Abstract
The natural selection of metabolism and mass can explain inter-specific body mass allometries from prokaryotes to mammals (Witting in Theor Popul Biol 117:23–42, https://doi.org/10.1016/j.tpb.2017.08.005, 2017a), with allometric exponents that depend on the selected metabolism and the packing of home ranges in predominately one (1D), two (2D), or three (3D) spatial dimensions. The predicted exponent for total metabolism for a 2D packing of home ranges increases from 3/4 to 7/4 when the fraction of the inter-specific body mass variation that follows from the natural selection of metabolism increases from zero to one. While a 3/4 exponent is commonly observed for inter-specific comparisons in mammals, a 7/4 exponent has so far not been reported. Yet, I detect the full range of exponents for evolution over time in the fossil record. There are no fossil data for allometric correlations between metabolism and mass, but I show that the allometry \( \left( {\dot{w} \propto w^{{\hat{\dot{w}}}} } \right) \) for the rate of evolution \( \left( {\dot{w} = {\text{ d }}w/{\text{ d }}t} \right) \) in mass (w) in physical time (t) is given by the underlying set of allometries for life history parameters, including mass-specific metabolism. The \(\hat{\dot{w}}\) exponent describes the curvature of body mass evolution in time, with predicted values including: 3/2 (2D) for within niche evolution in small horses over 54 million years. 5/4 (2D) and 9/8 (3D) for across niche evolution of maximum mass in four mammalian clades. 3/4 (2D) for fast evolution in large horses, and maximum mass in trunked and terrestrial mammals. And 1 for maximum mass across major lifeforms during 3.5 billion years of evolution along a metabolic bound. These results integrate the inter-specific allometries of existing species with a deeper understanding of their natural selection during evolutionary diversification over millions of years.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Body mass allometries are essential for our understanding of natural selection. They reveal how the life histories of biological organisms evolve in correlations with mass, providing a fingerprint of the underlying natural selection cause. Allometries are usually analysed in inter-specific comparisons, but another essential question is how they evolve over time in an evolutionary lineage that increases in size. I address the latter by developing theory for the natural selection of allometries in time, illustrating how to estimate the values of the selected exponents by the curvature of an increasing body mass trajectory. Given the prediction, I analyse fossil data in an attempt to identify the natural selection cause for some of the best-documented evolutionary trajectories in the fossil record.
One attempt to identify the cause for allometries explains metabolic scaling as a physiological adaptation where branching networks are optimised to supply the organism with energy for metabolism (West et al. 1997, 1999a, b; Banavar et al. 1999; Dodds et al. 2001; Dreyer and Puzio 2001; Rau 2002; Santillán 2003; Glazier 2010). This view is elaborated in the Metabolic theory of ecology, where physical and kinetic constraints on metabolism are influencing ecological processes like the rate of feeding and interaction (Gillooly et al. 2002; Brown et al. 2004; Sibly et al. 2012; Humphries and McCann 2014).
Where physical and biochemical laws may shape the ecology and physiology of biological organisms, the cause—for the evolution of large organisms with a metabolism that depends on mass—is the natural selection of metabolism and mass. It is, however, not self-evident how natural selection is able to select energy into mass at the cost of fast replication, and nor is it self-evident why metabolism is selected as an apparent function of mass.
Metabolic theory recognises the natural selection of mass in separate models on life history evolution (Brown and Sibly 2006; Bueno and López-Urrutia 2012). These models use an adaptive fitness optimisation like a multitude of other studies that argue for a variety of intrinsic and ecological causes for the evolution of mass (e.g., McLaren 1966; Schoener 1969; Stanley 1973; Roff 1981, 1986; Stearns and Crandall 1981; Stearns and Koella 1986; Gould 1988; Maurer et al. 1992; Charlesworth 1994; Bonner 2006; Caluset and Erwin 2008; Smith et al. 2010; Charnov 2011; DeLong 2012; Shoemaker and Caluset 2014; Baker et al. 2015). Basically all these hypotheses assume constant relative fitnesses, which implies a frequency-independent selection that operates by an increase in the rate of population dynamic growth (r) and/or carrying capacity (k). Fisher (1930) used this increase to formulate his fundamental theorem of natural selection (Witting 2000), a theorem that became the cornerstone of r/k selection theory (Anderson 1971; Charlesworth 1971; Roughgarden 1971; Clarke 1972). But body masses that are selected by an increase in r and/or k does not produce the observed inter-specific allometries, where an increase in mass is associated with a decline—and not an increase—in r and k (Fenchel 1974; Damuth 1981, 1987).
A frequency-dependent selection of mass, on the other hand, is able to reconcile an inter-specific decline in r and k with an increase in mass (Simpson 1953; Dawkins and Krebs 1979; Parker 1979, 1983; Haigh and Rose 1980; Maynard Smith and Brown 1986; Vermeij 1987; Witting 2000). The theory of Malthusian relativity (Witting 1995, 1997, 2008, 2017a, b) uses a density-frequency-dependent selection of mass and metabolism to explain the evolution of large body masses, including the values of allometric exponents for fitness related traits like net assimilated energy (\({\epsilon }\)), mass-specific metabolism (β), life-periods (t), reproduction (R), survival (p), population growth (r), abundance (n), and home range (h).
Fingerprint of Metabolic Selection
Malthusian relativity is an energy-based theory on the natural selection that follows from population growth and the associated levels of intra-specific interactive competition. It defines body mass by the net assimilated energy that is allocated into size instead of being used in metabolism or allocated to the replication of offspring. Mass is measured by the energy that is released from combustion, with an approximate proportional conversion between mass in joules and mass in grams.
Energetic trade-offs select for fast replication at the cost of body mass, yet Malthusian relativity finds that this frequency-independent selection for the absence of mass is counterbalanced by the density-frequency-dependence of interactive foraging that generates a population dynamic feedback selection for a non-negligible body mass, as illustrated in Fig. 1. The feedback selects net assimilated energy into mass by a density-dependent resource bias where the larger than average individuals monopolise resources in interactive competition.
Population dynamic feedback selection. A diagram of the population dynamic feedback selection of Malthusian relativity, with symbols that relate to the population average, and coloured circles that symbolize individual home ranges in two-dimensional space with interactive competition in zones of overlap. The winners (dominating colour) of interactive competition monopolize resources, and this generates a body mass biased resource access that is proportional to the slope of the multi-coloured bar in centrum, with the invariant interference (\({\iota }\)) of the selection attractor (**) determining the selection of this bias. The primary selection on resource handling (α) and mass-specific metabolism (β) generates an exponential increase in the net assimilated energy (\({\epsilon }\)), and this maintains relatively high population dynamic growth (r) and a continued feedback selection for an exponential increase in mass (w). The feedback attractor is illustrated by the outer ring of symbols [r: population growth \(\rightarrow \gamma \): density regulation \(\rightarrow n^*\): population abundance \(\rightarrow {\iota }\): interference level \(\rightarrow w\): selection on body mass \(\rightarrow r\): population growth]. Selection for a change in mass initiates the inner loop of mass-rescaling selection [w: mass \(\rightarrow \)\(t_j\): juvenile period \(\rightarrow \beta \): metabolic rate \(\rightarrow t_r\): reproductive period \(\rightarrow h\): home range \(\rightarrow {\iota }\): interference]. Modified from Witting (2017b) (Color figure online)
The ecological constraints—on the population-wide geometrical packing of foraging in home ranges and territories—are then selecting the allometric exponents between body mass and other fitness related traits (Witting 1995, 2017a). The ecological constraints select the numerical values of the allometric exponents through a mass-rescaling selection where a decline in mass-specific metabolism dilates the time-scale of natural selection during the selection of net energy into mass (Witting 2017a). This dilation of natural selection time is necessary to maintain the net energy of the average individuals during the selection of mass, i.e., necessary to maintain the energy that defines the population dynamic growth that maintains the level of interactive competition that selects net energy into mass.
The mass-rescaling selection is illustrated in Fig. 1 by the inner loop of symbols. The result is a set \(\mathbf{x}=\{ {\epsilon }, \beta , t, R, p, r, n, h \}\) of traits that evolve partial allometric correlations \(x_{w} = w^{\hat{x}_{w}}\) with mass (w), where \(x_{w} \in \mathbf{x}\) with allometric exponents \(\hat{x}_{w} \in \hat{\mathbf{x}}_{w}= \{ \hat{{\epsilon }}_{w}, \hat{\beta }_{w}, \hat{t}_{w}, \hat{R}_{w}, \hat{p}_{w}, \hat{r}_{w}, \hat{n}_{w}, \hat{h}_{w} \}\), with subscript w denoting mass-rescaling selection. When the population ecological constraints are formulated in mathematical equations, we can express the different traits by their allometric functions of mass, and solve the equation system for the unknown values of the exponents \(\hat{\mathbf{x}}_{w}\). This solution (Witting 1995) includes the well-known 1/4 and 3/4 exponents of Kleiber (1932) scaling from a two-dimensional (2D) packing of the foraging pattern, with alternative 1/6 and 5/6 exponents being selected by a three-dimensional (3D) packing.
The natural selection of allometries, however, is more complex because metabolism is selected not only by the allometric mass-rescaling selection, but also by a selection where an increase in mass-specific metabolism generates some of the net energy for the organism (Witting 2017a, b). This is because the net assimilated energy (\({\epsilon }\), SI unit J/s) is a product (\({\epsilon }={\alpha }\tilde{\beta }\)) between the mechanical/biochemical handling of resource assimilation (\({\alpha }= \grave{{\alpha }}\rho ^{**}\); resource handling in short; SI unit J; \(\grave{{\alpha }}\): intrinsic handling; \(\rho ^{**}\): resource density at population equilibrium), and the pace of handling (\(\tilde{\beta }=\beta /{W}\); SI unit 1/s), with pace being selected as a proxy for the rate of mass-specific metabolism (β; SI unit J/Js), with W (SI unit J/J) defined as the mass-specific work of handling from one joule metabolised per unit body mass (Witting 2017a).
The selection of metabolism by the pace of handling is likely the primary most basic form of natural selection on metabolism, with a direct connection to fitness by the net assimilated energy that it generates. The decline in mass-specific metabolism by mass-rescaling is instead a secondary selection response to the natural selection of mass. Both changes in metabolism impose a rescaling on the rate dependent traits; yet, the fraction of this rescaling that follows from the selection increase in mass is part of mass-rescaling, while the fraction that follows from the primary selection of metabolism is named metabolic-rescaling of rate dependent traits (Witting 2017a).
Metabolic-rescaling is transformed into partial correlations of metabolic-rescaling allometries (\(x_{\beta } = w^{\hat{x}_{\beta }}\), subscript β denotes metabolic-rescaling), i.e., the traits correlations with mass that evolve from metabolic-rescaling and the mass that is selected from the net energy that follows from the primary selection of mass-specific metabolism. The final allometries \(x=w^{\hat{x}}\) are products \(x=x_{\beta }x_{w}\) of the two partials of mass-rescaling and metabolic-rescaling selection. This implies exponents \(\hat{x}=\hat{x}_{\beta }+\hat{x}_{w}\) that depend on the relative importance of mass-specific metabolism for the evolution of net energy and mass, as described by a metabolic-rescaling exponent for mass-specific metabolism (\(\hat{\beta }_{\beta }\)) that varies from zero to one (Witting 2017a); with an associated final 2D exponent for mass-specific metabolism that varies from − 1/4 to 3/4.
Instead of focussing on the exponent of the final allometry, i.e., \(\hat{x}\) in \(x=x_{\beta }x_{w}=w^{\hat{x}}\), we may interpret the metabolic-rescaling component (\(x_{\beta } = w^{\hat{x}_{\beta }}\)) as the intercept of the mass-rescaling allometry (\(x_{w} = w^{\hat{x}_{w}}\)). A well-known case of this interpretation is the difference in the intercepts of the allometries for ectotherm and endotherm vertebrates. Both taxa have Kleiber-like mass-rescaling exponents across the species within each taxon, yet the allometric intercept for mass-specific metabolism is higher in endotherms. The higher intercept implies a higher mass-specific metabolism for similar sized species, reflecting that metabolism is more essential for resource consumption in endotherms.
For vertebrates with a similar mass-specific metabolism, endotherms have a larger downscaling of metabolism by a stronger mass-rescaling component, i.e., a larger mass as the mass-rescaling exponents are about the same. This illustrates that endotherms tend to have larger body masses than ectotherms because they generate more net energy from a higher rate of resource handling.
The primary selection of metabolism can explain a wide range of allometries across lifeforms from virus over prokaryotes and larger unicells to multicellular animals with sexual reproduction. The exponents of empirical allometries have been found to change in transitions across the major taxa (e.g., Makarieva et al. 2005, 2008; DeLong et al. 2010), and Witting (2017a, b) was able to shows how a directional decline—in the importance of metabolism for the selection of mass—can explain not only the evolution of the different allometric exponents, but also the evolutionary transitions between the lifeforms of the major taxa.
Allometries in Time
Allometries are usually used to describe relationships across existing species, and the study of Witting (2017a, b) is no exception. However, with this paper, I extend the analysis to describe the natural selection of metabolism, mass, and allometries in time.
The population dynamic feedback selection of mass may stabilise at different selection attractors that depend on the underlying selection of net energy (Witting 1997, 2018). The attractor is an evolutionary equilibrium with a stable mass when there are no evolutionary changes in net energy. Yet, for unconstrained selection in a stable environment there is selection for an exponential increase in net energy, with the attractor being an evolutionary steady state where mass is selected to increase exponentially on the per-generation time-scale of natural selection (Witting 1997, 2003, 2018). Hence, we may express the rate of change in mass as an allometric function of mass
with an exponent of unity and a log-linear increase on the per-generation time-scale (\(\tau \)) of natural selection. It is this base-case expectation for unconstrained exponential evolution in a stable environment that I use to analyse the selection of metabolism, mass, and allometries in time.
In across species comparisons, it is the amount of inter-specific variation in resource handling relative to the amount of variation in the primary selected component of mass-specific metabolism that determines the values of the selected allometric exponents (Witting 2017a, b). Yet, when we study the allometries of a body mass that is selected to increase over time, it follows that the allometric exponents that are selected at any given time will depend on the relative rates of evolution in the different traits. At the evolutionary steady state, the selected increase in mass \( \left( {r_{w} = {\text{ d }}\ln w/{\text{ d }}\tau } \right) \) is given by the selected increase in net energy \( \left( {r_{\epsilon } = {\text{d }}\ln \epsilon /{\text{d}}\tau } \right) \), with the latter being dependent on the selected increase in resource handling (\(r_{{\alpha }}\)) and mass-specific metabolism (\(r_{\beta _{\beta }}\), with sub-subscript \(\beta \) denoting the primary selected component of metabolism). This implies that the selected allometric exponents will dependent on the relative importance of metabolism for the selection of mass, as expressed e.g. by the \(r_{\beta _{\beta }}/r_{{\alpha }}\)-ratio.
We may thus use the allometric exponents of an evolutionary body mass trajectory in time to determine the \(r_{\beta _{\beta }}/r_{{\alpha }}\)-ratio of its underlying natural selection cause. However, to apply this method to fossils we need to solve the problem that the only life history trait that is commonly estimated for fossil animals is mass. How can we estimate the exponents of the underlying allometries, when there are no estimated traits to correlate with the observed evolutionary changes in mass?
I show that we can use the curvature in the rate of evolutionary change in mass in physical time to estimate the underlying allometries and thus the importance of mass-specific metabolism for the natural selection of mass. When the allometry for the rate of change in mass is determined empirically from fossils it is expressed in physical time. Hence, with \(\tau \) being time in generations, t being time in years, and \(t_g\propto w^{\hat{t}}\) being one generation in years at generation \(\tau \), we have
and a rate of change in mass in physical time
that depends on the allometric exponent for the evolution of generation time with mass (\(\hat{t}\)), with \(\hat{\dot{w}}= 1 - \hat{t}\) being a bending exponent that describes the curvature in the rate of change in mass in physical time.
The log-linear trajectory of Eq. 1 will thus bend into a curve in physical time (\(\hat{\dot{w}}\ne 1\)) whenever \(\hat{t} \ne 0\), i.e., whenever the time-scale of natural selection evolves with the evolutionary changes in mass (Witting 1997; Okie et al. 2013). When natural selection time dilates by a generation time that increases with mass (\(\hat{t}>0 \Rightarrow \hat{\dot{w}}<1\)) it bends body mass evolution downwards in a concave trajectory with a continuously declining \(\text{ d }\ln w / \text{ d }t\) derivative. Upward bending, with a convex trajectory and a continuously increasing derivative, occurs when the time-scale of natural selection contracts from a generation time that declines with mass (\(\hat{t}<0 \Rightarrow \hat{\dot{w}}>1\)).
I show theoretically that this bending—of a log-linear body mass trajectory on the time-scale of natural selection into a curved trajectory in physical time—is a fingerprint of the underlying primary selection of mass-specific metabolism. With an overall allometric scaling that depends on metabolic-rescaling, I find the curvature of body mass evolution to depend on the primary selected mass-specific metabolism, with the equations of natural selection predicting the \(\hat{\dot{w}}\) exponent of Eq. 3 as a continuous function of the \(r_{\beta _{\beta }}/r_{{\alpha }}\)-ratio.
In applying the theory to fossil data, I focus on four distinct types of natural selection that I identify by first-principle arguments to specific trajectories in the fossil record. Each of the four selections has its own theoretical \(r_{\beta _{\beta }}/r_{{\alpha }}\)-ratio and \(\hat{\dot{w}}\) exponents. The intriguing question is then whether the observed exponents falsify the theory, or coincide with the first-principle predictions as the latter may reveal natural selection causes behind some of the best-documented evolutionary trajectories in the fossil record.
The four selection types assume constant \(r_{\beta _{\beta }}/r_{{\alpha }}\)-ratios and selection at the evolutionary steady state with an exponentially increasing mass on the per-generation time-scale. We may generally not expect this level of idealised conditions for evolutionary lineages over timespans that are sufficiently long to allow for the detection of an exponential trajectory, especially not when data are sampled from a fossil record that is incomplete when it comes to the documentation of trajectories for individual evolutionary lineages. Yet, the predicted values of the \(\hat{\dot{w}}\) exponent pertain to these idealised conditions, and it is essential to select carefully among the fossil data in order to avoid a comparison of apples and oranges.
Methods
Theory
I extend the selection equations of Malthusian relativity to predict the bending exponent \(\hat{\dot{w}}\) for body mass evolution. The original deduction of the evolutionary steady state (Witting 1997) did not partition the selection of net energy into the selection of the handling of resource assimilation and the selection of the pace of this process, as represented by mass-specific metabolism. But, with allometries that depend in part on the primary selected metabolism (Witting 2017a), a distinction is necessary to predict the bending exponents of evolutionary trajectories.
Appendix 1 develops the exponential increase of the evolutionary steady state with primary selection on handling and mass-specific metabolism (model symbols in Table 1). The resulting equations are combined with the results from Witting (2017a) and solved in Appendix 2 to obtain the allometric exponents as a function of the \(r_{\beta _{\beta }}/r_{{\alpha }}\)-ratio and the spatial dimensionality of the foraging pattern. The theoretical exponent for generation time (\(\hat{t}\)) is then inserted into Eq. 3 to calculate the bending exponent (\(\hat{\dot{w}}\)) for body mass evolution in physical time.
The first of the four selection types that I relate to data is an unconstrained symmetrical selection across ecological niches. It has similar levels of heritable variation in resource handling and mass-specific metabolism, generating similar rates of increase in the two parameters with a \(r_{\beta _{\beta }}/r_{{\alpha }}\)-ratio of unity. Selection across niches may also cause a second non-symmetrical type of selection that is characterised by a \(r_{\beta _{\beta }}/r_{{\alpha }}\)-ratio that approaches zero. The latter may occur when fast improvements in resource handling are easy, and the maximum size is increasing fast by an increase in resource handling that outruns the background evolution of mass-specific metabolism. The underlying cause may be a resource density that increases across niches, or a resource handling efficiency that increases as a mechanistic function of the evolutionary increase in mass.
These across niche selections are likely to occur in the larger species of clades that diversifies in evolutionary radiations. Not only are the larger species likely to evolve from adaptations to ecological niches that allow for a larger resource consumption, but nor should their niche access be limited by competitively superior species. This contrasts to smaller species that may have their niche access and resource handling constrained by the inter-specific dominance of larger species. The trajectory for the maximum size of a clade is the obvious place to look for an unconstrained evolution of mass.
The third selection relates to the alternative scenario where a lineage evolves within a relatively stable ecological niche. This results in an adaptation that selects resource handling to an evolutionary optimum where \(r_{{\alpha }}\rightarrow 0\) and the \(r_{\beta _{\beta }}/r_{{\alpha }}\)-ratio approached infinity.
The final selection that I examine relates to macroevolution on the time-scale of major transitions; first from prokaryotes to unicellular eukaryotes, then by the multilevel selection transition to multicellular animals with sexual reproduction (Witting 2002, 2017b), and finally between the major lifeforms of multicellular animals that have taken the record of maximum size over time. Each of these transitions require a major reorganisation of the biochemical/mechanical/ecological mechanisms of resource handling (α). The rate of evolution in the macroevolutionary component of α and mass should thus be much smaller than the rates that are realised when a given lifeform adapts, by relatively simple phenotypic adjustments, to resources across ecological niches.
We may thus expect that the rate of macroevolutionary increase in resource handling will be smaller than the persistent primary selected increase in mass-specific metabolism (\(r_{{\alpha }}\ll r_{\beta _{\beta }}\)), so that the primary selected increase in metabolism should be able to outrun the decline from mass-rescaling on the macro evolutionary time-scale. Macroevolution should thus take mass-specific metabolism to an upper limit, as measured in the species with the highest metabolism, i.e., creating an invariant mass-specific metabolism across the smallest species of each lifeform. The result is macroevolution along a metabolic bound where the primary selected increase in metabolism is exactly outbalancing the mass-rescaling decline, i.e., where the product of metabolic-rescaling and mass-rescaling is zero \(\hat{\beta }_{\beta }+\hat{\beta }_{w}=0\).
The four selection types have specific \(r_{\beta _{\beta }}/r_{{\alpha }}\)-ratios; the ratios, however, may not be constant on a microevolutionary time-scale. The realised rates of increase in α and β will depend on available mutations, and these may not be homogeneous in time. Yet, when integrated over longer time-scales, we may expect the \(r_{\beta _{\beta }}/r_{{\alpha }}\)-ratio to stabilise at central values that reflect the underlying generation of heritable variance.
A central value, however, will also depend on fluctuations in the selection gradients, as caused by variation in climate, resource availability, and inter-specific competition (Witting 1997, 2008, 2018). The overall selection for a size increase may shift to a decline if the net assimilated energy is declining due to an environmental or inter-specific competitive crisis. The selection trend for an increase in size—and the associated bending exponents—will thus be somewhat hidden in a diverse and complex species distribution of step-wise increasing and declining body masses.
Data
The expectation of a complex size distribution on top of a trend for a general increase agrees with mammalian data for the past 100 million years of evolution. They show a widespread increase in size (Alroy 1998; Baker et al. 2015) in agreement with Cope’s (1887) rule. Around 70% of descendant species are larger than their ancestors (Baker et al. 2015), about 30% are smaller, and the distribution of long-term trajectories is far from a homogeneous exponential-like increase in mass.
While selection at the evolutionary steady state has been used to simulated the evolution of mammalian species distributions over the past 65 million years (Witting 2018), my focus is on the identification of specific body mass trajectories that approximate the four modes of selection over a timespan that allows for the estimation of the bending exponent \(\hat{\dot{w}}\). Hence, I aim to identify cases where the fossil record show a relatively homogeneous increase in size over timespans that are so long that the general increase is evident in relation to the variation of the data. I am focusing only on a few long-term trajectories that all have a relatively smooth exponential-like increase in size, as predicted by the evolutionary steady state.
The maximum size of a clade is often found to increase in a near-exponential manner during periods of phylogenetic radiation (e.g., Hayami 1978; Trammer 2005; Smith et al. 2010; Okie et al. 2013). Hence, I use the data from Smith et al. (2010) and Okie et al. (2013) on the evolution of maximum mass over millions of years in mammals and mammalian clades to resemble evolution across ecological niches. These trajectories originate partially by speciation during evolutionary radiation, and they do generally not represent the continued evolution of a single lineage. The trajectories reflect the overall bounds on the evolution of resource handling given an average rate of evolution in mass-specific metabolism across the larger species in a clade. The macroevolutionary estimates of the bending exponent are thus integrating across species to reflect the overall trend in the evolutionary variation in mass-specific metabolism and maximum mass within a clade.
Another case for an exponential-like increase in body mass is the evolution of fossil horses (Equidae) over the past 57 million years in North America (MacFadden 1986; Witting 1997; Shoemaker and Caluset 2014). During the Eocene and Oligocene, from about 57 to 25 million years ago (ma), horses were browsers with a relatively small increase in size (from about 25 to 50 kg) and an absence of a strong diversification of species. In the early and middle Miocene from about 25 to 10 ma horses had a major pulse of body mass evolution, with an associated strong taxonomic diversification of both browsing and grazing horses (MacFadden 1986). This lead to the evolution of large horses (up to about 1 ton for Equus giganteus around 0.012 ma, Eisenmann 2003), yet the evolution of small types continued with the 80 kg Nannippus peninsulatus around 3.5 ma being an example of the latter.
In order to capture this pattern, I divide the MacFadden (1986) data on body masses of fossil horses into small (below 100 kg) and large horses (above 100 kg) to consider their evolution separately. Small browsing horses may represent well-adapted lineages that evolve within a restricted niche space where \(r_{{\alpha }}\) is about zero and the \(r_{\beta _{\beta }}/r_{{\alpha }}\)-ratio approaches infinity. Yet, we expect a transition where the diversification of larger horses after 25 ma reflects selection across niches. This would imply a \(r_{\beta _{\beta }}/r_{{\alpha }}\)-ratio around unity, if not zero as the fast evolution of size indicates a resource handling that might outrun the background selection of mass-specific metabolism.
Finally, I use data from Bonner (1965) and Payne et al. (2009) on the evolution of maximum size in mobile organisms over 3.5 billion years, to test for a macroevolutionary invariance in mass-specific metabolism across major lifeforms. This hypothesis is also checked for a predicted invariance in the mass-specific metabolism of extant species across the major lifeforms of prokaryotes, unicellular eukaryotes, and the multicellular animals of aquatic ectotherm, terrestrial ectotherms, and terrestrial endotherms. This macroevolutionary invariance is already documented for species distributions as a whole (Makarieva et al. 2005, 2008; Kiørboe and Hirst 2014; Witting 2017b), and I use the data of Savage et al. (2004) and Makarieva et al. (2008) to select the three species with the highest mass-specific metabolism from each group to test for the existence of a mass invariance at the upper metabolic limit (Supplementary Table S7).
Having estimates of mass over time for each trajectory, I calculate rates of evolution \(\text{ d }w/\text{ d }t = (w_{t_d}-w_{t_a})/(t_{d}-t_{a})\) from adjacent data points at time \(t_a\) and \(t_d\) (Supplementary Tables S1 to S6; \(w_{t_a}\) is the ancestral, and \(w_{t_d}\) the descendant, mass. For horses, I use the ancestral-descendant species pairs that were identified by MacFadden 1986). The bending exponent (\(\hat{\dot{w}}\)) is then estimated as the slope of the linear regression of \(\ln (\text{ d }w / \text{ d }t)\) on \(\ln \bar{w}\), with \(\bar{w}=(w_{t_a}+w_{t_d})/2\).
Because the theoretical bending exponents are predicted from a selection increase in mass, I include only species comparisons where descendants are larger than ancestors in my data estimates of bending exponents. This should give me the best most unbiased estimates, as it removes cases where a climatic and/or inter-specific competitive fluctuation causes a short-term decline in the size of a taxa that is otherwise increasing steadily over time.
As the selection of data limits the number of data points for the regressions, I face a trade-off between solid regressions (from many data points) and a low number of taxa with data on one side, and more uncertain regressions (on fewer data points) and a higher number of taxa with data. I chose a balance with potentially somewhat uncertain regressions based on five or more data points, as this allowed me to obtain independent estimates for eight mammalian trajectories.
The predicted bending exponents depend not only on the \(r_{\beta _{\beta }}/r_{{\alpha }}\)-ratio, but also on the dominant dimensionality of the spatial packing of the home ranges and territories that the individuals of a species compete for. To obtain independent estimates of this dimensionality, I use the data based classification in Witting (2017a), where the dimensionality of different taxa follows from the average point estimate of the allometric exponent for mass-specific metabolism, as reported in the literature for inter-specific allometries among existing species. These estimates agree in most cases with first-principle expectations, where the individuals of many pelagic and tree living species (like cetaceans and primates) classify as 3D, with an extra vertical dimension relative to most mammals that compete for 2D home ranges only.
The spatial dimensionality of a taxon might be a decimal number between one and three, as the different species of a taxon might differ in their dimensionality, and as some species might have an intermediate home range packing that is, e.g., 2D in some habitats and 3D in others. Yet, I use the somewhat simpler discrete classification of 2D versus 3D, with 1D being a rare case that has not yet been identified in allometric correlations (Witting 2017a).
Results
Exponential Increase
Appendix 1 finds that primary selection on resource handling (α) and mass-specific metabolism (β) generates an exponential increase on the per-generation time-scale in the average resource handling, mass-specific metabolism, and net assimilated energy (\({\epsilon }\)) of the individuals in the population. The rate of exponential increase in net energy
is the sum of the rates of increase in handling (\(r_{{\alpha }}\)) and mass-specific metabolism (\(r_{\beta _{\beta }}\)).
The exponential increase in net energy generates a sustained population dynamic feedback selection. The net energy that is allocated into reproduction causes population growth and interactive competition, with the latter selecting net energy into mass
at the cost of a continued increase in the rate of reproduction, and density of, and interactive competition within, the population. The result is an evolutionary steady state, where an allometric selection \({\partial }\ln w / {\partial }\ln {\epsilon }= 1/\hat{{\epsilon }}\) selects an exponential increase in mass
from the allometric exponent for net energy (\(\hat{{\epsilon }}\)) and the underlying rates of increase in resource handling and mass-specific metabolism.
This selection of mass is associated with a mass-rescaling selection (Witting 2017a) that dilates the time-scale of reproduction (\(t_r\propto w^{\hat{t}_{w}}\), with \(\hat{t}_{w}>0\)) by a partial decline in mass-specific metabolism (\(1/t_r\propto \beta \propto w^{\hat{\beta }_{w}}\), with \(\hat{\beta }_{w}<0\)). This dilation of natural selection time maintains a balance between net energy and mass that maintains the population dynamic pressure on the interactive selection of net energy into mass. A larger offspring metabolises more energy during growth, and this will cause a decline in average reproduction (and population growth and interactive competition) with an increase in mass, unless the rate of mass-specific metabolism is selected to decline and the reproductive period is selected to increase (Witting 2017a). The result is mass-rescaling allometries, where exponents like
were shown by Witting (1995) to follow from the spatial dimensionality (d) of the interactive foraging behaviour.
Allometries in Time
The partial decline in mass-specific metabolism from mass-rescaling selection occurs independently of the primary selection for an exponential increase in mass-specific metabolism. The latter generates a metabolic-rescaling of the life history, with Appendix 2 showing that the evolutionary steady state implies a metabolic-rescaling exponent for mass-specific metabolism
that depends on the \(r_{\beta _{\beta }}/r_{{\alpha }}\)-ratio and ecological dimensionality (d). Combined with the mass-rescaling exponent of Eq. 7 it defines the final allometry (\(\hat{\beta }=\hat{\beta }_{w}+\hat{\beta }_{\beta }\)) as illustrated in Fig. 2.
The allometric scaling of mass-specific metabolism (\(\beta \propto w^{\hat{\beta }} \propto w^{\hat{\beta }_{\beta }} w^{\hat{\beta }_{w}}\)) as it evolves from the \(r_{\beta _{\beta }}/r_{{\alpha }}\)-ratio given 2D and 3D interactions. The blue curves are the exponents of metabolic-rescaling (\(\hat{\beta }_{\beta }\)), the green lines the exponents of mass-rescaling (\(\hat{\beta }_{w}\)), and the red curves the exponents of the final allometries (\(\hat{\beta }=\hat{\beta }_{\beta }+\hat{\beta }_{w}\)) (Color figure online)
A constant mass-rescaling intercept (\(x_{\beta } = w^{\hat{x}_{\beta }}\)) with no metabolic-rescaling (\(\hat{\beta }_{\beta }=0\)) is obtained only when there is no primary evolution in mass-specific metabolism and \(r_{\beta _{\beta }}/r_{{\alpha }}= 0\). More generally, the mass-rescaling intercept will increase with an evolutionary increasing body mass, because the mass increase follows partly, or fully, from an evolutionary increase in mass-specific metabolism. For symmetrical evolution where \(r_{\beta _{\beta }}/r_{{\alpha }}= 1\), the mass-rescaling intercept will increase to the 3/8 power of body mass in 2D, and the 5/12 power in 3D. At the limit where metabolism evolves unconstrained (\(r_{\beta _{\beta }}> 0\)) with a constraint on resource handling (\(r_{{\alpha }}= 0\), with \(r_{\beta _{\beta }}/r_{{\alpha }}\approx \infty \)), we find a mass-rescaling intercept for mass-specific metabolism that scales to the 3/4 power in 2D, and the 5/6 power in 3D.
Having the metabolic-rescaling exponents for mass-specific metabolism as a function of the \(r_{\beta _{\beta }}/r_{{\alpha }}\)-ratio, it is straightforward to transform the theoretical exponents from Table 3 in Witting (2017a) into the allometric exponents of Table 2. For the body mass trajectory of an evolutionary lineage in time, it gives the allometric exponents of the different traits as a function of the \(r_{\beta _{\beta }}/r_{{\alpha }}\)-ratio and the dimensionality of the foraging behaviour.
Mass in Physical Time
With equations for the evolution of handling, mass-specific metabolism and mass on a per-generation time-scale (Eqs. 29 and 31), and the associated allometric scaling with mass (Table 2), we have a predictive theory for life history evolution on the per-generation time-scale of natural selection. This evolution depends on the per-generation rate of change in mass (\(r_{w}\) from Eq. 6), with the remaining life history being a function of the evolving mass (Table 2), with a functional relationship that follows from the \(r_{\beta _{\beta }}/r_{{\alpha }}\)-ratio and d.
To transform the predicted body mass trajectories to the physical time-scale of the fossil record, we use Eq. 3 to scale the rate of change in mass with the evolutionary changes in generation time, with \(\hat{t}=-\hat{\beta }\). This predicts a natural selection time that dilates to the 1/4 power in 2D and the 1/6 power in 3D when there is no evolutionary increase in the mass-rescaling intercept for mass-specific metabolism. Natural selection time is body mass invariant when \(r_{\beta _{\beta }}/r_{{\alpha }}=1/2(d-1)\), i.e., 1/2 in 2D and 1/4 in 3D, and it contracts to the \(-1/8\) power in 2D and the \(-1/4\) power in 3D for symmetrical unconstrained evolution where \(r_{\beta _{\beta }}/r_{{\alpha }}=1\). At the \(r_{\beta _{\beta }}/r_{{\alpha }}\approx \infty \) limit, where mass is increasing exclusively because of increased metabolism, the contraction of natural selection time occurs to the \(-1/2\) power in 2D and the \(-2/3\) power in 3D.
By scaling Eq. 3 with the predicted \(\hat{t}\) we obtain the \(\dot{w} \propto w^{\hat{\dot{w}}}\) allometry, with the following exponent
for the rate of change in mass in physical time (Table 2).
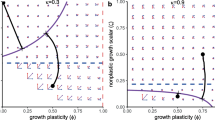
All of the predicted mass trajectories are log-linear on the per-generation time-scale of natural selection. Dependent on the \(r_{\beta _{\beta }}/r_{{\alpha }}\)-ratio and d, the corresponding trajectories in physical time are bent downward (\(\hat{\dot{w}}<1\)) by a dilation of natural selection time, or upward (\(\hat{\dot{w}}>1\)) by a contraction. This is illustrated in Fig. 3, with the evolutionary trajectory being log-linear in physical time (\(\hat{\dot{w}}=1\)) only when the \(r_{\beta _{\beta }}/r_{{\alpha }}\)-ratio is \(1/2(d-1)\) (yellow trajectory).
Evolutionary bending by contraction and dilation of natural selection time. A theoretical illustration of body mass evolution when the rate of exponential increase [\(r_{w}=(r_{{\alpha }}+r_{\beta _{\beta }})/\hat{{\epsilon }}\)] is constant on the per-generation time-scale of natural selection (2D interactions). The initial generation time is a year for all lineages, and the per-generation time-scale (right y-axis) is shown in physical time by the coloured x-axis. The evolutionary trajectories are the same for all lineages in generations, with the degree of bending in physical time following from the \(r_{\beta _{\beta }}/r_{{\alpha }}\)-ratio. There is only a downward bend from time dilation when \(r_{\beta _{\beta }}/r_{{\alpha }}=0\) (blue). Time dilation and contraction are equally strong (with no overall bend) when \(r_{\beta _{\beta }}/r_{{\alpha }}=1/2(d-1)\) (yellow). Contraction with upward bending is dominating for unconstrained symmetrical evolution (\(r_{\beta _{\beta }}/r_{{\alpha }}= 1\), green), and extreme for within niche evolution with resource handling at an evolutionary optimum (\(r_{\beta _{\beta }}/r_{{\alpha }}\approx \infty \), red) (Color figure online)
Theoretical examples for the evolution of maximum mammalian mass (w; top) and associated lifespan (t; middle) in physical time, given 2D and 3D interactions and constant rates of evolution on the per-generation time-scale. The \(r_{\beta _{\beta }}/r_{{\alpha }}\)-ratio is estimated by the slope (\(\hat{\dot{w}}\) exponent, Eq. 9) of the allometry (\(\dot{w} \propto w^{\hat{\dot{w}}}\)) for the rate of evolutionary increase in body mass (\(\dot{w}=\text{ d }w/\text{ d }t\), kg per million year) in physical time (bottom) (Color figure online)
These results are illustrated for placental mammals in Fig. 4. It shows, for illustrative purposes only, the span of potential exponential trajectories from an estimated 125 g ancestor at 65 ma (O’Leary et al. 2013) to a 10 tonnes terrestrial (2D) and 100 a tonnes pelagic (3D) mammal today. Included are also the \(\hat{\dot{w}}\) exponents for the evolutionary increase in mass (Eq. 9).
The shape of the trajectories depends on the \(r_{\beta _{\beta }}/r_{{\alpha }}\)-ratio. When the ratio is zero it follows that net energy and body mass are increasing exclusively because of improved resource handling. This implies that there is no evolutionary change in mass-specific metabolism except for the decline that follows from the rescaling of the life history in response to the evolutionary increase in mass. The log trajectory is then levelling off over time with a \(\hat{\dot{w}}\) exponent of 3/4 or 5/6, dependent upon dimensionality. At the other extreme, the \(r_{\beta _{\beta }}/r_{{\alpha }}\)-ratio is infinite, the energetic increase is exclusively due to enhanced metabolism, and the log trajectory is strongly convex with a \(\hat{\dot{w}}\) exponent of 3/2 or 5/3. In between, the trajectory is less convex when the \(r_{\beta _{\beta }}/r_{{\alpha }}\)-ratio is one, and it is linear when the ratio is 1/2 in 2D and 1/4 in 3D.
Empirical Evidence
The bending exponents and \(r_{\beta _{\beta }}/r_{{\alpha }}\)-ratios that are estimated from fossil data are shown in Figs. 5 and 6. The table in Fig. 5 list all mammalian estimates, with a colour code for the aggregation of data estimates around \(r_{\beta _{\beta }}/r_{{\alpha }}\)-ratios of infinity (red), unity (green), and zero (blue). The plots in Fig. 5 illustrate each selection type by an example, and the estimated \(r_{\beta _{\beta }}/r_{{\alpha }}\)-ratios resemble the selection expectations for the different taxa, as discussed below.
Mammalian evolution. Results for fossil horses (Equidae) and the maximum mass of other mammalian clades. Plots: Illustrative examples of theoretical (curves) and empirical (dots) body mass (w) trajectories (top). The curves are simulated from the \(r_{\beta _{\beta }}/r_{{\alpha }}\)-ratios that are calculated (by Eq. 9) from the slope (\(\hat{\dot{w}}\)-exponent and SE estimated by linear regression) of the allometry for the rate of increase in mass (\(\dot{w}=\text{ d }w/\text{ d }t\), kg per million year) in physical time (bottom plots). Table: The estimated \(r_{\beta _{\beta }}/r_{{\alpha }}\)-ratio approaches infinity for small fossil horses, indicating masses that are selected from an evolution increase in mass-specific metabolism (red). Most \(r_{\beta _{\beta }}/r_{{\alpha }}\)-ratios for maximum mass are around one, with a similar increase in mass-specific metabolism and resource handling (green). With a ratio around zero, the increase in large horses and the maximum mass for trunked (Proboscidea) and terrestrial mammals is dominated by increased resource handling/availability (blue). No taxa showed log-linear evolution along a metabolic bound. 2D-3D classification from Witting (2017a). n: Number of data points in regressions, with data from MacFadden (1986), Smith et al. (2010) and Okie et al. (2013), see Supplementary Tables S1 to S4. The trajectory plots include all data points, while the regressions are based only on cases where descendants are larger than ancestors (see methods for details) (Color figure online)
The maximum mass (length raised to third power) of mobile organisms over 3.5 billion years of evolution. The \(\text{ d }w / \text{ d }t\) exponent (\(\hat{\dot{w}}\)) with SE is estimated by linear regression on the allometry for the rate of increase in mass in physical time. Data from Bonner (1965), Supplementary Table S5
Evolution Across Niches \(\left( \frac{r_{\beta _{\beta }}}{r_{{\alpha }}}=1\right) \)
A symmetrical unconstrained selection across niches may be expected as a base case type of evolution for the maximum mass of taxonomic clades over time. This evolution should generate an average \(r_{\beta _{\beta }}/r_{{\alpha }}\)-ratio around one, with an associated \(\hat{\dot{w}}\) exponent of \((6d-3)/4d\), i.e., 9/8 (1.13) for 2D and 5/4 (1.25) for 3D. The resulting trajectories are convex in physical time due to a time contraction where natural selection time evolves as \(t_{g,\tau } \propto w_{\tau }^{(3-2d)/4d}\) (green curves in Fig. 3 and 4).
This evolution is approximated for maximum mass in four out of five mammalian clades (Fig. 5, table). 2D-like \(\hat{\dot{w}}\) exponents of 1.10 and 1.19 are observed for 64 million years of evolution in terrestrial carnivores (Carnivora/Fissipedia), and 50 million years in even-toed ungulates (Artiodactyla). 3D-like \(\hat{\dot{w}}\) exponents of 1.28 are observed for 30 million years of evolution in whales (Cetacea & Mysticeti) and 55 million years in primates (Primates).
Fast Evolution \(\left( \frac{r_{\beta _{\beta }}}{r_{{\alpha }}}=0\right) \)
An average \(r_{\beta _{\beta }}/r_{{\alpha }}\)-ratio around zero is expected for selection across niches when evolution in resource handling outruns evolution in mass-specific metabolism. This is usually associated with fast body mass evolution, making it a candidate for the evolution of the largest species during evolutionary radiations. It has a \(\hat{\dot{w}}\) exponent of \((2d-1)/2d\), and a concave trajectory in physical time due to a time dilation where the time-scale of natural selection evolves as \(t_{g,\tau } \propto w_{\tau }^{1/2d}\) (blue curves in Figs. 3 and 4).
This evolution is not observed in whales, but observed \(\hat{\dot{w}}\) exponents of 0.79 and 0.83 are only slightly larger than the predicted 0.75 for the 2D-evolution of the maximum mass of trunked mammals (Probocidae) over 49 million years, and for the maximum mass of terrestrial mammals over 100 million years of evolution (Fig. 5). With a 2D \(\hat{\dot{w}}\) exponent of 0.75, this evolution is indicated also for large horses during their evolutionary radiation over the last 25 million years.
Evolution Within Niches \(\left( \frac{r_{\beta _{\beta }}}{r_{{\alpha }}}\approx \infty \right) \)
An average \(r_{\beta _{\beta }}/r_{{\alpha }}\)-ratio that approaches infinity is expected for lineages that evolve within an ecological niche. When resource handling adapts to the optimum of the niche and \(r_{{\alpha }}\rightarrow 0\), it follows that net energy and mass can increase only by a selected increase in mass-specific metabolism, with \(r_{\beta _{\beta }}/r_{{\alpha }}\rightarrow \infty \). Such lineages will have a \(\hat{\dot{w}}\) exponent of \((2d-1)/d\), i.e., 1.5 for 2D and 1.67 for 3D. Their mass trajectories are strongly convex in physical time due to a time contraction where the time-scale of natural selection evolves as \(t_{g,\tau } \propto w_{\tau }^{(1-d)/d}\) (red curves in Fig. 3 and 4).
While the larger horses during their evolutionary radiation after 25 ma show typical across-niche selection, the evolution of smaller horses appears to be driven by within-niche selection with a \(\hat{\dot{w}}\) exponent of \(1.72 \pm 0.30\) and a \(r_{\beta _{\beta }}/r_{{\alpha }}\)-ratio around infinity (Fig. 5).
Evolution at Metabolic Limit \(\left( \frac{r_{\beta _{\beta }}}{r_{{\alpha }}}=\frac{1}{2d-2}\right) \)
When mass-specific metabolism is selected along an upper bound we have that \(\hat{\beta }_{\beta }=-\hat{\beta }_{w}=1/2d\), because the upward bend from the time contraction of metabolic acceleration is balanced against the downward bend from the time dilation of mass-rescaling. The result is an invariant mass-specific metabolism over time (\(\hat{\beta }=0\)), and a log-linear trajectory where \(\hat{\dot{w}}=1\) (yellow curves in Figs. 3 and 4). This evolution has a \(r_{\beta _{\beta }}/r_{{\alpha }}\)-ratio of \(1/2(d-1)\), i.e., 1/2 for 2D and 1/4 for 3D.
The increase in maximum mass across the major life history transitions of non-sessile organisms over 3.5 billion years of evolution was found to have a \(\hat{\dot{w}}\) exponent of \(1.07 \pm 0.02\) (Fig. 6) given data from Bonner (1965), and \(1.07 \pm 0.06\) across selected maximum masses (Supplementary Table S6) from Payne et al. (2009). This apparent log-linear evolution on the largest macroevolutionary scale is supported by a body mass invariant mass-specific metabolism across major lifeforms from prokaryotes to mammals (Makarieva et al. 2005, 2008; Kiørboe and Hirst 2014; Witting 2017b). The latter invariance is observed not only across species distributions in general, but also for the maximum observed mass-specific metabolism across prokaryotes, unicellular eukaryotes, aquatic multicellular ectotherms, terrestrial ectotherms, and terrestrial endotherms (linear regressions on double logarithmic scale estimate slopes between \(-0.02 \pm 0.01\) and \(0.01 \pm 0.02\), Supplementary Table S7).
Overall Comparison
All of the nine data estimates of the \(r_{\beta _{\beta }}/r_{{\alpha }}\)-ratio make sense in relation to the selection types that can be expected for the different taxa; with a pairwise comparison between the nine predicted and observed bending exponents (\(\hat{\dot{w}}\)) having a correlation coefficient of 0.98 (\(p=0.000004\); slope of linear regression 1.13 with SE = 0.09).
Some of the taxa, however, may also have other expectations, as it is somewhat difficult to draw a clear line between an expected maximum mass evolution with a \(r_{\beta _{\beta }}/r_{{\alpha }}\)-ratio of zero and unity. One alternative theoretical expectation would be that a fast evolution of mass, with a \(r_{\beta _{\beta }}/r_{{\alpha }}\)-ratio of zero, should apply only to maximum mass across all terrestrial mammals. This would imply a misclassification of large horses and trunked mammals, and a correlation coefficient of 0.77 (\(p=0.02\); slope of linear regression 1.17 with SE = 0.36) between the expected and observed bending exponents.
Another alternative would be that both trunked mammals and whales have fast body mass evolution just as all terrestrial mammals. This would imply a misclassification of large horses and whales, and a correlation coefficient of 0.72 (\(p=0.03\); slope of linear regression 0.88 with SE = 0.33) between the expected and observed bending exponents. In either case, there is a significant correlation between the predicted and observed bending exponents.
Relating to the connection between the dimensionality of the foraging behaviour and the bending exponent, it is only the set of empirical cases with an estimated \(r_{\beta _{\beta }}/r_{{\alpha }}\)-ratio around unity that contains taxa with 3D foraging. This set has two 3D cases and two 2D cases, and a correlation coefficient of 0.9 (\(p=0.09\); slope of linear regression 1.09 with SE = 0.36) between the predicted and observed bending exponents.
Discussion
Figure 6 plots the maximum limit on body mass evolution from the origin of the first self-replicating cells to whales. The theory of Malthusian relativity covers this huge scale of evolution. Starting from inert replicators with no metabolism, the theory predicts not only the increase in mass and metabolism, but also the evolutionary succession of lifeforms from a gradual unfolding of population dynamic feedback selection with a directional decline in the importance of metabolism for the natural selection of mass (Witting 2002, 2017a, b). This prediction includes the evolution of sexually reproducing multicellular animals by a complete multilevel population dynamic feedback selection as the fourth lifeform in the succession of major transitions. Multicellular animals are selected from larger unicells with an incomplete feedback selection, larger unicells are selected from small prokaryote-like self-replicating cells with no feedback selection, and these are selected from inert replicators with no metabolism, no cell, and practically no mass by an initial mass-dependence of mass-specific metabolism.
Evolution of Size Distributions
The main-focus of this paper is not on the major transitions, but on the underlying body mass trajectories that follow from an associated selection increase in net energy. Fossil data, on e.g. mammals (Alroy 1998; Baker et al. 2015), show a widespread increase in size in agreement with Cope’s (1887) rule. Yet the distribution of long-term trajectories is far from a homogeneous exponential-like increase in mass, and this has generated the hypothesis that an increase or decrease in mass is about equally likely, with both options having a multitude of potential causes (e.g., Schoener 1969; Stanley 1973; Gould 1988; Maurer et al. 1992; Bonner 2006; Brown and Sibly 2006; Caluset and Erwin 2008; Smith et al. 2010; Shoemaker and Caluset 2014). The selection of size has been argued to be so mechanistically diverse that it is best described as a neutral diffusion that produces an overall size increase by chance, simply because evolution was initiated at a lower size limit (Stanley 1973; Gould 1988; McKinney 1990; Jablonski 1997; Caluset and Erwin 2008; Shoemaker and Caluset 2014).
While a neutral diffusion may be sufficient for a statistical description for the evolution of size distributions, the hypothesis fails to explain rates of evolution in large versus small mammals (Baker et al. 2015), and it is insufficient from a mechanistic point of view. The quality-quantity trade-off [where parents can produce a few large or many small offspring from the same amount of energy (Smith and Fretwell 1974; Stearns 1992)] generates a constant frequency-independent selection for a continued decline in mass (Witting 1997, 2008, 2017a, b). This physiological selection for the near absence of mass implies, quite generally, that large animals can evolve only by a persistent frequency-dependent selection that is strong enough to out-balance the downward pull of the quality-quantity trade-off (Witting 2017b). The selection of species with non-negligible masses is thus a very active non-neutral process, where population dynamic feedbacks generate interactions between the frequency-independent selection of the physiology and the density-frequency-dependent selection of the intra-specific and inter-specific interactive competition.
To identify the underlying causes for the body mass distributions of the fossil record, it is essential to focus on more specific predictions than a statistical preference for an increase in size. As the latter is relatively easy to explain by a multitude of hypotheses, we need to identify specific patterns that are predicted only by a few, or a single, of the potential hypotheses. An obvious place to look for specific signals is on the allometric relations that evolve from the natural selection of mass.
Witting (2018) used Malthusian relativity to simulate the evolution of species distributions and inter-specific allometries across placental and marsupial terrestrial mammals. Starting from a single ancestor for each clade at the Cretaceous-Palaeogene boundary 65 ma, he simulated the evolution of present species distributions. Initial niche differentiations selected for a fast differentiation in net energy and mass, with a mass-rescaling selected Kleiber scaling where total metabolism increased to the 3/4 power of mass across species. While the selection of the major body mass variation from niche differentiation ceased around 50 to 30 ma, the estimated species distributions of body masses continued to evolve by an underlying background selection in mass-specific metabolism. This selection was strongest in placentals, where it bent the metabolic allometry over time and explained (Witting 2018) an observed curvature in the inter-specific allometry (Hayssen and Lacy 1985; Dodds et al. 2001; Packard and Birchard 2008; Kolokotrones et al. 2010; MacKay 2011). This created an approximate 3/4 exponent for the upper half of the size distribution, and an approximate 2/3 exponent for the lower half, providing a natural selection explanation for the 2/3 versus 3/4 controversy that has dominated the field of allometries for decades (e.g. Rubner 1883; Kleiber 1932; Heuser 1982; Feldman and McMahon 1983; Calder 1984; Dodds et al. 2001; White and Seymour 2003; Savage et al. 2004; Glazier 2010).
Where Witting (2018) focussed on the connection between body mass evolution over millions of years and the resulting inter-specific allometries today, I compared Malthusian relativity predictions with fossil data in an attempt to identify the allometries as they evolve over millions of years.
A Lack of Fossil Data
A direct test would compare the predicted exponents for body mass evolution in time with allometric exponents from fossil data. Yet these data are almost completely absent, as it is nearly impossible to estimate other life history traits than size from fossils.
There have been some attempts to estimate the age composition and maximal lifespan of fossil horses. Van Valen (1964), Hulbert (1984), and O’Sullivan (2005) used dental wear for age estimation. Yet, wear is rate dependent and there seems to be no straightforward way to convert dental wear rates in fossils into absolute age estimates. Other studies that use distinct dental wear-classes (Hulbert 1982) may be more promising, however, wear-classes may not necessarily represent year-classes (Hulbert 1984). For four species of fossil horses ranging back 18 million years (Hulbert 1982), the estimated lifespans varied from 70% to 75% of the expected given Kleiber scaling (as predicted by a \(\hat{\dot{w}}\) exponent of 0.75 in Fig. 5) and a 24 year lifespan for Burchell’s zebra (Equus bruchelli).
The Curvature of Evolution
With life history estimates for fossil species being too few for reliable estimates of allometries, I focused on the evolution of body mass in time. With mass-specific metabolism being selected as the rate of the handling that converts resource energy into replication (Witting 2017a, b), the evolutionary steady state predicts an exponential increase in net energy and mass-specific metabolism on the per-generation time-scale of natural selection. I found this increase to explain not only the population dynamic feedback selection for an exponential increase in mass, but also allometric exponents for body mass evolution in time.
These exponents depend on the importance of mass-specific metabolism for the selection of mass (Table 2), as described by the ratio for the rate of increase in mass-specific metabolism relative to the rate of increase in resource handling (\(r_{\beta _{\beta }}/r_{{\alpha }}\)-ratio). This allowed me to calculate the \(r_{\beta _{\beta }}/r_{{\alpha }}\)-ratio and the allometric exponents from the curvature of body mass evolution, as estimated from fossils by the bending exponent (\(\hat{\dot{w}}\)) for the rate of change in mass in physical time (\(\dot{w} \propto w^{\hat{\dot{w}}}\)).
The \(r_{\beta _{\beta }}/r_{{\alpha }}\)-ratio for steady state selection has a potential continuum from zero to infinity, with a bending exponent that is predicted to span from 3/4 to 3/2 given 2D interactions, and from 5/6 to 5/3 given 3D. Yet, on this continuum, I identified four types of selection that relate to four idealised conditions at different evolutionary scales.
The limit trajectory across major lifeforms (Fig. 6) has an almost log-linear trajectory in physical time (\(\hat{\dot{w}}=1\)). This corresponds with a \(r_{\beta _{\beta }}/r_{{\alpha }}\)-ratio of \(1/2(d-1)\) and evolution along a metabolic bound, where the selection increase in resource handling is so slow that the primary selection of mass-specific metabolism is able to outbalance the secondary decline in mass-specific metabolism that follows from the mass-rescaling selection of the evolutionary increase in size. The result is a mass-specific metabolism that is selected to, and maintained at, an upper bound over time.
A different limit is within-niche selection with optimal resource handling. When a lineage evolves in a stable niche over an extended time-period, we can expect a resource handling that adapts to the ecological conditions. The result is a rate of evolution in resource handling that approaches zero, and thus a \(r_{\beta _{\beta }}/r_{{\alpha }}\)-ratio that approaches infinity with a body mass that increases exclusively from a selection increase in mass-specific metabolism. This generates a \(\hat{\dot{w}}\) exponent around 3/2 for interactive competition in 2D, as approximated for small horses over 54 million years of evolution. Small horses persisted as browsers over the entire period, with a relatively slow increase in size from about 25 to 80 kg.
This contrasts to the fast increase from about 50 kg to 1 ton in the maximum size of grazing horses, which occurred from about 25 to 10 ma when horses had a major pulse of evolution during a period with strong taxonomic diversification (MacFadden 1986). This fast increase is associated with an alternative bending exponent of 0.75 (SE = 0.50) that corresponds with a \(r_{\beta _{\beta }}/r_{{\alpha }}\)-ratio of zero, and a body mass that increases from a rate of evolution in resource handling that outruns the background selection of mass-specific metabolism. This is the base-case expectation for fast body mass evolution, and it can occur when fast improvements in resource handling are easy, e.g., by a resource density that increases across niches, or by a resource handling efficiency that increases as a mechanistic function of the increase in size. This form of fast evolution was also identified for maximum mass in trunked and all terrestrial mammals.
The last idealised selection is symmetrical with a \(r_{\beta _{\beta }}/r_{{\alpha }}\)-ratio of unity. It relates to the base-case type of body mass evolution across ecological niches where the level of heritable variation is similar for resource handling and mass-specific metabolism. The selection is associated with \(\hat{\dot{w}}\) exponents of 9/8 in 2D and 5/4 in 3D, and it was identified as an approximation for the evolution of maximum mass in four out of five mammalian clades.
Several of the empirical bending exponents are estimated from relatively few data (table in Fig. 5), and they are somewhat uncertain as reflected by their standard error. It is, e.g., a bit surprising that the large size increase in whales is estimated with a symmetrical selection (\(r_{\beta _{\beta }}/r_{{\alpha }}\approx 1\)), instead of a fast selection that is dominated by an increase in resource handling or density (\(r_{\beta _{\beta }}/r_{{\alpha }}\approx 0\)). Yet, three potential predictions—which reflected the uncertainty on the classification of \(r_{\beta _{\beta }}/r_{{\alpha }}\)-ratios around unity and zero—had significant correlations between the predicted and observed bending exponents, indicating a high level of agreement between theory and data.
Natural Selection
How can a natural selection that is contingent and random at the basic level of genetic mutations predict existing lifeforms (Witting 2017b), the curvature of metabolic scaling in placental mammals (Witting 2018), and the curvature of body mass trajectories in the fossil record (this study)? This is because populations of self-replicators are dissipative systems that generate a deterministic-like selection from the use of energy in self-replication, population growth, and intra-population interactive competition. The use of selected energy generates a population dynamic feedback selection that unfolds almost inevitably like a monotonic function from the very origin of inert replicating molecules (Witting 2017b). With a steady influx of energy, there is a sustained evolution where a few general laws of selection are choosing a limited number of paths in the random space of mutations.
This implies a selection where a succession of lifeforms, exponential evolutionary trajectories, and inter-specific allometric variation follow more or less as a deterministic function of the selected variation in net energy across time and inter-specific space. A large fraction of the life history variation that were regarded as adaptations in the past are straightforwardly calculated from the selection balance between population growth and intra-population interactive competition.
An essential part of the feedback relates to selection on a per-generation time-scale, as observed for both morphological (Lynch 1990; Gingerich 1993; Okie et al. 2013) and molecular traits (Martin and Palumbi 1993; Gillooly et al. 2005; Nabholz et al. 2008; Galtier et al. 2009; Broham 2011). With natural selection selecting mass-specific metabolism as the rate of biological processes, it follows that each species has its own time-scale of natural selection that dilates and contracts relative to physical time dependent upon the evolutionary changes in metabolism and mass.
It is this dilation and contraction of the frequency of natural selection changes that bend the log-linear trajectories of exponential evolution in physical time. With the actual values of the predicted curvature agreeing with fossil data, I found that it is essential to include evolutionary changes in natural selection time in order to interpret evolutionary changes in time.
Data availability
The paper is based on data that are already available in the literature and on the net. Data recalculations are uploaded as supplementary information.
References
Alroy, J. (1998). Cope’s rule and the dynamics of body mass evolution in North American fossil mammals. Science, 280, 731–734.
Anderson, W. W. (1971). Genetic equilibrium and population growth under density-regulated selection. The American Naturalist, 105, 489–498.
Baker, J., Meade, A., Pagel, M., & Venditti, C. (2015). Adaptive evolution toward larger size in mammals. Proceedings of the National Academy of Sciences of the United States of America, 112, 5093–5098.
Banavar, J. R., Maritan, A., & Rinaldo, A. (1999). Size and form in efficient transportation networks. Nature, 399, 130–132.
Bonner, J. T. (1965). Size and cycle. Princeton: Princeton University Press.
Bonner, J. T. (2006). Why size matters. From bacteria to blue whales. Princeton: Princeton University Press.
Broham, L. (2011). The genome as a life-history character: Why rate of molecular evolution varies between mammal species. Philosophical Transactions of the Royal Society B, 366, 2503–2513.
Brown, J. H., Gillooly, A. P., Allen, V. M., & Savage, G. B. (2004). Towards a metabolic theory of ecology. Ecology, 85, 1771–1789.
Brown, J. H., & Sibly, R. M. (2006). Life-history evolution under a production constraint. Proceedings of the National Academy of Sciences of the United States of America, 103, 17595–17599.
Bueno, J., & López-Urrutia, A. (2012). The offspring-development-time/offspring-number trade-off. The American Naturalist, 179, E196–E203.
Calder, W. A. I. (1984). Size, function, and life history. Cambridge: Harvard University Press.
Caluset, A., & Erwin, D. H. (2008). The evolution and distribution of species body size. Science, 321, 399–401.
Charlesworth, B. (1971). Selection in density-regulated populations. Ecology, 52, 469–474.
Charlesworth, B. (1994). Evolution in age-structured populations (2nd ed.). Cambridge: Cambridge University Press.
Charnov, E. L. (2011). Body size is the history of life: Growth confronting landscapes of death. Evolutionary Ecology Research, 13, 553–555.
Clarke, B. (1972). Density-dependent selection. The American Naturalist, 106, 1–13.
Cope, E. D. (1887). The origin of the fittest. New York: D. Appleton and Co.
Damuth, J. (1981). Population density and body size in mammals. Nature, 290, 699–700.
Damuth, J. (1987). Interspecific allometry of population density in mammals and other animals: The independence of body mass and population energy-use. Biological Journal of the Linnean Society, 31, 193–246.
Dawkins, R., & Krebs, J. R. (1979). Arms races between and within species. Proceedings of the Royal Society London B, 205, 489–511.
DeLong, J. P. (2012). Experimental demonstration of a ‘rate-size’ trade-off governing body size optimization. Evolutionary Ecology Research, 14, 343–352.
DeLong, J. P., Okie, J. G., Moses, M. E., Sibly, R. M., & Brown, J. H. (2010). Shifts in metabolic scaling, production, and efficiency across major evolutionary transitions of life. Proceedings of the National Academy of Sciences of the United States of America, 107, 12941–12945.
Dodds, P. S., Rothman, D. H., & Weitz, J. S. (2001). Re-examination of the “3/4-law” of metabolism. Journal of Theoretical Biology, 209, 9–27.
Dreyer, O., & Puzio, R. (2001). Allometric scaling in animals and plants. Journal of Mathematical Biology, 43, 144–156.
Eisenmann, V. (2003). Gigantic horses. Paleobiology, 50, 57–73.
Feldman, H. A., & McMahon, T. A. (1983). The 3/4 mass exponent for energy metabolism is not a statistical artifact. Respiratory Physiology, 52, 149–163.
Fenchel, T. (1974). Intrinsic rate of natural increase: The relationship with body size. Oecologia, 14, 317–326.
Fisher, R. A. (1930). The genetical theory of natural selection. Oxford: Clarendon.
Galtier, N., Blier, P. U., & Nabholz, B. (2009). Inverse relationship between longevity and evolutionary rate of mitochondrial proteins in mammals and birds. Mitochon, 9, 51–57.
Gillooly, J. F., Allen, A. P., West, G. B., & Brown, J. H. (2005). The rate of DNA evolution: Effects of body size and temperature on the molecular clock. Proceedings of the National Academy of Sciences of the United States of America, 102, 140–145.
Gillooly, J. F., Charnov, E. L., West, G. B., Savage, V. M., & Brown, J. H. (2002). Effects of size and temperature on developmental time. Nature, 417, 70–73.
Gingerich, P. D. (1993). Quantification and comparison of evolutionary rates. American Journal of Science, 293A, 453–478.
Glazier, D. S. (2010). A unifying explanation for diverse metabolic scaling in animals and plants. Biological Reviews, 85, 111–138.
Gould, S. J. (1988). Trends as changes in variance: A new slant on progress and directionality in evolution. Journal of Paleontology, 62, 319–329.
Haigh, J., & Rose, M. R. (1980). Evolutionary game auctions. Journal of Theoretical Biology, 85, 381–397.
Hayami, I. (1978). Notes on the rates and patterns of size change in evolution. Paleobiology, 4, 252–260.
Hayssen, V., & Lacy, R. C. (1985). Basal metabolic rates in mammals: Taxonomic differences in the allometry of BMR and body mass. Comparative Biochemistry and Physiology, 81A, 741–754.
Heuser, A. A. (1982). Energy metabolism and body zise, I. Is the 0.75 mass exponent of Kleiber’s equation a statistical artifact? Respiratory Physiology, 48, 1–12.
Hulbert, R. C. (1982). Population dynamics of the three-toed horse Neohipparion from the late Miocene of Florida. Paleobiology, 8, 159–167.
Hulbert, R. C. (1984). Paleoecology and population dynamics of the early Miocene (Hemingfordian) horse Parahippus leonensis from the Thomas Farm site, Florida. Journal of Vertebrate Paleontology, 4, 547–558.
Humphries, M. M., & McCann, K. S. (2014). Metabolic constraints and currencies in animal ecology. Metabolic Ecology, 83, 7–19.
Jablonski, D. (1997). Body-size evolution in cretaceous molluscs and the status of cope’s rule. Nature, 385, 250–252.
Kiørboe, T., & Hirst, A. G. (2014). Shifts in mass scaling of respiration, feeding, and growth rates across life-form transitions in marine pelagic organisms. The American Naturalist, 183, E118–E130.
Kleiber, M. (1932). Body and size and metabolism. Hilgardia, 6, 315–353.
Kolokotrones, T., Savage, V., Deeds, E. J., & Fontana, W. (2010). Curvature in metabolic scaling. Nature, 464, 753–756.
Lynch, M. (1990). The rate of morphological evolution in mammals from the standpoint of the neutral expectation. The American Naturalist, 136, 727–741.
MacFadden, B. J. (1986). Fossil horses from “eohippus” (hyracotherium) to equus: scaling, cope’s law, and the evolution of body size. Paleobiology, 12, 355–369.
MacKay, N. J. (2011). Mass scale and curvature in metabolic scaling. Journal of Theoretical Biology, 280, 194–196.
Makarieva, A. M., Gorshkov, V. G., & Bai-Lian, L. (2005). Energetics of the smallest: Do bacteria breathe at the same rate as whales. Proceedings of the Royal Society B, 272, 2219–2224.
Makarieva, A. M., Gorshkov, V. G., Li, B., Chown, S. L., Reich, P. B., & Gavrilov, V. M. (2008). Mean mass-specific metabolic rates are strikingly similar across life’s major domains: Evidence for life’s metabolic optimum. Proceedings of the National Academy of Sciences of the United States of America, 105, 16994–16999.
Martin, A. P., & Palumbi, S. R. (1993). Body size, metabolic-rate, generation time, and the molecular clock. Proceedings of the National Academy of Sciences of the United States of America, 90, 4078–4091.
Maurer, B., Brown, J. H., & Rusler, R. (1992). The micro and macro in body size evolution. Evolution, 46, 939–953.
Maynard Smith, J., & Brown, R. L. W. (1986). Competition and body size. Theoretical Population Biology, 30, 166–179.
McKinney, M. L. (1990). Trends in body-size evolution. In K. J. McNamara (Ed.), Evolutionary trends (pp. 75–118). Tucson: The University of Arizona Press.
McLaren, I. A. (1966). Adaptive significance of large size and long life of the chaetognarth sagitta elegans in the arctic. Ecology, 47, 852–855.
Nabholz, B., Glemin, S., & Galtier, N. (2008). Strong variations of mitochondrial mutation rate across mammals the longevity hypothesis. Molecular Biology and Evolution, 25, 120–130.
Okie, J. G., Boyer, A. G., Brown, J. H., Costa, D. P., Ernest, S. K. M., Evans, A. R., et al. (2013). Effects of allometry, productivity and lifestyle on rates and limits of body size evolution. Proceedings of the Royal Society B, 280, 20131007.
O’Leary, M., Bloch, J. I., Flynn, J. J., Gaudin, T. J., Gillombardo, A., Giannini, N. P., et al. (2013). The Placental mammal ancestor and the Post-K-Pg radiation of Placentals. Science, 339, 662–667.
O’Sullivan, J. A. (2005). Population dynamics of Archaeohippus blackbergi (Mammalia; Equidae) from the Miocene Thomas Farm fossil site of Florida. Bulletin Journal, 45, 449–463.
Packard, G. C., & Birchard, G. F. (2008). Traditional allometric analysis fails to provide a valid perdictive model for mammalian metabolic rates. The Journal of Experimental Biology, 211, 3581–3587.
Parker, G. A. (1979). Sexual selection and sexual conflict. In M. S. Blum & N. A. Blum (Eds.), Sexual selection and reproductive competition in insects (pp. 123–166). New York: Academic Press.
Parker, G. A. (1983). Arms races in evolution-an ess to the opponent-independent cost game. Journal of Theoretical Biology, 101, 619–648.
Payne, J. L., Boyer, A. G., Brown, J. H., Finnegan, S., Kowalewski, M., Krause, R. A., et al. (2009). Two-phase increase in the maximum size of life over 3.5 billion years reflect biological innovation and environmental opportunity. Proceedings of the National Academy of Sciences of the United States of America, 106, 24–27.
Peters, R. H. (1983). The ecological implication of body size. Cambridge: Cambridge University Press.
Rau, A. R. P. (2002). Biological scaling and physics. Journal of Biosciences, 27, 475–478.
Robertson, A. (1968). The spectrum of genetic variation. In R. C. Lewontin (Ed.), Population biology evolution (pp. 5–16). New York: Syracuse University Press.
Roff, D. A. (1981). On being the right size. The American Naturalist, 118, 405–422.
Roff, D. A. (1986). Predicting body size with life history models. Bioscience, 836, 316–323.
Roughgarden, J. (1971). Density-dependent natural selection. Ecology, 5, 453–468.
Rubner, M. (1883). Über den einfluss der korper grosse auf stoff-und kraft-wechsel. Zeitschrift fur Biologie, 19, 535–562.
Santillán, M. (2003). Allometric scaling law in a simple oxygen exchanging network: Possible implications on the biological allometric scaling laws. Journal of Theoretical Biology, 223, 249–257.
Savage, V. M., Gillooly, J. F., Wooduff, W. H., West, G. B., Allen, A. P., Enquist, B. J., et al. (2004). The predominance of quarter-power scaling in biology. Functional Ecology, 18, 257–282.
Schoener, T. W. (1969). Models of optimal size for solitary predators. The American Naturalist, 103, 277–313.
Shoemaker, L., & Caluset, A. (2014). Body mass evolution and diversification within horses (family Equidae). Ecology Letters, 17, 211–220.
Sibly, R. M., Brown, J. I., & Kodric-Brown, A. (2012). Metabolic ecology: A scaling approach. Chichester: Wiley.
Simpson, G. G. (1953). The major features of evolution. New York: Columbia University Press.
Smith, C. C., & Fretwell, S. D. (1974). The optimal balance between size and number of offspring. The American Naturalist, 108, 499–506.
Smith, F. A., Boyer, A. G., Brown, J. H., Costa, D. P., Dayan, T., Ernest, S. K. M., et al. (2010). The evolution of maximum body size of terrestrial mammals. Science, 330, 1216–1219.
Stanley, S. M. (1973). An explanation for cope’s rule. Evolution, 27, 1–26.
Stearns, S. C. (1992). The evolution of life histories. Oxford: Oxford University Press.
Stearns, S. C., & Crandall, R. E. (1981). Quantitative predictions of delayed maturity. Evolution, 35, 455–463.
Stearns, S. C., & Koella, J. (1986). The evolution of phenotypic plasticity in life-history traits: Predictions for norms of reaction for age- and size-at-maturity. Evolution, 40, 893–913.
Taylor, P. D. (1996). The selection differential in quantitative genetics and ess models. Evolution, 50, 2106–2110.
Trammer, J. (2005). Maximum body size in a radiating clade as a function of time. Evolution, 59, 941–947.
Van Valen, L. (1964). Age in fossil horse populations. Acta Zoologica, 45, 93–106.
Vermeij, G. J. (1987). Evolution and escalation. Princeton: Princeton University Press.
West, G. B., Brown, J. H., & Enquist, B. J. (1997). A general model for the origin of allometric scaling laws in biology. Science, 276, 122–126.
West, G. B., Brown, J. H., & Enquist, B. J. (1999a). A general model for the structure and allometry of plant vascular systems. Nature, 400, 664–667.
West, G. B., Brown, J. H., & Enquist, B. J. (1999b). The fourth dimension of life: Fractal geometry and allometric scaling of organisms. Science, 284, 1677–1679.
White, C. R., & Seymour, R. S. (2003). Mammalian basal metabolic rate is proportional to body mass\(^{2/3}\). Proceedings of the National Academy of Sciences of the United States of America, 100, 4046–4049.
Witting, L. (1995). The body mass allometries as evolutionarily determined by the foraging of mobile organisms. Journal of Theoretical Biology, 177, 129–137. https://doi.org/10.1006/jtbi.1995.0231.
Witting, L. (1997). A general theory of evolution. By means of selection by density dependent competitive interactions. Århus: Peregrine Publisher.
Witting, L. (2000). Interference competition set limits to the fundamental theorem of natural selection. Acta Biotheoretica, 48, 107–120. https://doi.org/10.1023/A:1002788313345.
Witting, L. (2002). From asexual to eusocial reproduction by multilevel selection by density dependent competitive interactions. Theoretical Population Biology, 61, 171–195. https://doi.org/10.1006/tpbi.2001.1561.
Witting, L. (2003). Major life-history transitions by deterministic directional natural selection. Journal of Theoretical Biology, 225, 389–406. https://doi.org/10.1016/S0022-5193(03)00274-1.
Witting, L. (2008). Inevitable evolution: Back to The Origin and beyond the 20th Century paradigm of contingent evolution by historical natural selection. Biological Reviews, 83, 259–294. https://doi.org/10.1111/j.1469-185X.2008.00043.x.
Witting, L. (2017a). The natural selection of metabolism and mass selects allometric transitions from prokaryotes to mammals. Theoretical Population Biology, 117, 23–42. https://doi.org/10.1016/j.tpb.2017.08.005.
Witting, L. (2017b). The natural selection of metabolism and mass selects lifeforms from viruses to multicellular animals. Ecology and Evolution, 7, 9098–9118. https://doi.org/10.1002/ece3.3432.
Witting, L. (2018). The natural selection of metabolism explains curvature in allometric scaling. Oikos, 127, 991–1000. https://doi.org/10.1111/oik.05041.
Acknowledgements
I thank those who collect fossils, estimate their body sizes, and publish the data. I am also grateful to reviewers and editors that helped me improve the article.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Appendices
Appendix 1
Exponential Increase
Earlier studies (Witting 1997, 2003, 2017a, b, 2018) have found large body masses to be selected by a population dynamic feedback selection, where the net energy that is allocated into reproduction generates population growth and interactive competition. The latter creates a positive correlation between net energy and mass because the larger than average individuals monopolise resources during interactive competition. The positive correlation is then selecting net energy into mass at the cost of reproduction and a continued increase in the density of individuals.
As metabolism burns energy it follows that selection will optimise the physiological and ecological work that is essential for the life history. One implication of this (Witting 2017a), is a selection where the net energy
of the average individual (SI unit J/s) is selected as a product between the handling of resource assimilation [\({\alpha }= \grave{{\alpha }}\rho ^{**}\); SI unit J, given by intrinsic handling (\(\grave{{\alpha }}\)) at the equilibrium resource density \(\rho ^{**}\)], and the pace (\(\tilde{\beta }=\beta /{W}\); SI unit 1/s) of this process, with pace being selected as a proxy for mass-specific metabolism (\(\beta \); SI unit J/Js), with \({W}\) (SI unit J/J) being the mass-specific work of handling that is obtained by metabolising one joule per unit body mass, with mass (w; SI unit J) given as biotic (combustion) energy.
The average per-generation rate of replication
is one for populations that evolve over long periods of time, and it is proportional to net energy (\({\epsilon }\)), with p being the probability to survive to reproduce, \(R=t_r{\epsilon }/ \acute{\beta }w\) being lifetime reproduction (unitless number), \(t_r\) (SI unit s) the reproductive period in physical time, and \(\acute{\beta }\) a unitless scaling that accounts for energy that is metabolised by the offspring (Witting 2017a).
Given the constraints of Eqs. 10 and 11, the per-generation selection gradients on the three resource assimilation parameters \(x \in \{{\epsilon }, {\alpha }, \tilde{\beta }\}\) are unity on logarithmic scale
with \(r = \ln {\lambda }\). Then, from the secondary theorem of natural selection (\(\text{ d }x/\text{ d }\tau = {\sigma }{_{x}^{2}} {\partial }r / {\partial }x\), Robertson 1968; Taylor 1996) we expect an exponential increase
on the per-generation time-scale of natural selection, given unconstrained selection as defined by an invariant heritable variance (\(\sigma _{\ln x}^2\)). The rate of increase in net energy
is the sum of the rates of increase in the two subcomponents of resource handling and metabolic pace. When selection is constrained, e.g., by an \({\alpha }\) that is approaching the selection optimum of a niche, or a \(\beta \) that is approaching an upper limit, it follows that the heritable variance will approach zero and that the rate of increase will cease.
The metabolic increase of Eq. 13 is caused by the primary selection of metabolism (subscript \(\beta \)). This is to be distinguishes from the metabolic decline in the mass-rescaling component (\(\beta _{w}\)) of mass-specific metabolism, which is a secondary selection response to the selection changes in mass (Witting 2017a).
Resource handling (\({\alpha }\)), on the other hand, is a pure primary selected parameter that generates net energy independently of the evolutionary changes in metabolism. It has a mass-rescaling exponent of unity (Witting 2017a) that reflects that an energetic increase in mass is caused exclusively by an increase in \({\alpha }\) for cases with no primary evolution in mass-specific metabolism. Although resource handling is defined independently of the primary selection of metabolism, it does have an indirect metabolic-rescaling exponent (\(\hat{{\alpha }}_{\beta }=1-\hat{\beta }_{\beta }\)) that reflects the importance of \({\alpha }\), relative to \(\beta \), for the generation of the net energy that is selected into mass (see Eq. 25 in Witting 2017a).
Net energy in physical time (\({\epsilon }\)) is affected by mass-rescaling (\(\hat{{\epsilon }}_{w} = \hat{{\epsilon }} = (2d-1)/2d\), Witting 2017a). Yet, net energy on the per-generation time-scale (which is the relevant time-scale for population dynamics and natural selection) is a primary selected parameter, with an allometric exponent of unity that reflects the conversion of net energy into mass. The mass-rescaling selection of mass maintains the current primary selected net energy by a dilation of natural selection time that balances the decline in net energy in physical time (Witting 2017a).
The increase in net energy generates population growth with a density-dependent equilibrium, where Eq. 83 in Witting (2017a) determines the level of interactive competition
as a function of net energy, with \({\gamma _{\iota }}\) being the density dependence of interference competition, \({\gamma }\) the overall strength of density regulation, and \({\acute{{\epsilon }}}_{0}\) a measure of net energy on the per-generation time-scale. The population pressure of interference competition generates population dynamic feedback selection, where the resulting rate of increase in mass
is a product between the selection gradient [\({\partial r_i^* / \partial \ln {w}_i |_{{w}_i=w}}=({\psi }{\gamma _{\iota }}/{\gamma })\ln ({\acute{{\epsilon }}}_{0}/w)-1\), with subscript i denoting intra-population variation; from Eq. 28 in Witting 2017b] and the heritable variance (\({\sigma }{_{\ln w}^{2}}\)), with \({\psi }\) being the gradient in the fitness cost of interference across the body mass variation in the population [as the larger individuals monopolise resources in interactive competition; see Witting, 2017a; b for details].
As the \({\acute{{\epsilon }}}_{0}/w\) component of Eq. 16 is invariant of mass (Witting 2017a), the expected increase is exponential, and it may be rewritten as
with an invariant selection relation
Hence, we have
and a natural selection that defines mass
by the inverse of the net energy allometry
where \(\hat{{\epsilon }} = (2d-1)/2d\) is the predicted exponent from Witting (2017a), \({{\epsilon }_{\circ }}\) the intercept, and d the spatial dimensionality of the intra-specific interactions.
By setting Eq. 19 equal to Eq. 16, and noting the constraint of Eq. 11, we find the mass
of the evolutionary steady state with an exponential increase in mass. Then, from Eq. 16 we have
that confirm that the steady state is the unconstrained selection attractor for mass, with ** superscript denoting the attractor of unconstrained selection.
If we insert Eq. 15 into Eq. 16, and set \(r_{w}=r_{{\epsilon }}/\hat{{\epsilon }}\) from Eq. 19, we find the level of intra-specific interference competition
for an exponentially evolving mass to be somewhat higher than the level of interference for a stable mass at an equilibrium attractor (\(\overline{*}*\)) with an upper constraint on net energy, where \({\iota }{^{\overline{*}*}}=1/{\psi }\) (from Eq. 25 in Witting 2017b). As the parameters on the right hand side of Eq. 24 are expected to be body mass invariant, the theoretically deduced allometries (Witting 1995, 2017a) apply for body mass evolution at steady state. As noted already, this implies an energetic exponent of \(\hat{{\epsilon }}=(2d-1)/2d\), and with \(r_{{\epsilon }}={\sigma }{_{\ln {\epsilon }}^{2}}\) from Eq. 13, we find that the level of interference reduces to
for the symmetrical case where \({\sigma }{_{\ln w}^{2}}={\sigma }{_{\ln {\epsilon }}^{2}}\). The position of this selection attractor of the steady state is shown on the selection integral in Fig. 1f in Witting (2017b).
Nearly all the allometries considered here and in Witting (2017a) describe variation in the average traits across evolved populations; either across species (Witting 2017a) or along an evolutionary lineage in time (current paper). Yet, the steady state defines also an important intra-specific allometry that describes the correlation between reproduction and mass across the individuals in the evolving population. To obtain this allometry, insert Eq. 25 into the selection gradient \({\partial r_i^* / \partial \ln {w}_i |_{{w}_i=w}}={\psi }{\iota }{^{*}}-1\) for the multicellular animal in Witting (2017b), and integrate over \(\ln w_i\) to find that the within population variation in the pR-product scale as
As this exponent is 4/3 and 6/5 for organisms with intra-specific interactions in two and three spatial dimensions, we find a reproductive rate that is about proportional to mass when survival is relatively invariant. This proportionality is often observed in natural populations (Peters 1983), and it reflects a level of interactive competition that is so high that resource monopolization scales to the approximate second power of mass.
Allometries in Time
The predicted exponential increase in metabolism and mass on the per-generation time-scale is an essential finding for evolutionary biology in itself. To understand the implication in detail, we need to transform the prediction into physical time to compare with fossil data. Hence, we need to predict the correlated evolution between mass and the per-generation time-scale of natural selection.
This correlation is given by the allometries in this subsection, with allometries being predicted not only for the generation time that is needed for the time-transformation of body mass evolution, but also for a larger set of traits that allow for more general predictions of life history evolution in time.
The exponents of the final allometries evolve as the sum of the partial correlations of the metabolic-rescaling and mass-rescaling that follow from the primary selection of metabolism and mass (Witting 2017a). To deduce this evolution we have from Eqs.14 and 19 that
Now let
The exponential increase in α, \(\beta _{\beta }\), \({\epsilon }\) and w on the per-generation time-scale may then be expressed as a function of the exponential increase (\(r_{w}\)) in mass, i.e.,
From these equations we can construct time allometries. For this, solve the body mass relation for time \(\tau =\ln (w_\tau /w_0)/r_{w}\). Insert this time into the other relations and obtain
with intercepts \({{\alpha }_{\circ }}={\alpha }_0/w_0\), \({\beta _{\circ }}=\beta _{\beta , 0}/w_0\), and \({{\epsilon }_{\circ }}={\epsilon }_0/w_0\). Hence, from Eqs. 28, 30, and \(\hat{{\epsilon }}=(2d-1)/2d\), we find the following allometries for an evolutionary lineage in time
From the inverse relation between biotic time and metabolic pace, we find generation time to evolve with metabolic-rescaling as
and mass-rescaling as
The mass-rescaling of Eq. 33 dilates natural selection time as the body mass is increasing by natural selection. The mass-rescaling intercept, however, will decline by the metabolic-rescaling (Eq. 32) that evolves by the primary selected mass-specific metabolism, and this causes a contraction of natural selection time. Whether natural selection time will actually contract or dilate depends on the level of metabolic-rescaling relative to mass-rescaling with the final allometry for generation time evolving as
To transform the predicted trajectories to the physical time-scale of the fossil record, we have that the rate of change in mass in physical time is
Then, from Eq. 34 we obtain the following allometry
for the rate of evolutionary change in mass, where
and \({r_{w,\circ }}=r_{w}/{t_{g,\circ }}\).
Rights and permissions
About this article
Cite this article
Witting, L. The Natural Selection of Metabolism Explains Curvature in Fossil Body Mass Evolution. Evol Biol 47, 56–75 (2020). https://doi.org/10.1007/s11692-020-09493-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11692-020-09493-y