Abstract
DNA methylation, an epigenetic mechanism used by cells to control gene expression, has an important biological role in plant development and environmental fitness. Since plant DNA methylation is closely related to environmental conditions, variation during the day is expected. Here, in genetically identical plants of Populus nigra clone N46, DNA methylation changes in leaves over a 24 h period were detected using the methylation-sensitive amplification polymorphism method. The results showed different DNA methylation patterns in mature poplar leaves: not only in individuals at the same time, but also in samples at each of the six time during the day. In addition, night samples had a higher percentage of methylation than in morning samples. However, no statistically significant differences were found among the samples gathered at different times. Similar results were obtained for three other P. nigra clones with different genetic backgrounds. Real time qPCR showed that the DNA methyltransferase genes Pt-MET1 and Pt-SOM1 involved in CG DNA methylation in poplar were stable over a 24 h period in leaves of P. nigra N46 compared with circadian-controlled genes. That could be part of the reason that methylation of CCGG sites is stable in those leaves. That DNA methylation differed even in genetically identical plants indicates the specificity of DNA methylation changes in their genomes. No statistically significant differences in methylation changes were found between day and night, suggesting that DNA methylation is more stable than expected and is unlikely to be involved in circadian regulation in plants.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
DNA methylation is an important epigenetic modification in the genomes of eukaryotes such as fungi, plants, and animals (Martienssen and Colot 2001). In plant genomes, DNA methylation predominantly occurs at symmetric CG sites; it is also found in symmetric CHG and asymmetric CHH sites (in which H = A, T, or C) (Law and Jacobsen 2010). In Arabidopsis, it occurs at CG, CHG, and CHH sites at rates of approximately 24.0, 6.7, and 1.7%, respectively (Cokus et al. 2008). For equivalent sites in Populus trichocarpa, the rates are approximately 41.9, 20.9, and 3.25%, respectively (Feng et al. 2010). DNA methylation in plants is catalyzed by domains rearranged methyltransferase 2 (DRM2). The maintenance of DNA methylation at CG, CHG, and CHH sites is catalyzed by three DNA methyltransferases: CG methylation is maintained by DNA methyltransferase 1 (MET1), CHG methylation is maintained by chromomethylase 3 (CMT3), and CHH methylation by DRM2 (Law and Jacobsen 2010). Besides the DNA methyltransferases in plants, 5-methylcytosine DNA glycosylases can actively remove a methyl group (–CH3), indicating the dynamic nature of DNA methylation in plant genomes (Agius et al. 2006; Penterman et al. 2007; Vining et al. 2012; Zhu 2009).
Changes in DNA methylation have been reported during plant gametogenesis, flower patterning, and seed development (Calarco et al. 2012; Ikeda 2012; Martienssen and Colot 2001; Ma et al. 2015; Xing et al. 2015). In recent years, a number of studies have also investigated changes in plant DNA methylation under different environmental conditions (Mirouze and Paszkowski 2011). Most detected clear changes in DNA methylation (Dowen et al. 2012; Eichten and Springer 2015; Raj et al. 2011; Yaish et al. 2011). The main environmental factors that influence DNA methylation status are temperature and light. For example, hypomethylation was induced by cold stress in Antirrhinum, and the degree of transposon Tam3 DNA methylation was positively correlated with growth temperature (Hashida et al. 2006). In addition, the expression of the Arabidopsis gene At3g50770 was inversely correlated with promoter DNA methylation at an elevated temperature (Naydenov et al. 2015). Plants grown in the light, and those grown in the dark also have different methylation levels (Omidvar and Fellner 2015). Under natural conditions, environmental factors such as humidity, temperature, and light intensity change throughout the day, and therefore, changes in DNA methylation in plant genomes are expected. Although daily changes in DNA methylation have been found in human blood and in mouse liver (Bönsch et al. 2007; Xia et al. 2015), variations in DNA methylation during the day and at night have yet to be reported for plants.
Poplars (Populus L.) are model forest tree species, and one of the most commercially and ecologically important forest trees. Because they can rapidly reproduce asexually to produce genetically identical clones, poplars are ideal plants for studying methylation variation. Here, using the P. nigra clone N46, we analyzed variations in DNA methylation over 24 h using the methylation-sensitive amplification polymorphism (MSAP) method. We found different DNA methylation patterns in mature poplar leaves, not only among individuals at the same time, but also among samples at each of six times. A higher methylation percentage and variation rates in DNA methylation were observed in the night samples compared with the morning samples. However, statistical analyses showed that there were no significant differences among the samples from different times. Similar results were also obtained for three other P. nigra clones with different genetic backgrounds.
Materials and methods
Plant material and growth conditions
Four P. nigra clones (N46, N08, N15 and N77) were used in the experiments, initially sourced from Belgium, West China, Yugoslavia and Russia, respectively, in 2000 and preserved in a germplasm bank in Beijing since then (Chu et al. 2014). In early April 2012, homogeneous, dormant, 15 cm-long hardwood cuttings from 1 year-old stems of N46 were planted in standard nursery potting medium and placed in a greenhouse in Beijing, China (46°44′ N, 117°10′ W), under a controlled temperature (24–30 °C) and natural sunlight. Plants were watered, fertilized and rotated every 2 weeks. On August 8, 2012, 30 plants with homogeneous size were selected, and mature leaves (the 4th–6th leaves from the plant top) in each plant were collected at 8:00, 12:00, 16:00, 20:00, 24:00 and 4:00, five biological replicates each time (three for MSAP analysis and two for qPCR analysis). Daylight on the sample collection day was from 5:19 to 19:20. All leaves were frozen immediately in liquid nitrogen after collection and stored in −80 °C.
Dormant, 15 cm-long hardwood cuttings from 1 year-old stems of P. nigra clone N08, N15, and N77 were propagated as above and grown in the same greenhouse as N46 at the end of March 2013. On June 3, 2014, plants of homogeneous size from each clone were selected, and three biological replicates of mature leaves (4th–6th leaves from the top) were collected at 12:00 and at 24:00. Daylight on the sample collection day was from 4:47 to 19:38. All leaves were frozen immediately in liquid nitrogen after collection and stored at −80°C.
DNA extraction and MSAP analysis
Genomic DNA from leaves of each plant was isolated by standard CTAB method (Porebski et al. 1997) and the concentration of genomic DNA was checked with a Nanodrop 8000 spectrophotometer. Only qualified DNA samples (1.8 < A260/280 < 2.0, 2.0 < A260/230 < 2.3) were used for further analysis.
The MSAP procedures were performed following the general steps described by Cervera et al. with modifications (Cervera et al. 2002). To avoid experimental variation, the digestion, ligation and amplification reactions were carried out simultaneously in all the DNA samples of each genotype. For each sample, 450 ng samples of genomic DNA were digested by both restriction enzymes EcoRI/HpaII and EcoRI/MspI, respectively. EcoRI/HpaII digestions were performed in a 20 μL volume of 10× buffer 1 (New England Biolabs, NEB), 10 U EcoRI and 5 U HpaII for 12 h at 37 °C. EcoRI/MspI digestions were carried out in a 20 μL volume of 10× buffer 4 (New England Biolabs, NEB), 10 U EcoRI and 10 U MspI for 12 h at 37 °C. Digestion products were inactivated at 65 °C for 20 min. Two different adapters were ligated to the digested DNA in a 20 μL volume containing 2 μL 10× T4 DNA ligase buffer, 10 μL digested DNA, 1 μL T4 DNA ligase (400 U/μL), 1 μL EcoRI adapter (5 pmol/μL), 1 μL HpaII/MspI adapter (50 pmol/μL) for 16 h at 16 °C. Ligation DNA products were inactivated at 65 °C for 20 min.
Pre-amplification reaction was carried out in a Perkin Elmer 9600 thermocycler with a 20 μL volume of 2 μL 10× exTaq DNA polymerase buffer (20 mM Mg2+ Plus), 1.6 μL dNTP (2.5 mM each), 0.5 μL of each primer (EcoRI+A and HpaII/MspI+0), 0.2 μL exTaq DNA polymerase (5 U/μL, Takara, Dalian, China) and 4 μL of ligation DNA product. PCR procedure was 2 min at 94 °C, then for 20 cycles of 20 s at 94 °C, 30 s at 56 °C, and 2 min at 72 °C, and 2 min at 72 °C for extension.
Pre-amplification mixture was diluted 20-fold for use as templates for the selective amplification. The PCR was performed in a 20 μL volume of 2 μL 10× exTaq DNA polymerase buffer (Mg2+-free), 2.4 μL dNTP (2.5 mM each), 0.25 μL of fluorescently labeled EcoRI primers (10 pmol/μL), 0.25 μL HpaII/MspI primer (10 pmol/μL), 1.2 μL Mg2+ (25 mM), 0.2 μL exTaq DNA polymerase (5 U/μL), and 4 μL of diluted pre-amplified DNA. A touchdown PCR program was used in the selective amplification: 2 min at 94 °C; ten cycles of 20 s at 94 °C, 30 s at 66 °C (decrease by 1 °C in each cycle), and 2 min at 72 °C; then 20 cycles of 20 s at 94 °C, 30 s at 56 °C, 2 min at 72 °C; and a final extension of 30 min at 60 °C.
Selective amplification products of P. nigra clones were separated by capillary electrophoresison GeXP Genetic Analysis System (Beckman, American) and ABI3730 (Life technologies, American). Raw data was analyzed in fragment analysis module of GeXP software and GeneMarker V2.2.0. Disregarding bands under 60 bp for their fuzzy appearance, fragments ranging from 60 to 600 bp were marked, exported with 1 (presence) or 0 (absence), and used in data analysis.
Data analysis
The amplified DNA fragments were divided into four types: (1) type I, products amplified by both EcoRI/HpaII and EcoRI/MspI combinations, indicating the nonmethylated sites; (2) type II, products amplified by EcoRI/MspI but not by EcoRI/HpaII, indicating full and hemi-methylation of internal cytosine sites; (3) type III bands, products amplified only by EcoRI/HpaII, not EcoRI/MspI, indicating hemi-methylated external cytosine sites; (4) type IV, no products amplified by any enzyme combination, indicating full-methylation of external cytosine, or full methylation of both cytosines, or hemi-methylation of both cytosines, or an unknown mutation in CCGG site, further referred to as “uncertain” sites (Schulz et al. 2013). Percentage of methylation was calculated as (type II + type III)/ (type I + type II + type III) (Karan et al. 2012).
The polymorphic ratio (PR) was used to represent the difference in MSAP patterns among genetically identical plants collected at the same time. The detected sites that showed common EcoRI/HpaII and EcoRI/MspI patterns among replicates were called nonpolymorphic methylation sites (NMS), while those that differed among replicates were called polymorphic methylation sites (PMS). The PR was calculated as follows: \( {\text{PR }}\left( \% \right) \, = {\text{ PMS }}/ \, \left( {{\text{NMS }} + {\text{ PMS}}} \right) \times { 1}00\% \).
The msap package in the R environment was used to analyze the MSAP results to assess epigenetic variation (Perez-Figueroa 2013). The default parameters were used in the MSAP analysis, except for the parameter loci.per.primer, which is a vector providing the number of loci/fragments obtained per primer combination. To assess the methylation differences among the six time points, we considered samples collected at the same time as a population. To assess the methylation differences among different genotypes (N08, N15, and N77), we considered all samples of each genotype as a population. Epigenetic variation among different times and genotypes was assessed by an analysis of molecular variance (AMOVA) (Excoffier et al. 1992).
RNA isolation and real-time quantitative PCR
Total RNA of P. nigra N46 was extracted from mature leaves of two biological replicates using an RNeasy Plant Mini Kit (Qiagen, Germany) according to the manufacturer’s instructions. The RNA samples were treated with RNase-free DNase (Promega, WI, USA) for 30 min at 37 °C. The cDNA was synthesized with Superscript II RNase-Reverse Transcriptase (Invitrogen, CA, USA). Two poplar CG DNA methylation maintenance genes (Pt-MET1: Potri.018G138000, Potri.004G134000; Pt-SOM1: Potri.007G026700, Potri.019G129900) and one clock-controlled gene (Pt-LHCA2: Potri.003G171500 and Potri.T147200) were selected, and primers were designed by Primer3 web (http://primer3.ut.ee). Ubiquitin-like (Potri.005G198700) was used as reference genes.
The real-time quantitative PCR analysis was carried out in an ABI Prism 7500 sequence detector (Applied Biosystems, CA, USA). Each PCR (final volume 20 μL) contained 1 μL first-strand cDNA, 200 nM primers and 1× SYBR PCR mixture (TaKaRa Bio). The amplification conditions were: 10 s at 95 °C, followed by 40 cycles of 5 s at 95 °C and 35 s at 60 °C. Three to four replicates for each RNA sample were included. Relative quantification values were calculated by the \( 2^{{{{ - \Delta \Delta {\rm C}}}_{\text{T}} }} \) method (Livak and Schmittgen 2001).
Results
DNA methylation in genetically identical plants of P. nigra N46 within and among six sampling times
Genomic DNA was extracted from the leaves of 30 P. nigra N46 plants collected at six times (08:00, 12:00, 16:00, 20:00, 24:00 and 04:00) each day, with five replicates for each time. A total of 20 MSAP primer combinations were used to detect the methylation patterns at each time with an automatic sequencer. Each primer combination produced 32–83 bands varying in length from 58 to 600 bp, with most bands ranging from 100 to 400 bp. E2-HM15 produced the fewest bands (n = 32) and E7-HM17 produced the most (n = 83).
From the 30 samples collected at six different times, the 20 primer combinations detected a total of 1076 CCGG loci. DNA methylation varied among samples collected at the same time and also among samples collected at different times. The polymorphic sites among replicates of each time were analyzed, and PR was calculated. The results indicated that the 24:00 samples had the highest rate of polymorphic sites, with a PR of 6.23%, followed by those from 16:00, 20:00, 08:00, 12:00, and 04:00, with rates of 5.3, 4.65, 4.09, 3.54, and 1.66%, respectively. The average rates of methylation in samples from different times were similar. The samples from 12:00 had the lowest level of DNA methylation (48.88 ± 0.05%), while the samples from 24:00 had the highest (49.33 ± 0.22%) (Table 1).
To test the differences in methylation from samples among the six times, MSAP profiles were analyzed using the R package msap (Perez-Figueroa 2013). The AMOVA revealed that there were no significant differences in the samples among the six times (ϕ ST = 0.1588, P = 0.0516). This result indicated that DNA methylation did not change consistently depending on the time of day.
DNA methylation in three other genotypes of P. nigra
Additional analyses were performed with DNA samples isolated from the leaves of three clones of P. nigra to determine whether there were similar methylation variations in other genotypes. Based on the above results, two times (12:00 and 24:00) were selected, and the leaves of three replicates in every clone at each time point were collected. Ten randomly selected primer combinations were used to investigate the methylation patterns in those samples.
Our results confirmed that there were variations in methylation profiles both in the samples collected at each time, and the samples collected at different times in all three P. nigra genotypes (Table 2). Just like N46, samples from 24:00 in N77 had the highest rate of polymorphic sites. In N08 and N15, samples from 12:00 had the highest rate of polymorphic sites. The average methylation percentage in the samples from 24:00 was higher than for the 12:00 samples in all three P. nigra genotypes. The AMOVA analysis also revealed that there were no significant differences in the MSAP patterns between samples from the two times in the three P. nigra clones (N08: ϕ ST = 0.5946, P = 0.1014; N77: ϕ ST = 0.006897, P = 0.5023; N15: ϕ ST = 0.3095, P = 0.103).
Genetic variation was observed in terms of the DNA methylation of the three P. nigra clones. The AMOVA analysis revealed significant differences among the three genotypes (ϕ ST = 0.5686, P < 0.0001). The pairwise analysis showed a significant difference between two genotypes. Since the clones grew in the same environment during this experiment and they were kept in the same germplasm bank for over 10 years, the variation in DNA methylation reflects the genetic variation of these genotypes (Raj et al. 2011).
Expression of DNA methyltransferases
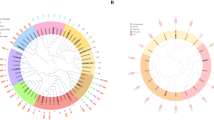
To elucidate the reason for the stable methylation status of the CCGG sites of P. nigra N46 during the day and night, the mRNA expression of two Arabidopsis DNA methyltransferases homologs in poplar involved in CG DNA methylation was analyzed by real time qPCR using poplar Pt-MET1 (Potri.018G138000, Potri.004G134000), homologs of Arabidopsis DNA METHYLTRANSFE-RASE 1 (MET1); Pt-SOM1 (Potri.007G026700, Potri.019G129900), homologs of DECREASE IN DNAMETHYLATION 1 (DDM1). One poplar homolog of the Arabidopsis clock-controlled gene Pt-LHCA2 (Potri.003G171500, Potri.T147200) was also detected. Compared with the clock-controlled gene, Pt-MET1 and Pt-SOM1 were stably expressed at all sampling times in leaves of P. nigra N46 during the day and night (Fig. 1).
Expression comparison of DNA methyltransferase genes and clock-controlled gene at six times. Pt-MET1 is poplar METHYLTRANSFERASE 1; Pt-SOM1 is poplar DECREASE IN DNAMETHYLATION 1; Pt-LHCA2 is poplar LHCA2. Pt-MET1 and Pt-SOM1 are the DNA methyltransferases involved in CG DNA methylation; LHCA2 is the clock-controlled gene. The qRT-PCR was performed on total RNA extracted from leaf samples (two biological replicates) at each of the six times. Experiments were done in three to four replicates, and the error bars in the figure represent standard errors
Discussion
In this study, we first used genetically identical plants from the P. nigra clone N46 to determine whether DNA methylation changed over time (day and night) using the MSAP method. Because the plants were genetically identical, it was expected that the same MSAP profiles would be obtained for those samples collected at each time. However, unexpectedly, methylation variation was found in all of the samples from the six times, with the polymorphic rate of detected CCGG sites ranging from 1.66 to 6.23%. This phenomenon was also observed for three other genotypes of P. nigra, suggesting that it is common in poplars. DNA methylation variation in genetically identical plants has been reported in both herbaceous plants and woody plants. In Arabidopsis, approximately 1% of CCGG sites were shown to differ in methylation status in the Ler ecotype under normal culture conditions (Cervera et al. 2002). In maize, differentially methylated regions (DMRs) were found to differ between individuals in either the control population or stressed populations (Eichten and Springer 2015). In addition, in the woody tree Pinus pinea, cytosine methylation polymorphism among ramets of each propagated tree ranged from 0.46 to 9.72% (Sáez-Laguna et al. 2014). The methylation variation in those genetically identical plants in P. pinea was considered to have resulted from different ontological stages, developmental stages, or microenvironmental variation among plants during their growth. These factors might also have contributed to the variation in methylation among samples from the same times. In our case, such variation might have been derived from the poplar buds that had formed the previous year.
Although different DNA methylation patterns were found among samples from the six times, no significant differences were obtained by statistical analysis, indicating that the methylation status in the mature leaves of poplars kept relatively stable throughout the day. Then, why didn’t daily changes in the environmental condition influence the methylation status in poplar? In Arabidopsis, the circadian clock regulates the expression of hundreds of genes, but evidently not DNA methyltransferases (Harmer et al. 2000). In our study, we found that mRNA transcript levels of MET1 and DDM1, that maintain the CG sites methylation in poplar, remained during the day and night, compared with the circadian-controlled genes. That could be part of the reason for the relatively stable methylation of CCGG sites in those leaves. In plants, DNA methylation was established by a RNA-directed DNA methylation process and maintained mainly by MET1 and DDM1. Under environmental stimuli, plants respond instantly by regulating the expression of genes through transcription regulation (Wilkins et al. 2009). And then epigenetic regulation such as histone modification might act in the short term (Kumar and Wigge 2010). DNA methylation change under stress might be through an accumulation of methylated sites over a longer time than 1 or 2 days (Rico et al. 2014). Because cytosine methylation is adding a methyl group (–CH3) from S-adenosyl-l-methionine on a cytosine, it is not an economical regulation mechanism for instant response. Up to now, the DNA methylation changes reported were all results of a long period of environmental stimuli (Gourcilleau et al. 2010; Rico et al. 2014). Therefore, the changes in DNA methylation at different times might be stochastic and less related to the time of day. Even under stressed conditions, only minimal evidence for consistent changes in maize DNA methylation patterns was found when using the data from all replicates (Eichten and Springer 2015). We thus suspected that, even under identical conditions, the changes in methylation might not occur in synchrony in all individual plants with the same genetic background. This might be the result of the dynamic nature of DNA methylation (Zhu 2009).
The MSAP method is limited for quantifying changes in DNA methylation (Pecinka and Scheid 2012). However, it is a simple, cheap tool for studying DNA methylation and has been successfully used to detect natural variation in methylation in many plant species, including poplars (Salmon et al. 2008; Herrera and Bazaga 2011; Li et al. 2011; Song et al. 2012; Herrera et al. 2013; Ma et al. 2013; Yu et al. 2013; Lira-Medeiros et al. 2010; Sáez-Laguna et al. 2014). Our results obtained using the MSAP method showed significant differences in DNA methylation among different P. nigra genotypes, which again indicates that this is a powerful method for detecting natural variation in methylation in plants with different genetic backgrounds.
Genome resequencing revealed that the P. nigra genome contained approximately 14,000 EcoRI sites (unpublished data). Theoretically, there are 28,000 EcoRI/HpaII or EcoRI/MspI fragments that could be amplified using different selective nucleotides (Cervera et al. 2002). In this study, only a small proportion of the CCGG sites were detected in the 4 clones. However, besides the CCGG sites, substantial methylation occurred at the CHG and CHH sites in plants. Owing to the limitations of the sites detected, we could only conclude that poplar genomic DNA varies in terms of methylation, even in genetically identical plants. However, the lack of any significant changes in methylation among the six sampling times within 24 h suggested that methylation status of plant CCGG sites in mature leaves is relatively stable throughout the day. This work also provides scientific bases for sample collection in studies of epidemic variations in plants.
References
Agius F, Kapoor A, Zhu JK (2006) Role of the Arabidopsis DNA glycosylase/lyase ROS1 in active DNA demethylation. Proc Natl Acad Sci USA 103(31):11796–11801. doi:10.1073/pnas.0603563103
Bönsch D, Hothorn T, Krieglstein C, Koch M, Nehmer C, Lenz B, Reulbach U, Kornhuber J, Bleich S (2007) Daily variations of homocysteine concentration may influence methylation of DNA in normal healthy individuals. Chronobiol Int 24(2):315–326. doi:10.1080/07420520701290565
Calarco JP, Borges F, Donoghue MT, Van Ex F, Jullien PE, Lopes T, Gardner R et al (2012) Reprogramming of DNA methylation in pollen guides epigenetic inheritance via small RNA. Cell 151(1):194–205. doi:10.1016/j.cell.2012.09.001
Cervera MT, Ruiz-Garcia L, Martinez-Zapater JM (2002) Analysis of DNA methylation in Arabidopsis thaliana based on methylation-sensitive AFLP markers. Mol Genet Genomics 268(4):543–552. doi:10.1007/s00438-002-0772-4
Chu Y, Huang Q, Zhang B, Ding C, Su X (2014) Expression and molecular evolution of two DREB1 genes in black poplar (Populus nigra). PLoS ONE 9(6):e98334. doi:10.1371/journal.pone.0098334
Cokus SJ, Feng S, Zhang X, Chen Z, Merriman B, Haudenschild CD, Pradhan S, Nelson SF, Pellegrini M, Jacobsen SE (2008) Shotgun bisulphite sequencing of the Arabidopsis genome reveals DNA methylation patterning. Nature 452(7184):215–219. doi:10.1038/nature06745
Dowen RH, Pelizzola M, Schmitz RJ, Lister R, Dowen JM, Nery JR, Dixon JE, Ecker JR (2012) Widespread dynamic DNA methylation in response to biotic stress. Proc Natl Acad Sci USA 109(32):e2183–2191. doi:10.1073/pnas.1209329109
Eichten SR, Springer NM (2015) Minimal evidence for consistent changes in maize DNA methylation patterns following environmental stress. Front Plant Sci 6:308. doi:10.3389/fpls.2015.00308
Excoffier L, Smouse PE, Quattro JM (1992) Analysis of molecular variance inferred from metric distances among DNA haplotypes: application to human mitochondrial DNA restriction data. Genetics 131(2):479–491
Feng S, Cokus SJ, Zhang X, Chen PY, Bostick M, Goll MG, Hetzel J, Jain J, Strauss SH, Halpern ME, Ukomadu C, Sadler KC, Pradhan S, Pellegrini M, Jacobsen SE (2010) Conservation and divergence of methylation patterning in plants and animals. Proc Natl Acad Sci USA 107(19):8689–8694. doi:10.1073/pnas.1002720107
Gourcilleau D, Bogeat-Triboulot M-B, Le Thiec D, Lafon-Placette C, Delaunay A, El-Soud WA, Brignolas F, Maury S (2010) DNA methylation and histone acetylation: genotypic variations in hybrid poplars, impact of water deficit and relationships with productivity. Ann For Sci 67(2):208
Harmer SL, Hogenesch JB, Straume M, Chang HS, Han B, Zhu T, Wang X, Kreps JA, Kay SA (2000) Orchestrated transcription of key pathways in Arabidopsis by the circadian clock. Science 290(5499):2110–2113
Hashida SN, Uchiyama T, Martin C, Kishima Y, Sano Y, Mikami T (2006) The temperature-dependent change in methylation of the Antirrhinum transposon Tam3 is controlled by the activity of its transposase. Plant Cell 18(1):104–118. doi:10.1105/tpc.105.037655
Herrera CM, Bazaga P (2011) Untangling individual variation in natural populations: ecological, genetic and epigenetic correlates of long-term inequality in herbivory. Mol Ecol 20(8):1675–1688. doi:10.1111/j.1365-294X.2011.05026.x
Herrera CM, Medrano M, Bazaga P (2013) Epigenetic differentiation persists after male gametogenesis in natural populations of the perennial herb Helleborus foetidus (Ranunculaceae). PLoS ONE 8(7):e70730. doi:10.1371/journal.pone.0070730
Ikeda Y (2012) Plant imprinted genes identified by genome-wide approaches and their regulatory mechanisms. Plant Cell Physiol 53(5):809–816. doi:10.1093/pcp/pcs049
Karan R, DeLeon T, Biradar H, Subudhi PK (2012) Salt stress induced variation in DNA methylation pattern and its influence on gene expression in contrasting rice genotypes. PLoS ONE 7(6):e40203. doi:10.1371/journal.pone.0040203
Kumar SV, Wigge PA (2010) H2A. Z-containing nucleosomes mediate the thermosensory response in Arabidopsis. Cell 140(1):136–147
Law JA, Jacobsen SE (2010) Establishing, maintaining and modifying DNA methylation patterns in plants and animals. Nat Rev Genet 11(3):204–220. doi:10.1038/nrg2719
Li A, Hu BQ, Xue ZY, Chen L, Wang WX, Song WQ, Chen CB, Wang CG (2011) DNA methylation in genomes of several annual herbaceous and woody perennial plants of varying ploidy as detected by MSAP. Plant Mol Biol Rep 29:784–793. doi:10.1007/s11105-010-0280-3
Lira-Medeiros CF, Parisod C, Fernandes RA, Mata CS, Cardoso MA, Ferreira PC (2010) Epigenetic variation in mangrove plants occurring in contrasting natural environment. PLoS ONE 5(4):e10326. doi:10.1371/journal.pone.0010326
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(-delta delta C(T)) method. Methods 25(4):402–408. doi:10.1006/meth.2001.1262
Ma K, Song Y, Yang X, Zhang Z, Zhang D (2013) Variation in genomic methylation in natural populations of Chinese white poplar. PLoS ONE 8(5):e63977. doi:10.1371/journal.pone.0063977
Ma N, Chen W, Fan T, Tian Y, Zhang S, Zeng D, Li Y (2015) Low temperature-induced DNA hypermethylation attenuates expression of RhAG, an AGAMOUS homolog, and increases petal number in rose (Rosa hybrida). BMC Plant Biol 15:237. doi:10.1186/s12870-015-0623-1
Martienssen RA, Colot V (2001) DNA methylation and epigenetic inheritance in plants and filamentous fungi. Science 293(5532):1070–1074. doi:10.1126/science.293.5532.1070
Mirouze M, Paszkowski J (2011) Epigenetic contribution to stress adaptation in plants. Curr Opin Plant Biol 14(3):267–274. doi:10.1016/j.pbi.2011.03.004
Naydenov M, Baev V, Apostolova E, Gospodinova N, Sablok G, Gozmanova M, Yahubyan G (2015) High-temperature effect on genes engaged in DNA methylation and affected by DNA methylation in Arabidopsis. Plant Physiol Biochem 87:102–108. doi:10.1016/j.plaphy.2014.12.022
Omidvar V, Fellner M (2015) DNA methylation and transcriptomic changes in response to different lights and stresses in 7B-1 male-sterile tomato. PLoS ONE 10(4):e0121864. doi:10.1371/journal.pone.0121864
Pecinka A, Scheid OM (2012) Stress-induced chromatin changes: a critical view on their heritability. Plant Cell Physiol 53(5):801–808
Penterman J, Zilberman D, Huh JH, Ballinger T, Henikoff S, Fischer RL (2007) DNA demethylation in the Arabidopsis genome. Proc Natl Acad Sci USA 104(16):6752–6757. doi:10.1073/pnas.0701861104
Perez-Figueroa A (2013) MSAP: a tool for the statistical analysis of methylation-sensitive amplified polymorphism data. Mol Ecol Resour 13(3):522–527. doi:10.1111/1755-0998.12064
Porebski S, Bailey LG, Baum BR (1997) Modification of a CTAB DNA extraction protocol for plants containing high polysaccharide and polyphenol components. Plant Mol Biol Report 15(1):8–15
Raj S, Brautigam K, Hamanishi ET, Wilkins O, Thomas BR, Schroeder W, Mansfield SD, Plant AL, Campbell MM (2011) Clone history shapes Populus drought responses. Proc Natl Acad Sci USA 108(30):12521–12526. doi:10.1073/pnas.1103341108
Rico L, Ogaya R, Barbeta A, Penuelas J (2014) Changes in DNA methylation fingerprint of Quercus ilex trees in response to experimental field drought simulating projected climate change. Plant Biol 16(2):419–427. doi:10.1111/plb.12049
Sáez-Laguna E, Guevara MA, Diaz LM, Sanchez-Gomez D, Collada C, Aranda I, Cervera MT (2014) Epigenetic variability in the genetically uniform forest tree species Pinus pinea L. PLoS ONE 9(8):e103145. doi:10.1371/journal.pone.0103145
Salmon A, Clotault J, Jenczewski E, Chable V, Manzanares-Dauleux MJ (2008) Brassica oleracea displays a high level of DNA methylation polymorphism. Plant Sci 174(1):61–70
Schulz B, Eckstein RL, Durka W (2013) Scoring and analysis of methylation-sensitive amplification polymorphisms for epigenetic population studies. Mol Ecol Resour 13(4):642–653. doi:10.1111/1755-0998.12100
Song Y, Ma K, Bo W, Zhang Z, Zhang D (2012) Sex-specific DNA methylation and gene expression in andromonoecious poplar. Plant Cell Rep 31(8):1393–1405. doi:10.1007/s00299-012-1255-7
Vining KJ, Pomraning KR, Wilhelm LJ, Priest HD, Pellegrini M, Mockler TC, Freitag M, Strauss SH (2012) Dynamic DNA cytosine methylation in the Populus trichocarpa genome: tissue-level variation and relationship to gene expression. BMC Genomics 13:27. doi:10.1186/1471-2164-13-27
Wilkins O, Waldron L, Nahal H, Provart NJ, Campbell MM (2009) Genotype and time of day shape the Populus drought response. Plant J 60(4):703–715
Xia L, Ma S, Zhang Y, Wang T, Zhou M, Wang Z, Zhang J (2015) Daily variation in global and local DNA methylation in mouse livers. PLoS ONE 10(2):e0118101. doi:10.1371/journal.pone.0118101
Xing MQ, Zhang YJ, Zhou SR, Hu WY, Wu XT, Ye YJ, Wu XX, Xiao YP, Li X, Xue HW (2015) Global analysis reveals the crucial roles of DNA methylation during rice seed development. Plant Physiol 168(4):1417–1432. doi:10.1104/pp.15.00414
Yaish MW, Colasanti J, Rothstein SJ (2011) The role of epigenetic processes in controlling flowering time in plants exposed to stress. J Exp Bot 62(11):3727–3735. doi:10.1093/jxb/err177
Yu Y, Yang X, Wang H, Shi F, Liu Y, Liu J, Li L, Wang D, Liu B (2013) Cytosine methylation alteration in natural populations of Leymus chinensis induced by multiple abiotic stresses. PLoS ONE 8(2):e55772. doi:10.1371/journal.pone.0055772
Zhu JK (2009) Active DNA demethylation mediated by DNA glycosylases. Annu Rev Genet 43:143–166. doi:10.1146/annurev-genet-102108-134205
Author information
Authors and Affiliations
Corresponding authors
Additional information
Project funding: This study is supported by National Nonprofit Institute Research Grant of Chinese Academy of Forestry (TGB 2013010).
The online version is available at http:// www.springerlink.com
Corresponding editor: Zhu Hong
Rights and permissions
About this article
Cite this article
Diao, S., Wang, Y., Ding, C. et al. No consistent daily variation in DNA methylation detected in Populus nigra leaves by methylation-sensitive amplification polymorphism analysis. J. For. Res. 28, 653–660 (2017). https://doi.org/10.1007/s11676-016-0357-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11676-016-0357-4