Abstract
Heat stress transcription factors (HSFs) play an essential role in the adjustment of plants to high temperatures. These molecules have evolved complicated mechanisms that rely on interactions between different HSFs and other heat stress-related genes [such as bZIP28, multiprotein bridging factor 1c (MBF1c), calmodulin-binding protein kinase 3 (CBK3)] in response to different heat stresses (such as occasional or successive high temperatures). In the present study, phenotypic, gene expression and yeast two-hybrid assays revealed that HSFA2 and HSFA3 function in the same heat regulation pathway. The single mutants, hsfa2 and hsfa3 as well as double mutant hsfa2 and hsfa3, exhibited heat-sensitive phenotypes in acquired thermotolerance after a long recovery time (ATLR) but not in basic thermotolerance and acquired thermotolerance after a short recovery time (ATSR). The expression of HSP18.1-CI and HSP25.3-P was down-regulated in single and double mutants of hsfa2 and hsfa3 under successive heat stress in ATLR assays. In addition, HSFA2 interacted with HSFA3 at the protein level in yeast two-hybrid assays. These results demonstrated dynamic alterations in the expression of HSFA2, HSFA3 and other heat-related genes in ATLR assays, providing new insights into the relationship between HSFA2 and HSFA3; this information will refine the HSF network in the regulation of heat stress response.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
As sessile autophytic organisms, plants have an advantage regarding food-seeking but struggle against adverse environment stresses, such as high temperature. High temperatures impair plant growth and development, and extreme thermal stress even promotes programmed cell death (Vacca et al. 2004). For survival, plants have developed delicate mechanisms to overcome this adversity. The heat stress transcript factor (HSF) family is one of the major regulation groups that help plants to adapt to elevated high temperatures. Compared with the small HSF family in Drosophila or vertebrates, plants possess large HSF families, and more than 21 HSFs have been identified in Arabidopsis, while 24 HSFs have been identified in tomato, and 25 HSFs have been identified in rice (Scharf et al. 2011). Based on the presence of the conserved DNA-binding domain (DBD) and oligomerization domain (OD), HSF family proteins can be divided into A, B and C groups. In tomato, three main HSFs (HSFA1, HSFA2 and HSFB1) regulate the expression of heat responsive genes and plant thermotolerance, and these proteins are modulated through HSP70 and HSP90 (Hahn et al. 2011). In Arabidopsis, at least ten HSFs participate in the heat stress response, and most of these molecules have recently been summarized (see reviews Scharf et al. 2011; von Koskull-Doering et al. 2007). Previous studies have reported that HSFA1s acts as master regulator in the early heat response (Liu and Charng 2012), while HSFA2 and HSFA3 play important roles in prolonged heat stress in Arabidopsis (Schramm et al. 2008).
Potential heat shock elements (HSEs) have been identified in the promoter regions of all HSFs, indicating that HSFs may be auto-regulated or regulated through other HSFs (Nover et al. 2001). Indeed, HSFA1d and HSFA1e showed the transient transcriptional activation of the HSE1 motif localized at 188 bp upstream of the HSFA2. Interestingly, HSFA2 was suppressed only in hsfa1e and hsfa1d mutants, not in hsfa1a or hsfa1b single mutants or the hsfa1a hsfa1b double mutant, indicating that HSFA2 can only be regulated through hsfa1e and hsfa1d (Nishizawa-Yokoi et al. 2011). However, another study reported that HSFA1a and HSFA1b interact with HSFA2 via bimolecular fluorescence complementation (BiFC), indicating that these factors may assist HSFA2 in heat regulation at the protein level (Li et al. 2010). HsfA2 was not only mediated through HSFA1s but also other heat-related genes that participate in heat regulation. Mitogen-activated protein kinase 6 (MPK6) phosphorylate the 249 threonine of HSFA2, which is indispensable for the nuclear localization of HSFA2, and recent studies have revealed that MPK6 was highly induced under heat stress (Evrard et al. 2013). A mutant of fk506-binding protein 62 (fkbp62, also named as rof1) was sensitive to acquired thermotolerance after long recovery time (ATLR). Interestingly, the expression of small HSPs, located downstream of HSFA2, were dramatically suppressed in fkbp62 in response to stress and during the recovery time in ATLR assays. The ROF1-HSP90.1 complex showed nuclear localization after heat stress, which depends on the interaction between HSP90.1 and HSFA2, indicates that ROF1 participates in the HSFA2-sHSP pathway (Meiri and Breiman 2009). Unlike rof1, the rof2 mutant enhances plant thermotolerance, which also required HSFA2 during the recovery time in the ATLR assay. However, ROF2 hetero-polymerizes with ROF1 to participate in the function of the ROF1-HSP90.1-HSFA2 complex. Moreover, the transient co-expression of ROF2, ROF1 and HSFA2 suppresses the expression of sHSPs, suggesting that ROF2 functions as a negative feedback regulator of HSFA2 in plant thermotolerance (Meiri et al. 2010).
In addition to HSFA2, the regulation network of HSFA3 was also well characterized. The dehydration responsive element binding protein 2 (DREB2) was highly induced under high temperatures and salt and drought stresses (Liu et al. 1998; Sakuma et al. 2006a). Transiently expressed DREB2A or DREB2B directly binds to the DRE1 and DRE2 motifs in the promoter region of HSFA3, and both factors can intensively activate the expression of HSFA3. Furthermore, HSFA3 was remarkably promoted in DREB2A CA and DREB2C overexpression seedlings (Sakuma et al. 2006b; Chen et al. 2010), and the down-regulation of HSP18.1-CI and HSP25.3-P in dreb2a and dreb2c depends on HSFA3 (Chen et al. 2010; Schramm et al. 2008). Both mutant strains and the DREB2A, DREB2C and HSFA3 overexpression lines exhibited similar performances on basic heat resistance and seed germination after heat stress (Yoshida et al. 2008; Chen et al. 2010). These data revealed that DREB2 s directly controls HSFA3 in the heat regulation pathway.
Although intensive studies on the individual regulation networks of HSFA2 and HSFA3 have been reported, the interaction between these factors has only been hypothesized in a previous study (Schramm et al. 2008). In the present study, the direct interaction between HSFA2 and HSFA3 was confirmed through genetic analysis, gene expression analysis and yeast two-hybrid assays. The potential roles of these factors in heat regulation pathways were also discussed in the present study. These results provide valuable information for constructing a fine heat regulation network in Arabidopsis, which is useful and referable in crop research and breeding.
Materials and methods
Plant materials and growth conditions
The Arabidopsis thaliana Col-0 ecotype was used in the present study, and hsfa2 (SALK_008978), hsfa3 (SALK_011107) and hsp101 (CS16284) mutants were obtained from the Salk Institute. The double mutant hsfa2 hsfa3 was constructed by crossing hsfa2 with hsfa3. T-DNA insertions were examined as previously described (http://signal.salk.edu/tdnaprimers.2.html), and the primers are listed in Table S1. The plant growth conditions were maintained according to Li et al. (2012). Briefly, these mutants and wild-type seedlings were grown on nutrient composites (Pei Lei, China) or plates containing half-strength Murashige and Skoog medium (Sigma-Aldrich, St. Louis, MO, USA) with 1% (w/v) sucrose and 0.8% (w/v) agar. All materials were cultivated in a greenhouse at 22 °C with a 16-h light/8-h dark photoperiod and a relative humidity of 70% under an illumination density of 230–300 µEm−2 s−1.
Thermotolerance assays
Thermotolerance assays were conducted as previously described (Larkindale et al. 2005). The newly harvested dry seeds of each line were surface sterilized with ethanol for 1 min, followed by commercial bleach for 3 min, and rinsed 3–4 times with sterile H2O. The sterilized seeds were subsequently planted on plates and stratified for 3 days. For the basal thermotolerance (BT) assay, the seeds (plates) were heated at 45 °C in a water bath for the indicated time. For the acquired thermotolerance (AT) assays, 3- or 7-day-old seedlings were first acclimated from 38 °C for 90 min, recovered at 22 °C for 2 h, and subsequently treated at 45 °C for 150 min (ATSR). Alternatively, the plants were recovered at 22 °C for 2 days and subsequently treated at 45 °C for 1 h (ATLR). Five days after treatment, the expansion of the hypocotyl out of the seed coat was considered germination, and plants that remained green and producing new leaves were scored as survived (Larkindale et al. 2005). The data were expressed as the mean ± standard error (SE) (n = three biological replicates, 30 seeds per plant were analyzed for each replicate).
RNA preparation and RT-PCR examination
For RNA extraction, fresh seedlings (root included) were sampled from 3-day-old plants grown on plates after heat stress for the indicated times. Total RNA was prepared using TRIZOL reagent (Invitrogen, USA). For each sample, 5 μg of RNA was digested with 1 μl of DNase (Thermo, USA) to exclude residual DNA and subsequently used for reverse transcription using the TransScript First-Strand cDNA Synthesis Super Mix (TransGen, China). Reverse transcription PCR (RT-PCR) was performed using 2 × Taq PCR MasterMix (RTC3104-03, Tiangen, Beijing). The following thermal cycle conditions were used: 94 °C for 4 min, followed by 26–40 cycles at 94 °C for 30 s, 58 °C for 30 s, and 72 °C for 30 s, with a final extension at 72 °C for 5 min and a gradual decrease in the temperature to 25 °C. A. thaliana Actin 7 (Act7) was used as the internal control. The gene-specific primer sequences are listed in Table S1.
Vector construction and yeast two-hybrid assay
The Matchmaker Two-Hybrid System (Clontech, USA) was used for the yeast two-hybrid assay. Fragments of HSFA2, HSFA3, HSFA2 OD, HSFA3 OD, HSFA1a OD and HSFA1b OD were amplified from an Arabidopsis cDNA library using the corresponding primers as listed in Table S1, and these fragments were subsequently inserted into the pGBKT7 and pGADT7 (Clontech, USA) vectors using the restriction sites indicated in Table S1. The yeast two-hybrid assay was performed with co-transformation following the manufacturer’s instructions. Briefly, the yeast cells of strain AH109 were co-transformed with the bait and prey vectors using a lithium acetate-based protocol and grown on synthetic dextrose media (SD). The transformants were first spread onto SD media lacking leucine and tryptophan (SD/-Leu/-Trp), and subsequently the co-transformed positive colonies were grown on SD media lacking leucine, tryptophan, adenine and histidine (SD/-Leu/-Trp/-Ade/-His) to detect the activation of the HIS3 and ADE2 reporter genes. A prey vector containing the SV-40 large T-antigen (pGAD-T) and a bait vector containing murine p53 (pGBKT7-53) were used as a positive control system (Clontech, USA), and a bait vector containing human lamin C (pGBKT7-Lam) was used as a negative control (Clontech, USA).
The β-galactosidase assay was performed according to the colony-lift filter assay (Laporte et al. 1999). Yeast cells grown on SD/-Leu/-Trp/-Ade/-His plates within 4 days were imprinted onto a filter (Whatman No. 5). After two cycles of freeze/thawing in liquid nitrogen, the filter was incubated on another filter presoaked with Z buffer/X-gal solution (45 mM Na2HPO4·7H2O, 40 mM NaH2PO4·H2O, 10 mM KCl, 1 mM MgSO4·7H2O, 0.8 mM X-gal and 0.04 mM β-mercaptoethanol) at 30 °C. The photographs were captured 2 h after incubation.
Statistical analysis
Significant differences in the germination and survival rates between the mutant and wild-type seedlings were determined using independent samples t tests. Differences were considered statistically significant at the P < 0.05 and P < 0.01 levels.
Results
HSFA3 and HSFA2 regulate plant ATLR in the same pathway
HSFA2 is acquired for the extension of acquired thermotolerance (Charng et al. 2007), and the potential regulation between HSFA2 and HSFA3 has been hypothesized in a previous study (Schramm et al. 2008). However, little evidence of how these two genes interact and the pathway involving these genes has been reported. To this end, we constructed an hsfa2 hsfa3 double mutant by crossing hsfa2 with hsfa3, and all mutants were confirmed as homozygous through the detection of T-DNA insertion fragments (Fig. 1). The newly produced double mutant, the hsfa2 and hsfa3 single mutants, wild-type plants, and the well-studied heat-sensitive mutant hsp101, which was served as a positive control for heat treatment, were submitted to a series different heat stress strategies, including seed germination against BT, seedling survival against ATLR and ATSR. The seeds from these plants were sown on the same plates as indicated (Fig. 2a, including BT and AT assays). For the BT assay, all plants exhibited similar seed germination rates under normal conditions (Fig. 2b, c). Compared with the control, germination was significantly delayed in all plants following heat stress; however, most of these plants can survive with similar germination rates, except hsp101, which was decreased to 10% of the wild-type (Fig. 2d, e).
Genotyping of hsfa2, hsfa3 and the hsfa2 hsfa3 double mutant. BP represents the T-DNA border primer, LP represents the left T-DNA border primer and RP represents the right genomic primer. BP + RP were used to identify homozygous mutants, while BP + LP were used to identify wild-type seedlings; when both results were positive, the mutant was heterozygous
Phenotypes of hsfa2, hsfa3 and hsfa2 hsfa3 after the BT test. a A diagrammatic drawing showing the positions of all plant seedlings (Fig. 3, Fig. S1). b, c Seed germination and statistical analysis of all mutants under normal conditions. d, e Seed germination and statistical analysis of all mutants after heating at 45 °C for 5 h. Photographic and statistical analyses were performed at 5 days after treatment. The data were collected from three independent biological repeats and expressed as the mean ± SE (standard error). Significant differences between wild-type and mutant plants at **P < 0.01 levels, based on a t test
Similar to the germination rate, no significant difference was detected in the survival rate among the mutant and wild-type seedlings under normal conditions (Fig. 3a, b). The mutants and wild-type seedlings were harmed at varying degrees in ATSR and ATLR treatments. However, hsfa2, hsfa3 and hsfa2 hsfa3 were indistinguishable from wild-type plants in ATSR, while the survival rate of the positive control, hsp101, decreased to 17.8% of wild-type seedlings (Fig. 3c, d). An obvious difference between the hsf mutants and wild-type seedlings was observed in the ATLR assays. Compared with the wild-type, the hsfa2, hsfa3, hsp101 and hsfa2 hsfa3 double mutants showed impaired heat resistance, with more yellow leaves and reduced survival rates (Fig. 3e, f). The survival rates of hsfa2, hsfa2 hsfa3 and hsp101 decreased to almost half that of wild-type; however, hsfa3 was not significantly different from the wild-type seedlings (Fig. 3e, f). To confirm the impaired performance of the mutants that underwent ATLR, plants at different ages were densely planted, and the results could be reproduced in a planting density-independent manner when tested with 3 day- and 7 day-old seedlings (Fig. S1a–f). Taken together, the phenotype of the hsfa2 hsfa3 double mutant was similar to that of the hsfa2 and hsfa3 single mutants after heat stress, similar to hsfa2.
Phenotypes of hsfa2, hsfa3 and hsfa2 hsfa3 after ATSR and ATLR assays. a Seedling survival rate and statistical analysis b of all the seedlings under normal conditions. c Seedling survival rate and statistical analysis d of all the seedlings after ATSR. e Seedling survival rate and statistical analysis f of all seedlings after ATLR. Photographic and statistical analyses were performed 15 days after sowing in control and ATSR and 17 days after sowing in ATLR. The data were collected from three independent biological repeats and expressed as the mean ± SE. Significant differences between wild-type and mutant plants at **P < 0.01 levels, based on a t test
Expression analysis of heat stress responsible genes in hsfa2 and hsfa3 mutants
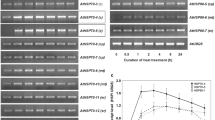
HSFA2 and HSFA3 co-regulated a subset of genes in transcriptional profiling studies (Table S2) (Nishizawa et al. 2006; Yoshida et al. 2008). Among these genes, HSP18.1-CI and HSP25.3-P are regulated by HSFA2 during the recovery time or prolonged heat stress in ATLR (Charng et al. 2007). We subsequently monitored the expression of these two genes with HSP101 in the 3-day-old seedlings of hsfa2, hsfa3, hsfa2 hsfa3 and hsp101 at different time points using an ATLR assay (Fig. 4a). RT-PCR revealed that the transcript levels of HSP18.1-CI and HSP25.3-P were dramatically increased within 30 min of the first heat stress treatment, and subsequently maintained for 2 h in all mutants (Fig. 4b A–C); the levels of both genes returned to normal after 48 h of recovery, and no significant difference was detected in all mutants (Fig. 4b D). In response to a second heat stress, both HSP18.1-CI and HSP25.3-P were induced to a slightly weaker degree in wild-type than in the first heat stress. However, during the long recovery time and the second heat stress, the transcript levels of HSP18.1-CI and HSP25.3-P were hardly induced in hsfa2 and hsfa2 hsfa3 and was relatively lower in hsfa3 than in the wild-type; this effect lasted to the end of the heat treatment (Fig. 4b E, F). The results indicated that the decreased expression of HSP18.1-CI and HSP25.3-P may directly correlate with the thermotolerance deficiency in hsfa2, hsfa3, hsfa2 hsfa3 mutants during the long recovery time.
Changes in the expression of heat-related genes in hsf mutants. a Schematic diagram showing the heat stress regimen. The arrows indicate the time points when the samples were harvested (A–F). b RT-PCR analysis of HSPs in the seedlings of WT and hsfa2, hsfa3, hsfa2 hsfa3 and hsp101 mutants. A–F indicate the time points described in a. Actin 7 was used as the internal control
However, the expression patterns of HSP18.1-CI and HSP25.3-P in hsp101 were similar to that of wild-type (Fig. 4b A–F), suggesting that the changes in the expression of these genes did not induce the heat-resistance deficiency in hsp101. Moreover, the HSP101 transcript levels were not changed in hsfa2 and hsfa3, and the levels of HSFA2 and HSFA3 were not altered in hsp101, indicating that HSP101 may not be required for heat sensitivity in hsfa2 and hsfa3 mutants (Fig. 4b).
HSFA2 interacts with HSFA3, HSFA1a and HSFA1b in vitro
In most of the sampling points of the heat stress treatment (Fig. 4a), HSFA3 expression was higher in the hsfa2 mutant than in wild-type, regardless of heat treatment (Fig. 4b A–F), suggesting a potential direct interaction between HSFA2 and HSFA3. To provide more confident evidence, a yeast two-hybrid assay was performed. The full-length and truncated coding sequences of the oligomerization domains (OD) of HSFA2, HSFA3, HSFA1a and HSFA1b were cloned into pGBKT7 and pGADT7, respectively. The interactions were examined through co-transformation and β-galactosidase staining. Both nutritional deficient screening and β-galactosidase staining revealed a strong interaction between HSFA2 and HSFA3 (Fig. 5), and the OD fragments of HSFA2 and HSFA3 were sufficient for the interaction of these two proteins (Fig. 5). The yeast two-hybrid results provided solid proof of the interaction between HSFA2 and HSFA3. Moreover, in the HSF family, the HSFA1 subfamily plays a dominant role in heat regulation; thus, to investigate whether HSFA2 directly interacts with the central factor HSFA1, yeast two-hybrid analysis was conducted with the combination of BD-HSFA2 with AD-HSFA1a OD and AD-HSFA1b OD, respectively. HSFA2 also exhibited a strong interaction with HSFA1a and HSFA1b in nutrition deficiency screening and β-galactosidase staining assays (Fig. 5).
Yeast two-hybrid assays. Nutrition deficiency screening (a) and β-galactosidase staining assays (b). Each combination is indicated beside the clones in (a), and the staining (b) was imprinted from (a). BD-A2OD, AD-A3OD, BD-A2, AD-A3, AD-A1aOD and AD-A1bOD represent pGBKT7-HSFA2 Oligomerization Domain, pGADT7-HSFA3 Oligomerization Domain, pGBKT7-HSFA2, pGADT7-HSFA3, pGADT7-HSFA1a Oligomerization Domain and pGADT7-HSFA1b Oligomerization Domain, respectively. The images of the nutrition deficiency screening were obtained at 3 days after inoculating on SD-Trp/-Leu/-Ade/-His media, and β-galactosidase staining was imaged at 2 h after incubation at 30 °C
Discussion
Genetic analysis with mutants is a useful method to study the relationship of two or more different genes. When the genes function in the same regulatory pathway, their double mutant should exhibit a similar phenotype and gene expression profile as the single mutants; in contrast, the phenotype of the double mutant may be a combination of the two single mutants (Luo et al. 2005). In the present study, the single and double mutants of HSFA2 and HSFA3 showed similar phenotypes in response to heat stress (Fig. 3, S1), indicating that the two genes function in the same heat regulation pathway. Genetic and expression analyses further suggested that HSFA3 may be downstream of HSFA2, reflecting its weaker heat-sensitive phenotype and co-regulation of the same heat-related genes in acquired thermotolerance (Fig. 4). Moreover, the direct interaction between HSFA2 and HSFA3 at the protein level is consistent with the results of the phenotypic and expression analyses (Fig. 5).
However, when a gene family has multiple members in the whole genome and these factors have similar gene functions, studying double or multiple mutants is an effective strategy to evaluate genetic redundancy. Success stories have previously been reported in the analysis of HSFA1 genes. The hsfa1a hsfa1b double mutants were insensitive in basic thermotolerance and slightly weaker in acquired thermotolerance assays (Lohmann et al. 2004); the survival rate of the hsfa1d hsfA1e double mutants decreased after an additional acquired thermotolerance treatment (Nishizawa-Yokoi et al. 2011). In addition, the quadruple mutant, hsfa1a hsfa1b hsfa1d hsfa1e completely lost the capacity for heat resistance (Liu et al. 2011). This information suggests that the four members of HsfA1 genes function redundantly. In the present study, the double mutant hsfa2 hsfa3 did not show more heat sensitivity than the hsfa2 or hsfa3 single mutant (Fig. 3, S1), suggesting that the functions of these genes are not completely redundant. The transcription profiles showed a tight relationship with the phenotype, and only 4% of heat responsive genes were affected in hsfa1a hsfa1b double mutants (Lohmann et al. 2004), while more than half of these genes were affected in the quadruple mutants (Liu et al. 2011). This effect was not observed for the hsfa2 hsfa3 double mutant, as the induction of HSP18.1-CI and HSP25.3-P in the double mutant was similar to that in the hsfa2 and hsfa3 single mutants after heat stress (Fig. 4b D–F). These results are consistent with the results showing that HSFA2 and HSFA3 function in the same pathway, and HSFA2 plays a dominant role over HSFA3.
HSFA1 is considered the master regulator in response to heat stress in Arabidopsis (Liu et al. 2011); another important HSF gene, HSFA2, regulates plant ATLR (Charng et al. 2007). The results of a genetic complementary analysis revealed that HSFA2 functions downstream of HSFA1 (Liu and Charng 2013). However, whether other HSF genes are associated with the HSFA1a-HSFA2 regulatory pathway remains unknown. In the present study, the results of the yeast two-hybrid assay revealed intense interactions between HSA1s-HSFA2 and HSFA2-HSFA3 (Fig. 5), confirming the results of a previous study showing that HSFA2 may work in the same pathway as HSFA1s (Liu and Charng 2013). However, the OD region is sufficient for the interaction between HSFA2 and HSFA3, HSFA1a, HSFA1b, respectively; they may be the key domain for the oligomerization between different HSFs, as similar results have also been reported in studies of HSF4 and HSF5 (Baniwal et al. 2007).
However, according to previous studies, none of the single or double mutants of HSFA2 and HSFA3 exhibited a more sensitive phenotype than the hsfa1s quadruple mutants, which also confirmed the leading role of HSFA1 in plant heat regulation. While the expression analysis revealed the independence between HSP101 and HSFA2 or HSFA3 compared with wild-type, neither HSFA2 nor HSFA3 transcripts were affected in hsp101, and the HSP101 transcript in hsfa2 and hsfa3 remained unaffected (Fig. 4). Thus, HSFA2 and HSFA3 may function in parallel pathways with HSP101 in regulating heat stress.
The expression of HSFAs was not only suppressed by HSFBs (Ikeda et al. 2011) but also by HSFAs, e.g., the expression of HSFA4a and HSFA4c can be specifically suppressed by HSFA5 (Baniwal et al. 2007). In the present study, the expression of HSFA3 was enhanced in the hsfa2 mutant, particularly after heat treatment at 37 °C for 1 h (Fig. 4b B). Interestingly, this effect seemed independent of heat stress treatment because at most sample points (including the normal condition, Fig. 4b A) of the heat stress treatment, HSFA3 was up-regulated at varying degrees in the hsfa2 mutant, suggesting that HSFA2 may suppress the expression of HSFA3. Alternatively, HSFA3 may have a complimentary effect to the hsfa2 mutant. However, the results of the functional analysis of the hsfa3 single mutant in the present study conflicted with those of previous studies (Schramm et al. 2008; Chen et al. 2010). A transcriptional profiling study reported that HSFA3 was down-regulated 9.4-fold in a 4-week-old hsfa2 mutant line after successive heat stress treatments (42 °C for 3 h, 20 °C for 21 h, 42 °C for 1 h) (Schramm et al. 2006), while in the present study, HSFA3 was enhanced in hsfa2 at different sample points of ATLR assays. These contradictory results may reflect the different development stages and different heat stress methods, as heat resistance differed with plant development stages and heat stress regimen, even with the same material (Yu et al. 2014). However, at the end of the second heat stress treatment, both HSFA2 and HSFA3 transcripts were too negligible for detection using reverse transcript PCR analysis. Moreover, HSFA3 was reported to regulate seed germination under high temperatures and plant ATSR (Schramm et al. 2008). However, no significant differences were observed in basic thermotolerance (Fig. 2) and ATSR assays (Fig. 3c, d) in the present study. In contrast, we observed a reduction of HSFA3 in plant ATLR (Fig. 3d, e). Interestingly, the expression of HSP18.1-CI and HSP25.3-P in hsfa2, hsfa3 and the hsfa2 hsfa3 double mutant was indistinguishable from that in wild-type seedlings in the first heat stress of ATLR but was significantly suppressed in the second heat stress (Fig. 4b B–F), suggesting that HSFA3 may primarily function in ATLR regulation. Thus, these results showed dynamic alterations in the expression of HSFA2 and HSFA3 and related HSP genes in ATLR assays, making the heat regulation network of HSFs more explicit.
Author contribution statement
X. L. and E. Y. designed the experiment, X. L., W. W. J. W. and Y. C. performed the experiments and analyzed the data, and X. L., E. Y., W. L. and B. M. drafted the manuscript. All authors participated in the discussion.
References
Baniwal SK, Chan KY, Scharf KD et al (2007) Role of heat stress transcription factor HsfA5 as specific repressor of HsfA4. J Biol Chem 282:3605–3613. doi:10.1074/jbc.M609545200
Charng YY, Liu HC, Liu NY et al (2007) A heat-inducible transcription factor, HsfA2, is required for extension of acquired thermotolerance in Arabidopsis. Plant Physiol 143:251–262. doi:10.1104/pp.106.091322
Chen H, Hwang JE, Lim CJ et al (2010) Arabidopsis DREB2C functions as a transcriptional activator of HsfA3 during the heat stress response. Biochem Biophys Res Commun 401:238–244. doi:10.1016/j.bbrc.2010.09.038
Evrard A, Kumar M, Lecourieux D et al (2013) Regulation of the heat stress response in Arabidopsis by MPK6-targeted phosphorylation of the heat stress factor HsfA2. Peer J 1:e59
Hahn A, Bublak D, Schleiff E et al (2011) Crosstalk between Hsp90 and Hsp70 chaperones and heat stress transcription factors in tomato. Plant Cell 23:741–755
Ikeda M, Mitsuda N, Ohme-Takagi M (2011) Arabidopsis HsfB1 and HsfB2b act as repressors of the expression of heat-inducible Hsfs but positively regulate the acquired thermotolerance. Plant Physiol 157:1243–1254
Laporte SA, Oakley RH, Zhang J et al (1999) The β2-adrenergic receptor/βarrestin complex recruits the clathrin adaptor AP-2 during endocytosis. Proc Natl Acad Sci USA 96:3712–3717
Larkindale J, Hall JD, Knight MR et al (2005) Heat stress phenotypes of Arabidopsis mutants implicate multiple signaling pathways in the acquisition of thermotolerance. Plant Physiol 138:882–897
Li M, Berendzen KW, Schoeffl F (2010) Promoter specificity and interactions between early and late Arabidopsis heat shock factors. Plant Mol Biol 73:559–567. doi:10.1007/s11103-010-9643-217
Li X, Yu E, Fan C et al (2012) Developmental, cytological and transcriptional analysis of autotetraploid Arabidopsis. Planta 236:579–596
Liu H, Charng Y (2012) Acquired thermotolerance independent of heat shock factor A1 (HsfA1), the master regulator of the heat stress response. Plant Signal Behav 7:547–550
Liu HC, Charng YY (2013) Common and distinct functions of Arabidopsis class A1 and A2 heat shock factors in diverse abiotic stress responses and development. Plant Physiol 163:276–290
Liu Q, Kasuga M, Sakuma Y et al (1998) Two transcription factors, DREB1 and DREB2, with an EREBP/AP2 DNA binding domain separate two cellular signal transduction pathways in drought-and low-temperature-responsive gene expression, respectively, in Arabidopsis. Plant Cell 10:1391–1406
Liu HC, Liao HT, Charng YY (2011) The role of class A1 heat shock factors (HSFA1s) in response to heat and other stresses in Arabidopsis. Plant Cell Environ 34:738–751. doi:10.1111/j.1365-3040.2011.02278.x
Lohmann C, Eggers-Schumacher G, Wunderlich M et al (2004) Two different heat shock transcription factors regulate immediate early expression of stress genes in Arabidopsis. Mol Genet Genom 271:11–21. doi:10.1007/s00438-003-0954-8
Luo M, Dennis ES, Berger F et al (2005) MINISEED3 (MINI3), a WRKY family gene, and HAIKU2 (IKU2), a leucine-rich repeat (LRR) KINASE gene, are regulators of seed size in Arabidopsis. Proc Natl Acad Sci USA 102:17531–17536. doi:10.1073/pnas.0508418102
Meiri D, Breiman A (2009) Arabidopsis ROF1 (FKBP62) modulates thermotolerance by interacting with HSP90.1 and affecting the accumulation of HsfA2-regulated sHSPs. Plant J 59:387–399. doi:10.1111/j.1365-313X.2009.03878.x
Meiri D, Tazat K, Cohen-Peer R et al (2010) Involvement of Arabidopsis ROF2 (FKBP65) in thermotolerance. Plant Mol Biol 72:191–203. doi:10.1007/s11103-009-9561-3
Nishizawa A, Yabuta Y, Yoshida E et al (2006) Arabidopsis heat shock transcription factor A2 as a key regulator in response to several types of environmental stress. Plant J 48:535–547. doi:10.1111/j.1365-313X.2006.02889.x
Nishizawa-Yokoi A, Nosaka R, Hayashi H et al (2011) HsfA1d and HsfA1e involved in the transcriptional regulation of HsfA2 function as key regulators for the Hsf signaling network in response to environmental stress. Plant Cell Physiol 52:933–945. doi:10.1093/pcp/pcr045
Nover L, Bharti K, Doering P et al (2001) Arabidopsis and the heat stress transcription factor world: how many heat stress transcription factors do we need? Cell Stress Chaperones 6:177–189. doi:10.1379/1466-1268(2001)006<0177:aathst>2.0.co;2
Sakuma Y, Maruyama K, Osakabe Y et al (2006a) Functional analysis of an Arabidopsis Transcription Factor, DREB2A, involved in drought-responsive gene expression. Plant Cell 18(5):1292–1309. doi:10.1105/tpc.105.035881
Sakuma Y, Maruyama K, Qin F et al (2006b) Dual function of an Arabidopsis transcription factor DREB2A in water-stress-responsive and heat-stress-responsive gene expression. Proc Natl Acad Sci USA 103:18822–18827. doi:10.1073/pnas.0605639103
Scharf KD, Berberich T, Ebersberger I et al (2011) The plant heat stress transcription factor (Hsf) family: structure, function and evolution. BBA Gene Regul Mech 1819:104–119
Schramm F, Ganguli A, Kiehlmann E et al (2006) The heat stress transcription factor HsfA2 serves as a regulatory amplifier of a subset of genes in the heat stress response in Arabidopsis. Plant Mol Biol 60:759–772. doi:10.1007/s11103-005-5750-x
Schramm F, Larkindale J, Kiehlmann E et al (2008) A cascade of transcription factor DREB2A and heat stress transcription factor HsfA3 regulates the heat stress response of Arabidopsis. Plant J 53:264–274. doi:10.1111/j.1365-313X.2007.03334.x
Vacca RA, De Pinto MC, Valenti D et al (2004) Production of reactive oxygen species, alteration of cytosolic ascorbate peroxidase, and impairment of mitochondrial metabolism are early events in heat shock-induced programmed cell death in tobacco Bright-Yellow 2 cells. Plant Physiol 134:1100–1112
Von Koskull-Doering P, Scharf KD, Nover L (2007) The diversity of plant heat stress transcription factors. Trends Plant Sci 12:452–457. doi:10.1016/j.tplants.2007.08.014
Yoshida T, Sakuma Y, Todaka D et al (2008) Functional analysis of an Arabidopsis heat-shock transcription factor HsfA3 in the transcriptional cascade downstream of the DREB2A stress-regulatory system. Biochem Biophys Res Commun 368:515–521. doi:10.1016/j.bbrc.2008.01.134
Yu E, Fan C, Yang Q et al (2014) Identification of heat responsive genes in Brassica napus siliques at the seed-filling stage through transcriptional profiling. PLoS One 9:e101914. doi:10.1371/journal.pone.0101914
Acknowledgements
This work was financially supported through a grant from the Special Foundation from Guizhou Academy of Agriculture Science ([2013] 003), the Provincial Natural Science Foundation of Guizhou Province (Qian J [2015]2080), and the Provincial Natural Science Foundation of Guizhou Province (Qian J LKN [2013] 04).
Author information
Authors and Affiliations
Corresponding authors
Additional information
Communicated by A Chandra.
X. Li and X. Wang contributed equally to the work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
11738_2017_2351_MOESM1_ESM.tif
Fig. S1 Phenotypes of hsfa2, hsfa3 and hsfa2 hsfa3 in ATLR assays with different plant densities at different growth stages. (a-c) Seedlings planted using spaced sowing. (d-f) Seedlings planted using bunch sowing. (g) Schematic drawing showing the positions of all plant seedlings. (a, d) Plants grown under normal conditions. (b, e) Heat stress was performed at 3 d after sowing. (c, f) Heat stress was performed 7 d after sowing. The images were captured 13 d after sowing in control and ATSR and 17 d after sowing in ATLR (TIFF 4494 kb)
Rights and permissions
About this article
Cite this article
Li, Xd., Wang, Xl., Cai, YM. et al. Arabidopsis heat stress transcription factors A2 (HSFA2) and A3 (HSFA3) function in the same heat regulation pathway. Acta Physiol Plant 39, 67 (2017). https://doi.org/10.1007/s11738-017-2351-7
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11738-017-2351-7