Abstract
Onion (Allium cepa) is an important globally cultivated crop and is known to be susceptible to purple blotch caused by Alternaria porri. The causal pathogens of blight symptoms from onion in Myanmar were isolated and identified. In addition to Stemphylium vesicarium, a large-spored Alternaria with unique morphology as well as a small-spored Alternaria were obtained. To identify the two Alternaria fungal pathogens, morphological characteristics and molecular phylogenies based on multigene sequence analysis of the internal transcribed spacer of ribosomal DNA (ITS) region, glyceraldehyde-3-phosphate dehydrogenase (GAPDH), Alternaria major allergen (ALT), translation-elongation factor 1 (EF1-α), and RNA polymerase second largest subunit (RPB2) genes were used. This revealed the presence of a small-spored Alternaria, A. burnsii, and a new large-spored species here described as A. cepae sp. nov. The novel species is morphologically distinct from its closely related species of A. montanica. Pathogenicity assays revealed that Stemphylium vesicarium, A. burnsii, and A. cepae were the causal agents of the onion leaf blight of the current study, and that A. cepae exhibited the most virulence. However, A. porri, which has been reported as the most important onion pathogen worldwide, was absent during this investigation.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Onion (Allium cepa L., Alliaceae) is indigenous to central Asia and is widely cultivated globally. Its production is reduced by numerous diseases at various growth stages. Among these, purple blotch caused by Alternaria porri (Ellis) is one of the most destructive diseases, occurring on the foliage and reducing crop quality and yield (Gupta and Gupta 2013; Woudenberg et al. 2014). It typically appears as a brownish-purple lesion with concentric rings, leading to plant death when severe (Black et al. 2012). Another pathogen, Stemphylium vesicarium (Wall.), is common in warm and moist environments, causing damage on its own or in conjunction with A. porri (Aveling et al. 1993).
The genus Alternaria, for which there are about 589 legitimate species epithets, currently contains 366 accepted and recognizable species (Wijayawardene et al. 2020). These species, commonly found as plant pathogens, lead to substantial economic losses by causing leaf blight or leaf spot on various crops and as post-harvest pathogens (Andersen et al. 2001; Thomma 2003). Advanced analytical methods using molecular approaches have become essential for separating Alternaria species into sections and identifying them to species level. These analyses use multiple gene loci of the internal transcribed spacer regions 5.8S rDNA (ITS), 18S rDNA (SSU), 28S rDNA (LSU), glyceraldehyde-3-phosphate dehydrogenase (GAPDH), RNA polymerase second largest subunit (RPB2), translation-elongation factor 1 (EF1-α), Alternaria major allergen gene (ALT), endopolygalacturonase (endoPG), and an anonymous gene region (OPA10-2) (Pryor and Gilbertson 2000; Hong et al. 2005; Lawrence et al. 2011, 2013; Woudenberg et al. 2013, 2014, 2015). Stemphylium. was proposed by Wallroth (1833) with Stemphylium botryosum as the type species. It is a monophyletic genus of filamentous ascomycetes comprising pathogens and saprobes with a wide range of host plants (Farr et al. 1989; Köhl et al. 2009; Crous et al. 2016; Woudenberg et al. 2017). There are approximately 200 reported names currently described as recognizable taxa of Stemphylium (Woudenberg et al. 2017).
In studies on onion diseases in India, Alternaria purple blotch and Stemphylium blight have been considered important foliage diseases in most cultivation regions (Gupta and Pathak 1988; Suheri and Price 2000; Mathur and Sharma 2006). In contrast, Myanmar has recorded few fungal diseases on cultivated crops. Thaung (1970) reported plant diseases on various hosts in Myanmar, and A. porri was identified as a pathogen causing onion purple blotch nationwide. However, to our knowledge, onion diseases have not been reported in Myanmar since then. Furthermore, yield losses caused by fungal diseases have not been well documented in Myanmar. During the 2018–2019 onion growing season, a disease outbreak occurred in Ywarthitgyi village in Naypyidaw, the capital of Myanmar. The typical symptoms were dark brown necrotic lesions, chlorotic foliage, and foliage dieback, distinct from purple blotch disease. As a consequence, 80% of the cultivated onion in that village was affected by the disease and production was shut down in 2019. The study was carried out to identify the causal agents of this onion disease based on morphological characters, molecular analyses, and pathogenicity tests.
Materials and methods
Sample collection and fungal isolation
In February 2019, leaves from six symptomatic plants were randomly sampled from three onion plantations in Ywarthitgyi village, Naypyidaw, Myanmar. Small leaf segments with disease lesions were placed in Petri dishes on moist filter paper and kept at 25℃ in the dark for 1 − 2 days to observe the causal fungal pathogens. Single spores emerging from the margins of the disease lesions were isolated using a sterile glass needle under a stereoscopic microscope and then transferred onto potato dextrose agar (PDA, Difco™, Detroit, MI, USA). Fifteen pure cultures representative of all plants were obtained (Table 1) and deposited in the Fungal Herbarium at Yangtze University, Jingzhou, China, and in the Agricultural Culture Collection of China (ACCC), Beijing, China.
Morphological characterization
Two Alternaria strains, based on the colony and conidial characters, YZU 191023 representing large-spored and YZU 191042 representing small-spored, were selected for the further identification process. Colony characteristics were determined using PDA at 25℃ in the dark for 1 week (Deng et al. 2018). Morphology was then determined and characterized using a mycological color chart (Rayner 1970). The strains, grown on potato carrot agar (PCA) at 22℃ under an 8 h photoperiod for 1 week (Simmons 2007), were used to determine conidial characteristics. The conidia (n = 50) were mounted in lactophenol picric acid solution to be photographed for characterization, using a Nikon ECLIPSE Ni-U microscopic system (Nikon, Japan).
DNA extraction, PCR amplification, and sequencing
Total genomic DNA extraction was carried out using fresh mycelia grown on PDA, according to Cenis (1992). PCR amplification and sequencing for Alternaria were performed to amplify genes of ITS region with primers ITS5/ITS4 (White et al. 1990), EF1-α with EF1-728F/EF1-986R (Carbone and Kohn 1999), GAPDH with GPD1/GPD2 (Berbee et al. 1999), ALT with Alt-for/Alt-rev (Hong et al. 2005), and RPB2 with RPB2-6F/RPB2-7cR (Liu et al. 2019; Sung et al. 2007). Additionally, the cmdA gene using the primer pair CaldF1/CaldR1 (Lawrence et al. 2013), ITS and GAPDH genes, which were phylogenetically informative for species resolution within the Pleospora clade (Inderbitzin et al. 2009; Puig et al. 2015; Woudenberg et al. 2017), were used to identify Stemphylium species. PCR reactions were performed in a BIORAD T100 thermocycler (BIO-RAD, USA) with a total volume of 25 μL, comprising 12.5 μL 2 × Taq PCR Starmix (Genstar, Beijing, China), 1.25 μL of each primer, 2 μL template DNA, and 8 μL sterile distilled water. The resulting products were electrophoresed in 1% agarose gel and visualized under UV transillumination. Successfully amplified fragments were sequenced in both directions by BGI (Beijing Genomics Institute). The resulting sequences were viewed using BioEdit v7.0.9 (Hall 1999) and assembled in PHYDIT 3.2 (Chun 1995). Consensus sequences were deposited in GenBank (Table 1).
Phylogenetic analyses
Newly generated gene sequences were preliminarily subjected to BLASTn search in NCBI (https://www.blast.ncbi.nlm.nih.gov/). Subsequently, related sequences and reference sequences (Woudenberg et al. 2014, 2015) were retrieved from the GenBank database (https://www.ncbi.nlm.nih.gov/genbank/). Phylogeny based on ITS, GAPDH, ALT, EF1-α, and RPB2 gene sequences of the concatenated dataset was aligned using MEGA 6.0 (Tamura et al. 2013). In addition, the sequences of three different loci (ITS, GAPDH, cmdA) were concatenated and subjected to phylogenetic analysis for Stemphylium strains, YZU 191366 and YZU 191367. Phylogenetic analyses of each alignment were performed using maximum likelihood (ML), maximum parsimony (MP), and Bayesian inference (BI) methods. The best-fit model was GTR + I + G, as recommended by MRMODELTEST 2.3 (Nylander 2004). ML analyses were performed in RAxML 7.0.3 (Stamatakis et al. 2008) using the GTR + I + G model, with 1000 bootstrap replicates. MP analyses were conducted in PAUP 4.0b10 (Swofford 2003) using heuristic searches involving random sequence additions, with the tree bisection-reconnection branch-swapping algorithm. Other supportive tree scores, namely tree length, consistency index, retention index, and the rescaled consistency index, were also calculated (Table 2). Gaps within the alignments were treated as missing data. BI analyses were done in MrBayes 3.1.2 (Ronquist and Huelsenbeck 2003), implemented for 1,000,000 generations of Markov chain Monte Carlo (MCMC) searches, to determine the posterior probability, branch length, and substitution parameters using the previously mentioned best model. We discarded the first 25% of the samples as burn-in and calculated a majority-rule consensus tree. The yielded trees were visualized in FigTree 1.3.1 (Rambaut and Drummond 2010).
Pathogenicity tests
Pathogenicity tests were conducted on local onion obtained from Ywarthitgyi village by using a spore suspension inoculation (20 µL drop with the concentration of 1 × 105 conidia mL−1) technique in parallel with the colonized agar plug (2 mm in diameter from 5-day-old PDA cultures) technique. For the spore suspension assay, the strains were cultured on PCA for 7 days and then flooded with sterilized distilled water. The conidia were then dislodged into the water by rubbing the colony surface with a sterile glass rod. The number of conidia was counted using a hemocytometer and adjusted to the final concentrations. Sterilized distilled water and uncolonized agar plugs were used as negative controls. The tests were repeated three times. All inoculated plants were covered with a clean polythene bag to maintain the moisture content and were kept in the greenhouse at 25℃. Disease development was observed daily. We calculated disease severity (the disease index) on a scale of 0–5: 0, no disease symptoms; 1, lesion diameter < 10 mm; 2, lesion diameter 10–20 mm; 3, lesion diameter 20–30 mm; 4, lesion diameter 30–40 mm; and 5, complete drying or leaves split from the center. The disease index was determined as DI = (0n0 + 1n1 + 2n2 + 3n3 + 4n4 + 5n5)/5 N × 100, where n0–5 represent the number of leaves with each scale (0–5), and N represents the total number of leaves. Re-isolation and re-identification were attempted to fulfill Koch’s postulates.
Results
Phylogenetic analysis
Phylogeny of the combined ITS, GAPDH, ALT, EF1-α, and RPB2 gene sequences was constructed to determine a more accurate placement of newly collected Alternaria strains. The phylogenetic tree information was presented in Table 1. The full-length alignment of the combined dataset was stored in TreeBASE (Study no. 26265). The tree topology generated by ML was identical to that generated using MP and BI analyses and was therefore used as the basal tree (Fig. 1). All large-spored strains recovered in this study formed a distinct clade with high support values of 1.0 (BI), 100% (ML), and 99% (MP) that did not include any reference strains, as a sister clade to A. montanica and A. scorzonerae, suggesting that this is a new species. The small-spored strains formed a well-supported clade (BI, 1.0; MP, 100; ML, 100) that contained reference sequences of A. burnsii and A. tomato. However, morphological differences between A. burnsii and A. tomato suggested that the small-spored strains recovered during the current study are A. burnsii (see “Taxonomy” section below). In PCR amplification of Stemphylium, the ITS region, GAPDH, and cmdA gene resulted in sequences of 538 nt, 525 nt, and 667 nt, respectively. Sequences were deposited in GenBank with the accession number MW052760–MW052764 (Supplementary Table S1). Phylogenetic analyses of the combined dataset confirmed that two representative strains grouped together with S. vesicarium reference strains, CBS 322.49 and CBS 133905, with high support values of 1.0 (PP), 100% (ML), and 99% (BS) (Supplementary Fig. S1).
ML phylogenetic tree combined from ITS, GAPDH, ALT, EF1-α, and RPB2 gene sequences of Alternaria species from Allium cepa and the related taxa. Bayesian posterior probabilities (PP) > 0.70, maximum likelihood (ML) > 70%, and parsimony bootstrap values (BS) > 70% are indicated above/below the branches (PP/ML/BS). Taxon names, strain numbers, host, and geographic origins are provided. The scale bar represented the number of nucleotide substitutions. Strains from the present study are shown in bold. Ex-type strains (T) and representative strains (R) are noted in superscript
Taxonomy
Alternaria cepae A. A. Htun & J. X. Deng, sp. nov.
MycoBank No: MB 835551
Etymology. The specific epithet refers to the species of the host plant, Allium cepa.
Descriptions. Colonies on PDA in the dark at 25℃ (Fig. 2B), surface smooth, pale luteous to bay at the margin, colony reverse chestnut, scarlet, and rust pigmentation, 53–54 mm in diam., and colonies on PCA under fluorescent light/dark cycle of 8/16 h at 22℃ (Fig. 2C), surface ochreous, cinnamon, amber pigmentation, sulfur-yellow in reverse, 59‒61 mm diam. Conidiophore macronematous, solitary, single conidiogenous locus arising directly from the aerial hyphae, 26–75 (–124) × 5–10 μm, normally with 3‒5 septate, brown, smooth-walled. Conidia on PCA (Fig. 2D, E), solitary, narrow obclavate, internal cell formation retains distosepta throughout conidial enlargement, 53–85 (–90) × 12–30 μm, 5–7 transverse septa but rarely with a longitudinal euseptum, some conidia having a short-to-medium beak (8–30 μm) around 4.5–10 μm wide, occasionally up to 38 μm long, and similar conidia on host, 43–75 (–82) × 16–24 (–26) µm at 22℃, normally with short blunter beaks 5–24 (–40) µm long, and 49–79 × 15–23 (–26) µm at 25℃, with 10–26(–46) µm beaks.
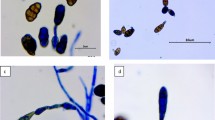
Morphological characteristics of Alternaria cepae and its causal symptoms on Allium cepa. Symptoms in the field (A); colony on PDA (B) and on PCA (C) for 7 days; sporulation patterns (D) and conidia (E) on PCA; pathogenicity test symptoms on living leaves inoculated with mycelium plug method (F) and conidial suspension (G) (4 days after inoculation). Scale bars: D = 50 µm, E = 25 µm
Holotype. MYANMAR. NAYPYIDAW: Ywarthitgyi village (194,751), 19°52′ 27.372″N, 96°11′34.332″E, 115 m, isolated from leaf blight of onion, single spore isolation, colonies grown on PDA and PCA for 7 days, 12 Feb 2019, A. A. Htun. (Holotype YZU-H-0035). Ex-type culture: YZU 191023 = ACCC39717.
Additional specimen examined. MYANMAR. NAYPYIDAW: Ywarthitgyi village, 12 Feb 2019, A. A. Htun (YZU 191024 = ACCC39718, YZU191025 = ACCC39719).
Habitat. Leaf blight on Allium cepa.
Distribution. Naypyitaw (Central Myanmar).
Notes: Phylogenetically, the species is the closest to A. montanica in section Porri based on the ITS, GAPDH, ALT, EF1-α, and RPB2 gene sequences. In conidial morphology, it differs significantly from A. montanica, which produces conidia with a long, narrow, and tapered apical beak that becomes a short, broad secondary conidiophore near the conidial apex (Table 3).
Alternaria burnsii Uppal, Patel & Kamat, Indian J.Agric.Sci.8:61 (1938).
Descriptions. Colonies on PDA for 7 days in darkness (Fig. 3A), surface buff to honey, cottony to vinaceous buff, united margin, 72‒73 mm in diameter. Colonies on PCA (Fig. 3B) incubated for 7 days under fluorescent light/dark cycle of 8/16 h at 22℃, surface pale-smoky gray, vinaceous buff, grayish sepia, fuscous black in reverse, 59‒63 mm diam. Conidia narrow ovoid to long ovoid or long ellipsoid, 20‒50 × 8‒15 μm, with beaks 3‒30 μm, rarely forming longitudinal septa and 4‒7 transverse septa, normally 5‒9 catenated conidia in a chain (Fig. 3C, D).
Morphological characteristics of Alternaria burnsii and its pathogenicity on Allium cepa. Colony on PDA (A) and on PCA (B) for 7 days; sporulation patterns (C) and conidia (D) on PCA; pathogenicity tests on living leaves inoculated with mycelium plug method (E) and conidial suspension (F) (7 days after inoculation). Scale bars: C = 50 µm, D = 25 µm
Notes. Alternaria burnsii was first described in India from Cuminum cyminum (Uppal et al. 1938). In the present study, the conidia of this species were similar to the morphological description by Simmons (2007) (Table 3). It is phylogenetically near to A. tomato; however, this species differs from A. burnsii by having ellipsoid to long-ovoid conidia (39–65 × 13‒22 μm), with a single beak 60‒105 μm, and no evidence of catenation (Table 3). The host range of A. burnsii is reported as Apiaceae: Cuminum cyminum (Uppal et al. 1938), Bunium persicum (Mondal et al. 2002), Apium graveolens (Zhang 2003; Zhuang 2005), and Cucurbitaceae: Cucurbita maxima (Paul et al. 2015). In the present study, A. burnsii was isolated from Liliaceae: Allium cepa.
Pathogenicity tests
The new species of A. cepae and A. burnsii were pathogenic to inoculated plants, regardless of the inoculation technique. A small necrotic spot was first observed on the inoculated leaves 1 day after inoculation (dpi) for the new species. The spot expanded aggressively and transformed into a water-soaked lesion with concentric rings at the edge, similar to the symptoms observed in the field (Fig. 2A, F, G). All of the inoculated leaves eventually collapsed at 3–5 dpi. For A. burnsii strains, water-soaked lesions appeared 3 dpi and gradually converted into sunken lesions with a creamy margin of 5 dpi (Fig. 3E, F). Using both inoculation methods, the disease incidence for the new species was 100%, with a severity of 80%; for A. burnsii, it was 40%, with a severity of 25%. The disease symptom of S. vesicarium on the inoculated leaves initially comprised a small, brown, and water-soaked lesion. Finally, the center of the lesion turned from brown to black with the concentric ring at the margin (Supplementary Fig. S2), which was very similar to that of A. cepae.
Discussion
Alternaria, an omnipresent fungus consisting of hundreds of species, leads to pre- and post-harvest crop losses and has been recorded as a critical fungal pathogen because of its global dissemination on various hosts (Lawrence et al. 2016; Meena et al. 2017). Alternaria profoundly attacks Allium species at two growth stages, when the leaves mature and before the bulb develops (Stavely and Slana 1971). There is a long history of studies on Alternaria species on Allium cultivars, specifically A. allii, A. palandui, A. porri (purple blotch), A. cepulicola, A. ascaloniae, A. iranica, A. prasonis, A. vanuatuensis, A. alternata, and A. tenuissima (Simmons 2007; Vélez-Rodríguez and Rivera-Vargas 2007).
Woudenberg et al. (2013, 2014, 2015), via multilocus phylogenetic analysis, indicated the misidentification of some of these Alternaria species in the past based mainly on morphological and host data. Among the Alternaria species infecting onions, A. ascaloniae has been synonymized as A. solani-nigri, A. iranica as A. thunbergiae, A. vanuatuensis as A. allii, and A. tenuissima and A. palandui as A. alternata. Gene sequence information for A. cepulicola is not available from GenBank, consequently, its phylogenetic position and possible synonymy with some of these species remain uncertain. In addition, A. cepulicola were obviously different from A. cepae by producing large conidia in a short chain (Table 3). In the current study, multigene phylogeny was employed alongside morphological characteristics to identify two Alternaria species that had not previously been associated with onions globally.
A new Alternaria species, named as A. cepae in the current study, was found to be associated with leaf blight in onion plantations in Naypyidaw, Myanmar. The taxonomy of Alternaria strains from this onion leaf blight was examined based on morphological characters and multilocus DNA sequence data. The combined gene phylogeny revealed that the strains formed a distinct lineage with well-supported bootstrap values. The other Alternaria species recovered from onion in this study, A. burnsii, was distinguished from its closest phylogenetic relative, A. tomato. Although these species were reciprocally monophyletic in the seven gene phylogeny of Woudenberg et al. (2015), support for the A. burnsii clade in that phylogeny was relatively low (0.88 Bayesian posterior probability, 68% maximum likelihood bootstrap support). Al-Nadabi et al. (2018) also failed to distinguish between A. burnsii and A. tomato using multigene phylogeny of the same five genes used in this study and resorted to referring to these species as the A. burnsii–A. tomato species complex. Woudenberg et al. (2015) described that in most cases, Alternaria species cannot be fully resolved using single-gene phylogenies and reiterate the conclusion of Al-Nadabi et al. (2018) that additional gene regions might be needed to differentiate between A. burnsii and A. tomato.
Furthermore, pathogenic Stemphylium vesicarium was isolated from the diseased tissue (Supplementary Figs. S1‒S2). This species has been recorded as an onion pathogen in Myanmar (https://www.ippc.int/static/media/files/pestreport/2016/12/01/Pests_of_Onion_in_Myanmar.pdf). Although most previous studies report significant destruction of onion crops when S. vesicarium co-occurs with A. porri (Aveling et al. 1993, Suheri and Price 2001, Mathur and Sharma 2006), A. porri was not obtained in the current study, even though it is a recognized onion fungal pathogen in Myanmar (Thaung 1970). Rather, A. cepae, A. burnsii, and S. vesicarium were recorded as the causal pathogens of onion leaf blight, leading to plant death. This study shed new light on the pathogenic fungi of onion, Alternaria and Stemphylium, based on their morphology, molecular data, and pathogenicity. Furthermore, A. cepae was described as a new taxon in Alternaria. To the best of our knowledge, this is the first report of onion leaf blight caused by A. burnsii in Myanmar, as well as the first report of the onion being a host for A. burnsii.
Data availability
All data generated or analyzed during this study are included in this published article (and its supplementary files). All sequences data generated in this study are available in NCBI GenBank.
References
Al-Nadabi H, Maharachchikumbura S, Agrama H, Al-Azri M, et al. (2018) Molecular characterization and pathogenicity of Alternaria species on wheat and date palms in Oman. Eur J Plant Pathol 152.https://doi.org/10.1007/s10658-018-1550-4
Andersen B, Krøger E, Roberts RG (2001) Chemical and morphological segregation of Alternaria alternata, A. gaisen and A. longipes. Mycol Res 105:291–299. https://doi.org/10.1017/S0953756201003446
Aveling TAS, Snyman HG, Naude SP (1993) Evaluation of seed treatments for reducing Alternaria porri and Stemphylium vesicarium on onion seed. Plant Dis 77:1009–1011. https://doi.org/10.1094/PD-77-1009
Berbee ML, Pirseyedi M, Hubbard S (1999) Cochliobolus phylogenetics and the origin of known, highly virulent pathogens, inferred from ITS and glyceraldehyde-3-phosphate dehydrogenase gene sequences. Mycologia 91:964–977. https://doi.org/10.1080/00275514.1999.12061106
Black L, Conn K, Gabor B, Kao J, Lutton J (2012) Purple blotch. In: Conn KE, Lutton JS, Rosenberger SA (eds) Onion disease guide. Seminis Vegetable Seeds Inc., St. Louis
Carbone I, Kohn ML (1999) A method for designing primer sets for speciation studies in filamentous ascomycete. Mycologia 91:553–556. https://doi.org/10.1080/00275514.1999.12061051
Cenis JL (1992) Rapid extraction of fungal DNA for PCR amplification. Nucleic Acids Res 20:2380. https://doi.org/10.1093/nar/20.9.2380
Chun J (1995) Computer assisted classification and identification of actinomycetes. Dissertation, Unversity of Newcastle. http://theses.ncl.ac.uk/jspui/handle/10443/410
Crous PW, Wingfield MJ, Richardson DM et al (2016) Fungal planet description sheets: 400–468. Persoonia 36:316–458. https://doi.org/10.3767/003158516X692185
Deng JX, Li MJ, Paul NC, Oo MM, Lee HB, Oh SK, Yu SH (2018) Alternaria brassicifolii sp. nov. isolated from Brassica rapa subsp. pekinensis in Korea. Mycobiology 46:172–176. https://doi.org/10.1080/12298093.2018.1468054
Farr DF, Bills GF, Chamuris GP, Rossman AY (1989) Fungi on plants and plant products in the United States. APS Press, St. Paul, p 1252
Gupta RC, Gupta RP (2013) Effect of integrated disease management packages on diseases incidence and bulb yield of onion (Allium cepa L.). SAARC J Agric 11:49–59. https://doi.org/10.3329/sja.v11i2.18401
Gupta RBL, Pathak VN (1988) Yield losses in onions due to purple blotch disease caused by Alternaira porri. Phytophylactica 20:21–23 https://hdl.handle.net/10520/AJA03701263_1214
Hall TA (1999) BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp Ser 41:95–98
Hong SG, Cramer RA, Lawrence CB, Pryor BM (2005) Alt a 1 allergen homologs from Alternaria and related taxa: analysis of phylogenetic content and secondary structure. Fungal Genet Biol 42:119–129. https://doi.org/10.1016/j.fgb.2004.10.009
Inderbitzin P, Mehta YR, Berbee ML (2009) Pleospora species with Stemphylium anamorphs: a fourlocus phylogeny resolves new lineages yet does not distinguish among species in the Pleospora herbarum clade. Mycologia 101:329–339. https://doi.org/10.3852/08-071
Köhl J, Groenenboom-De HB, de Goossen-Van G (2009) Pathogenicity of Stemphylium vesicarium from different hosts causing brown spot in pear. Eur J Plant Pathol 124:151–162. https://doi.org/10.1007/s10658-008-9402-2
Lawrence DP, Park MS, Pryor BM (2011) Nimbya and Embellisia revisited, with nov. comb for Alternaria celosiae and A. perpunctulata. Mycol Prog 11:799–815. https://doi.org/10.1007/s11557-011-0793-7
Lawrence DP, Gannibal PB, Peever TL, Pryor BM (2013) The sections of Alternaria: formalizing species-group concepts. Mycologia 105:530–546. https://doi.org/10.3852/12-249
Lawrence DP, Rotondo F, Gannibal PB (2016) Biodiversity and taxonomy of the pleomorphic genus Alternaria. Mycol Prog 15:1–22. https://doi.org/10.1007/s11557-015-1144-x
Liu HF, Liao J, Chen XY, Liu QK, Yu ZH, Deng JX (2019) A novel species and a new record of Alternaria isolated from two Solanaceae plants in China. Mycol Prog 18:1005–1012. https://doi.org/10.1007/s11557-019-01504-3
Mathur K, Sharma SN (2006) Evaluation of fungicides against Alternaria porri and Stemphylium vesicarium disease of onion in Rajasthan. J Mycol Plant Pathol 36:323–324
Meena M, Gupta SK, Swapnil P, Zehra A, Dubey MK, Upadhyay RS (2017) Alternaria toxins: potential virulence factors and genes related to pathogenesis. Front Microbiol 8:1451. https://doi.org/10.3389/fmicb.2017.01451
Mondal KK, Rana SS, Sood P, Singh Y (2002) Kalazira: a new host for Alternaria burnsii. Indian Phytopathol 55:532
Nolla JAB (1927) A new Alternaria disease of onions (Allium cepa L.). Phytopathology 17:115–132
Nylander JAA (2004) MrModeltest v2. Program distributed by the author. Evolutionary Biology Centre, Uppsala University
Paul NC, Deng JX, Lee HB, Yu SH (2015) Characterization and pathogenicity of Alternaria burnsii from seeds of Cucurbita maxima (Cucurbitaceae) in Bangladesh. Mycobiology 43:384–391. https://doi.org/10.5941/MYCO.2015.43.4.384
Pryor BM, Gilbertson RL (2000) Molecular phylogenetic relationships amongst Alternaria species and related fungi based upon analysis of nuclear ITS and mt SSU rDNA sequences. Mycol Res 104:1312–1321. https://doi.org/10.1017/S0953756200003002
Puig M, Ruz L, Montesinos E, Moragrega C, Llorent I (2015) Combined morphological and molecular approach for identification of Stemphylium vesicarium inoculum in pear orchards. Fungal Biol 119:136–134. https://doi.org/10.1016/j.funbio.2014.11.006
Rambaut A, Drummond A (2010) FigTree v.1.3.1. Institute of Evolutionary Biology. University of Edinburgh, Edinburgh
Rao VG (1963) Two new species of Alternaria on economic hosts from India. Sydowia 17:70–73
Rayner RW (1970) A mycological colour chart. Commonwealth Mycological Institute, Kew
Ronquist F, Huelsenbeck JP (2003) MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics:1572–1574. https://doi.org/10.1093/bioinformatics/btg180
Simmons EG (2007) Alternaria an identification manual. CBS Fungal Biodiversity Centre, Utrecht, pp 10–12
Stamatakis A, Hoover P, Rougemont J (2008) A rapid bootstrap algorithm for the RAxML web servers. Syst Biol 57:758–771. https://doi.org/10.1080/10635150802429642
Stavely JR, Slana LJ (1971) Relation of leaf age to the reaction of tobacco to Alternaria alternata. Phytophathology 61:73–78
Suheri H, Price TV (2000) Infection of onion leaves by Alternaria porri and Stemphylium vesicarium and disease development in controlled environments. Plant Pathol 49:375–382
Suheri H, Price TV (2001) The epidemiology of purple leaf blotch on leeks in Victoria, Australia. Eur J Plant Pathol 107:503–510. https://doi.org/10.1046/j.1365-3059.2000.00458.x
Sung GH, Sung JM, Hywel-Jones NL, Spatafora JW (2007) A multi-gene phylogeny of Clavicipitaceae (Ascomycota, fungi): identification of localized incongruence using a combinational bootstrap approach. Mol Phylogenet Evol 44:1204–1223. https://doi.org/10.1016/j.ympev.2007.03.011
Swofford DL (2003) PAUP*: Phylogenetic analysis using parsimony, * and other methods. Version 4.0b10. Sinauer Associates, Sunderland. Open J Mar Sci 7:1
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S (2013) MEGA6: molecular evolutionary genetics analysis version 6.0. Mol Biol Evol 30:2725–2729. https://doi.org/10.1093/molbev/mst197
Thaung MM (1970) New records of plant disease in Burma. PANS Pest Articles & News Summaries 16:638–640
Thomma BP (2003) Alternaria spp.: from general saprophyte to specific parasite. Mol Plant Pathol 4:225–236. https://doi.org/10.1046/j.1364-3703.2003.00173.x
Uppal BN, Patel MK, Kamat MN (1938) Alternaria blight of cumin. Indian J Agric Sci 8:49–62
Vélez-Rodríguez L, Rivera-Vargas LI (2007) Recent studies of fungal pathogens of onion in Puerto Rico. J Agric Univ Puerto Rico 91:31–4. https://doi.org/10.46429/jaupr.v91i1-2.2651
Wallroth FG (1833) Flora Cryptogamica Germaniae, pars. post. Nürnberg: J. L. Schrag. pp 923
White TJ, Burns T, Lee S, Taylor J (1990) Amplification and direct sequencing of fugal ribosomal RNA genes for phylogenetics. In: Innis MA, Gelfand DH, Sninsky JJ (eds) PCR protocols: a guide to methods and applications. Academic Press Inc, New York, pp 315–322
Wijayawardene NN, Hyde KD, Al-Ani LKT, Tedersoo L, Haelewaters D, Rajeshkumar KC, Zhao RL, Aptroot A, Leontyev DV (2020) Outline of fungi and fungus-like taxa. Mycosphere 11:1060–1456. https://doi.org/10.5943/mycosphere/11/1/8
Woudenberg JHC, Groenewald JZ, Binder M, Crous PW (2013) Alternaria redefined. Stud Mycol 75:171–212. https://doi.org/10.3114/sim0015
Woudenberg JHC, Truter M, Groenewald JZ, Crous PW (2014) Large-spored Alternaria pathogens in section Porri disentangled. Stud Mycol 79:1–47. https://doi.org/10.1016/j.simyco.2014.07.003
Woudenberg JHC, Seidl MF, Groenewald JZ, de Vries M, Stielow JB, Thomma BP, Crous PW (2015) Alternaria section Alternaria: species, formae speciales or pathotypes? Stud Mycol 82:1–21. https://doi.org/10.1016/j.simyco.2015.07.001
Woudenberg JHC, Hanse B, Van Leeuwen GCM, Groenewald JZ, Crous PW (2017) Stemphylium revisited. Stud Mycol 87:77–103. https://doi.org/10.1016/j.simyco.2017.06.001
Zhang TY (2003) Flora Fungorum Sinicorum, Vol. 16: Alternaria. Science Press, Beijing, p 284
Zhuang WY (2005) Fungi of northwestern China. Mycotaxon, Ithaca, pp 1–43
Acknowledgements
The authors sincerely thank Prof. Jian Ma for the assistance on the nomenclature of the new species.
Funding
This study was funded by the National Natural Science Foundation of China (31400014).
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. The samples collection were carried out by Aye Aye Htun. Deng Jian Xin supported scientific guidance during laboratory and field studies. The initial fungal isolation was performed by Aye Aye Htun, who led the entire research work with Liu Hai Feng and He Lin. Xia Zhen Zhou and Sein Lai Lai Aung contributed to data analysis. The manuscript was written by Aye Aye Htun, and all authors provided critical feedback and helped shape the research, analysis, and manuscript. Deng Jian Xin supervised the final version as well. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Section editor: Gerhard Rambold
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
ESM 1
Fig. S1 Phylogenetic tree combined dataset of ITS, GAPDH and cmdA sequences of Stemphylium vesicarium and its closest relative taxa. Bayesian posterior probabilities (PP) > 0.70, maximum likelihood (ML) >70% and parsimony bootstrap values (BS) > 70% are indicated above/below the branches (PP/ML/BS). Taxa names, strain numbers and geographic origins are provided. The scale bar represented the number of nucleotide substitutions. Strains from the present study were shown in bold. Type strain (T) is noted in superscript (PPTX 60 kb)

ESM 2
Fig. S2 Morphological characteristics of Stemphylium vesicarium and the pathogenicity on Allium cepa. Colony on PDA for 7 days at 25 °C (A); Sporulation patterns (B); Conidiophore (C) and conidia (D) on SNA; Pathogenicity on living leaves with inoculated mycelium plug method after 4 days inoculation (E). Scale bars = 25 μm (PNG 1957 kb)
Rights and permissions
About this article
Cite this article
Htun, A.A., Liu, H.F., He, L. et al. New species and new record of Alternaria from onion leaf blight in Myanmar. Mycol Progress 21, 59–69 (2022). https://doi.org/10.1007/s11557-021-01765-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11557-021-01765-x