Abstract
Transforming growth factor-betas (TGF-βs) and their family members that include bone morphogenic proteins and activins have been implicated in the regulation of proliferation, hibernation, quiescence and differentiation of hematopoietic stem cells (HSCs). Increasing evidence suggests that the superfamily of TGF-βs play an integral role in the intercellular cross-talk between the stem cells and their microenvironment as well as within the cells at an intracellular level. Active sites of hematopoiesis, such as fetal liver and bone marrow are known to have abundant presence of TGF-β indicating their importance in the maintenance and regulation of hematopoiesis. One of the striking features of TGF-β superfamily is the variety of effects they evoke, contingent on the developing history of the responding cells. In the present review, we discuss the Smad-dependent and Smad-independent TGF-β signaling pathways in order to understand and underscore their role in the regulation of HSCs.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The transforming growth factor-beta (TGF-β) family contains three closely related mammalian isoforms—TGF-β1, β2, and β3 that arose by duplication of a common ancestor. Similarity is the most striking in the C-terminal domain (64–82 %), with nine conserved cysteine residues forming four intra-chain and one inter-chain disulfide bonds. Despite this high sequence homology, analysis of the in vivo functions of the three isoforms by gene knockouts reveal striking differences, illustrating their non-redundancy. Overall, TGF-β1 is the most abundant isoform, with the largest sources of TGF-β1 being platelets (20 mg/kg) and bone (200 μg/kg) (Janssens et al. 2005).
TGF-β1 is a ubiquitous, multifunctional growth factor that regulates a broad range of biological processes, including cell proliferation, cell survival, cell differentiation, cell migration, and production of extracellular matrix (ECM) molecules (Han et al. 2000). The combined actions of these cellular responses mediate the global effects of TGF-β1 on immune responses, angiogenesis, wound healing, development, and bone formation (Janssens et al. 2005). Regarding the diversity of processes in which TGF-β1 is involved, it is not surprising that this cytokine is of major importance both during embryogenesis and in maintaining tissue homeostasis during life (Godár et al. 1999). Although the role of TGF-β1 in normal physiologic processes is vital, several lines of evidence implicate the role of this cytokine in the pathogenesis of autoimmune diseases, malignancy, impaired wound healing and experimental and human fibrotic conditions (Border and Noble 1995). Involvement of TGF-β1 in these diseases likely occurs via its diverse effects on a number of factors important in fibrosis, such as synthesis and deposition of various ECM molecules. TGF-β1 stimulates fibroblasts to synthesize collagen, fibronectin and glycosaminoglycans (Ignotz and Massague 1986; Sporn et al. 1987); it enhances neovascularization and modulates the production of a variety of proteases and their inhibitors (Ghahary et al. 1999). Collectively, these reports suggest that TGF-β may function as a double-edged sword with both therapeutic and pathological potential.
TGF-β signaling pathway
TGF-β is stored in the ECM as a latent complex with its prodomain. The extracellular concentration of TGF-β activity is primarily regulated by the conversion of latent TGF-β to active TGF-β. Although tissues contain significant quantities of latent TGF-β, only a fraction of this latent TGF-β is activated to generate a cellular response. The secretion of members of the TGF-β family as latent complexes necessitates the existence of regulated activation process, which is most probably mediated through the activities of proteases that preferentially degrade the TGF-β pro-segments and thereby release the highly stable and active TGF-β dimer (Annes et al. 2003). A commonality among these activators is that they are all indicative of ECM perturbations. Indeed, given the profound effects of TGF-β on matrix homeostasis, the primary change that the TGF-β sensor detects may be alterations in the matrix. To name a few, latent TGF-β can be activated by plasmin, matrix metalloproteases 9 and 2 (MMP-9 and 2), thrombospondin and αvβ6 integrin (Derynck et al. 2001).
The TGF-β receptors
TGF-β can interact with its receptor to induce signaling. All members of the TGF-β superfamily signal through a dual receptor system of type I and type II transmembrane serine/threonine kinases. These receptors belong to a family of glycoproteins characterized by a cysteine-rich extracellular region, a single transmembrane α-helix, and a cytoplasmic domain with a kinase domain (Fukuda et al. 1998). Although there is only one type II TGF-β receptor (TβRII), there are three type I receptors, namely activin receptor like kinase 1, 2 and 5 (ALK1, 2 and 5) (Derynck et al. 2001). For members of the TGF-β family, TβRII is the sole type II receptor shown to mediate signaling. This is reflected by the phenotypic identity of the tgfbr2 and those tgfb1 knockout mice that die in utero (Sanford et al. 1997). Of the type I receptors, ALK5, ALK1, and possibly ALK2, can transmit TGF-β signals. ALK5 (TβRI) is the most important type I receptor for TGF-β, which is underscored by the comparable (although not identical) phenotypes of tgfb1 and alk5 knockout mice: histological examination of the yolk sacs of tgfbr1−/− embryos shows an image very similar to that of tgfb1−/− embryos that die during embryogenesis (Larsson et al. 2001). In bone cells, TβRI seems to be the only type I receptor involved in signaling (Dallas et al. 2002). Recently, it has been shown that transcription intermediary factor 1γ (Tif1γ) controls TβRI turnover and promotes physiological aging of hematopoietic stem cells (HSCs) (Quéré et al. 2014).
Betaglycan and endoglin are the so called type III or accessory receptors, which are indirectly involved in signaling through the modulation of ligand-binding specificity. Betaglycan (TβRIII) can bind all three TGF-β isoforms and is implicated in the presentation of TGF-β to TβRII (Lopez-Casillas et al. 1993). For TGF-β2, which has a low intrinsic affinity for TβRII, both in vitro and in vivo data have demonstrated its signaling being dependent on the presentation of this isoform by betaglycan (Sankar et al. 1995; Stenvers et al. 2003). Endoglin can bind TGF-β1 and β3 in the presence of TβRII (Cheifetz et al. 1992). Mutations in ENG, the gene encoding endoglin, lie at the basis of the human disease, hereditary hemorrhagic telangiectasia, an autosomal dominant disorder characterized by multi-system vascular dysplasia (McAllister et al. 1994). A murine model of this disorder presents with a phenotype that is remarkably similar to that of tgfb1 and tgfbr2 knockout mice, suggesting an in vivo requirement for endoglin in TGF-β1 signaling (Bourdeau et al. 1999). Bone marrow stromal cells (BMSCs) and mature osteoblasts express the two types of type III receptors, whereas osteoclasts seem to lack betaglycan (Walsh et al. 2003).
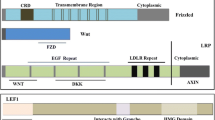
In the absence of ligand, both type I and type II receptors are present as homodimers. Upon ligand binding TβRII recruits and phosphorylates TGF-β receptor-I (TβRI) kinase activity. The ligand-induced multimerization of the receptor is followed by a trans-phosphorylation of the conserved glycine- and serine-rich (GS) domain of TβRI by the constitutively phosphorylated TβRII kinase, resulting in the activation of TβRI (Massague 1998). This trans-phosphorylation is the first step in the intracellular transmission of the signal (Fig. 1).
Signaling by TGF-β family members through the Smad-dependent and Smad-independent pathways: TGF-β induces its response by multimerization of TGF-β receptor II (TβRII) and TGF-β receptor I (TβRI). Both the receptors have the same overall domain structure—an extracellular cysteine rich domain (ED), a single transmembrane helix and an intracellular serine-threonine kinase domain (KD). TGβRII activates TβR1 by transphosphorylation of its glycine- and serine-rich (GS) domains. The Smad-dependent signaling is initiated by the binding of the Mad homology (MH) 2 domain of R-Smad to an adaptor protein (SARA). Eventually R-Smad dissociates from TβRI and oligomerizes with Smad 4 to form a heterodimeric complex which translocates into the nucleus thereby regulating the cellular response. The Smad-independent signaling is initiated by activation of the mitogen-activated protein kinase (MAPK) pathways—the p38 MAPK, c-Jun amino terminal kinase (JNK) and p44/42 or the extracellular regulated kinase (ERK)1/2. The MAPK pathways may or may not be regulated by the binding of MAPK kinase kinase (MAPKKK), TGF-β-associated kinase (TAK) 1 with TAK1-binding protein (TAB) 1 leading to the activation of transcription factors—ATF, cJun and STAT proteins. Depending upon the type of TGF-β signaling pathway initiated, the cell is directed to undergo either proliferation or differentiation or apoptosis
Smad-dependent signaling
It was almost 25 years ago that Smads were first discovered to be key intracellular mediators of the transcriptional responses to TGF-β (Raftery et al. 1995; Sekelsky et al. 1995). Soon after the genetic studies in Drosophila melanogaster provided a breakthrough in our understanding of intracellular TGF-β signaling through the identification of mothers against dpp (Mad). Its protein product plays a role in mediating the function of decapentaplegic (dpp), the D. melanogaster ortholog of bone morphogenic protein 2 or 4 (BMP2 or 4) (Sekelsky et al. 1995). This discovery was followed by the genetic identification of the homologous Sma genes in Caenorhabditis elegans and subsequently the Smad genes (for Sma and Mad related) in vertebrates (Akiyoshi et al. 1999). The Smads turned out to play a central role in the transmission of signals from all receptors activated by TGF-β superfamily members to the target genes in the nucleus (Wrana 2009).
Smads can be classified into three groups: receptor-mediated Smads (R-Smads), common-partner Smad (Co-Smads) and inhibitory Smads (I-Smads). Of the eight Smad family members in humans, five (Smads 1, 2, 3, 5 and 8) function as receptor substrates. Smads 2 and 3 do so as substrates of TGF-β, nodal and activin receptors, and Smads 1, 5, and 8 as substrates of the receptors for BMPs, myostatin and antimuellerian hormone. The only Co-Smad identified so far in mammals is Smad 4, which is commonly used by all TGF-β superfamily members. The class of the I-Smads comprises Smad 6 and 7. Smad 6 is an inhibitor of BMP signaling, whereas Smad7 inhibits both TGF-β/activin and BMP signaling (He et al. 2006).
The R and Co-Smads share a similar structure with conserved amino- and carboxyterminal domains, the Mad homology (MH)-1 and MH2 domains, connected by a more divergent linker region. In addition, the R-Smads contain a carboxyterminal phosphorylation site, the SSXS motif. Lacking any recognizable enzyme activity, Smads achieve their signaling capacity mainly through protein–protein or DNA protein interactions, exerted by the different domains. The MH1 domain can mediate direct DNA binding, whereas the MH2 domain is implicated in receptor interaction, Smad oligomerization, and transcriptional activation. Both domains further drive nuclear import and allow binding to various transcription factors and cofactors. The divergent linker region contains multiple phosphorylation sites, allowing fine tuning of Smad functioning by many different signaling pathways in the cell, which converge on phosphorylation of this region (Kretzschmar and Massague 1998; Janssens et al. 2005).
Through their MH2 domain, R-Smads can bind to the GS domain of TβRI, an interaction promoted by adaptor proteins such as Smad anchor for receptor activation (SARA). SARA specifically interacts with TβRI and functions to recruit Smad 2 and Smad 3 (R-Smads) to the activated receptor complex, presumably in the endocytotic compartment (Tsukazaki et al. 1998; Wu et al. 2000; Di Guglielmo et al. 2003). Binding of the R-Smad to TβRI causes phosphorylation of the former at its carboxy-terminal SSXS motif by the TβRI kinase domain, which causes R-Smad to dissociate from the receptor complex and oligomerize with Smad 4 to form a heterodimeric complex that is then translocated into the nucleus (Wu et al. 2001; Qin et al. 2002; Inman and Hill 2002; Chacko et al. 2004). R-Smad/Smad 4 complexes have been shown to interact directly with specific Smad-binding elements (SBEs), GC-rich regions in the promoter of TGF-β target genes, as well as with transcription factors, co-activators and co-repressors to regulate transcription of target genes in both cell-type-specific and ligand dose-dependent manner (Heldin et al. 1997; Derynck and Zhang 2003; Ten Dijke and Hill 2004). R-Smads and Smad 4 bind to specific DNA sequences with a 100-fold lower affinity than the interacting high-affinity, DNA-binding transcription factors, yet their DNA binding (except Smad 2) is required for transcriptional activation (Derynck and Zhang 2003).
The Smad complexes accumulate in the nucleus and remain there for hours (Ten Dijke and Hill 2004). The levels of the Smad complexes in the nucleus, therefore, determine the nature and the duration of the signal. Upon discontinuation of the signal, the R-Smads get dephosphorylated and disassociated from Smad 4, and are exported from the nucleus. If the receptors are active, Smad signaling continues, but if the receptors are inactive, the dephosphorylated Smads accumulate over time in the cytoplasm and the signaling stops (Inman and Hill 2002; Xu et al. 2002).
The I-Smads act as inhibitors of Smad-mediated signal transduction by interacting with the type I receptor and inhibiting the phosphorylation of R-Smads (Nakao et al. 1997), by recruiting E3-ubiquitin ligases to degrade the activated type I receptors or by direct dephosphorylation and subsequent inactivation of the type I receptor (Shi and Massague 2003; Ten Dijke and Hill 2004). Alternatively, I-Smads may compete with Smad 4 in binding R-Smads and thereby prevent the formation of the R-Smad/Smad 4 complex. This general mechanism underlies a large number of TGF-β gene responses controlling cell proliferation, organization and fate (Larsson and Karlsson 2005).
Smad-independent signaling
Although the Smads are critical mediators in the TGF-β signaling pathway, a substantial body of evidence illustrates the existence of additional, Smad-independent pathways (Aubin et al. 2004). First, a partial preservation of TGF-β signaling in Smad 4-deficient cells is highly suggestive of a Smad-independent signaling (Dai et al. 1999; Hocevar et al. 1999; Sirard et al. 2000). In addition, several other lines of evidence point to the involvement of mitogen-activated protein kinase (MAPK) signaling pathways in transmitting the TGF-β signals from receptor to nucleus.
In vitro kinase assays have demonstrated that TGF-β can activate all three MAPK pathways—leading to extra-cellular regulated kinase (ERK), c-Jun amino terminal kinase (JNK) and p38 MAPK activation (Mulder 2000) and phosphorylation of members of the Jun, Fos and activating transcription factor (ATF) families, which homo and heterodimerize to form the activator protein-1 (AP1) (Johnson and Lapadat 2002). Also the observation that MAPK consensus sites are found in all R-Smads further adds to the complexity of TGF-β signaling and also suggests that TGF-β and MAPK crosstalk may constitute an important mechanism regulating the cellular outcome of TGF-β signals (Aubin et al. 2004).
Crosstalk can be obtained through physical interaction between Smad 2, 3 and 4 and members of the Jun, Fos and ATF families bound to their AP-1 site in the promoter of the target genes (Zhang et al. 1998; Liberati et al. 1999). In addition, JNK (activated by TGF-β) can phosphorylate Smad 3, thus facilitating activation and nuclear translocation of the latter in response to TGF-β (Engel et al. 1999). When the profile of hundreds of TGF-β-controlled genes in fibroblasts deficient in Smad 2, Smad 3 or ERK signaling respectively was investigated, Smad 3 was demonstrated to be the critical mediator for expression of immediate early genes (IEGs). Smad 2 and the ERK pathways were found to function predominantly in the trans-modulation of immediate early and intermediate gene regulation (Janssens et al. 2005). Despite ample in vitro evidence in the literature for the involvement of MAPKs in the TGF-β signaling cascade, data that unequivocally demonstrate the need for MAPK pathways in the in vivo TGF-β-mediated responses are lacking. Although knockout and transgenic mouse models of numerous MAPK signaling intermediates are available (Wada and Penninger 2004) none of them are scored for defects in TGF-β signaling. However, keratinocytes derived from MAPK kinase (MAPKK)—MEKK1 deficient mice show no migration in response to TGF-β1 (Zhang et al. 2003) and MAPKK (MKK) 3(−/−) mesangial cells are defective in TGF-β1-induced vascular endothelial growth factor (VEGF) expression (Wang et al. 2004). TGF-β-induced activation of hyaluronan synthases is mediated through the activation of p38 MAPK and blocking of p38 MAPK inhibits TGF-β effect by 90 % (Stuhlmeier and Pollaschek 2004). These observations clearly show the requirement for MAPK-dependent signaling in transmitting the TGF-β signals.
TGF-β-associated kinase (TAK) 1 is a member of the MAPKK kinase (MAPKKK) family (which functions in TGF-β signaling pathways in mammalian cells) and is activated by various cytokines including TGF-β family ligands and IL1 (Kishimoto et al. 2000). Upon stimulation by IL1, TAK1 constitutively associates with TAK1-binding protein (TAB 1) through a tumor necrosis factor receptor-associated factor 6 (TRAF6)-dependent mechanism, which further leads to the activation of JNK and nuclear factor (NF)-κB pathways suggesting a role of TAB 1 as an adaptor that links TAK1 to TRAF6 in response to IL1, thereby mediating TAK1 activation (Takaesu et al. 2001). The association of the TAK1-TAB 1 complex also leads to p38 kinase activation. Although TAB 1 was originally thought to interact with and activate TAK1 directly, recent research shows that TAB 1 can bind directly to p38 and promote MKK-independent p38 autophosphorylation (Lu et al. 2006). This demonstrates that the stress kinases can be also regulated by an indirect mechanism involving the TAK1-TAB 1 pathway either in a TGF-β-dependent/independent mechanism further adding to the complexity of the Smad-independent TGF-β signaling.
To summarize, JNK, ERK, and p38 MAPK all contribute considerably to the whole of the TGF-β-induced responses, but further characterization is required to assess their importance in relation to the Smad dependent and other TGF-β-induced signaling pathways.
Role of TGF-β signaling in HSCs
A critical role for TGF-β in the regulation of HSCs and progenitor cells was demonstrated more than 15 years ago. The original findings showed a potent inhibition by TGF-β1 on the growth of early multiple progenitor populations (MPPs), while more mature progenitors were unaffected (Ohta et al. 1987; Keller et al. 1988). A large number of studies on both human and murine cells have supported these original findings of potent growth inhibitory actions on early hematopoietic progenitors (Ottmann and Pelus 1988; Sing et al. 1988; Keller et al. 1990; Jacobsen et al. 1991). Although the mechanism of TGF-β action on hematopoietic progenitors is not fully understood, certain studies reveal that the effects are in part due to down regulation of cytokine receptors [like receptors of interleukin (IL) 1, granulocyte monocyte-colony stimulating factor (GM-CSF), IL3, granulocyte-colony stimulating factor (G-CSF) and stem cell factor (SCF)] and modulation of genes involved in cell cycle (Karlsson et al. 2007). A study by Scandura et al. (2004) has shown that TGF-β induces cell cycle arrest in human hematopoietic cells by an upregulation of the cyclin-dependent kinase inhibitor, p57KIP2. This is supported by the findings demonstrating reversibility in the growth inhibitory actions of TGF-β, suggesting that TGF-β delays the proliferation rather than exerting an irreversible negative effect such as induction of apoptosis (Sitnicka et al. 1996; Batard et al. 2000). However, a number of reports have shown the involvement of TGF-β in apoptosis of bone marrow (BM) progenitors. In fact, both apoptotic and anti-apoptotic effects of TGF-β have been described (Jacobsen et al. 1995; Veiby et al. 1996; Dybedal et al. 1997). Thus, TGF-β may regulate growth of hematopoietic progenitors through effects on both cell cycling and apoptosis. Furthermore, neutralization studies using TGF-β monoclonal antibodies has shown to recruit early progenitor cells into cell cycle determining the role of endogenous TGF-β signaling in maintaining the quiescence of HSCs (Scandura et al. 2004). TGF-β is now documented as a potent inhibitor of HSC proliferation in vitro, while its role in vivo is largely unknown (Söderberg et al. 2009). In a recent study conducted by Park et al. (2014), they showed that Mushashi-2 (Msi2) is an important regulator of the HSC translatome that controls cell fate, lineage bias and TGF-β signaling in HSCs.
On the other hand, it has been difficult to determine the effects of TGF-β on more mature progenitor cells due to the expression of several receptors by them and because of their dependence upon growth factors. For example TGF-β inhibits IL3-induced granulocyte–macrophage (GM) colony formation; while GM-CSF-induced GM colony formation is stimulated (Ruscetti and Bartelmetz 2001). These studies demonstrate that the effects of TGF-β in these in vitro systems are dependent on the differentiation stage of the target cells and the actions of other cytokines.
Working at a common level of convergence for all TGF-β superfamily signals, Smad 4 is the key player in orchestrating these effects. Thus, the role of Smad 4 in the regulation of HSCs has been keenly looked upon (Wrana 2009). Several groups have demonstrated that conventional Smad−/− embryos die at embryonic day 7.5 because of the impaired proliferation of ectoderm, resulting in a lack of mesoderm formation (Sirard et al. 1998; Yang et al. 1998), while adult Smad 4 heterozygote mice develop polyps and tumors of the gastrointestinal tract (Takaku et al. 1999; Xu et al. 2000).
Such findings have also been observed in human patients with juvenile polyposis syndrome caused by Smad 4 mutations (Howe et al. 1998). Hence, because of the early embryonic lethality caused by the homozygous knockout of the Smad 4 gene, its role in HSC function has remained elusive. But Karlsson et al. (2007), were successful in studying the complete role of Smad 4-dependent signaling in HSCs and hematopoiesis by means of inducible MxCre/Smad 4−/− mice, indicating that this Smad is critical for the self-renewal of HSCs. Simultaneously, other works have identified the nuclear protein intermediary factor-1 γ (TIF1γ) as a competitor to Smad 4 for R-Smad binding in human CD34+ cells, demonstrating additional complexities in the interaction between canonical Smad pathway and other regulatory circuits in hematopoietic cells (He et al. 2006).
One of the properties of the Smad-signaling pathway is that they are inherently redundant and as a result, shared by numerous ligands (Chadwick et al. 2005). Therefore, to investigate whether a simultaneous blocking of Smad-signaling branches in HSCs might affect fate decisions, such as self-renewal and differentiation, Blank et al. (2006) carried out a one of a kind study wherein they blocked the entire Smad network in murine HSCs using over expression of the I-Smad 7 by a retroviral gene transfer approach. Through the construction of such a model they were able to successfully show that blocking of the entire Smad pathway led to increased self renewal in vivo, as assessed by both phenotypic and functional assays in primary and secondary recipients. Most importantly, Smad 7-overexpressing HSCs could give rise to both myeloid and lymphoid cell compartments at normal distributions, providing evidence for an unperturbed differentiation capacity. Furthermore, gene-expression analysis of purified HSCs from BM of recipient mice revealed decreased levels of p21 in parallel with an increase of Bmi-1, thus suggesting a plausible mechanism downstream of Smad 7. Despite the success using gene knockout models, the overall molecular mechanisms of receptor/Smad signal transduction in HSCs still remain poorly characterized, mainly due to their rarity in the BM and the requirement of large numbers of cells for the functional analysis of signal transduction (Utsugisawa et al. 2006).
In a study, the molecular mechanisms by which stromal-derived factor (SDF) 1 exhibited a cell cycle promoting effect and interacted with TGF-β’s negative effects on cell cycle orchestration of human hematopoietic CD34+ progenitor cells was analyzed. They showed that a cross-talk between SDF1 and TGF-β signaling pathways in the control of CD34+ cell cycling was mediated through phospho-inositide 3-kinase (PI3K)/AKT and Smad 3 signaling pathways. These results shed new light on the intracellular mechanisms of HSC hibernation (by TGF-β) and activation (by SDF1) in maintaining hematopoietic homeostasis (Chabanon et al. 2008).
Work carried out with both vertebrate and invertebrate model systems indicates that TGF-β1 can serve as a potent morphogen, acting across the developing tissues in a graded fashion to specify a patterned array of cell fate. A defining feature of the morphogens is their ability to specify multiple cell types over a range of concentrations (Fortunel et al. 2000). Active sites of hematopoiesis, such as fetal liver and red marrow, have been shown to have an abundant presence of TGF-β1 (Schmid et al. 1991; Kale and Limaye 1999), indicating that this pleiotrophic growth factor indeed has an important role in the maintenance and the regulation of all stages of hematopoiesis (Ruscetti and Bartelmetz 2001). In the Dexter-type long-term cultures (LTBMC), copious amounts of TGF-β1 gets accumulated over time and the cultures become static (Cashman et al. 1990; Eaves et al. 1991). However, neither its exact function in maintaining the active hematopoiesis nor its mode of action has been clearly defined (Kale 2004). TGF-β1 has been reported to preserve the immaturity of the primitive hematopoietic stem progenitor cells (HSPCs) by increasing the expression of CD34 antigen on them (Batard et al. 2000; Pierelli et al. 2002). It has also been demonstrated in cell line models that an exogenous addition of TGF-β1 causes upregulation of the CD34 antigen in the primitive cell lines like TF1 and KG1a, but not in the more differentiated cell lines like HL60 and K562 (Batard et al. 2000). Studies using ex vivo expansion assays have further shown that in liquid culture system supplemented with growth factors, TGF-β1 preserves the long-term culture initiating cells (Garbe et al. 1997). These observations suggest that TGF-β1 may protect the early HSPCs from excessive differentiation signals.
Discussion
Multiple signaling networks orchestrate the development and the differentiation of embryonic as well as adult stem cells into functional cells of neuronal, hematopoietic, mesenchymal, and epithelial lineages. Among these, the signaling mechanisms activated by TGF-β family proteins have emerged as key players in various aspects of the stem cell development such as the self-renewal and maintenance of stem cells in their undifferentiated state (stem cell pool), their commitment towards a specific differentiation lineage, and the progression of differentiation along an individual lineage (functional differentiation and maturation). As an outcome of the gene knockout experiments carried out in the ES cells, the TGF-β family proteins have emerged as bifunctional regulators of the maturation of cells in various lineages mentioned above (Mishra et al. 2005). TGF-β is also known to play an essential role in regulating the homeostasis of cells in the lymphoid lineage. Although, the TGF-β signaling is not required for normal thymopoiesis, it is essential for regulating expansion, activation, as well as for maintaining effector functions of the mature CD4+ and CD8+ T cells in the peripheral lymphoid organs and the target tissues (Letterio 2005).
Kale et al. have demonstrated that TGF-β1 evokes a biphasic dose-dependent response in the HSCs. They reported that low concentrations induced p44/42 MAPK activation whereas high inhibitory concentrations induced p38 activation in KG1a cells and a bidirectional or biphasic effect of TGF-β1 on colony formation (CFU) of hematopoietic cells as a function of its concentration. Furthermore, they showed that the signaling mechanisms induced by high inhibitory concentrations of TGF-β1 was mediated by Smad 3—TAB 1-TAK 1—p38 MAPK—ATF2—c-Jun pathway, whereas low concentrations induced proliferation through activation of the p44/42 MAPK-STAT pathway (Kale and Limaye 1999; Kale 2004; Kale and Vaidya 2004). Additionally, Challen et al. (2010) showed that TGF-β1 appeared to be a general stimulatory factor for the myeloid-biased HSC (My-HSC) proliferation, while having an inhibitory effect on the lymphoid-biased HSC (Ly-HSCs), thereby supporting the theory that distinct HSC subtypes are indeed differentially regulated by TGF-β1.
Most of the studies have labeled TGF-β1 as a well-known inhibitor of hematopoiesis, its action largely dependent upon the differentiation status of the cells (Hu and Zuckerman 2001; Kim and Letterio 2003). Whereas the early HSPCs seem to be sensitive to the inhibition by TGF-β1, more differentiated progenitors get stimulated by this factor (Fortunel et al. 2000; Ruscetti and Bartelmetz 2001). A study by Yamazaki et al. (2009) uncovered a critical role of TGF-β as a candidate niche signaling molecule in the control of HSC hibernation or quiescence. Since hibernation of HSCs is indispensable for the HSC maintenance, it is quite possible that TGF-β could be used as a novel tool for an ex vivo modeling of the HSC niche. It is also possible that since TGF-β maintains the undifferentiated state of HSCs—an important property of ‘stemness’, other studies may have misinterpreted the role of TGF-β1 as a negative regulator of early HSPC proliferation! Figuring out the precise molecular mechanism(s) underlying HSC hibernation may be needed to elucidate this issue.
References
Akiyoshi S, Inoue H, Hanain JI et al (1999) c-Ski acts as a transcriptional co-repressor in transforming growth factor-β signaling through interaction with Smads. J BiolChem 274(49):35269–35277
Annes JP, Munger JS, Rifkin DB (2003) Making sense of latent TGFβ activation. J Cell Sci 116:217–224
Aubin J, Davy A, Soriano P (2004) In vivo convergence of BMP and MAPK signaling pathways: impact of differential Smad1 phosphorylation on development and homeostasis. Genes Dev 18:1482–1494
Batard P, Monier MN, Fortunel N et al (2000) TGF-(beta)1 maintains hematopoietic immaturity by a reversible negative control of cell cycle and induces CD34 antigen up-modulation. J Cell Sci 113(Pt 3):383–390
Blank U, Karlsson G, Moody JL et al (2006) Smad7 promotes self-renewal of hematopoietic stem cells. Blood 108(13):4246–4254
Border WA, Noble NA (1995) Targeting TGF-β for treatment of disease. Nat Med 1:1000–1001
Bourdeau A, Dumont DJ, Letarte M (1999) A murine model of hereditary hemorrhagic telangiectasia. J Clin Invest 104:1343–1351
Cashman JD, Eaves AC, Raines EW et al (1990) Mechansims that regulate the cell cycle status of very primitive hematopoietic cells in long-term human marrow cultures. I. Stimulatory role of a variety of cell activators and inhibitory role of TGF β1. Blood 75:96–101
Chabanon A, Desterke C, Rodenburger E et al (2008) A cross-talk between SDF-1 and TGF-β controls the quiescence/cycling switch of CD34+ progenitors through FoXO3 and mTOR. Stem Cells 26(12):3150–3161
Chacko BM, Qin BY, Tiwari A et al (2004) Structural basis of heteromericSmad protein assembly in TGF-β signaling. Mol Cell 5:813–823
Chadwick K, Shojaei F, Gallacher L et al (2005) Smad7 alters cell fate decisions of human hematopoietic repopulating cells. Blood 105(5):1905–1915
Challen GA, Boles NC, Chambers SM et al (2010) Distinct hematopoietic stem cell subtypes are differentially regulated by TGF-β1. Cell Stem Cell 6:265–278
Cheifetz S, Bellon T, Cales C et al (1992) Endoglin is a component of the transforming growth factor-β receptor system in human endothelial cells. J Biol Chem 267:19027–19030
Dai JL, Schutte M, Bansal RK et al (1999) Transforming growth factor-β responsiveness in DPC4/SMAD4-null cancer cells. Mol Carcinog 26:37–43
Dallas SL, Rosser JL, Mundy GR, et al (2002) Proteolysis of latent transforming growth factor-β (TGF-β)-binding protein-1 by osteoclasts. A cellular mechanism for release of TGF-β from bone matrix. J Biol Chem 277:21352–21360
Derynck R, Zhang YE (2003) Smad-dependent and Smad-independent pathways in TGF-beta family signalling. Nature 425(6958):577–584
Derynck R, Akhurst R, Balmain A (2001) TGF-β signaling in tumor suppression and cancer progression. Nat Gen 29:117–129
Di Guglielmo GM, Le Roy C, Goodfellow AF et al (2003) Distinct endocytic pathways regulate TGF-β receptor signaling and turnover. Nat Cell Biol 5:410–421
Dybedal I, Guan F, Borge OJ et al (1997) Transforming growth factor-beta1 abrogates Fas induced growth suppression and apoptosis of murine bone marrow progenitor cells. Blood 90(9):3395–3403
Eaves CJ, Cahsmna JD, Kay RJ et al (1991) Mechansims that regulate the cell cycle status of very primitive hematopoietic cells in long-term human marrow cultures. II. Analysis of positive and negative regulators produced by stromal cells within the adherent layer. Blood 78:110–117
Engel ME, McDonnell MA, Law BK et al (1999) Interdependent SMAD and JNK signaling in transforming growth factor-β-mediated transcription. J Biol Chem 274:37413–37420
Fortunel N, Hatzfeld J, Aoustin L et al (2000) Specific dose-response effects of TGF-beta1 on developmentally distinct hematopoietic stem/progenitor cells from human umbilical cord blood. Hematol J 1:126–135
Fukuda K, Kawata S, Tamura S et al (1998) Transforming growth factor-β1-induced degradation of activated Src tyrosine kinase in rat fibroblasts. Oncogene 16:3349–3356
Garbe A, Spyridonidis A, Mobest D et al (1997) Transforming growth factor beta 1 delays formation of granulocyte macrophage colony-forming cells, but spares more primitive progenitors during ex vivo expansion of CD34+ hematopoietic progenitor cells. Br J Hematol 99:951–958
Ghahary A, Tredget EE, Mi L et al (1999) Cellular responses to latent TGF-β1 is facilitated by insulin-like growth factor-II/mannose-6-phosphate receptors on MS-9 cells. Exp Cell Res 251:111–120
Godár S, Hořejši V, Weidle UH et al (1999) M6P/IGFII-receptor complexes urokinase receptor and plasminogen for activation of transforming growth factor-β1. Eur J Immunol 29:1004–1013
Han J, Hajjar DP, Tauras JM et al (2000) Transforming growth factor-β1 (TGF-β1) and TGF-β2 decrease expression of CD36, the type B scavenger receptor, through mitogen-activated protein kinase phosphorylation of peroxisome proliferators-activated receptor-γ. J BiolChem 275(2):1241–1246
He W, Dorn DC, Erdjument-Bromage H et al (2006) Hematopoiesis controlled by distinct TIF1γ and Smad4 branches of the TGFβ pathway. Cell 125:929–941
Heldin CH, Miyazono K, ten Dijke P (1997) TGF-beta signalling from cell membrane to nucleus through SMAD proteins. Nature 390(6659):465–471
Hocevar BA, Brown TL, Howe PH (1999) TGF-β induces fibronectin synthesis through a c- Jun N-terminal kinase-dependent, Smad4-independent pathway. EMBO J 18:1345–1356
Howe JR, Roth S, Ringold JC et al (1998) Mutations in the SMAD4/DPC4 gene in juvenile polyposis. Science 280:1086–1088
Hu X, Zuckerman KS (2001) Transforming growth factor: signal transduction pathways, cell cycle mediation, and effects on hematopoiesis. J Hematother Stem Cell Res 10:67–74
Ignotz RA and Massague J (1986) Transforming growth factor-β stimulates the expression of fibronectin and collagen and their incorporation into the extracellular matrix. J Biol Chem 261:4337–4345
Inman GJ and Hill CS (2002) Stoichiometry of active Smad-transcription factor complexes on DNA. J Biol Chem 277:51008–51116
Jacobsen SE, Keller JR, Ruscetti FW et al (1991) Bidirectional effects of transforming growth factor beta (TGF-beta) on colony-stimulating factor-induced human myelopoiesisin vitro: differential effects of distinct TGF-beta isoforms. Blood 78(9):2239–2247
Jacobsen FW, Stokke T, Jacobsen SE (1995) Transforming growth factor-beta potently inhibits the viability-promoting activity of stem cell factor and other cytokines and induces apoptosis of primitive murine hematopoietic progenitor cells. Blood 86(8):2957–2966
Janssens K, ten Dijke P, Janssens S et al (2005) Transforming growth factor-β1 to the bone. Endocr Rev 26(6):743–774
Johnson GL, Lapadat R (2002) Mitogen-activated protein kinase pathways mediated by ERK, JNK, and p38 protein kinases. Science 298:1911–1912
Kale VP (2004) Differential activation of MAPK signaling pathways by TGFβ1 forms the molecular mechanism behind its dose dependent bi-directional effects on hematopoiesis. Stem Cells Dev 13:27–38
Kale VP, Limaye LS (1999) Stimulation of adult human bone marrow by factors secreted by fetal liver hematopoietic cells: in vitro evaluation using semisolid clonal assay system. Stem Cells 17(2):107–116
Kale VP, Vaidya AA (2004) Molecular mechanisms behind the dose-dependent differential activation of MAPK pathways induced by transforming growth factor-β1 in hematopoietic cells. Stem Cells Dev 13:536–547
Karlsson G, Blank U, Moody JL et al (2007) Smad4 is critical for self-renewal of hematopoietic stem cells. J Exp Med 204(3):467–474
Keller JR, Mantel C, Sing GK et al (1988) Transforming growth factor beta 1 selectively regulates early murine hematopoietic progenitors and inhibits the growth of IL-3-dependent myeloid leukemia cell lines. J Exp Med 168(2):737–750
Keller JR, McNiece IK, Sill KT et al (1990) Transforming growth factor beta directly regulates primitive murine hematopoietic cell proliferation. Blood 75(3):596–602
Kim SJ, Letterio J (2003) Transforming growth factor-beta signaling in normal and malignant hematopoiesis. Leukemia 17:1731–1737
Kishimoto K, Matsumoto K, Ninomiya-Tsuji J (2000) TAK1 mitogen-activated protein kinase kinasekinase is activated by autophosphorylation within its activation loop. J Biol Chem 275(10):7359–7364
Kretzschmar M, Massague J (1998) SMADs: mediators and regulators of TGF-beta signaling. Curr Opin Genet Dev 8:103–111
Larsson J, Karlsson S (2005) The role of Smad signaling in hematopoiesis. Oncogene 24:5676–5692
Larsson J, Goumans MJ, Sjostrand LJ et al (2001) Abnormal angiogenesis but intact hematopoietic potential in TGF-β type I receptor-deficient mice. EMBO J 20:1663–1673
Letterio JJ (2005) TGF-β signaling in T cells: roles in lymphoid and epithelial neoplasia. Oncogene 24:5701–5712
Liberati NT, Datto MB, Frederick JP et al (1999) Smads bind directly to the Jun family of AP-1 transcription factors. PNAS 96:4844–4849
Lopez-Casillas F, Wrana JL, Massague J (1993) Betaglycan presents ligand to the TGF β signaling receptor. Cell 73:1435–1444
Lu G, Kang YJ, Han J et al (2006) TAB-1 modulates intracellular localization of p38 MAP kinase and downstream signaling. J Biol Chem 281(9):6087–6095
Massague J (1998) TGF-β signal transduction. Annu Rev Biochem 67:753–791
McAllister KA, Grogg KM, Johnson DW et al (1994) Endoglin, a TGF-β binding protein of endothelial cells, is the gene for hereditary haemorrhagic telangiectasia type 1. Nat Genet 8:345–351
Mishra L, Derynck R, Mishra B (2005) Transforming growth factor–β signaling in stem cells and cancer. Science 310:68–71
Mulder KM (2000) Role of Ras and Mapks in TGFβ signaling. Cytokine Growth Factor Rev 11:23–35
Nakao A, Afrakhte M, Moren A et al (1997) Identification of Smad7, a TGF beta-inducible antagonist of TGF-beta signalling. Nature 389(6651):631–635
Ohta M, Greenberger JS, Anklesaria P et al (1987) Two forms of transforming growth factor-beta distinguished by multipotentialhaematopoietic progenitor cells. Nature 329(6139):539–541
Ottmann OG, Pelus LM (1988) Differential proliferative effects of transforming growth factor betaon human hematopoietic progenitor cells. J Immunol 140(8):2661–2665
Park SM, Deering RP, Lu Y et al (2014) Musashi-2 controls cell fate, lineage bias, and TGF-β signaling in HSCs. J Exp Med 211(1):71–87
Pierelli L, Marone M, Bonanno G et al (2002) Transforming growth factor-beta 1 causes transcriptional activation of CD34 and preserves haematopoietic stem/progenitor cell activity. Br J Haematol 118:627–637
Qin BY, Lam SS, Correia JJ et al (2002) Smad3 allostery links TGF-β receptor kinase activation to transcriptional control. Genes Dev 16:1950–1963
Quéré R, Saint-Paul L, Carmignac V et al (2014) Tif1γ regulates the TGF-β1 receptor and promotes physiological aging of hematopoietic stem cells. PNAS 111(29):10592–10597
RafteryL A, Twombly V, Wharton K et al (1995) Genetic screens to identify elements of the decapentaplegic signaling pathway in Drosophila. Genetics 139:241–254
Ruscetti FW, Bartelmetz SH (2001) Transforming growth factor beta, pleiotropic regulator of hematopoietic stem cells: potential physiological and clinical relevance. Int J Hematol 74:18–25
Sanford LP, Ormsby I, Gittenberger-de Groot AC et al (1997) TGFβ2 knockout mice have multiple developmental defects that are nonoverlapping with other TGFβ knockout phenotypes. Development 124:2659–2670
Sankar S, Mahooti-Brooks N, Centrella M et al (1995) Expression of transforming growth factor type III receptor in vascular endothelial cells increases their responsiveness to transforming growth factor β2. J Biol Chem 270:13567–13572
Scandura JM, Boccuni P, Massague J et al (2004) Transforming growth factor β-induced cell cycle arrest of human hematopoietic cells requires p57KIP2 up-regulation. PNAS 101(42):15231–15236
Schmid P, Cox D, Bilbe G et al (1991) Differential expression of TGF-β1, β2 and β3 genes during mouse embryogenesis. Development 111:117–130
Sekelsky JJ, Newfeld SJ, Raftery LA et al (1995) Genetic characterization and cloning of mothers against dpp, a gene required for decapentaplegic function in Drosophila melanogaster. Genetics 139:1347–1358
Shi Y, Massague J (2003) Mechanisms of TGF-beta signaling from cell membrane to the nucleus. Cell 113(6):685–700
Sing GK, Keller JR, Ellingsworth LR et al (1988) Transforming growth factor beta selectively inhibits normal and leukemic human bone marrow cell growth in vitro. Blood 72(5):1504–1511
Sirard C, de la Pompa JL, Elia A et al (1998) The tumor suppressor gene Smad4/Dpc4 is required for gastrulation and later for anterior development of the mouse embryo. Genes Dev 12:107–119
Sirard C, Kim S, Mirtsos C et al (2000) Targeted disruption in murine cells reveals variable requirement for Smad4 in transforming growth factor β-related signaling. J Biol Chem 275:2063–2070
Sitnicka E, Ruscetti FW, Priestley GV et al (1996) Transforming growth factor beta 1 directly and reversibly inhibits the initial cell divisions of long-term repopulating hematopoietic stem cells. Blood 88(1):82–88
Söderberg SS, Karlsson G, Karlsson S (2009) Complex and content dependent regulation of hematopoiesis by TGF-beta superfamily signaling. Ann N Y Acad Sci 1176:55–69
Sporn MB, Roberts AB, Wakefield LM et al (1987) Some recent advances in the chemistry and biology of transforming gowth factor-beta. J Cell Biol 105:1039–1045
Stenvers KL, Tursky ML, Harder KW et al (2003) Heart and liver defects and reduced transforming growth factor β2 sensitivity in transforming growth factor β type III receptor deficient embryos. Mol Cell Biol 23:4371–4385
Stuhlmeier KM, Pollaschek C (2004) Differential effect of transforming growth factor β (TGF-β) on the genes encoding hyaluronan synthases and utilization of the p38 MAPK pathway in TGF-β-induced hyaluronan synthase 1 activation. J Biol Chem 279(10):8753–8760
Takaesu G, Ninomiya-Tsuji J, Kishida S et al (2001) Interleukin-1 (IL-1) receptor-associated kinase leads to activation of TAK1 by inducing TAB 2 translocation in the IL-1 signaling pathway. Mol Cell Biol 21(7):2475–2484
Takaku K, Miyoshi H, Matsunaga A et al (1999) Gastric and duodenal polyps in Smad4 (Dpc4) knockout mice. Cancer Res 59:6113–6117
Ten Dijke P, Hill CS (2004) New insights into TGF-beta-Smadsignalling. Trends BiochemSci 29(5):265–273
Tsukazaki T, Chiang TA, Davison AF et al (1998) SARA, a FYVE domain protein that recruits Smad2 to the TGFβ receptor. Cell 95:779–791
Utsugisawa T, Moody JL, Aspling M et al (2006) A road map toward defining the role of Smad signaling in hematopoietic stem cells. Stem Cells 24:1128–1136
Veiby OP, Jacobsen FW, Cui L et al (1996) The flt3 ligand promotes the survival of primitive hemopoietic progenitor cells with myeloid as well as B lymphoid potential. Suppression of apoptosis and counteraction by TNF-alpha and TGF-beta. J Immunol 157(7):2953–2960
Wada T, Penninger JM (2004) Mitogen-activated protein kinases in apoptosis regulation. Oncogene 23:2838–2849
Walsh S, Jefferiss C, Stewart K et al (2003) TGFbeta1 limits the expansion of the osteoprogenitor fraction in cultures of human bone marrow stromal cells. Cell Tissue Res 311:187–198
Wang L, Kwak JH, Kim SI et al (2004) Transforming growth factor-β1 stimulates vascular endothelial growth factor 164 via mitogen-activated protein kinase kinase 3-p38α and p38δ mitogen-activated protein kinase-dependent pathway in murine mesangial cells. J BiolChem 279:33213–33219
Wrana JL (2009) The secret life of Smad4. Cell 136:13–14
Wu G, Chen YG, Ozdamar B et al (2000) Structural basis of Smad2 recognition by the Smad anchor for receptor activation. Science 287:92–97
Wu JW, Fairman R, Penry J et al (2001) Formation of a stable heterodimer between Smad2 and Smad4. J Biol Chem 276:20688–20694
Xu X, Brodie SG, Yang X et al (2000) Haploid loss of the tumor suppressor Smad4/Dpc4 initiates gastric polyposis and cancer in mice. Oncogene 19(15):1868–1874
Xu L, Kang Y, Col S et al (2002) Smad2 nucleocytoplasmic shuttling by nucleoporins CAN/Nup214 and Nup153 feeds TGFbeta signaling complexes in the cytoplasm and nucleus. Mol Cell 10(2):271–282
Yamazaki S, Iwama A, Takayanagi S et al (2009) TGF-β as a candidate bone marrow niche signal to induce hematopoietic stem cell hibernation. Blood 113(6):1250–1256
Yang X, Li C, Xu X et al (1998) The tumor suppressor SMAD4/DPC4 is essential for epiblast proliferation and mesoderm induction in mice. PNAS 95:3667–3672
Zhang Y, Feng XH, Derynck R (1998) Smad3 and Smad4 cooperate with c-Jun/c-Fos to mediate TGF-β-induced transcription. Nature 394:909–913
Zhang L, Wang W, Hayashi Y et al (2003) A role for MEK kinase 1 in TGF-β/activin-induced epithelium movement and embryonic eyelid closure. EMBO J 22:4443–4454
Conflict of interest
The authors declare that they have no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Vaidya, A., Kale, V.P. TGF-β signaling and its role in the regulation of hematopoietic stem cells. Syst Synth Biol 9, 1–10 (2015). https://doi.org/10.1007/s11693-015-9161-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11693-015-9161-2