Abstract
Key message
This work identifies a new rose NAM/CUC subgroup member, RhNAC31. Overexpression of RhNAC31 in Arabidopsis confers salt, cold, and drought tolerance along with enhanced ABA sensitivity.
Abstract
Plant-specific NAM, ATAF1/2, and CUC (NAC) transcription factors serve essential functions in plant development and plant responses to environmental cues. Yet, transcription factors specific to the rose (Rosa hybrida) NAM/CUC3 subfamily are poorly understood. Here, we identify a novel NAM/CUC3-subfamily transcription factor, RhNAC31, that is associated with flower opening and can be induced by increased salt, cold, and dehydration stress treatment. RhNAC31 has a transactivation region in its C-terminal region, and its overexpression is associated with enhanced cold tolerance in Arabidopsis, conferring a higher survival rate and reduced reactive oxygen (H2O2 and O2−) levels. Under salt stress conditions, plants overexpressing RhNAC31 displayed increased germination rates and lower levels of H2O2, malondiadehyde (MDA), peroxidase (POD), and superoxide dismutase (SOD). Moreover, RhNAC31 conferred enhanced drought resistance by reducing the rate of water loss through leaves. Further characterization revealed a higher sensitivity of RhNAC31 transgenic plants to abscisic acid (ABA) both during and post-germination, causing lower germination and root inhibition rates under ABA treatment. Quantitative PCR experiments showed that numerous abiotic stress-related genes were activated by RhNAC31 overexpression. Our results highlight RhNAC31 as a positive transcriptional regulator of tolerance to multiple abiotic pressures, and we conclude that RhNAC31 has potential for use in the molecular breeding of stress-tolerant crops.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Plants have evolved numerous strategies to adapt to biotic and abiotic stressors. Sessile plant development, maturation, and productivity are influenced by aversive environmental cues, such as changes in temperature, reduced access to water, and salt stress. Numerous regulatory/transport proteins and functional proteins participate in response to continually changing environmental conditions.
Over the last decade, research has shown that many transcription factors (TFs) genes that act as regulators of downstream gene expression facilitate abiotic stress resistance in plants. In particular, NAC TFs, including NAM, ATAF1/2, and CUC, are widespread in many plant species (Olsen et al. 2005; Lim et al. 2007; Kim et al. 2009; Balazadeh et al. 2010; Liu et al. 2019). Comprehensive genomic analyses have found different NAC protein numbers in woody and herbaceous plants. For example, 117, 151, 152, 163, and 80 NAC members were found in the genomes of Arabidopsis, rice, soybean, poplar, and buckwheat, respectively (Hu et al. 2010; Pinheiro et al. 2009; Nuruzzaman et al. 2010). These proteins play pivotal roles in plant development, senescence, and tolerance of abiotic stressors (Lee et al. 2012; Nakashima et al. 2012; Puranik et al. 2012; Fang et al. 2015; Yu et al. 2018).
The N-terminus of NAC contains a conserved domain associated with DNA binding as well as a divergent C-terminus that regulates proximal transcription (Olsen et al. 2005). Comprehensive phylogenic analysis has revealed that NAC proteins can be divided into several major sub-families according to their domain architecture. These include no apical meristem/cup-shaped cotyledon (NAM/CUC3), vascular-related NAC domain (VND), secondary wall-associated NAC domain (SND), and stress-associated NAC (SNAC) proteins. Members of the SNAC protein subgroup have been identified as positive regulators of abiotic stress responses in both dicots and monocots. Arabidopsis RD26/ANAC072 is involved in drought, salt, and pathogen response signaling, and is modulated by plant hormones such as jasmonate (JA) and abscisic acid (ABA) (Fujita et al. 2004). Rice SNAC3 has been found to enhance resistance to high temperatures and drought by regulating reactive oxygen species levels (Fang et al. 2015), and banana MaNAC1 participates in the low-temperature response by interacting with a key regulator of cold acclimation, MaICE1 (Shan et al. 2014). Recently, several studies have suggested that NAM/CUC3 subfamily NAC proteins are involved in boundary maintenance, secondary wall synthesis, cell division, leaf senescence (Mao et al. 2017), and organ development (Zhang et al. 2018). Although NAC TFs have attracted extensive attention, to date, it is unknown whether NAM/CUC3 subfamily members act as master regulators of abiotic stress responses.
As one of the most well-known ornamental plants worldwide, rose (Rosa hybrida) is of high economic value. Most rose cultivars encounter environmental pressures that restrict their growth and affect their economic value. Therefore, exploiting genes that enhance rose stress resistance may offer considerable economic potential. Recently, various dehydration-related TFs, including NACs, have been examined in rose using cDNA microarray analyses (Dai et al. 2012). Several NAC TFs have been found to be transcriptionally induced by dehydration, or by ethylene released from cut rose flowers, but few of these TFs have been functionally characterized. It has been shown that RhNAC3, a member of the SNAC family, exhibits better performance under drought stress. RNA-silenced RhNAC3 lines showed decreased dehydration tolerance and reduced osmotic stress-related gene expression (Jiang et al. 2014). RhNAC2 belongs to the NAM/CUC3 subfamily and has been found to increase dehydration tolerance by regulating downstream genes related to cell expansion (Dai et al. 2012). Another NAM/CUC3 subfamily TF, RhNAC100, the closest homolog of AtNAC100, plays a role in petal cell expansion by fine-tuning the effects of miRNA164 and RhNAC100 on ethylene signaling (Pei et al. 2013). However, to date, there is no clear information regarding the involvement of NAC TFs in the development of rose plants. In this study, a novel development-related NAC transcription activator, RhNAC31, is upregulated in rose by several abiotic stressors, including dehydration, salt, and low temperatures. Overexpressing RhNAC31 in Arabidopsis exhibited enhanced tolerance to increased salinity, cold, and drought treatments without affecting biomass. Moreover, RhNAC31 transgenic plants were sensitive to ABA application both during and post-germination. Accordingly, numerous genes associated with responses to abiotic stress were upregulated by RhNAC31 overexpression. These findings indicate that RhNAC31 affects abiotic stress response and development in rose, and thereby may be a regulator of interest for horticultural plant engineering.
Materials and methods
Plant material and treatments
For gene expression analysis (Ma et al. 2006), Rosa hybrida ‘Samantha’, opening stage 2, was used. For salt plus ABA treatment, flower petals from the middle whorl were soaked in a solution of 100 μM ABA in 60 mM NaCl for 0, 6, 12, and 24 h, respectively. For the cold plus ABA treatment, rose petals were incubated in 100 μM ABA solution at 4 °C for 0, 6, 12, and 24 h, respectively. As a negative control, 0.1% DMSO was administered to mock samples in the absence of phytohormones. Samples of the petals, sepals, leaves, and stems of rose flowers at opening stage 2 used for RNA isolation were frozen at − 80 °C. An RNAprep Pure Plant kit (Tiangen, Beijing, China) was used to isolate RNA. The rose Ubiquitin1 gene (RhUbi1, Gene ID. JK622648) served as an internal control. A minimum of three biological samples was used in each experiment. Primer sequences can be found in Table S1.
Sequence and phylogenic analysis of RhNAC31
The predicted sequence of RhNAC31 was obtained from the rose flower suppression subtractive hybridization library (Dai et al. 2012). Specific primers (5′-ATGGGTCTGAGGGACATTGGA-3′ and 5′-TCATAGCAAAACCATGCTGTTCTC-3′) were used to amplify the full length of the RhNAC31-coding sequence by PCR. Moreover, we used thermal asymmetric interlaced PCR (TAIL-PCR) to isolate the regulatory sequence upstream of RhNAC31 (Liu and Chen 2007). Products were then subcloned into pMD-18 vectors (TaKaRa, Dalian, China). The resulting recombinant plasmid DNA was validated by sequencing and transformed into DH5α competent Escherichia coli cells. ClustalW was used to generate multiple sequence alignments of RhNAC31 and other NAC proteins (Chenna et al. 2003). MEGA5.0, using the Neighbor-Joining method and 500 bootstrap replicates (Tamura et al. 2011), was used to produce a phylogenic tree. Plant cis-acting Regulatory DNA Elements (PlantCARE) was used for the analysis and annotation of RhNAC31 (Lescot et al. 2002). Primer sequences can be found in Table S1.
RhNAC31 transactivation assays in yeast
For transactivation assays in yeast (Saccharomyces cerevisiae), the full-length RhNAC31 protein, as well as versions of the protein with N- and C-terminal truncations, were PCR amplified and subcloned into empty pBD vectors (Stratagene, La Jolla, CA, USA). The empty vector pBD, as well as the positive vector pGAL4, pBD-RhNAC31, pBD-RhNAC31-N, and pBD-RhNAC31-C were then transformed into the yeast host strain YRG-2. For a detailed protocol, see the Yeast Protocols Handbook PT3024-1 (Clontech). The resulting transformants were grown on synthetic dextrose (SD) media deficient in tryptophan (SD/Trp−) or tryptophan and histidine (SD/Trp−/His−). Transformed yeast cells were incubated at 30 °C for 3–8 h in the presence of X-gal. The formation of blue colonies indicated the presence of β-galactosidase. All primer sequences can be found in Table S1.
Vector construction and Arabidopsis transformation
The RhNAC31 open-reading frame (primers RhNAC31-FL-Xba I and RhNAC31-FL-Pst I) was subcloned into the pCAMBIA 1300 vector (Jiang et al. 2014). The floral dip method was used to transform the recombinant plasmid into wild-type (WT) Arabidopsis (Columbia), using Agrobacterium tumefaciens strain GV3101 (Clough and Bent 1998). The resulting seeds were grown in MS media including 50 mg/L hygromycin and those exhibiting 100% germination were used thereafter.
Seed germination in the presence of NaCl and ABA
To determine the effect of RhNAC31 on salt tolerance, approximately 20 seeds each of three lines of RhNAC31-overexpressing plants (OE# 2, 4, and 7), WT, and vector control (VC) were added to MS media containing 0, 50, 100, or 150 mM NaCl. To assess ABA sensitivity of seeds during germination, a second set of seeds (~ 20 seeds per line) were sown in MS plates saturated with 0, 0.2, 0.4, or 0.8 μM ABA. Following 3 days of stratification, seeds were exposed to 40–60% relative humidity and of 80–100 μmol/m2/s light intensity within a closed chamber. With a 16 h light/8 h dark cycle, seed germination proceeded at 23 ± 1 °C. After 7 days, we recorded the emergence rate of free radicals and the rate of cotyledon greening. Three independent biological replicates were used.
Root growth phenotype
To determine root growth in response to high salt and ABA treatments, the root growth of RhNAC31-overexpressing (OE# 2, 4, and 7), WT, and VC seedlings were examined. Plants grown in MS media for 7 days were transferred to MS plates containing NaCl (0, 50, 100, 150, or 200 mM) or ABA (0, 5, 10, or 30 μM). After 10 days of vertical growth, ImageJ (http://rsbweb.nih.gov/ij/) was used to measure the primary root length and the lateral root number. Plant dry weights were recorded for each NaCl or ABA treatment. Ten technical and three independent biological replicates were used for each treatment.
In situ histochemical localization of hydrogen peroxide (H2O2) and superoxide anion (O2 −)
H2O2 and O2− levels in plant tissues were detected in situ using diaminobenzidine (DAB) and nitroblue tetrazolium (NBT) staining as described by Shi et al. (2010). After 10 days of growth, H2O2, and O2− were detected in RhNAC31 transgenic, WT, and VC seedlings by immersion in either water (CK) or 200 mM NaCl for 1 h. For H2O2 detection, seedlings were vacuum-infiltrated in the dark using fresh DAB (1 mg/mL solution, pH 3.8) until brown sediment appeared. For O2− detection, samples were vacuum-infiltrated in natural light with fresh NBT in phosphate buffer (1 mg/mL, pH 7.8) until blue sediment were seen. Seedlings were then treated with concentrated and 90% ethanol for destaining prior to photographing using a Stemi DV4 light microscope (Carl Zeiss, Gottingen, Germany). H2O2 and O2− were quantitatively analyzed as per Wang et al. (2017). Ten technical and three biological replicates were used for each condition tested.
Determination of drought tolerance of RhNAC31 transgenic Arabidopsis
After 3 weeks, seedlings from the RhNAC31 overexpressing (OE# 2, 4, and 7), WT, and VC lines were grown in trapezoidal pots (10 cm wide at the top, 5 cm wide at the bottom, and 6 cm deep) in a mixture of vermiculite and humus (v/v 1:1), and were maintained at 23 ± 1 °C under a 16-h light/8-h dark cycle. All plants in the same growth conditions experienced drought for 15 days, and survival rates were measured 7 days after irrigation. More than 50 plants were used for each comparison. The degree of water loss in aboveground portions of the 3-week-old seedlings was assayed at designated timepoints on the laboratory bench (100 µmol/m2/s light intensity, 30–40% relative humidity, and 23–25 °C). Six seedlings from each independent line were used. The experiment included three independent biological replicates.
Quantification of malondialdehyde (MDA), peroxidase (POD), and superoxide dismutase (SOD)
MDA content was detected as described by Jouve et al. (2007) with little modification. For all lines, 0.2 g of fresh leaf tissue was ground and blended with 5 mL of 10% (w/v) trichloroacetic acid (TCA). Two mL of supernatant was diluted one-to-one with 0.6% (w/v) thiobarbituric acid (TBA) and mixed well. The mixture was then incubated for 30 min at 100 °C. The optical density (OD) value was measured at 450, 532, and 600 nm. MDA concentration was determined using the formula: C = 6.45(A532–A600) − 0.56A450, where C represents the soluble MDA concentration (µmol/L) and A532, A600, and A450 represent the OD values at 532, 600, and 450 nm, respectively. MDA content is expressed in units of μmol/g FW.
Enzymes were extracted as per Meng et al. (2014). In brief, 0.2 g of fresh shoot tissue was homogenized in 2 mL of 50 mM ice-cold buffer (pH 7.8) containing 50 mM K2HPO4 and 50 mM KH2PO4. The tissue was pelleted at a temperature of 4 °C at 12,000g for 20 min and the supernatant was subjected to POD and SOD activity examination. POD activity was determined using guaiacol (Nakano and Asada 1981). Enzyme extracts were mixed with 100 mM sodium acetate buffer (pH 6.0), 0.3% guaiacol solution, and 0.3% H2O2. To assess guaiacol oxidation, the OD value was measured at 470 nm over 1 min, with a unit of POD activity defined as a 0.01 unit change in absorbance per minute. SOD activity was determined using the photochemical NBT method (Beauchamp and Fridovich 1971); in this method, a unit is the amount of enzyme causing 50% inhibition at 560 nm. Enzyme extracts were mixed with 50 mM phosphate buffer (pH 7.8), 130 mM methionine, 750 μM NBT, 20 μM riboflavin, 100 μM EDTA Na2, and deionized water. Each sample included at least ten leaves or six seedlings from different independent lines.
Quantitative reverse transcription PCR analysis
Quantitative reverse transcription PCR (qRT-PCR) was performed using total RNA isolated from leaves of 3-week-old seedlings from the RhNAC31 transgenic and VC plant lines. RNA (1 µg) was treated with DNase and first-strand cDNA synthesis was performed using a PrimeScript RT reagent Kit with gDNA Erase (TaKaRa, Dalian, China). Using a 1–10 dilution of cDNA, qRT-PCR was conducted with the TaKaRa™ SYBR® FAST qPCR Kit (TaKaRa, Dalian, China). Actin 2 (GenBank: NM_112764) from Arabidopsis thaliana served as an internal control. Nine genes were selected. Each qRT-PCR assay used at least three biological replicates. Primer sequences can be found in Table S1.
Statistical analysis
We performed analyses of variance (ANOVAs) followed by Tukey’s range tests to assess the statistical significance of differences in mean values. In all analyses, a p value less than 0.05 was indicative of statistical significance. All analyses were carried out using SPSS Statistics v17.0 (IBM SPSS, Chicago, IL, USA).
Results
Isolation and sequence analysis of RhNAC31
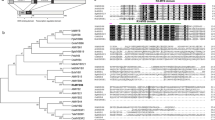
Numerous transcription factors (TFs) were obtained from a cDNA microarray analysis of rose flowers subjected to dehydration and rehydration treatments. A single cDNA encoding a full-length NAC protein was identified and named RhNAC31 (GenBank accession number: MF576436). RhNAC31 is a putative protein 286 amino acids long with an isoelectric point of approximately 5.60 and theoretical molecular weight of 32.8 kD. Phylogenic analysis revealed that RhNAC31 is highly homologous to Arabidopsis CUC3, which belongs to the NAM/CUC3 subgroup of NAC proteins (Fig. 1a). Multiple sequence alignment analysis with other reported rose NAC proteins revealed that RhNAC31 had 26.12% identity with RhNAC2 (GenBank: AFQ21786.1), 26.41% identity with RhNAC3 (Genbank: JK617768), and 23.12% identity with RhNAC100 (GenBank: AFS95065.1). Motif scan analysis showed that RhNAC31 has distinct N- and C-terminal regions. The N-terminal region contains five different heavily conserved motifs that serve together as a DNA-binding domain (Figs. 1b, S1). The highly divergent C-terminal domain functions in the regulation of downstream gene transcription. Bioinformatics analysis suggests that the promoter of the RhNAC31 gene (1.0 kb upstream of the transcription start site) contains numerous stress-response-related cis-elements, including MYB and HSE recognition sites, as well as one low-temperature-responsive (LTR) element (Fig. 1c). These results indicate that RhNAC31 may function in plant stress response.
Phylogenetic, multiple sequence alignment, and cis-acting element analysis of RhNAC31. a Phylogenic relationships between RhNAC31 and other plant NAC protein subgroup members. A neighbor-joining tree was produced using MEGA 5.0 and 500 bootstrap replicates. GenBank accession numbers are: RhNAC3 (JK617768), GmNAC4 (AAY46124.1), ANAC072 (AT4G27410), ANAC019 (AT1G52890), ANAC055 (AT3G15500), ATAF1 (AT1G01720), ATAF2 (AT5G08790), SNAC1 (NC_029258.1), TANAC67 (AHB32901.1), SNAC3 (LOC_Os01g09550), GmNAC11 (ACC66315.1), ANAP (AT1G06590), TANAC69 (AY625682), RhNAC2 (AFQ21786.1), RhNAC100 (AFS95065.1), CUC1 (AT3G15170), CUC2 (AT5G53950), CUC3 (AT1G76420), RhNAC31 (MF576436), VND1 (AT2G18060), VND2 (AT4G36160), VND3 (AT5G66300), VND4 (AT1G12260), VND5 (AT1G62700), NST1 (AT2G46770), NTL8 (AT2G27300), SND2 (AT4G28500), and SND3 (AT1G28470). The RhNAC31 protein is shown in bold. The subgroup names are shown on the right of the square. b RhNAC31 sequence compared to known NAC proteins. The alignment was generated with DNAMAN. Colored boxes are used to denote subdomains of the NAC binding domain consensus sequence (A–E). Amino acid residues are depicted by white letters on a black background. Conserved and comparable residues are shown in gray. c Distribution of cis-acting elements in RhNAC31 promoter regions. DNA sequence of stress-associated cis-acting elements is indicated as follows: open rectangle; MYB, open triangles; HSE, close rectangle, LTR
Expression of RhNAC31 in rose
We examined the expression patterns of RhNAC31 in response to environmental stimuli and at different stages of rose flower opening. Quantitative RT-PCR analysis showed that RhNAC31 expression is associated with the rose flower-opening process. Expression increased from its lowest level at flowering stage 0. Upon flower opening, RhNAC31 expression reached a peak level that was approximately 39 times higher than at stage 0 (Fig. S2a). Under dehydration treatment, the increase in RhNAC31 mRNA expression peaked after 12 h, and thereafter declined (Fig. S2b). Next, we examined RhNAC31 expression levels in response to salt or cold stress plus ABA treatment, respectively. We found that a 24 h NaCl treatment caused increased RhNAC31 expression relative to the control condition. When ABA was administered simultaneously with the 24 h NaCl treatment, RhNAC31 transcript abundance was significantly higher than it was at 0, 6, and 12 h (Fig. 2a). Moreover, compared to the control condition, 12 h cold treatment also caused a significant increase in RhNAC31 expression. Administration of ABA with cold treatment did not significantly change RhNAC31 gene expression (Fig. 2b). RhNAC31 mRNA levels were also evaluated in different rose organs. The expression of RhNAC31 was much greater in sepals than in leaves, stems, or petals (Fig. 2c). This work indicates that RhNAC31 expression is influenced by environmental stress, and that ABA is associated with increased RhNAC31 expression specifically in response to NaCl or cold treatments.
Expression patterns and transactivation analysis of RhNAC31. a qRT-PCR analysis of RhNAC31 in salt treatment with exogenous ABA. Petals of rose flowers at opening stage 2 were soaked in 60 mM NaCl with or without 100 μM ABA supplemented for 0, 6, 12, and 24 h, respectively. b qRT-PCR analysis of RhNAC31 in cold treatment with exogenous ABA. Petals were immersed with 100 μM ABA at 4 °C for 0, 6, 12, and 24 h, respectively. Controls were immersed with 0.1% dimethylsulfoxide for the same time durations. c RhNAC31 expression levels of rose plant organs. Total RNA was isolated from leaves, stems, petals, and sepals of rose plants at flowering stage 2. Each qRT-PCR analysis reports the mean of three independent technical replicates for three biological replicates. The results show the mean ± SE. RhUbi1 was used as the internal control. ANOVA and Tukey HSD tests were used to evaluate statistical significance. d Yeast β-galactosidase transactivation assay of RhNAC31. The full-length, N-, and C-terminal truncations of RhNAC31 were subcloned into a pBD vector and transformed into YRG-2 yeast. Transformants were assessed for growth on SD plates with and without histidine and were stained by X-gal. Shown are: pBD and a negative control; pGAL4 and a positive control. pBD-RhNAC31, pBD-RhNAC31-N and pBD-RhNAC31-C represent full length, N-, and C-termini truncated RhNAC31 fused to GAL4-BD, respectively
RhNAC31 is a transcriptional activator
The N-terminal (RhNAC31-N lacking 158–286 AA) and C-terminal (RhNAC31-C lacking 1–157 AA) elements of RhNAC31 were fused with the GAL4 DNA-binding domain and subcloned into a pBD vector, yielding information about transcriptional activity. All transformed yeasts grew well in SD/Trp− media, but only transformants carrying pBD-RhNAC31, pBD-RhNAC31-C, and vectors positive for pGAL4 grew and showed β-galactosidase activity in SD/Trp−His− media (Fig. 2d). Yeast carrying pBD-RhNAC31-N and the empty pBD vector did not survive on SD/Trp−His− media. Taken together, these data suggest that RhNAC31 is a transcriptional activator possessing a C-terminal transactivation domain.
Heightened salt tolerance in RhNAC31 transgenic Arabidopsis plants
For further functional characterization, RhNAC31 was overexpressed in Arabidopsis driven by a constitutive promoter (Jiang et al. 2014). The T3 generation of three independent RhNAC31-overexpression lines (OE# 2, 4, and 7) were selected for subsequent study (Fig. 3b). We found no obvious morphological changes in leaves and flowers between the WT, VC, and RhNAC31-overexpressing lines (Fig. S3). Seeds of the RhNAC31-overexpressing (OE# 2, 4, and 7), WT, and VC lines were sown on MS media containing 0, 50, 100, and 150 mM NaCl. The rate of seed germination between RhNAC31-OE and control lines showed no significant differences when grown on 0 and 50 mM NaCl plates (Fig. 3a). However, the germination rates of all plants decreased when grown on plants with concentrations of 100 and 150 mM NaCl. When supplemented with 100 mM NaCl, the germination rates of OE# 2, 4, and 7 (91.2, 95.1, and 98.3%, respectively) were increased relative to the germination rates found in the WT and VC lines (86.5% and 84.2%, respectively). When stressed with 150 mM NaCl, OE# 2, 4, and 7 plants again showed significantly increased germination rates relative to the WT and VC lines (Fig. 3a, c).
RhNAC31-OE lines show enhanced resistance to salt stress in Arabidopsis. a Seed germination phenotype of RhNAC31-OE and control lines. Homozygous T3 seeds of RhNAC31-OE (OE #2, 4, and 7), wild type (WT), and vector plants (VC) were grown on MS plates containing 0, 50, 100, or 150 mM NaCl and incubated at 0 °C for 3 days followed by maintenance at 23 °C for germination. Seedlings were photographed 7 days after planting. Bars indicate 1 cm. b Quantitative RT-PCR analysis of RhNAC31-OE plants. A probe for RhNAC31 was used to study transgene levels in three of the RhNAC31-OE lines (OE#2, 4, and 7). Leaves of 3-week-old seedlings from VC and RhNAC31 transgenic plants were used. AtActin2 was used as a normalization control. c Germination rates of RhNAC31-OE and control lines for the experiment, as shown in Fig. 3a. Data represent mean ± SE of three replicates (20 seeds per line). d Root phenotypes of RhNAC31-OE and control plants. RhNAC31-OE (OE#2, 4, and 7), WT, and VC lines were grown for 7 days and then added to MS agar plates containing 0, 50, 100, 150, and 200 mM NaCl. Photographs were made 10 days after planting. Bars indicate 1 cm. Also shown are increment root length analysis (e), lateral root number analysis (f), and dry weight analysis (g). The increment root length, lateral root number, and dry weight were measured after 10 days of vertical growth. Significant differences between RhNAC31-OE and VC control plants are indicated by bars with lower case text. Three biological replicates were obtained using ten plants per treatment. Standard error (n = 3) is represented by error bars
We next examined root phenotypes of NaCl-treated RhNAC31-OE plants. Seven-day-old seedlings were transferred to fresh MS media containing 0, 50, 100, 150, and 200 mM NaCl and were then grown vertically for 10 days. We found no significant differences between RhNAC31-OE lines and controls (WT and VC) in terms of lateral root number with or without NaCl treatment (Fig. 3f). At 0, 50, and 100 mM NaCl and the primary root lengths of RhNAC31-OE plants were similar to those of control plants. However, roots were significantly longer (a 1.54-fold increase) in plants exposed to the 150 mM NaCl condition. Higher doses of NaCl (i.e., 100, 150, and 200 mM) resulted in reduced plant dry weight for all plants, with 200 mM showing the strongest effect (Fig. 3d, e). The dry weights of RhNAC31-overexpressing seedlings were significantly higher than those of control seedlings (Fig. 3g). Thus, overexpressing RhNAC31 resulted in improved salt tolerance, including a moderate increase in primary root length and biomass production at high salinity.
Overexpression of RhNAC31 enhanced antioxidant capacity under salt stress
Next, we explored the accrual of H2O2 with diaminobenzidine (DAB) and O2− with nitroblue tetrazolium (NBT) staining. Ten-day-old seedlings and separated leaves from RhNAC31 transgenic (OE #2, 4, and 7) and control (WT and VC) plants were exposed to a water (CK) or 200 mM NaCl solutions for 1 h (Figs. 4a, S4). Under normal conditions, no significant differences in staining were observed between RhNAC31-OE plants and controls. When supplemented with 200 mM NaCl, all plants showed a marked elevation in ROS levels, and in RhNAC31-OE plants brown (DAB) and blue sediment (NBT) levels were much lower (Fig. 4a). We also measured H2O2 and O2− levels in the leaves of 10-day-old seedlings treated with 0 and 200 mM NaCl solutions. Three RhNAC31-OE lines under 200 mM NaCl showed O2− levels that were 60.4, 55.8, and 46.9% of those found in the VC line, respectively (Fig. 4b). Similarly, three RhNAC31-OE lines under 200 mM NaCl showed H2O2 levels that were 89.4, 84.2, and 68.4% of those found in the VC line, respectively (Fig. 4c). Thus, both H2O2 and O2− levels were found to be significantly lower. Moreover, histochemical staining of detached leaves in both RhNAC31 transgenic plants and control plants showed similar results (Fig. S4). These results demonstrate that oxidation levels can be lowered by RhNAC31 under salt stress, suggesting that RhNAC31 plays a role in regulating reactive oxygen species.
ROS levels in RhNAC31-OE and control Arabidopsis plants. a DAB and NBT staining of RhNAC31-OE and control plants. RhNAC31-OE lines (OE #2, 4, and 7), WT, and VC plants were grown for 10 days before treatment with water or 200 mM NaCl for 1 h, after which plants would be stained with DAB and NBT solutions to quantify H2O2 and O2− accumulation in leaves. Pictures were taken with a stereomicroscope. Bars indicate 1 cm. b O2− activity in RhNAC31-OE and control plants. c H2O2 levels in RhNAC31-OE and control plants. Data represent mean ± SE of three replicates. Letters above the columns indicate significant (p < 0.05) differences when analyzed using a Tukey HSD test
Overexpression of RhNAC31 increases tolerance to freezing stress
Since RhNAC31 promoter regions contain a low-temperature-responsive (LTR) element, which is essential for cold tolerance (Nordin et al. 1993), we also examined the cold tolerance of RhNAC31 transgenic plants relative to control plants. To examine the effect of RhNAC31-overexpression on freezing tolerance, 12-week-old seedlings from the RhNAC31-OE (OE#2, 4, and 7), WT, and VC lines were exposed to − 4 °C for 1 day, and then recovered at 4 °C. After 1 day of recovery under normal growth conditions, most WT and VC seedlings’ leaves became white and did not grow, while transgenic plants were still green and regrew well (Fig. 5a). RhNAC31-OE plants also showed a higher survival rate (i.e., 95.3%) at low temperatures relative to WT and VC plants (70.2% and 69.4%; Fig. 5b). We next investigated the MDA content, SOD activity, and POD activity of RhNAC31 overexpressing plants and control plants under both normal and freezing conditions. No significant differences between RhNAC31-OE plants and control plants were detected at normal temperatures. However, in the − 4 °C treatment, the MDA content of RhNAC31-OE plants was less than 45.7% of the VC plants, and the SOD and POD activities were significantly higher than those of control plants (Fig. 5c–e). Moreover, the accumulation of H2O2 and superoxides at low temperatures was also observed through diaminobenzidine (DAB) and nitroblue tetrazolium (NBT) staining. Under normal conditions, no differences in histochemical staining were observed between controls and the three RhNAC31 transgenic lines. After the freezing treatment, brown (DAB) and blue (NBT) staining of the RhNAC31-OE plants was weaker than in control plants (Fig. 5f, g). Thus, O2− production and H2O2 content were lowered by overexpression of RhNAC31 under freezing temperatures, indicating that RhNAC31 transgenic plants accumulated fewer reactive oxygen species and incurred reduced cell membrane damage under cold stress.
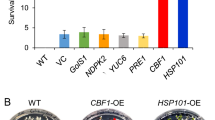
Enhancement of cold tolerance on RhNAC31-OE Arabidopsis. a RhNAC31-OE plant phenotypes under low-temperature stress. Two-week-old plant seedlings were grown under normal conditions before being transferred to − 4 °C for 1 day, followed by 1 day of recovery at 4 °C. Pictures were photographed at the indicated time points. b Survival rates of RhNAC31-OE, WT, and VC plants following cold stress. Data represent mean ± SE (n = 5). Shown are data for: MDA content (c), SOD activity (d), and POD activity (e) of RhNAC31-OE seedlings under normal and cold stress conditions. Data represent mean ± SE (n = 5). f NBT staining for O2− activity in RhNAC31-OE, WT, and VC plants. g DAB staining for H2O2 activity in RhNAC31-OE, WT, and VC plants. DAB and NBT staining used for measuring the accumulation of H2O2 and O2− in 2-week-old plant seedlings leaves of plants, respectively, before and after the 4 °C recovery. Data represent the mean ± SE (n = 5). Bars indicate 1 cm
Performance of RhNAC31-transgenic plants under drought stress
Drought is a crucial stressor for crops. Here, the performance of RhNAC31-overexpressing Arabidopsis plants in response to water deprivation was studied in soil during the seedling stage. In the first few days without irrigation, no morphological differences were seen between RhNAC31-OE and control lines. After 15 days of water deprivation, WT and VC plants exhibited strong symptoms of water loss and severe wilting in leaves. In comparison, water loss in the RhNAC31-OE lines was reduced, and a small amount of green color was visible in some leaves. After 7 days of re-watering, a large proportion of RhNAC31-OE plants recovered and thereafter grew normally, while all VC and WT plants died (Fig. 6a). Survival rates of the three transgenic lines ranged from 65.2 to 81.5% (Fig. 6b), and these values were significantly higher than the survival rates of control line plants. Changes in physiological indices including POD and SOD activity were also assessed. Fifteen days after drought exposure, RhNAC31-OE lines showed significantly higher POD and SOD activity than did the controls (Fig. 6c, d). Water protection was evaluated by determining the fresh leaf weights of the VC and WT lines as well as the three RhNAC31 transgenic lines at designated time points. After 5 h of dehydration, the water loss rate of the three independent transgenic lines was 43.2, 39.8, and 36.9%, respectively, and these values were much lower than the water loss rates of the controls (Fig. 6e). These data, therefore, suggest that RhNAC31 can also improve drought tolerance in Arabidopsis.
RhNAC31-overexpressing Arabidopsis plants show improved drought tolerance. a Response of RhNAC31 transgenic Arabidopsis to water deprivation. Following 3 weeks of growth, RhNAC31-OE and control plants were grown for and additional 15 days without irrigation. Survival rates were determined 7 days after reinitiating irrigation. b Survival rates of WT, VC, and RhNAC31-OE plants under drought stress. Also shown are the POD activity (c) and SOD activity (d) values for the rosette leaves of WT, VC, and RhNAC31-OE plants after 14 days of drought. Vertical bars represent means, and error bars represent SE based on three independent experiments. Statistically significant differences as determined by Tukey HSD tests (p < 0.05) are indicated by different letters. FW fresh weight of rosette leaves. e Kinetics of water loss in detached leaves. The detached leaves from 3-week-old plant seedlings from WT, VC and three RhNAC31 transgenic plants were dehydrated for 5 h. Error bars indicate SE (n = 6). **p < 0.01
Sensitivity to ABA during germination and growth of RhNAC31-overexpressing Arabidopsis
Since ABA serves a critical function in drought tolerance, we next studied the impact of ABA application on the germination and growth of RhNAC31-OE plants. Both the RhNAC31 transgenic and control seeds were put on MS plates supplemented with 0, 0.2, 0.4, and 0.8 μM ABA. As shown in Fig. 7a, 100% of seeds germinated for both RhNAC31 transgenic plants and controls. After exposure to exogenous ABA, seed germination in all plants was inhibited, but the effect on seeds from the transgenic lines was comparatively weaker (Fig. 7b). With the addition of 0.2 μM ABA, approximately 58.1% of control seeds germinated, whereas 39.1, 35.7, and 30.4% of seeds from the three RhNAC31-OE lines germinated, and these seeds displayed both cotyledons and true leaves (Fig. 7a, b). Similarly, the seed germination rates of the RhNAC31-OE lines were far less than those of the control lines when supplemented with 0.4 μM and 0.8 μM ABA. We also noted the root phenotype of RhNAC31-OE and control plants treated with 5, 10, or 30 μM ABA and in the absence of ABA. With respect to lateral root numbers, no significant differences were observed between lines, regardless of ABA supplementation (Fig. 7c, d). However, primary root length was inhibited in the presence of ABA. Below 30 μM of ABA, the root lengths of the three RhNAC31-OE lines were 5.11, 3.25, and 3.07 mm, respectively (Fig. 7e). Moreover, plant dry weights of the RhNAC31-OE plants were significantly lower than the controls when exposed to 5 μM and 10 μM ABA (Fig. 7f). These results indicate that RhNAC31 enhances plant ABA sensitivity during and after germination.
RhNAC31 overexpression enhanced ABA sensitivity of the transgenic plants. a Germination of WT, VC and RhNAC31-OE plants under ABA treatment. Bars indicate 1 cm. b Seed germination rates of WT, VC, and RhNAC31-OE plants after ABA treatment. Data represent mean ± SE (n = 5). c Vertical root growth phenotype of WT, VC, and RhNAC31-OE plants seedlings under different doses of ABA. RhNAC31-OE, WT, and VC plants were germinated on MS agar plates for 1 week and then moved to a fresh MS agar plate containing 0, 5, 10, or 30 μM ABA. The picture was obtained 10 days after planting. Bars indicate 1 cm. Lateral root number (d), increment root length (e), and dry weight (f) of WT, VC, and RhNAC31 transgenic plants under different doses of ABA. The lateral root number, increment root length, and dry weight were recorded after 10 day vertical growth. Data represent the mean ± SE (n = 10). Statistically significant differences as determined by Tukey HSD tests (p < 0.05) are indicated by different letters
Upregulation of abiotic stress-response genes in RhNAC31 transgenic plants
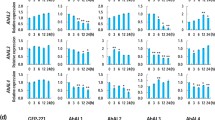
Since RhNAC31 enhances plant tolerance to increased salinity, reduced temperatures, and drought stresses, and also increases ABA sensitivity, we further assessed the molecular regulation of downstream abiotic stress responsive genes by RhNAC31. The nine selected genes included two stress-related marker genes, RD29A (AT1G12610) (Msanne et al. 2011) and RAB18 (AT1G43890) (Lång and Palva 1992); two ABA synthesis and/or response genes, AB12 (AT5G57050) (Merlot et al. 2001) and ABF4; two cold signaling or response genes, CIPK3 (AT2G26980) and CBF4 (AT5G51990); and three salt-associated genes, NHX (AT5G27150), SOS1 (AT2G01980), and SOS3 (AT5G24270). These genes were chosen to determine the influence of RhNAC31 overexpression on their level of transcription. Our qRT-PCR results indicate higher gene expression levels in RhNAC31-OE lines than in control lines (Fig. 8). In addition, the promoter sequences of these upregulated abiotic stress-related genes were evaluated using the Arabidopsis genome database (http://www.arabidopsis.org/), and the Plant Care database (http://bioinformatics.psb.ugent.be/webtools/plantcare/html/) for cis-element identification. The genes tested contained NAC recognition sites CGT[G/A] (Tran et al. 2004; Ernst et al. 2004) within their promoter regions, suggesting that a variety of stress-response genes may be involved in RhNAC31-induced abiotic stress tolerance.
q-RT-PCR analysis of abiotic stress-related genes in the VC control and RhNAC31-transgenic lines. Leaves from 3-week-old RhNAC31-OE and control lines were used to isolate RNA. qRT-PCR was used to analyze the transcript expression levels of stress-related marker genes (i.e., RD29A, RAB18, CBF4), ABA-responsive genes (i.e., ABI2, ABF4, CIPK3), and SOS pathway-related genes (i.e., NHX, SOS1, SOS3). Our results indicate the fold change of each gene relative to the VC plants. The internal control gene AtActin2 was used to normalize the mean values of three biological replicates. To assess statistical differences in means, ANOVAs were followed by Tukey’s range tests
Discussion
We assessed the expression of RhNAC31 at different stages of flower opening using quantitative RT-PCR. RhNAC31 transcript levels increased substantially in petals as flowering progressed, suggesting that the function of this gene may be related to petal expansion or petal senescence. Moreover, RhNAC31 expression is strongly induced by dehydration, cold, and salt treatment (Figs. 2, S2). RhNAC31 transcript levels were especially enhanced when exposed to salt or cold treatment in combination with ABA, which implies that ABA is involved in salt and cold signal transduction pathways. These stress induced expression patterns are consistent with those of a previously reported rose SNAC subgroup member, RhNAC3, which functions in dehydration response by affecting osmotic stress-related genes (Jiang et al. 2014). Therefore, the specific inductive patterns of RhNAC31 relative to other NAC proteins suggest that it functions in flower development and abiotic stress-response signaling in rose. Moreover, three RhNACs in rose, i.e., RhNAC2, RhNAC3, and RhNAC100, have been functionally studied. RhNAC2 and RhNAC3 are involved in resistance to drought conditions (Dai et al. 2012; Jiang et al. 2014), and RhNAC100 modulates cell expansion in flower petals via ethylene signaling (Pei et al. 2013). Thus, we believe that our report is the first to describe a rose NAM/CUC3 subgroup member TF, RhNAC31, that regulates salt, cold, and drought tolerance.
Mounting evidence supports the hypothesis that NAM/CUC3 subgroup members are mainly involved in plant development processes, and that overexpressing NAM/CUC3 subgroup members can cause morphological changes in transgenic plants. For example, overexpression of OsNAC6 in Oryza saliva (Nakashima et al. 2007) and NAM in Petunia hybrida (Souer et al. 1996) results in abnormal leaf growth and flower development. However, overexpression of RhNAC31 in Arabidopsis did not cause visible phenotypic changes relative to WT and VC in standard growth conditions (Fig. S3), and this may be an advantage for molecular breeding.
In plants, several NAC TFs serve as key regulators of ABA-mediated stress-response signaling. These include Arabidopsis RD26 (Fujita et al. 2004), and SNAC2 in rice (Hu et al. 2008). Relative to control plants, RhNAC31 transgenic plants exhibited increased ABA sensitivity both during and post-germination. Interestingly, two ABA synthesis and/or response genes, ABI2 (Merlot et al. 2001) and ABF4, were upregulated in RhNAC31 transgenic plants (Fig. 8). ABI2 encodes a protein homologous to phosphatases 2C that participates in ABA signal transduction (Leung et al. 1997) and ABF4 is a master TF that cooperatively regulates ABRE-dependent ABA pathways (Yoshida et al. 2010). The upregulation of these genes implies that RhNAC31 may interact with ABRE and activate ABRE-driven downstream genes.
The phytohormone ABA coordinates a complicated genetic network that is responsive to environmental conditions and functions positively in plant tolerance to environmental stressors. Overexpression of RhNAC31 showed improved salinity tolerance, a higher survival rate, enhanced root growth, and enhanced cold resistance (Figs. 3, 5). ABA and cold induced genes, including RAB18 and CBF4, are known regulators of ABA-dependent pathways (Haake et al. 2002; Lång and Palva 1992). CIPK3 is a calcium sensor-associated kinase, and is responsible for modulating ABA and cold signal transduction (Kim et al. 2003). Expression levels of RAB18, CBF4, and CIPK3 in RhNAC31 transgenic plants were much higher than in controls, and several copies of NAC recognition sites were found within their promoter regions (Table S2). These findings strongly suggest that RhNAC31 participates in crosstalk between salt, cold, and drought responses.
Changes in cell membranes are early plant responses to abiotic stresses. Membrane integrity and stability maintenance is a major indicator of stress resistance in plants. In addition, MDA content, SOD activity, and POD activity are all widely used to assess plant tolerance to high salinity, low temperature, and dehydration. Relative to control plants, the RhNAC31 transgenic lines showed better performance with respect to MDA content, SOD activity, and POD activity in the presence of increased salt concentration, reduced temperature, and drought conditions (Figs. 5, 6). This demonstrates that the overexpression of RhNAC31 can significantly enhance physiological protection under adverse environmental stresses.
Drought, salt, and other abiotic stresses often trigger an abnormal abundance of reactive oxygen species (ROS). This disequilibrium affects ROS homeostasis and leads to oxidative stress. This in turn can cause reduced photosynthetic efficiency, lower cell membrane stability, severe cellular damage, leaf wilting, and protein denaturing (Benjamin and Nielsen. 2006; Cruz de Carvalho 2008; Hanin et al. 2011; Choudhury et al. 2013, 2017). To prevent these phenomena, plants have developed sophisticated regulatory networks to tightly monitor ROS homeostasis and can initiate scavenging pathways when needed in cells. The ability to regulate the ROS balance is effectively correlated with the capacity of plants to thrive in high salinity and dehydration conditions. Our results demonstrate that RhNAC31 is a beneficial regulator of the plant response to dehydration, salt, and cold in rose. The fact that RhNAC31-OE leaves showed significantly lower H2O2 and MDA contents and higher SOD and POD activities relative to control leaves indicated that RhNAC31 can strengthen ROS scavenging and reduce membrane lipid peroxidation. Accordingly, RhNAC31 overexpression led to the upregulation of ROS-associated genes, including NHX (Yokoi et al. 2002), SOS1, and SOS3 (Qiu et al. 2002), all of which are important for ion homeostasis. Therefore, RhNAC31 likely regulates ROS scavenging gene expression to protect plant membranes and help to maintain redox homeostasis.
The development-related NAC protein RhNAC31 confers salt, cold, and drought resistance by controlling ROS, abiotic stresses, and ABA-related gene expression to coordinate ROS homeostasis. Moreover, RhNAC31 modulates ABA sensitivity and may function through an ABA-dependent pathway. Taken together, we speculate that RhNAC31 is a promising candidate gene for the future generation of improved crop varieties.
Author contribution statement
JXQ designed the experiments; DAQ, LSC, LW, and HQ performed the experiments; WKL, LQC, and LQH analyzed data; JXQ wrote the paper. All authors read and approved the final manuscript.
Abbreviations
- ABA:
-
Abscisic acid
- DAB:
-
Diaminobenzidine
- H2O2 :
-
Hydrogen peroxide
- MDA:
-
Malondialdehyde
- MS:
-
Murashige and Skoog
- NBT:
-
Nitroblue tetrazolium
- ORF:
-
Open reading frame
- O2 − :
-
Superoxide anion
- POD:
-
Peroxidase
- SOD:
-
Superoxide dismutase
- TFs:
-
Transcription factors
- qRT-PCR:
-
Quantitative reverse transcription PCR
- WT:
-
Wild type
References
Balazadeh S, Siddiqui H, Allu AD, Matallana-Ramirez LP, Caldana C, Mehrnia M, Zanor MI, Köhler B, Mueller-Roeber B (2010) A gene regulatory network controlled by the NAC transcription factor ANAC092/AtNAC2/ORE1 during salt-promoted senescence. Plant J 62:250–264
Beauchamp C, Fridovich I (1971) Superoxide dismutase: improved assays and an assay applicable to acrylamide gels. Anal Biochem 44:276–287
Benjamin JG, Nielsen DC (2006) Water deficit effects on root distribution of soybean, field pea and chickpea. Field Crops Res 97:248–253
Chenna R, Sugawara H, Koike T, Lopez R, Gibson TJ, Higgins DG, Thompon JD (2003) Multiple sequence alignment with the clustal series of programs. Nucleic Acids Res 31:3497–3500
Choudhury S, Panda P, Sahoo L, Panda SK (2013) Reactive oxygen species signaling in plants under abiotic stress. Plant Signal Behav 8:e23681
Choudhury FK, Rivero RM, Blumwald E, Mittler R (2017) Reactive oxygen species, abiotic stress and stress combination. Plant J 90:856–867
Clough SJ, Bent AF (1998) Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J 16:735–743
Cruz de Carvalho MH (2008) Drought stress and reactive oxygen species: production, scavenging and signaling. Plant Signal Behav 3:156–165
Dai F, Zhang C, Jiang X, Kang M, Yin X, Lu P, Zhang X, Zheng Y, Gao J (2012) RhNAC2 and RhEXPA4 are involved in the regulation of dehydration tolerance during the expansion of rose petals. Plant Physiol 160:2064–2082
Ernst HA, Olsen AN, Larsen S, Lo Leggio L (2004) Structure of the conserved domain of ANAC, a member of the NAC family of transcription factors. EMBO Rep 5:297–303
Fang Y, Liao K, Du H, Xu Y, Song H, Li X, Xiong L (2015) A stress-responsive NAC transcription factor SNAC3 confers heat and drought tolerance through modulation of reactive oxygen species in rice. J Exp Bot 66:6803–6817
Fujita M, Fujita Y, Maruyama K, Seki M, Hiratsu K, Ohme-Taraqi M, Tran LS, Yamaquchi-Shinozaki K, Shinozaki K (2004) A dehydration-induced NAC protein, RD26, is involved in a novel ABA-dependent stress-signaling pathway. Plant J 39:863–876
Haake V, Cook D, Riechmann JL, Pineda O, Thomashow MF, Zhang JZ (2002) Transcription factor CBF4 is a regulator of drought adaptation in Arabidopsis. Plant Physiol 130:639–648
Hanin M, Brini F, Ebel C, Toda Y, Takeda S, Masmoudi K (2011) Plant dehydrins and stress tolerance: versatile proteins for complex mechanisms. Plant Signal Behav 6:1503–1509
Hu H, You J, Fang Y, Zhu X, Qi Z, Xiong L (2008) Characterization of transcription factor gene SNAC2 conferring cold and salt tolerance in rice. Plant Mol Biol 67:169–181
Hu R, Qi G, Kong Y, Kong D, Gao Q, Zhou G (2010) Comprehensive analysis of NAC domain transcription factor gene family in Populus trichocarpa. BMC Plant Biol 10:145
Jiang X, Zhang C, Lv P, Jiang G, Liu X, Dai F, Gao J (2014) RhNAC3, a stress-associated NAC transcription factor, has a role in dehydration tolerance through regulating osmotic stress-related genes in rose petals. Plant Biotechnol J 12:38–48
Jouve L, Jacques D, Douglas GC, Hoffmann L, Hausman JF (2007) Biochemical characterization of early and late bud flushing in common ash (Fraxinus excelsior L.). Plant Sci 172:962–969
Kim KN, Cheong YH, Grant JJ, Pandey GK, Luan S (2003) CIPK3, a calcium sensor-associated protein kinase that regulates abscisic acid and cold signal transduction in Arabidopsis. Plant Cell 15:411–423
Kim JH, Woo HR, Kim J, Lim PO, Lee IC, Choi SH, Hwang D, Nam HG (2009) Trifurcate feed-forward regulation of age-dependent cell death involving miR164 in Arabidopsis. Science 323:1053–1057
Lång V, Palva ET (1992) The expression of a rab-related gene, rab18, is induced by abscisic acid during the cold acclimation process of Arabidopsis thaliana (L.) Heynh. Plant Mol Biol 20:951–962
Lee S, Seo PJ, Lee HJ, Park CM (2012) A NAC transcription factor NTL4 promotes reactive oxygen species production during drought-induced leaf senescence in Arabidopsis. Plant J 70:831–844
Lescot M, Dehais P, Thijs G, Marchal K, Moreau Y, Van de Peer Y, Rouzé P, Rombauts S (2002) PlantCARE, a database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences. Nucleic Acids Res 30:325–327
Leung J, Merlot S, Giraudat J (1997) The Arabidopsis ABSCISIC ACID-INSENSITIVE2 (ABI2) and ABI1 genes encode homologous protein phosphatases 2C involved in abscisic acid signal transduction. Plant Cell 9:759–771
Lim PO, Kim HJ, Nam HG (2007) Leaf senescence. Annu Rev Plant Biol 58:115–136
Liu Y, Chen Y (2007) High-efficiency thermal asymmetric interlaced PCR for amplification of unknown flanking sequences. Biotechniques 43:649–656
Liu M, Ma Z, Sun W, Huang L, Wu Q, Tang Z, Bu T, Li C, Chen H (2019) Genome-wide analysis of the NAC transcription factor family in Tartary buckwheat (Fagopyrum tataricum). BMC Genom 20:113
Ma N, Tan H, Liu X, Xue J, Li Y, Gao J (2006) Transcriptional regulation of ethylene receptor and CTR genes involved in ethylene-induced flower opening in cut rose (Rosa hybrida) cv. Samantha. J Exp Bot 57:2763–2773
Mao C, Lu S, Lv B, Zhang B, Shen J, He J, Luo L, Xi D, Chen X, Ming F (2017) A rice NAC transcription factor promotes leaf senescence via ABA biosynthesis. Plant Physiol 174:1747–1763
Meng JF, Xu TF, Wang ZZ, Fang YL, Xi ZM, Zhang ZW (2014) The ameliorative effects of exogenous melatonin on grape cuttings under water-deficient stress: antioxidant metabolites, leaf anatomy, and chloroplast morphology. J Pineal Res 57:200–212
Merlot S, Gosti F, Guerrier D, Vavasseur A, Giraudat J (2001) The ABI1 and ABI2 protein phosphatases 2C act in a negative feedback regulatory loop of the abscisic acid signaling pathway. Plant J 25:295–303
Msanne J, Lin J, Stone JM, Awada T (2011) Characterization of abiotic stress-responsive Arabidopsis thaliana RD29A and RD29B genes and evaluation of transgenes. Planta 234:97–107
Nakano Y, Asada K (1981) Hydrogen peroxide is scavenged by ascorbate-specific peroxidase in spinach chloroplasts. Plant Cell Physiol 22:867–880
Nakashima K, Tran LS, Van Ngunan D, Fujita M, Maruyama K, Todaka D, Ito Y, Hayashi N, Shinozaki K, Yamaguchi-Shiniozaki K (2007) Functional analysis of a NAC-type transcription factor OsNAC6 involved in abiotic and biotic stress-responsive gene expression in rice. Plant J 51:617–630
Nakashima K, Takasaki H, Mizoi J, Shinozaki K, Yamaguchi-Shinozaki K (2012) NAC transcription factors in plant abiotic stress responses. Biochim Biophys Acta 1819:97–103
Nordin K, Vahala T, Palva ET (1993) Differential expression of two related, low-temperature-induced genes in Arabidopsis thaliana (L.) Heynh. Plant Mol Biol 21:641–653
Nuruzzaman M, Manimekalai R, Sharoni A, Satoh K, Kondoh H, Ooka H, Kikuchi S (2010) Genome-wide analysis of NAC transcription factor family in rice. Gene 465:30–44
Olsen AN, Ernst HA, Leggio LL, Skriver K (2005) NAC transcription factors: structurally distinct, functionally diverse. Trends Plant Sci 10:79–87
Pei H, Ma N, Tian J, Luo J, Chen J, Li J, Zheng Y, Chen X, Fei Z, Gao J (2013) An NAC transcription factor controls ethylene-regulated cell expansion in flower petals. Plant Physiol 163:775–791
Pinheiro GL, Marques CS, Costa MD, Reis PA, Alves MS, Carvalho CM, Fietto LG, Fontes EP (2009) Complete inventory of soybean NAC transcription factor: sequence conservation and expression analysis uncover their distinct roles in stress response. Gene 444:10–23
Puranik S, Sahu PP, Srivastava PS, Prasad M (2012) NAC proteins: regulation and role in stress tolerance. Trends Plant Sci 17:369–381
Qiu QS, Guo Y, Dietrich MA, Schumaker KS, Zhu JK (2002) Regulation of SOS1, a plasma membrane Na+/H+ exchanger in Arabidopsis thaliana, by SOS2 and SOS3. Proc Natl Acad Sci USA 99:8436–8441
Shan W, Kuang JF, Lu WJ, Chen JY (2014) Banana fruit NAC transcription factor MaNAC1 is a direct target of MaICE1 and involved in cold stress through interacting with MaCBF1. Plant Cell Environ 37:2116–2127
Shi J, Fu XZ, Peng T, Huang XS, Fan QJ, Liu JH (2010) Spermine pretreatment confers dehydration tolerance of citrus in vitro plants via modulation of antioxidative capacity and stomatal response. Tree Physiol 30:914–922
Souer E, van Houwelingen A, Kloos D, Mol J, Koes R (1996) The no apical meristem gene of Petunia is required for pattern formation in embryos and flowers and is expressed at meristem and primordia boundaries. Cell 85:159–170
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731–2739
Tran LS, Nakashima K, Sakuma Y, Simpson SD, Fujita Y, Maruyama K, Fujita M, Seki M, Shinozaki K, Yamaguchi-Shinozaki K (2004) Isolation and functional analysis of Arabidopsis stress-inducible NAC transcription factors that bind to a drought-responsive cis-element in the early responsive to dehydration stress 1 promoter. Plant Cell 16:2481–2498
Wang K, Zhong M, Wu Y, Bai Z, Liang Q, Liu Q, Pan Y, Zhang L, Jiang B, Jia Y, Liu G (2017) Overexpression of a chrysanthemum transcription factor gene DgNAC1 improves the salinity tolerance in chrysanthemum. Plant Cell Rep 36:571–581
Yokoi S, Quintero FJ, Cubero B, Ruiz MT, Bressan RA, Haseqawa PM, Pardo JM (2002) Differential expression and function of Arabidopsis thaliana NHX Na+/H+ antiporters in the salt stress response. Plant J 30:529–539
Yoshida T, Fujita Y, Sayama H, Kidokoro S, Maruyama K, Mizoi J, Shinozaki K, Yamaquchi-Shinozaki K (2010) AREB1, AREB2, and ABF3 are master transcription factors that cooperatively regulate ABRE-dependent ABA signaling involved in drought stress tolerance and require ABA for full activation. Plant J 61:672–685
Yu S, Huang A, Li J, Gao L, Feng Y, Pemberton E, Chen C (2018) OsNAC45 plays complex roles by mediating POD activity and the expression of development-related genes under various abiotic stress in rice root. Plant Growth Regul 84:519–531
Zhang J, Huang GQ, Zou D, Yan JQ, Li Y, Hu S, Li XB (2018) The cotton (Gossypium hirsutum) NAC transcription factor (FSN1) as a positive regulator participates in controlling secondary cell wall biosynthesis and modification of fibers. New Phytol 217:625–640
Acknowledgements
This work was supported by the Promotive Research Fund for Excellent Young and Middle-Aged Scientists of Shandong Province (No. BS2014SW032), National Key Research and Development Project of China (2018YFD1000400), and the National Natural Science Foundation of China (No. 31501798).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Additional information
Communicated by J. Huang.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Ding, A., Li, S., Li, W. et al. RhNAC31, a novel rose NAC transcription factor, enhances tolerance to multiple abiotic stresses in Arabidopsis. Acta Physiol Plant 41, 75 (2019). https://doi.org/10.1007/s11738-019-2866-1
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11738-019-2866-1











