing the language of the original manuscript.
Funding
National Key R&D Program of China (2018YFD0600105); Sichuan Science and Technology Program (2018JY0211); Sichuan Science and Technology Program (2017N20008); Sichuan Science and Technology Program (2016NYZ20038).
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by J. Huang.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Chen, X., Gao, S., Shen, P. et al. Genetic diversity analysis of intraspecific hybridization between Plumbago auriculata and Plumbago auriculata f. alba based on horticultural traits and molecular markers. Acta Physiol Plant 43, 31 (2021). https://doi.org/10.1007/s11738-020-03188-9
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11738-020-03188-9