Abstract
Tumor necrosis factor-alpha (TNF-α) inhibits osteogenic differentiation of murine bone marrow stromal cells, and transcription factor Runx2 serves as an essential regulation target in the process. The underlying mechanism may involve the regulation of Runx2 expression and the Runx2 activity in downstream gene transcription, which has not been fully elucidated. In this study, ST2 murine bone marrow-derived stromal cells were treated with bone morphogenetic protein-2 (BMP-2) and/or TNF-α in osteogenic medium, and the expression of Runx2 was estimated. Cells were transfected with Runx2, p65, inhibitor of κBα (IκBα), 9.0 kb bone sialoprotein (BSP) promoter-luciferase or osteoblast-specific cis-acting element 2 (OSE2)-luciferase reporter vectors, and then real time-PCR and dual luciferase analysis were used to investigate the effect of TNF-α on Runx2-activated osteogenic gene transcription and the molecular mechanism. We found that TNF-α inhibited BMP-2-induced osteogenic marker expression and both the spontaneous and BMP-2-induced Runx2 expression. TNF-α stimulation or overexpression of nuclear factor-kappa B (NF-κB) p65 subunit repressed the Runx2-activated BSP and osteocalcin (OC) transcriptions. The Runx2-induced 9.0 kb BSP promoter activity was attenuated by TNF-α or p65, while the OSE2 activity was not affected. Besides, blockage of NF-κB by IκBα overexpression eliminated these inhibitory effects of TNF-α on Runx2 signaling. These results suggest that in murine bone marrow stromal cells undergoing osteogenic differentiation, TNF-α and it activated NF-κB pathway inhibit the expression of Runx2 gene, and suppress the Runx2-mediated osteogenic gene transcription via the 9.0 kb BSP promoter.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Tumor necrosis factor-alpha (TNF-α) is a potent inflammatory cytokine and is acknowledged to play an essential role in chronic inflammatory bone loss, such as rheumatoid arthritis (RA) and periodontitis. Physiological endogenous levels of TNF-α are sufficient to reduce the maximum peak bone mineral density (BMD) and bone mass without affecting bone resorption (Li et al. 2007). Anti-TNF-α therapy has been proven to be efficacious in improving the BMD and bone turn over markers in patients with RA and ankylosing spondylitis (Henderson and Davis 2006; Seriolo et al. 2006; Sakthiswary and Das 2013), further approving the negative effect of TNF-α on osteogenesis. TNF-α has been shown to inhibit bone matrix synthesis by osteoblasts in early researches (Bertolini et al. 1986; Canalis 1987; Gilbert et al. 2000). Our previous study have demonstrated that TNF-α at concentrations ranging from 1 ng/ml to 100 ng/ml inhibits osteogenic differentiation of ST2 murine bone marrow stromal cells in a dose- and time-dependent manner without impairing cell survival (Huang et al. 2011). The inhibitory effect of TNF-α on osteogenic differentiation is independent of cell apoptosis (Gilbert et al. 2005). To date, multiple osteogenic signaling including bone morphogenetic protein (BMP) and Wnt have been shown to be down-regulated by TNF-α (Li et al. 2010; Huang et al. 2014; Liu et al. 2014), but the underlying molecular mechanism remains to be further elucidated.
Runx2 is an important transcription factor required for expressions of multiple osteogenic genes, including collagen I, osteopontin (OPN), alkaline phosphatase (ALP), bone sialoprotein (BSP) and osteocalcin (OC) (Ducy 2000). Runx2 functions by binding to specific sites in the osteogenic gene promoters and activating transcription. Osteoblast-specific cis-acting element 2 (OSE2) in promoters of multiple osteogenic genes is activated by Runx2 binding and enhances the gene transcription (Ducy et al. 1997; Jiang et al. 1999; Xiao et al. 1999). And the 9.0 kb BSP promoter containing several Runx2 binding sites contributes to transcription activation of BSP by Runx2 in vivo (Roca et al. 2005; Tu et al. 2008). Runx2 mutant mice show a complete lack of ossification because of the maturational arrest of osteoblasts, revealing that Runx2 is essential for osteoblast differentiation and bone formation (Komori et al. 1997). In vitro studies show that Runx2 expression is regulated at multiple levels during osteoblast differentiation, including transcription, mRNA stabilization and translation (Banerjee et al. 2001; Prince et al. 2001; Sudhakar et al. 2001). Interestingly, these studies also show discordance between Runx2 expression and activity (Banerjee et al. 2001; Prince et al. 2001), and in some cases the transactivation ability of Runx2 rather than Runx2 expression makes sense. For example, disturbed extracellular matrix-α2-intergrin interaction inhibits osteoblast differentiation through downregulation of Runx2 binding to OSE2 without a change in Runx2 mRNA or protein level (Xiao et al. 1998). Regulation of Runx2 activity in osteogenic gene transcription independent of Runx2 expression, also plays an essential role in osteoblast lineage commitment of MSCs (Shui et al. 2003).
TNF-α has been reported to inhibit Runx2 expression and promote Runx2 degradation in osteoblasts or MSCs during osteogenic differentiation (Gilbert et al. 2002; Kaneki et al. 2006; Lacey et al. 2009; Marupanthorn et al. 2015). However, the effects of TNF-α on Runx2 downstream signaling such as OSE2 and BSP promoter activities, have not been elucidated yet. Besides, nuclear factor-kappa B (NF-κB) activated by TNF-α is found to mediate the inhibitory effect of TNF-α on osteogenic differentiation (Hayden and Ghosh 2014). In our previous study, TNF-α activates NF-κB signaling by increasing the p65 subunit nuclear expression, which is inhibited by overexpression of inhibitor of NF-κBα (IκBα) (Huang et al. 2011). Whether the NF-κB pathway is involved in TNF-α-induced Runx2 signaling regulation is also unclear.
In this study, we performed a series of in vitro experiments to explore the changes of Runx2 expression and Runx2-induced transcription activity in ST2 murine marrow stromal cells with TNF-α treatment. In addition, the role of NF-κB in the regulation of Runx2 signaling was also preliminarily investigated.
Methods and materials
Materials and reagents
Expression plasmids PBABE-puro-IκBα-mut, RelA/p65 cFlag pcDNA3 and empty vectors were obtained from Addgene (Cambridge, MA). pGL3-BSP9.0luc reporter vector was a generous gift from Prof. Chen (Tufts University School of Dental Medicine), while OSE2-luciferase reporter and PCMV5-Runx2/PCMV5 vectors were provided by Dr. Mikami (Nihon University School of Dentistry).
Cell culture
Murine bone marrow-derived stromal cell line ST2 (Riken BioResource Center, Tsukuba, Ibaraki, Japan) were maintained in α-minimum essential medium (α-MEM; HyClone, Logan, UT) supplemented with 10 % (v/v) fetal bovine serum (FBS; HyClone), 100 U/ml penicillin (Invitrogen, Carlsbad, CA) and 100 μg/ml streptomycin (Invitrogen) in humidified 5 % CO2 air at 37 °C. For osteogenic differentiation, osteogenic medium containing additional 10 nM dexamethasone, 10 mM β-glycerophosphate and 50 μg/ml ascorbic acid (Sigma-Aldrich, St. Louis, MO) was used as previously described (Huang et al. 2011; Bakhtina et al. 2014).
Cell transfection and dual luciferase assay
ST2 cells were transfected with RelA/p65 cFlag pcDNA3 or empty vector using Fugene HD transfection reagent (Roche Diagnostics, Indianapolis, IN) according to the manufacturers’ instruction. Cells were then incubated with 400 μg/ml G418 (Sigma-Aldrich) for 14 d. The stably transfected cells named ST2-p65 and ST2-control1 respectively were further identified by western blot and used in this study. Retroviral stocks containing pBABE-puro-IκBα-mut and the pBABE-puro empty viral vector were prepared and infected the cells as stated in our previous study (Huang et al. 2011). After selection and identification, the stably transfected cells named ST2-IκBα and ST2-control2 were used in the following experiments. In experiments with Runx2 overexpression, cells were transiently transfected with the pCMV5-Runx2 or pCMV5 plasmid using Fugene HD. For luciferase assays, cells were co-transfected with pGL3-BSP9.0luc or OSE2-luciferase reporter and Renilla luciferase vectors (20:1), and then incubated with or without TNF-α. Quantitative analyses of relative luciferase activities were performed using Dual-Luciferase Reporter Assay System (Promega, Madison, WI) 48 h after transfection.
Real time reverse transcription-polymerase chain reaction (RT-PCR)
Real time RT-PCR analysis was performed as previously stated (Mikami et al. 2008). Briefly, total RNA was isolated with TRIzol Reagent (Invitrogen) and reverse-transcribed to cDNA with a RevertAid First Strand cDNA Synthesis Kit (Fermentas, Shenzhen, China). Real time PCR was performed using THUNDERBIRD SYBR qPCR Mix (TOYOBO, Osaka, Japan) on a Light Cycler II 480 (Roche Diagnostics, Indianapolis, IN). Sequences of the primers for murine Runx2, BSP, OC and GAPDH were listed as following:
-
5′-CCCAGCCACCTTTACCTACA-3′ and 5′-TATGGAGTGCTGCTGGTCTG-3′ (Runx2); 5′-GAAGCAGGTGCAGAAGGAAC-3′ and
-
5′-ACTCAACGGTGCTGCTTTTT-3′ (BSP); 5′-AAGCAGGAGGGCAATAAGGT-3′ and 5′-TAGGCGGTCTTCAAGCCATA-3′ (OC);
-
5′-ACCACAGTCCATGCCATCAC-3′ and 5′-TCCACCACCCTGTTGCTGTA-3′
(GAPDH). GAPDH was used as an internal control.
Western blot
Whole cell lysates were prepared as described previously (Huang et al. 2011). Protein samples were quantified by a BCA Protein Assay Kit (Beyotime, Jiangsu, China), separated using 10–12 % Bis-Tris gels (Beyotime, Jiangsu, China) and subsequently transferred onto a 0.45 μm Immobilon-NC Transfer Membrane (Merck Millipore, Darmstadt, Germany). Antibody for IκBα (1:500) was obtained from Sigma-Aldrich. Antibodies for Runx2 (1:200), β-actin (1:1000) and p65 (1:300) were purchased from Santa Cruz Biotechnologies (Dallas, Texas). The secondary antibodies were horseradish peroxidase (HRP)-linked goat anti-rabbit IgG (H + L) and goat anti-mouse IgG (H + L) (Zhongshan Golden Bridge, Beijing, China). Blots were visualized using ECL chemiluminescence reagents from Pierce Biotechnology (Rockford, IL).
Statistical analysis
Data were shown as means ± standard error of the mean (SEM) from at least three experiments. T-test and One-way Analysis of Variance (ANOVA) were used to evaluate significance using SPSS 19.0 Statistics Software (SPSS Inc., Chicago, IL). P < 0.05 was considered statistically significant.
Results
Protein expression in stably transfected cells
The ST2 cells transfected with pBABE-puro-IκBα-mut or pBABE-puro were collected and subjected to western blot to evaluate the protein levels of IκBα. Protein expression of IκBα was increased by 2.8 folds in ST2-IκBα cells compared with ST2-control2 cells, as shown in our previous study (data not shown) (Huang et al. 2011). The protein expression of p65 protein was also estimated in ST2-p65 and ST2-control1 cells by western blot, which showed that p65 protein was significantly elevated in ST2-p65 cells compared with that in ST2-control1 cells (Fig. 1). The target genes were successfully transduced and the proteins were stably expressed in transfected cells.
Protein expression in stably transfected cells. ST2 cells transfected with p65 or empty vector were named ST2-p65 and ST2-control1 cells respectively. After selection, the protein expression of p65 protein was estimated in ST2-p65 and ST2-control1 cells by Western blot. *, p < 0.05 vs. control group.
TNF-α inhibited BMP-2-induced osteogenic differentiation through NF-κB
To determine the effects of TNF-α on BMP-2-induced osteogenic gene expression, ST2-control2 and ST2-IκBα cells were treated with 100 ng/ml BMP-2 and/or 100 ng/ml TNF-α in osteogenic medium for 3 d, and real time PCR was used to evaluate the mRNA expressions of BSP and OC. ST2-IκBα and ST2-control2 cells without any treatment were used as control respectively. The mRNA expressions of BSP and OC were elevated in BMP-2-treated cells, which were reduced by additional TNF-α in ST2-control2 cells (Fig. 2a, b ). In the ST2-IκBα cells, BMP-2 showed similar positive effects on the expressions of BSP and OC as in the ST2-congtrol2 cells, while additional TNF-α did not exhibit inhibition on the BMP-2-induced osteogenic gene expressions (Fig. 2a, b ). We then identified the effects of p65 subunit using ST2-p65 and ST2-control1 cells treated with 100 ng/ml BMP-2 in osteogenic medium. After 3 d, BSP and OC expressions were increased in both groups of cells, but in the p65 overexpressed cells, the positive effects were weaker than those in the control cells (Fig. 2c, d ). TNF-α inhibited BMP-2-induced osteogenic gene expression through activation of NF-κB pathway.
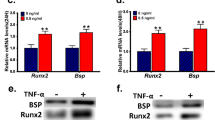
TNF-α inhibited BMP-2-induced osteogenic gene expression. (a, b) ST2-IκBα and ST2-control2 cells were treated with 100 ng/ml BMP-2 and/or 100 ng/ml TNF-α in osteogenic medium, and real time PCR was used to evaluate the mRNA expressions of BSP (a) and OC (b) after 3 d of treatment. (c, d) ST2-p65 and ST2-control1 cells were treated with or without BMP-2 for 3 d, and real time PCR was used to detect the mRNA expressions of BSP (c) and OC (d). *, p < 0.05 vs. control group; #, p < 0.05 vs. BMP-2 group.
TNF-α inhibited spontaneous and BMP-2-induced Runx2 expression though NF-κB
The ST2 cells were incubated with or without 100 ng/ml TNF-α in osteogenic medium for 3 d. Real time RT-PCR result showed significant decreased expression of Runx2 gene in cells treated with TNF-α compared to those without TNF-α treatment, which was consistent with our previous finding (Fig. 3a ). And the protein expression of Runx2 showed a similar change in cells treated with TNF-α by western blot analysis (Fig. 3b ).
TNF-α inhibited spontaneous and BMP-2-induced Runx2 expression. (a, b) ST2 cells were treated with 100 ng/ml TNF-α in osteogenic medium for 3 d, and mRNA (a) and protein (b) levels of Runx2 were analyzed by real time PCR and Western blot. *, p < 0.05 vs. control group. (c) ST2-IκBα and ST2-control2 cells were treated with 100 ng/ml BMP-2 and/or 100 ng/ml TNF-α for 3 d, and mRNA expression of Runx2 was analyzed. *, p < 0.05 vs. control group; #, p < 0.05 vs. BMP-2 group.
To investigate the effect of TNF-α on BMP-2-induced Runx2 expression, ST2-control2 and ST2-IκBα cells were treated with 100 ng/ml BMP-2 and/or 100 ng/ml TNF-α in osteogenic medium and real time RT-PCR was performed after 3 d of treatment. ST2-IκBα and ST2-control2 cells without any treatment were used as control respectively. BMP-2 treatment increased the Runx2 mRNA level in both cell lines. Additional TNF-α significantly reduced the mRNA expression of Runx2 induced by BMP-2 in ST2-control2 cells, while TNF-α did not show negative effect on the BMP-2-induced Runx2 expression in ST2-IκBα cells (Fig. 3c ). TNF-α inhibited BMP-2-induced Runx2 expression, and blockage of NF-κB signaling eliminated this inhibitory effect of TNF-α.
TNF-α inhibited Runx2-activated osteogenic gene expression through NF-κB
To determine the effects of TNF-α on Runx2-activated osteogenic gene expression, the ST2-IκBα and ST2-control2 cells were transiently transfected with pCMV5-Runx2/pCMV5 and treated with or without TNF-α. Real time RT-PCR was performed to analyze the mRNA expressions of BSP and OC, and the ST2-IκBα and ST2-control2 cells without any treatment were used as control respectively. Runx2 overexpression by gene transfection enhanced BSP and OC mRNA expression, and TNF-α treatment remarkably depressed the positive effects of Runx2 on BSP and OC mRNA expression in the ST2-control2 cells (Fig. 4a, b ). Moreover, Runx2 overexpression showed similar effects on osteogenic gene transcription in ST2-IκBα cells as in ST2-control2 cell. However, no difference was found in the osteogenic gene transcriptions between ST2-IκBα cells with and without TNF-α treatment (Fig. 4a, b ). Our results showed that TNF-α decreased Runx2-activated osteogenic gene expression though NF-κB pathway.
TNF-α inhibited Runx2-induced osteogenic differentiation through NF-κB. (a) The ST2-IκBα and ST2-control2 cells were transiently transfected with pCMV5-Runx2/pCMV5 and then treated with 100ng/ml TNF-α for 3 d. Real time PCR was used to evaluate the mRNA levels of osteogenic gene BSP (a) and OC (b). *, p < 0.05 vs. pCMV5 group; #p < 0.05 vs. pCMV5-Runx2 group.
TNF-α inhibited Runx2-induced 9.0 kb BSP promoter-mediated transcription activity rather than OSE2-mediated transcription activity via NF-κB
In view of our findings that TNF-α inhibited Runx2-activated osteogenic gene expression, we used the pGL3-BSP9.0luc and OSE2-luciferase reporters to explore the downstream signaling of Runx2 affected by TNF-α. Luciferase reporter vectors and pCMV5-Runx2/pCMV5 vector were transiently co-transfected into the ST2-control2 cells as well as the ST2-IκBα cells. Then cells were treated with or without 100 ng/ml TNF-α in osteogenic medium. Relative luciferase activity was analyzed 48 h after transfection to evaluate the promoter regions mediated transcription activities. The results showed that the 9.0 kb BSP promoter-mediated transcription was activated by Runx2 overexpression, which was significantly inhibited by TNF-α treatment in ST2-control2 cells (Fig. 5a ). Furthermore, inhibition of NF-κB signaling by IκBα overexpression completely eliminated the inhibitory effects of TNF-α on Runx2-activated 9.0 kb BSP promoter. To further identify the role of NF-κB pathway, the ST2-p65 and ST2-control1 cells were co-transfected with pCMV5-Runx2 and pGL3-BSP9.0luc reporter vectors, and cultured in osteogenic medium for luciferase activity analysis. P65 overexpression depressed the Runx2-induced 9.0 kb BSP promoter activity (Fig. 5b ). Our results indicate that TNF-α inhibits the activation of 9.0 kb BSP promoter by Runx2 through the NF-κB pathway.
TNF-α inhibited Runx2-induced 9.0 kb BSP promoter activity through NF-κB. (a) The ST2-IκBα and ST2-control2 cells were transiently co-transfected with pCMV5-Runx2/pCMV5, pGL3-BSP9.0luc reporter and Renilla luciferase vectors, and then cells were treated with or without 100 ng/ml TNF-α. Dual luciferase assay was used to analyze the relative luciferase activities 48 h after transfection. *, p < 0.05 vs. pCMV5 group; #, p < 0.05 vs. pCMV5-Runx2 group. (b) ST2-p65 and ST2-control1 cells were transiently transfected with BSP9.0luc reporter and Renilla luciferase vectors. Relative luciferase activities were analyzed after 48 h. *, p < 0.05 vs. control group.
On the other hand, Runx2 overexpression was capable to induce OSE2-mediated transcription, but the TNF-α did not exhibit negative effect on the Runx2-induced OSE2 activity in ST2-control2 cells (Fig. 6a ). Neither activation nor blockage of NF-κB signaling led to differences in OSE2-mediated luciferase expression (Fig.6a, b ). Runx2/OSE2-mediated transcription was not affected by TNF-α.
TNF-α and NF-κB did not affect the Runx2-induced OSE2 activity. (a) The ST2-IκBα and ST2-control2 cells were transiently co-transfected with pCMV5-Runx2/pCMV5, OSE2-luciferase reporter and Renilla luciferase vectors, and then cells were treated with or without 100 ng/ml TNF-α. Dual luciferase assay was used to analyze the relative luciferase activities 48 h after transfection. *, p < 0.05 vs. pCMV5 group. (b) ST2-p65 and ST2-control1 cells were transiently transfected with OSE2-luciferase reporter and Renilla luciferase vectors. Relative luciferase activities were analyzed after 48 h.
Discussion
Osteogenic differentiation is guided physiologically by multiple signaling pathways, including BMP and Wnt pathways. The activated signaling cascades converge on the transcription factor Runx2 to regulate osteogenic gene expression (Franceschi et al. 2003). Blockage of Runx2 signaling substantially antagonizes the anabolic effects of osteogenic growth factors (Gori et al. 1999; Kook et al. 2015). In inflammatory microenvironment like periodontitis, inflammatory cytokines such as TNF-α influence the function of adult MSCs in tissue repair. TNF-α has been proved to be a potent inhibitor of osteogenic differentiation (Gilbert et al. 2000). According to our previous study, TNF-α at the concentration of 100 ng/ml inhibits osteogenic differentiation of ST2 cells, evidenced by the reduced expression of osteogenic marker, decreased alkaline phosphatase activity and mineralization (Huang et al. 2011). In this study, the anabolic effects of growth factor BMP-2 on osteogenic gene expression were also significantly depressed by TNF-α in murine bone marrow stromal cells, which had been previously reported in murine pre-osteoblasts and myoblasts (Mukai et al. 2007; Huang et al. 2014). And then we focused on the changes of transcription factor Runx2 in response to TNF-α, in the expression level and regulation of downstream signaling. TNF-α inhibited spontaneous and BMP-2-induced Runx2 expression, and the inhibition of Runx2 expression by TNF-α was eliminated by the NF-κB inhibitor IκBα. Combined with our previous findings, it can be indicated that NF-κB mediates the inhibitory effects of TNF-α on spontaneous and BMP-2-induced Runx2 expression.
As previous studies showed, osteogenic differentiation was primarily associated with increase in Runx2 activity rather than synthesis of Runx2 in human bone marrow-derived MSCs (Shui et al. 2003). Similar regulation pattern also exists in murine osteoblasts (Xiao et al. 1998; Tarapore et al. 2016). We first found in murine ST2 cells that besides the negative effect on Runx2 expression, TNF-α also inhibited Runx2-activated transcription of osteogenic gene BSP and OC. Using promoter-luciferase reporter vector to transfect cells, we further identified that TNF-α repressed the 9.0 kb BSP promoter-mediated transcription activation by Runx2. There are two possible mechanisms for the phenomenon and firstly, TNF-α affects the binding ability of BSP promoter to Runx2. Other transcription factor such may change the promoter conformation after binding, and this transformation impairs the affinity of Runx2 for its binding sites (Franceschi et al. 2003). And secondly, Runx2 shows repressive effect on BSP promoter (Javed et al. 2001). By animal studies using transgenic or gene knock-out mouse, researchers found that Runx2-binding sites in proximal region (within 4.8 kb) of BSP promoter exhibited enhancer activity (Roca et al. 2005), while Runx2-binding sites in distal region (between 4.8 kb and 9.0 kb) of BSP promoter repressed transcription (Tu et al. 2008). Co-repressors interacted with Runx2 may facilitate the inhibitory effect of Runx2 on transcription activity through the distal region of BSP promoter. For example, Histone deacetylase 3 (HDAC3) was shown to act as a Runx2 co-repressor to suppress BSP gene expression in Saos-2 human osteosarcoma cells (Lamour et al. 2007). From the above evidences, TNF-α inhibited Runx2-activated BSP transcription by repressing the 9.0 kb BSP promoter activity. The regulation of Runx2 downstream signaling in BSP transcription involves direct and indirect mechanisms, and NF-κB may serve as a major regulator in response to TNF-α.
OSE2 is the main Runx2-binding site in OC promoter that regulates OC transcription (Lian et al. 2003). Previous study showed a post-translational regulation of Runx2 activity in OSE2-mediated transcription, such as phosphorylation of Runx2 protein (Shui et al. 2003). But in this study, TNF-α and it activated NF-κB signal inhibited Runx2-activated OC transcription without suppressive effect on Runx2 up-regulated OSE2 activity. It is concluded that the inhibition of Runx2 activity in OC transcription needs the rest regions in OC promoter and is not through the post-translational mechanism. Other transcriptional regulatory elements within OC promoter may take part in the inhibitory effects of TNF-α and NF-κB signaling on Runx2-activated OC expression, which needs future investigation for confirmation.
In summary, TNF-α activates NF-κB signaling, which inhibits both the Runx2 expression and Runx2-activated transcription of osteogenic genes during osteogenic differentiation of murine bone marrow stromal cells. The down-regulation of 9.0 kb BSP promoter activity induced by Runx2 contributes to the inhibitory effects of TNF-α and NF-κB. Further research on the interactions among Runx2, NF-κB and the promoter of osteogenic gene will be helpful in understanding of the definite mechanism of Runx2 activity regulation by TNF-α within osteogenic differentiation.
Reference
Bakhtina A, Tohfafarosh M, Lichtler A, Arinzeh TL (2014) Characterization and differentiation potential of rabbit mesenchymal stem cells for translational regenerative medicine. In Vitro Cell Dev Biol Anim 50:251–260
Banerjee C, Javed A, Choi JY, Green J, Rosen V, van Wijnen AJ, Stein JL, Lian JB, Stein GS (2001) Differential regulation of the two principal Runx2/Cbfa1 n-terminal isoforms in response to bone morphogenetic protein-2 during development of the osteoblast phenotype. Endocrinology 142:4026–4039
Bertolini DR, Nedwin GE, Bringman TS, Smith DD, Mundy GR (1986) Stimulation of bone resorption and inhibition of bone formation in vitro by human tumour necrosis factors. Nature 319:516–518
Canalis E (1987) Effects of tumor necrosis factor on bone formation in vitro. Endocrinology 121:1596–1604
Ducy P (2000) Cbfa1: a molecular switch in osteoblast biology. Dev Dyn Off Publ Am Assoc Anatomists 219:461–471
Ducy P, Zhang R, Geoffroy V, Ridall AL, Karsenty G (1997) Osf2/Cbfa1: a transcriptional activator of osteoblast differentiation. Cell 89:747–754
Franceschi RT, Xiao G, Jiang D, Gopalakrishnan R, Yang S, Reith E (2003) Multiple signaling pathways converge on the Cbfa1/Runx2 transcription factor to regulate osteoblast differentiation. Connect Tissue Res 44(Suppl 1):109–116
Gilbert L, He X, Farmer P, Boden S, Kozlowski M, Rubin J, Nanes MS (2000) Inhibition of osteoblast differentiation by tumor necrosis factor-alpha. Endocrinology 141:3956–3964
Gilbert L, He X, Farmer P, Rubin J, Drissi H, van Wijnen AJ, Lian JB, Stein GS, Nanes MS (2002) Expression of the osteoblast differentiation factor RUNX2 (Cbfa1/AML3/Pebp2alpha a) is inhibited by tumor necrosis factor-alpha. J Biol Chem 277:2695–2701
Gilbert LC, Rubin J, Nanes MS (2005) The p55 TNF receptor mediates TNF inhibition of osteoblast differentiation independently of apoptosis. Am J Phys Endocrinol Metab 288:E1011–E1018
Gori F, Thomas T, Hicok KC, Spelsberg TC, Riggs BL (1999) Differentiation of human marrow stromal precursor cells: bone morphogenetic protein-2 increases OSF2/CBFA1, enhances osteoblast commitment, and inhibits late adipocyte maturation. J Bone Miner Res Off J Am Soc Bone Miner Res 14:1522–1535
Hayden MS, Ghosh S (2014) Regulation of NF-kappaB by TNF family cytokines. Semin Immunol 26:253–266
Henderson C, Davis JC (2006) Drug insight: anti-tumor-necrosis-factor therapy for ankylosing spondylitis. Nat Clin Pract Rheumatol 2:211–218
Huang H, Zhao N, Xu X, Xu Y, Li S, Zhang J, Yang P (2011) Dose-specific effects of tumor necrosis factor alpha on osteogenic differentiation of mesenchymal stem cells. Cell Prolif 44:420–427
Huang RL, Yuan Y, Tu J, Zou GM, Li Q (2014) Opposing TNF-alpha/IL-1beta- and BMP-2-activated MAPK signaling pathways converge on Runx2 to regulate BMP-2-induced osteoblastic differentiation. Cell Death Dis 5:e1187
Javed A, Barnes GL, Jasanya BO, Stein JL, Gerstenfeld L, Lian JB, Stein GS (2001) Runt homology domain transcription factors (Runx, Cbfa, and AML) mediate repression of the bone sialoprotein promoter: evidence for promoter context-dependent activity of Cbfa proteins. Mol Cell Biol 21:2891–2905
Jiang H, Sodek J, Karsenty G, Thomas H, Ranly D, Chen J (1999) Expression of core binding factor Osf2/Cbfa-1 and bone sialoprotein in tooth development. Mech Dev 81:169–173
Kaneki H, Guo R, Chen D, Yao Z, Schwarz EM, Zhang YE, Boyce BF, Xing L (2006) Tumor necrosis factor promotes Runx2 degradation through up-regulation of Smurf1 and Smurf2 in osteoblasts. J Biol Chem 281:4326–4333
Komori T, Yagi H, Nomura S, Yamaguchi A, Sasaki K, Deguchi K, Shimizu Y, Bronson RT, Gao YH, Inada M, Sato M, Okamoto R, Kitamura Y, Yoshiki S, Kishimoto T (1997) Targeted disruption of Cbfa1 results in a complete lack of bone formation owing to maturational arrest of osteoblasts. Cell 89:755–764
Kook SH, Heo JS, Lee JC (2015) Crucial roles of canonical Runx2-dependent pathway on Wnt1-induced osteoblastic differentiation of human periodontal ligament fibroblasts. Mol Cell Biochem 402:213–223
Lacey DC, Simmons PJ, Graves SE, Hamilton JA (2009) Proinflammatory cytokines inhibit osteogenic differentiation from stem cells: implications for bone repair during inflammation. Osteoarthritis Cartilage / OARS, Osteoarthritis Res Soc 17:735–742
Lamour V, Detry C, Sanchez C, Henrotin Y, Castronovo V, Bellahcene A (2007) Runx2- and histone deacetylase 3-mediated repression is relieved in differentiating human osteoblast cells to allow high bone sialoprotein expression. J Biol Chem 282:36240–36249
Li W, Yu B, Li M, Sun D, Hu Y, Zhao M, Cui CB, Hou S (2010) NEMO-binding domain peptide promotes osteoblast differentiation impaired by tumor necrosis factor alpha. Biochem Biophys Res Commun 391:1228–1233
Li Y, Li A, Strait K, Zhang H, Nanes MS, Weitzmann MN (2007) Endogenous TNFalpha lowers maximum peak bone mass and inhibits osteoblastic Smad activation through NF-kappaB. J Bone Miner Res Off J Am Soc Bone Miner Res 22:646–655
Lian JB, Stein JL, Stein GS, van Wijnen AJ, Montecino M, Javed A, Gutierrez S, Shen J, Zaidi SK, Drissi H (2003) Runx2/Cbfa1 functions: diverse regulation of gene transcription by chromatin remodeling and co-regulatory protein interactions. Connect Tissue Res 44(Suppl 1):141–148
Liu W, Konermann A, Guo T, Jager A, Zhang L, Jin Y (2014) Canonical Wnt signaling differently modulates osteogenic differentiation of mesenchymal stem cells derived from bone marrow and from periodontal ligament under inflammatory conditions. Biochim Biophys Acta 1840:1125–1134
Marupanthorn K, Tantrawatpan C, Tantikanlayaporn D, Kheolamai P, Manochantr S (2015) The effects of TNF-alpha on osteogenic differentiation of umbilical cord derived mesenchymal stem cells. J Med Assoc Thail = Chotmaihet Thangphaet 98(Suppl 3):S34–S40
Mikami Y, Takahashi T, Kato S, Takagi M (2008) Dexamethasone promotes DMP1 mRNA expression by inhibiting negative regulation of Runx2 in multipotential mesenchymal progenitor, ROB-C26. Cell Biol Int 32:239–246
Mukai T, Otsuka F, Otani H, Yamashita M, Takasugi K, Inagaki K, Yamamura M, Makino H (2007) TNF-alpha inhibits BMP-induced osteoblast differentiation through activating SAPK/JNK signaling. Biochem Biophys Res Commun 356:1004–1010
Prince M, Banerjee C, Javed A, Green J, Lian JB, Stein GS, Bodine PV, Komm BS (2001) Expression and regulation of Runx2/Cbfa1 and osteoblast phenotypic markers during the growth and differentiation of human osteoblasts. J Cell Biochem 80:424–440
Roca H, Phimphilai M, Gopalakrishnan R, Xiao G, Franceschi RT (2005) Cooperative interactions between RUNX2 and homeodomain protein-binding sites are critical for the osteoblast-specific expression of the bone sialoprotein gene. J Biol Chem 280:30845–30855
Sakthiswary R, Das S (2013) The effects of TNF alpha antagonist therapy on bone metabolism in rheumatoid arthritis: a systematic review. Curr Drug Targets 14:1552–1557
Seriolo B, Paolino S, Sulli A, Ferretti V, Cutolo M (2006) Bone metabolism changes during anti-TNF-alpha therapy in patients with active rheumatoid arthritis. Ann N Y Acad Sci 1069:420–427
Shui C, Spelsberg TC, Riggs BL, Khosla S (2003) Changes in Runx2/Cbfa1 expression and activity during osteoblastic differentiation of human bone marrow stromal cells. J Bone Miner Res Off J Am Soc Bone Miner Res 18:213–221
Sudhakar S, Li Y, Katz MS, Elango N (2001) Translational regulation is a control point in RUNX2/Cbfa1 gene expression. Biochem Biophys Res Commun 289:616–622
Tarapore RS, Lim J, Tian C, Pacios S, Xiao W, Reid D, Guan H, Mattos M, Yu B, Wang CY, Graves DT (2016) NF-kappaB has a direct role in inhibiting bmp- and Wnt-induced matrix protein expression. J Bone Miner Res Off J Am Soc Bone Miner Res 31:52–64
Tu Q, Zhang J, Paz J, Wade K, Yang P, Chen J (2008) Haploinsufficiency of Runx2 results in bone formation decrease and different BSP expression pattern changes in two transgenic mouse models. J Cell Physiol 217:40–47
Xiao G, Wang D, Benson MD, Karsenty G, Franceschi RT (1998) Role of the alpha2-integrin in osteoblast-specific gene expression and activation of the Osf2 transcription factor. J Biol Chem 273:32988–32994
Xiao ZS, Hinson TK, Quarles LD (1999) Cbfa1 isoform overexpression upregulates osteocalcin gene expression in non-osteoblastic and pre-osteoblastic cells. J Cell Biochem 74:596–605
Acknowledgment
This study was supported by Natural Science Foundation of China (No. 81271141).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Editor: Tetsuji Okamoto
Rights and permissions
About this article
Cite this article
Ye, X., Huang, H., Zhao, N. et al. Inhibition of Runx2 signaling by TNF-α in ST2 murine bone marrow stromal cells undergoing osteogenic differentiation. In Vitro Cell.Dev.Biol.-Animal 52, 1026–1033 (2016). https://doi.org/10.1007/s11626-016-0068-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11626-016-0068-3