Abstract
Research was undertaken to clarify the taxonomic identity of leaf rust (Pucciniales) fungi on bioenergy switchgrass in the Eastern and Central U.S. We integrated internal transcribed spacer 2 (ITS2) and partial 28S ribosomal RNA gene sequence data from collections taken from cultivated switchgrass and herbarium specimens, including purported aecial and telial states of Puccinia graminicola and Puccinia pammelii. Maximum likelihood and Bayesian analyses revealed four monophyletic clades: Puccinia emaculata sensu stricto (s.s.), P. pammelii, P. graminicola, and Puccinia novopanici. Results also indicated that P. emaculata s.s. was not affecting cultivated, bioenergy switchgrass. Aecidium pammelii and P. pammelii were distinct phylogenetically from P. emaculata s.s. and grouped within a well-supported clade, demonstrating aecial-telial host alternation for P. pammelii between Euphorbia corollata and switchgrass. Aecidium stillingiae on queen’s delight (Stillingia sylvatica)—a purported aecial state host for P. graminicola—shared identical sequences with the recently described species Puccinia pascua. The latter fungus, however, was recovered within a subclade of P. graminicola. Hence, queen’s delight likely is not an aecial host to P. graminicola s.s. Additional molecular studies are warranted to determine species boundaries within the P. graminicola complex. The majority of contemporary collections from cultivated switchgrass were recognized as P. novopanici. Collectively, bioenergy switchgrass is host to at least three phylogenetically distinct species, presenting a significant challenge to the future selection and breeding of switchgrass with improved rust resistance.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Switchgrass (Panicum virgatum L.) is a native warm-season (C4) perennial grass that is distributed widely across eastern North America, wherein it exhibits considerable genotypic and phenotypic variability as well as wide ecological adaptation to environmental clines (Das et al. 2004; Casler et al. 2007; Cortese and Bonos 2013; Lowry et al. 2014). In addition to its robust genetic variation and geographic range, switchgrass has a demonstrated capability of persistence and growth on marginal quality land and, hence, has been considered as a model feedstock for the production of biomass and cellulosic biofuels in the USA (McLaughlin and Adams Kszos 2005; Schmer et al. 2008; Davis et al. 2009; Kiniry et al. 2012; Parish et al. 2012; Casler et al. 2015; Sanford et al. 2016). The switchgrass breeding community has placed a high priority on the development of improved cultivars with maximized biomass yield and conversion potential (Parrish et al. 2012; Casler and Vogel 2014; Lipka et al. 2014) that are also adapted to regional abiotic and biotic stresses (Casler et al. 2011, 2015; Arias Aguirre et al. 2012; Parish et al. 2012; Casler and Vogel 2014; Lipka et al. 2014; Lowry et al. 2014). With the expansion (i.e., scale-up) of switchgrass cultivation, plant breeders and growers alike anticipate that increased acreage dedicated to switchgrass will also result in a corresponding increase in diseases, particularly those incited by phytopathogenic fungi (Crouch et al. 2009; Stewart and Cromey 2011; Uppalapati et al. 2013; Zhu et al. 2013; Serba et al. 2015). One-hundred and twenty (120) species of fungi have been determined to occur on switchgrass in the USA (Farr and Rossman 2018), 82 of these fungi cause disease on switchgrass as well as allied species of Panicum. However, from a historical perspective, the rust fungi (Pucciniales) present the most obvious and significant threat to the sustainable and long-term production of switchgrass (Cornelius and Johnston 1941; Uppalapati et al. 2013; Serba et al. 2015; Sykes et al. 2016), as they are among the most notorious and economically damaging pathogens in intensive agriculture systems (McDonald and Linde 2002; Strange and Scott 2005). Moreover, Sykes et al. (2016) demonstrated that severe leaf rust disease, caused purportedly by Puccinia emaculata, can reduce the downstream conversion of switchgrass to cellulosic ethanol by as much as 58%.
The order Pucciniales consists of more than 6000 species across approximately 120 genera, wherein Puccinia and Uromyces—the two largest genera of rust fungi—possess some 4000 and 600 described species, respectively (Cummins and Hiratsuka 2003). Taxa within Puccinia and Uromyces are highly specialized obligate biotrophic pathogens of vascular plants that often have complex and variable life cycles, possessing up to five spore stages (O-IV: O = spermatium, I = aeciospore, II = urediniospore, III = teliospore, and IV = basidiospore). Heteroecious species can alternate between unrelated aecial and telial hosts (Cummins and Hiratsuka 2003). While rust fungi within these genera display a diversity of life cycles, their classification has been based principally on the morphology of teliospores and, to a lesser extent, the uredinial, spermogonial, and aecial spore stages (Dietel 1928; Arthur 1934; Cummins 1971; Cummins and Hiratsuka 2003). Though important historically for family- and genus-level classification, the utilization of teliospore morphology (e.g., cell number, dimensions, wall thicknesses) for species delineation—either alone or in combination with urediniospore morphology (i.e., dimensions and surface ornamentation)—has resulted in the grouping of morphological similar, yet phylogenetically discrete and, sometimes, distantly related taxa as congeneric (Zambino and Szabo 1993; Wingfield et al. 2004; Aime 2006; Minnis et al. 2012). The adoption of molecular-based phylogenetic approaches, in lieu of or integrated with morphologic comparisons, over the last two decades has led to the increasing discovery of cryptic rust fungi, indicating that phenotypic similarities can be due to morphological convergence rather than genetic inheritance from a shared ancestor (synapomorphies). Thus, morphological-based classification can be misinformative for inference(s) of evolutionary relationships and delimiting species (Roy et al. 1998; Szabo 2006; van der Merwe et al. 2007; Maier et al. 2003; Liu and Hambleton 2010, 2013; Bennett et al. 2011; Yun et al. 2011; Kenaley et al. 2014; Vialle et al. 2013; Liu and Hambleton 2010, 2013). This is exemplified by the genus Uromyces, whereby numerous taxa within the genus have long been considered one-celled, teliospore variants of the Puccinia spp. and, over the past decade, the paraphyletic nature of Uromyces has been demonstrated (Aime 2006; van der Merwe et al. 2007; Aime et al. 2018).
The muddled and uncertain taxonomy of rust fungi is particularly true for less-studied taxa affecting native perennial grasses, such as switchgrass, with recently realized agronomic potential in biomass and lignocellulosic ethanol production (Kenaley et al. 2016; Demers et al. 2017). According to historical reports, at least four species of rust fungi have been implicated in switchgrass leaf rust in North America, including Puccinia emaculata Schwein., Puccinia graminis Pers., Puccinia huberi Henn., and Uromyces graminicola Burrill (Burrill 1884; Arthur 1934; Ramachar and Cummins 1963, 1965; Anonymous 1970; Cummins 1971; Lenne 1990). Puccinia emaculata was described by Schweinitz in 1834 from specimens on witchgrass (Panicum capillare), and thereafter, the known telial host range of P. emaculata expanded considerably to include an additional six Panicum spp. (P. amarum Elliot, P. amarulum Elliot, P. gattingeri Nash, P. hallii Vasey, P. miliaceum L., and P. philadelphicum Bernh ex. Trin.) as well as barnyard grass (Echinochloa crus-galli [L.] P. Beauv.), decorative millet (Echinochloa holciformis [Kunth] Nash), and variable panicgrass (Dichanthelium commutatum [Schult.] Gould). Moreover, in the early- and mid-twentieth centuries, two switchgrass-infecting rust fungi—Puccinia pammelii Arthur. and Puccinia panici Dietel—were reclassified under P. emaculata (Dietel 1895; Arthur 1905, 1934; Ramachar and Cummins 1965). Arthur (1905) placed P. panici in synonymy with P. pammelii; however, in his manual of rust flora, Arthur (1934) maintained P. panici over P. pammelii. Thereafter, Ramachar and Cummins (1965) followed the convention of Arthur (1934) and reclassified P. panici—including P. pammelii and its aecial state Aecidium pammelii Trel.—under P. emaculata. Justification for the former reclassification of P. panici under P. pammelii likely was based on the lack of morphologic discontinuities and, purportedly, their shared aecial state, A. pammelii, on spurges (Euphorbiaceae Juss.: Euphorbia corollata L. and Euphorbia marginata Pursh) (Stuart 1901; Arthur 1905, 1906). The circumscription of both P. pammelii and P. panici under P. emaculata by Ramachar and Cummins (1965), however, complicated the taxonomy and presumed biology of P. emaculata sensu lato as Arthur (1934) defined P. emaculata sensu stricto (s.s.) as a principal parasite of witchgrass, not switchgrass, and clearly indicated past failures to demonstrate telial-to-aecial host alternation for P. emaculata s.s. between witchgrass and euphorbs as well as other native plant species outside of Euphorbiaceae (Arthur 1902, 1903, 1905, 1907, 1908, 1915, 1917). Thus, Ramachar and Cummins (1965) revision of P. emaculata to include P. panici (and syn. P. pammelii) likely created a compound species, consisting of at least two distinct taxa—P. emaculata s.s. and P. pammelii or P. panici—and, wherein, the telial-aecial host association between switchgrass and Euphorbia was solely based on circumscribed taxa, not P. emaculata s.s.
Although switchgrass leaf rust has received considerable attention in the Central and Eastern U.S. (Tiffany et al. 1990; Gravert et al. 2000; Gravert and Munkvold 2002; Gustafson et al. 2003; Zale et al. 2008; Hirsch et al. 2010; Gilley et al. 2013; Frazier et al. 2013; Uppalapati et al. 2013; Sykes et al. 2016), investigations integrating morphologic and DNA-based analyses to determine the inter- and intraspecific diversity of switchgrass rust fungi were not pursued until recently (Kenaley et al. 2016; Demers et al. 2017). Kenaley et al. (2016) utilized teliospore morphology, uredinial group (Ramachar and Cummins 1965), and phylogenetic analyses based on partial internal transcribed spacer (ITS) region data to delimit rust taxa affecting important bioenergy and ornamental switchgrass cultivars in the North Central and Eastern U.S. Results of this investigation clearly demonstrated two morphologically and phylogenetically discrete taxa—P. emaculata and U. graminicola—as well as one isolate (PUC-24) that was morphologically similar to P. emaculata yet was more closely related genetically to isolates of U. graminicola. Kenaley and colleagues utilized P. emaculata EU915294 for the purpose of phylogenetic placement and species determination; a full-length ITS sequence was obtained from rust-infected switchgrass in Tennessee (Zale et al. 2008). Likewise, prior to their report, no ITS sequence data existed in public genetic databases for U. graminicola; however, all isolates determined to be U. graminicola formed a well-supported monophyletic clade and possessed only one-celled teliospores that were consistent with past species descriptions (Arthur 1934; Ramachar and Cummins 1963). Kenaley et al. (2016), therefore, concluded that at least two phylogenetically distinct species were capable of inciting leaf rust on bioenergy and/or ornamental switchgrass; yet, given the uncertain ecology and taxonomy of P. emaculata and evidence of a possible second P. emaculata-like taxon, they suggested that a broader phylogenetic assessment of P. emaculata sensu lato (s.l.)—including its purported aecial state A. pammelii—was warranted. Moreover, Kenaley et al. (2016) also discussed the scant evidence supporting the aecial-telial host association reported previously for U. graminicola. Similar to P. emaculata s.l., the aecial hosts of U. graminicola include euphorbs as prairie tea (Croton monanthogynus Michx.), linear rushfoil (Crotonopsis linearis Michx.), Gulf Sebastian bush (Ditrysinia fruticosa [Bartram] Govaerts & Frodin; syn. Sebastiana ligustrina [Michx.] Müll.Arg.), and queen’s delight (Stillingia sylvatica L.; syn. Stillingia angustifolia [Torr.] Wats.) (Arthur 1934). However, as indicated by Kenaley et al. (2016), no inoculation test(s) confirmed host alteration for U. graminicola (Arthur 1906, 1909, 1910, 1917).
The recent taxonomic revision of Panicum rust fungi by Demers et al. (2017) clarified the species assemblages on switchgrass and allied panicgrasses across native landscapes in eastern North America. Using spore morphology (teliospore and urediniospore) and separate phylogenetic analyses for three nuclear genes (ITS2, partial 28S, and intergenic spacer (IGS) region), Demers and colleagues delimited six species of switchgrass-infecting rust fungi including Puccinia graminicola (Burrill) Demers & Castl. (comb. nov., syn. U. graminicola), P. pammelii (syn. P. panici), and four new species (Puccinia amari Demers, Puccinia cumminsii Liu and Hambleton, Puccinia novopanici Demers, Liu & Hambleton, and Puccinia pascua Demers, Liu & Hambleton). Notably absent from their species summary for switchgrass was P. emaculata s.s., which they determined to be a principal parasite of witchgrass. The majority of fungus material examined on switchgrass was classified to be P. novopanici—a species nearly morphologically identical and most closely related to P. emaculata s.s. Demers et al. (2017) also utilized their molecular analyses to confirm the aecial-telial host association of P. pammelii between switchgrass and the euphorb E. corollata and, conversely, to determine that Aecidium crotonopsidis Burrill (syn. Aecidium stillingiae Tracy and Earle 1899) was unlikely the aecial state of U. graminicola as previously suggested in the historical literature (Jackson 1917).
The primary objective of the research presented herein was to construct and analyze a broad phylogenetic dataset, integrating ITS2 and partial 28S sequence data taken from cultivated switchgrass and representative ribosomal DNA (rDNA) data from Demers et al. (2017) to reassess the species assemblage(s) of rust fungi affecting bioenergy and ornamental cultivars of switchgrass first provided in Kenaley et al. (2016). New for this study, we generated a combined phylogenetic dataset based on ITS2 and partial 28S sequences procured from single-sorus samples excised from historical herbarium specimens of P. emaculata on witchgrass, P. pammelii on E. corollata, and A. stillingiae on S. sylvatica—the purported aecial state of P. graminicola. Large subunit rDNA (28S) sequence data was also generated for the 35 isolates utilized previously by Kenaley et al. (2016) in phylogenetic tree reconstruction.
Materials and methods
Fungi and plant material
Switchgrass foliage with evident uredinia and/or telia was collected in 2011–2013 across ten localities in the North Central and Eastern U.S., including sites in Alabama, Iowa, Nebraska, New York, Pennsylvania, and South Dakota (Kenaley et al. 2016; Table 1). Kenaley et al. (2016) utilized this fungus-plant material for an exploratory survey of the rust fungi affecting common bioenergy and ornamental cultivars of switchgrass, subjecting this material to morphologic and DNA analyses based on only partial ITS data (partial ITS1, complete 5.8S and ITS2, partial 28S). Voucher information of the aforementioned collections—including accession number, plant host, and localities—is provided in Table 1. Specific for the present study, herbarium specimens possessing Aecidium-type aecia on the purported alternate hosts of P. graminicola and P. pammelii as well as the telial states of P. emaculata s.s. on witchgrass and P. graminicola on switchgrass were examined (Table 2).
DNA extraction
Genomic DNA from herbarium, single-sorus samples (uredinium or telium) on switchgrass and witchgrass was extracted using a DNeasy Plant Mini Kit (Qiagen, Valencia, CA) with modifications to the kit-provided protocol as described in Kenaley et al. (2016). A similar approach was utilized for procuring DNA from single-aecium samples taken from the purported alternate host of P. pammelii. and P. graminicola on E. corollata and Stillingiae sylvatica, respectively. To do this, each aecium was excised with leaf tissue using a sterile surgical scalpel, transferred to a separate sterile 1.5-mL safe-lock centrifuge tube, and ground to a fine powder using liquid nitrogen and mini-pestle before adding 500 μL AP1 (lysis) buffer and 10 μL proteinase K (10 mg mL−1; Qiagen) per tube. Thereafter, the tubes of lysed tissue were incubated overnight (55 °C at 1100 rpm) in a thermomixer (model 5350; Eppendorf, Hauppauge, NY) and, following a 12–14-h incubation, were placed in boiling water for 5 min. Following the bath in boiling water, the tubes were centrifuged briefly (3000 rpm for 5 s) to collect the lysis mixture and 150 μL of the kit-provided neutralization buffer (P3 buffer) was added immediately to each tube. All tubes were then centrifuged (1 min at 13,000 rpm) and placed at − 20 °C for 10 min. Lastly, before proceeding with steps 9–19 of the DNeasy Plant Mini protocol (Qiagen), each tube was removed from − 20 °C and centrifuged again for 5 min (13,000 rpm) to pelletize cell debris and ease transfer of the supernatant (~ 660 μL) to a QIAshredder mini spin column.
PCR amplification and DNA sequencing
Partial ITS sequence data (partial ITS1, complete 5.8S ribosomal RNA (rRNA) gene and ITS2, and partial 28S rRNA gene) from 2011–2013 collections and produced by Kenaley et al. (2016) were obtained via PCR amplification and DNA sequencing using the primer pair ITS1rustF10d (Barnes and Szabo 2007) and RUST1 (Kropp et al. 1995) or PRITS1f (5′-AGTGCACTTTATTGTGGCTCGA-3′; L. Castlebury, pers. comm.) and ITS4-B (Gardes and Brun 1993). The former primer pair yielded complete, full-length ITS amplicons (partial 18S; complete ITS1, 5.8S, and ITS2; and partial 28S), and hence, the 5′ ad 3′ ends of these sequences were pruned to a total of 747–759 nucleotides with identical coverage of the PRIT1f-ITS4B amplicon. Polymerase chain reaction mixtures (25 μL total volume) included 2.5 mL AccuPrime PCR buffer II (Invitrogen, Carlsbad, CA; 600 mM Tris-SO4 [pH 8.9], 180 mM [NH4]2SO4, 20 mM MgSO4, 2 mM dNTPs, thermostable AccuPrime protein, 10% glycerol), 1.5 μL of 50 mM MgSO4, 0.5 μL (10 mM) of each primer, 0.1 μL AccuPrime Taq DNA Polymerase High Fidelity (Invitrogen) with 3′ to 5′ exonuclease activity, 20.4 μL ultrapure PCR grade water (Ambion, Austin, TX), and 1.0 μL undiluted, DNA template. Thermal cycling conditions for the ITS PCRs consisted of the following: initial denaturation cycle for 3 min at 95 °C; 35 cycles for 30 s at 95 °C, 30 s at 50 °C, and 2 min at 72 °C; and a final extension step of 10 min at 72 °C. A similar approach to direct PCR amplification of the ITS2 (partial 5.8S, complete ITS2, and partial 28S) was first applied to whole-genomic DNA extractions from herbarium specimens, modified only to include 1.25 μL (20 mg/mL) bovine serum albumin (BSA; New England Biolabs, Beverly, MA) and primers RUST2inv (Aime 2006) and RUST1. If the first PCR attempt failed, the volume of undiluted DNA template was increased to 2.0 or 2.5 μL within the PCR mixture. Thereafter, for DNA extractions yielding no ITS2 amplicon after two PCR attempts, a semi-nested PCR was performed as follows: first-step amplification used 2.5 μL of undiluted DNA template and primer the pair RUST2inv (Aime 2006) and RUST1 whereas the second step used a 2.0–2.5 μL undiluted PCR product from the first-step reaction and the primer pair RUST2inv and ITS4-B (Gardes and Brun 1993). First- and second-step thermal cycling programs and reaction mixtures were identical to that for direct ITS amplification, adjusting only water volume to accommodate additional DNA template. A larger partial-28S fragment was PCR amplified from field collections and herbarium specimens with primers RUST2inv and LR6 (Vilgalys and Hester 1990) in 25 μL reaction mixtures (total volume) consisting of 2.5 μL undiluted DNA template, 5 μL 5× MyTaq reaction buffer (Bioline, Taunton, MA), 1.25 μL BSA (20 mg/mL), 0.5 μL each primer (20 mM), 0.5 μL MyTaq DNA polymerase (Bioline, Taunton, MA), and 14.75 μL ultrapure PCR grade water. Controls (i.e., minus genomic DNA) were included in each PCR run.
Across all PCRs, amplicon size was checked via electrophoresis and transillumination using 1.2% (w/v) agarose gels amended with GelRed (Biotium, Hayward, CA) or post-electrophoresis stained with ethidium bromide (1.2 mg/L 0.5× Tris-acetate-EDTA (TAE)) and visualization under ultraviolet fluorescence. ITS2 and 28S PCR products were purified using a DNA Clean and Concentrator Kit 5 (Zymo Research, Irvine, CA) according to the manufacturer’s protocol. Purified PCR products were normalized (130 ng per reaction) and sequenced bidirectionally—forward and reverse—using an ABI 3730 DNA sequencer (Applied Biosystems, Foster City, CA), BigDye terminator DNA sequencing kit (Applied Biosystems), and the referred primers.
Several ITS2 (N = 5) and 28S (N = 4) PCR products for historically important herbarium specimens (e.g., cultures and type material) were also cloned and sequenced (see Figs. 1, 3, and 4, indicated by asterisks). To do this, secondary PCR, electrophoresis, and PCR product purification were executed as described previously and, thereafter, cleaned ITS2/28S PCR products were ligated into the cloning vector pCR4-TOPO (Invitrogen) and utilized in the transformation of Escherichia coli (One Shot TOP10 chemically competent E. coli; Invitrogen), following the manufacturer’s instructions. Three randomly chosen sequenced-derived clones per ITS2/28S plasmid prep were extracted using the Wizard SV Miniprep DNA Purification System (Promega, Madison, WI) according to the kit-provided protocol and sequenced as described above using the universal primer pair M13.
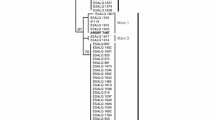
Majority-rule consensus tree based on maximum likelihood (ML) and Bayesian inference (BI) analyses using ITS2 sequence data (571 total characters across partial 5.8S rRNA gene, complete ITS2, and partial 28S rRNA gene). Numbers above branches indicate ML bootstrap values > 70% (after 104 replicates), and numbers below indicate BI clade credibility values > 90 (after 5.0 × 106 generations). Branches with conflicting ML and BI support were pruned. Taxon labels for Panicum rust fungi determined by Demers et al. (2017) are highlighted entirely in bold font
Phylogenetic analysis
New ITS2 and 28S sequence data were assembled and edited in CodonCode Aligner v5.1.5 (CodonCode Corporation, Dedham, MA) and confirmed to belong to the genus Puccinia by BLASTn searches (http://blast.ncbi.nlm.nih.gov) as well as comparison to authenticated sequences (Demers et al. 2017). All newly generated ITS2 and 28S sequences were deposited into GenBank (Tables 1 and 2). Thereafter, a primary ITS2 dataset was compiled by combining the newly generated ITS2 data from herbarium specimens with ITS sequences used previously in Kenaley et al. (2016) (Table 1) as well as representative sequences for Panicum rust fungi determined by Demers et al. (2017). The ITS2 dataset was then amended further to include Puccinia andropogonis Schwein (GenBank accession: DQ344516), Puccinia asparagi D.C. (AY217137), P. graminis Pers. (AF468044), Puccinia horiana Henn. (EU816912), and Puccinia striiformis Westend. (GQ457306) as ingroup taxa and Puccinia coronata var. coronata (KY426917) as the outgroup taxon.
A total of 81 ITS/ITS2 sequences were then aligned with the multiple sequence alignment option implemented in Clustal X v2.1 ((Larkin et al. 2007), pruned to include only 571 characters (partial 5.8S, complete ITS2, and partial 28S), and edited by eye in Geneious® v.9.0.5 (Biomatters Ltd., Auckland, New Zealand) to remove obvious misalignments (TreeBASE: http://purl.org/phylo/treebase/phylows/study/TB2:S22223).
A 28S data matrix was compiled as described previously for the ITS2 dataset, consisting of 28S sequences from isolates resequenced from Kenaley et al. (2016), herbarium specimens, and rust fungi of switchgrass and native panicgrasses examined by Demers et al. (2017). The final 28S alignment consisted of 73 taxa and spanned 979 characters. For complete analysis of the sequence data, ITS2 and 28S sequences per isolate or specimen were concatenated in Mesquite v3.04 (Maddison and Maddison 2015) and aligned in Clustal X v2.1, yielding 73 total taxa and consisting of 1388 characters. The number of ingroup taxa was reduced in the 28S and ITS2-28S dataset when compared to the ITS2 matrix because seven ingroup taxa utilized in the ITS2 did not have corresponding 28S sequence data.
Data matrices—ITS2, 28S, and combined IT2-28S—were subjected to maximum likelihood (ML) and Bayesian inference (BI) of phylogeny. ML analyses were executed via MetaPIGA v2.0 (Helaers and Milinkovitch 2010), whereas BI was performed using the Markov chain Monte Carlo (MCMC) method in MrBayes v3.2.6 (Ronquist et al. 2012). For the ML and BI analyses, the best model of sequence evolution for each of the three alignments was selected by the Akaike information criterion (AIC; Akaike 1974) implemented in jModelTest 2.1.7 (Guindon and Gascuel 2003; Darriba et al. 2012). ML analysis in MetaPIGA was performed using the bootstrapping heuristic search option consisting of 10,000 pseudoreplicates, while including all nucleotides and treating gaps as missing characters. Bayesian posterior probabilities for clade (node) support were calculated in MrBayes using one cold and three heated Markov chains, sampling every 100 generations over 5.0 × 106 generations for a total of 50,000 sampled generations. The potential scale reduction factor (PSRF) for each of the model parameters was > 1.0 when MrBayes was terminated. Chain convergence was also determined by examining average standard deviations of split frequencies and likelihood values. A 10% burn-in value was determined using Tracer v1.5 (Rambaut and Drummond 2009), removing 5000 sampled generations.
The genetic variability within and among switchgrass rust fungi was also examined by comparing mean pairwise genetic distances, nucleotide differences (N), and nucleotide similarities (%). Custom distance matrices were generated separately for ITS2, 28S, and combined ITS-28S sequence data for rust taxa assigned phylogenetically by ML and BI analyses to P. cumminsii, P. graminicola, P. novopanici, P. pammelii, and/or P. pascua using PAUP* (Swofford 2003); the best-fit model of sequence evolution; and the statistical software JMP Pro 13 (SAS Institute Inc., Cary, NC). Nucleotide differences and percent similarities among the isolates/specimens assigned to the aforementioned taxa were calculated in Geneious.
Results
Phylogenetic analyses and genetic statistics
ITS2
The ITS2 data matrix alignment consisted of 571 total characters of which 395 were constant, 74 were variable and parsimony-uninformative, and 102 were parsimony-informative. A consensus tree reconstructed based on ML and BI analyses of the ITS2 data is shown in Fig. 1, wherein four clades (I–IV) received significant support with ML bootstrap values and Bayesian posterior probabilities of > 95% after 10,000 pseudoreplicates and 5.0 × 106 generations, respectively. Clade I consisted of the type specimen for the recently described species P. novopanici (BPI 747673; Demers et al. 2017) as well as all isolates (PUC-1–PUC-23), and corresponding collections, previously determined to be P. emaculata by Kenaley et al. (2016). Puccinia emaculata s.s. was recovered in clade II. This clade included sequence data from the epitype (BPI 851570; Demers et al. 2017) as well as historically important culture no. 300 (PUR 18651) and no. 405 (PUR 18654) on witchgrass provided by H. H. Whetzel and likely used by J.C. Arthur for inoculation testing to determine host alternation of P. emaculata (Arthur 1934). Clade II-P. emaculata taxa were found only on witchgrass and were also more closely related, phylogenetically and genetically, to taxa within clade I-P. novopanici (Figs. 1 and 2a). Interestingly, ITS2 sequences for the 13 taxa assigned to clade II-P. emaculata, including the epitype, were identical (Fig. 2a). The lack of interspecific genetic variation for the ITS2 among these taxa was confirmed by direct-sequencing ITS2-derived clones from specimens PUR 18651 and PUR 18673 collected 22 years apart and at distant geographic localities (Table 2).
Combined matrices of pairwise intra- and inter-specific genetic distances, nucleotide similarities (%), and nucleotide differences based on the ITS2 gene region (a), ITS2, partial 28S rRNA gene (b), and concatenated ITS2-28S loci (c) for Puccinia taxa on switchgrass. 1Inspection of Puccinia amari BPI 064435 (Demers et al. 2017) revealed a seven-nucleotide insertion within ITS2 that was not present in P. amari BPI 893097
Puccinia pammelii and its aecial state formed a well-supported clade, clade III, and consisted of all single-aecium samples examined from E. corollata, including one procured from the isotype of A. pammelii (NY 611274) collected by L.H. Pammel in 1883 in Wisconsin. The P. emaculata-like isolate PUC-24 (CUP 068153) collected in 2013 from the switchgrass cultivar ‘Summer’ grown in Lincoln, Nebraska, also resolved to clade III, providing a species determination for this previously unknown isolate and herbarium accession (Kenaley et al. 2016). Collectively, the ITS2 sequence(s) for taxa in clade III-P. pammelii shared little homology to ITS2 sequences obtained from taxa determined to be P. emaculata and P. novopanici as well as taxa recovered in clade IV. The latter clade, clade IV, included three taxa presently treated as discrete species (Demers et al. 2017), wherein P. cumminsii and P. pascua resolved to a well-supported subclade within P. graminicola. These three species shared ≥ 95.7% nucleotide identity across the ITS2, differing by only 22.0–22.5 nucleotides among them. Aecidium stillingiae (PUR 11681 and PUR 11682) on queen’s delight—a reported aecial host of P. graminicola—was found in clade IV within the P. pascua group. Rust taxa corresponding to P. amari, P. cumminsii, and Puccinia crotonopsidis were not recovered from field collections performed by Kenaley et al. (2016), nor from herbarium specimens examined herein.
The 28S region and combined ITS-28S
Maximum likelihood and BI analyses using 28S sequence data clearly exhibited less phylogenetic resolution than those using the ITS2 gene region (Figs. 2b and 3). Of the aforementioned clades defined using ITS2 sequence data, all four clades—clade I-P. novopanici, clade II-P. emaculata, clade III-P. pammelii, and clade IV-P. graminicola complexes—collapsed to a single group that included P. amari, P. crotonopsidis, and Puccinia setariae. Puccinia cumminsii DAOM 114236 and DAOM 114238 were not included within the 28S data matrix as publicly available sequence data for this species included only ≤ 310 nucleotides of the 28S (Demers et al. 2017). These results were not unexpected as the 28S alignment (979 total characters) possessed 885 constant characters and, of the remaining 94 characters, only 18 characters were parsimony-informative.
Majority-rule consensus tree based on maximum likelihood (ML) and Bayesian inference (BI) analyses of 28S sequence data (979 total characters). Numbers above branches indicate ML bootstrap values > 70% (after 104 replicates), and numbers below indicate BI clade credibility values > 75 (after 107 generations). Branches with conflicting ML and BI support were pruned. Taxon labels for Panicum rust fungi determined by Demers et al. (2017) are highlighted entirely in bold font
In contrast, ML and BI analyses of the concatenated ITS-28S data strongly supported the existence of four monophyletic clades (Fig. 4) with each corresponding to clades I–IV recovered previously using ITS2 sequence data (Fig. 1). The concatenated ITS-28S dataset included 1388 aligned characters (1182 constant; 95 variable, parsimony-uninformative; 111 parsimony-informative). However, inspection of ITS-28S genetic statistics revealed that the resolution of the ITS-28S phylogeny was not improved markedly when compared to ITS2 data (Fig. 2a, c) as mean interspecific genetic distances and corresponding 95% confidence intervals were slightly smaller in comparison.
Majority-rule consensus tree based on maximum likelihood (ML) and Bayesian inference (BI) analyses using concatenated ITS2 and 28S sequence data (1388 total characters across partial 5.8S rRNA gene, complete ITS2, and partial 28S rRNA gene). Numbers above branches indicate ML bootstrap values > 70% (after 104 replicates), and numbers below indicate BI clade credibility values > 90 (after 5.0 × 106 generations). Branches with conflicting ML and BI support were pruned. Taxon labels for Panicum rust fungi determined by Demers et al. (2017) are highlighted entirely in bold font
Assignment of switchgrass rust species combinations
The primary objective of the present study was to reassess the species determinations for rust fungi affecting economically important bioenergy and ornamental cultivars previously characterized by Kenaley et al. (2016) (Table 1). According to phylogenetic analyses of the ITS2 and combined ITS2-28S datasets, P. novopanici, rather than P. emaculata, was the most commonly encountered rust fungus (105/121, 86.8% total isolates; CUP 068048–CUP 06152) across the ten sites examined by Kenaley and colleagues in 2011–2013. Switchgrass cultivars from which P. novopanici was identified included ‘Cave-in-Rock’, ‘Dallas Blue’, ‘Kanlow’, ‘Summer’, and ‘Sunburst’. Likewise, the previously unknown P. emaculata-like isolate (PUC-24, CUP 68153) from Lincoln, NE, was determined to be P. pammelii.
Discussion
The work presented suggests that P. novopanici—a recently described species—likely is the most common switchgrass rust fungus on bioenergy switchgrass in the North Central and Eastern U.S. This result is well supported by Demers et al. 2017 who, as part of their taxonomic assessment of Panicum rust fungi, also reported that P. novopanici was the most frequently encountered species across switchgrass rust specimens with origins in five states: Connecticut, Florida, Maryland, New York, and Virginia. Here, based on the past (Zale et al. 2008; Kenaley et al. 2016) and present data, we can now expand the geographic distribution of P. novopanici to include 12 states with the addition of Alabama, Iowa, Nebraska, Pennsylvania, South Dakota, Tennessee, and, West Virginia. Moreover, Arkansas (Hirsch et al. 2010), Mississippi (Gilley et al. 2013), and Oklahoma (Orquera-Tornakian et al. 2017) should also be added to where P. novopanici has been reported on cultivated switchgrass; rust species determination(s) within these reports/studies was achieved via BLASTn comparisons to P. emaculata EU915294 (= P. novopanici; Zale et al. 2008) or P. novopanici prior to or after the publication of Demers et al. (2017), respectively. Although the geographic range of P. novopanici likely consists of the eastern two thirds of North America, as it was also confirmed in southeastern Canada (Demers et al. 2017), there remains considerable work to be done to clarify its telial host range and biology—critical first steps in identifying current year and overwintering inoculum reservoirs. At present, authenticated telial hosts native to the USA include only bitter panicgrass (Demers et al. (2017) and, as demonstrated herein, upland and lowland ecotypes of switchgrass (e.g., ‘Cave-in-Rock’ and ‘Kanlow’, respectively; Table 1). Although a daunting task at first glance, decoupling the telial host range of P. novopanici from its morphologically similar relative P. emaculata, however, may be straightforward as P. emaculata may, in fact, be restricted to witchgrass. Demers et al. (2017) suggested that the telial host range of P. emaculata may be limited to witchgrass and our phylogenetic analyses agree with their findings in so far as P. emaculata was recovered from only witchgrass. Yet, the diversity of telial hosts to which P. emaculata s.l. was once assigned was underrepresented in the analyses by Demers and colleagues, and certainly, the same was true in the present study. Our sampling plan was skewed toward rust fungi on switchgrass, rather than rust taxa on other Panicum spp. and allied grasses common to landscapes in central and eastern North America, such as barnyard grass (Echinochloa crus-galli), Gattinger’s panicgrass (Panicum gattingeri), Hall’s panicgrass (Panicum hallii), Philadelphia panicgrass (Panicum philadelphicum), and variable panicgrass (Dichanthelium commutatum) (Arthur 1934; Ramachar and Cummins 1965; USDA-NRCS 2018). Taxonomic studies of rust fungi affecting the aforementioned panicgrasses, including switchgrass, therefore, could be an effective and efficient approach to further delineate the telial host range and geographic distributions of P. emaculata s.s. and P. novopanici. Moreover, because telial and aecial hosts of Puccinia spp. are often found in close proximity, such exploratory studies could yield valuable insight into possible telial-aecial host associations for P. emaculata and P. novopanici.
The heteroecious, macrocyclic life cycle of P. emaculata and P. novopanici remains unclear. Puccinia emaculata PUR 18651 and PUR 18654 were recovered in clade II-P. emaculata of the ITS2 and ITS-28S consensus trees. These specimens were collected by H. H. Whetzel and received by J.C. Arthur in 1904 and 1906 during the same time period. J.C. Arthur was performing numerous inoculation tests to determine the existence of telial-to-aecial host switching for P. emaculata (Arthur 1902, 1903, 1905, 1907, 1908). Throughout the early 1900s, J.C. Arthur inoculated over 24 native plants, including flowering spurge (E. corollata), in an attempt to identify the alternate host of P. emaculata, but without success. The absence of flowering spurge within the P. emaculata and P. novopanici groups confirms that E. corollata is not an alternate host to P. emaculata nor P. novopanici, as proposed by Demers et al. 2017. Considering the close phylogenetic relationship between P. emaculata and P. novopanici as well as the influence of aecial hosts on the evolutionary trajectory of rust fungi (Aime 2006; Szabo 2006; van der Merwe et al. 2008; Alaei et al. 2009; Liu and Hambleton 2013; Vialle et al. 2013; Aime et al. 2018), these species likely share an ancestral or extant related aecial host. Thus, elucidating the aecial host for either P. emaculata or P. novopanici could provide a definite starting point for identifying the aecial host(s) for the other.
Of the three rust fungi identified on bioenergy cultivars of switchgrass, only P. pammelii has the characteristic heteroecious, macrocyclic life cycle commonly associated with rust fungi in tall prairie ecosystems (Szabo 2006). Herein, as also reported by Demers et al. (2017), P. pammelii was resolved to a well-supported monophyletic group (clade III), consisting of one isolate (PUC-24) and the aecial state, Aecidium pammelii, on Euphorbia corollata (Figs. 1 and 4). Host alternation for P. pammelii was supported further by the inclusion of the 135-year-old type specimen, A. pammelii NY 611274, in clade II of the ITS2 consensus tree. Unfortunately, although we were successful in PCR amplifying the ITS2, we were unable to produce a 973-bp 28S sequence for A. pammelii NY 611274 as was completed for PUC-24, P. pammelii PUR 18699, P. pammelii PUR 50594, and P. pammelii PUR N2082 that were collected 40–109 years ago. The successful production of ITS2 and 28S sequences from PUC-24 was also important, because, until the present study, the species identity of this isolate remained unknown. Teliospore morphology for PUC-24 was consistent with species descriptions of P. emaculata (Arthur 1934; Ramanchar and Cummins1965); however, it possessed a unique partial ITS sequence (partial ITS1, complete 5.8S and ITS2, partial 28S), unlike isolates of P. emaculata (= P. novopanici) and Uromyces graminicola (= P. graminicola) reported in Kenaley et al. (2016). Moreover, to the best of our knowledge, this species determination represents the first report of P. pammelii affecting switchgrass in Nebraska. According to historical records (Farr and Rossman 2018) and the specimen summary for P. pammelii (Demers et al. 2017), the majority of reports related to the geographic distribution of this switchgrass rust are almost entirely based on the occurrence of A. pammelii. Thus, the geographic distribution of P. pammelii includes much of the South Central, Central, and Eastern U.S., corresponding approximately to the geographic distribution of E. corollata. In addition to the latter euphorb, Arthur (1908) demonstrated telial-to-aecial host alternation for P. pammelii s.s. on switchgrass to snow on the mountain (Euphorbia marginata Pursh), which purportedly has a significantly larger geographic range than E. corollata (USDA-NRCS 2018). The geographic distribution of snow on the mountain extends throughout much of the eastern two thirds of the USA from the central Rocky Mountains to the Atlantic Coast. However, as noted previously, Kenaley et al. (2016) rarely found P. pammelii in switchgrass in Nebraska, where both E. marginata and native switchgrass are known to co-occur.
We were particularly interested in phylogenetic comparisons of field isolates of P. graminicola (= U. graminicola) to P. pascua as both species possess aseptate teliospores and teliospore measurements (i.e., length, width, and cell wall thickness) that overlap (Kenaley et al. 2016; Demers et al. 2017). Collectively, based on our phylogenetic analyses of the ITS2 and ITS2-28S, neither P. pascua nor P. cumminsii (Demers et al. 2017) was isolated from bioenergy switchgrass or selected ornamentals at any of the sites examined in 2011–2013 (Table 1, Fig. 1). Moreover, ML and BI analyses using ITS2 data from P. graminicola, P. cumminsii, and P. pascua placed all three species into a single clade (clade II; Fig. 1) wherein P. cumminsii and P. pascua grouped into a discrete subclade among isolates and historical specimens of P. graminicola. Hence, these taxa appear to represent a morphologically cryptic compound species. This result agrees with phylogenetic analysis of ITS data presented by Demers et al. (2017), wherein P. cumminsii, P. graminicola, and P. pascua were recovered in the monophyletic group referred to as the P. graminicola complex. Demers and colleagues, however, justified species designation based solely on IGS data, which, according to their analyses, clearly separated these two taxa. Thus, additional molecular and morphological data—the telial state for P. cumminsii and uredinial state of P. pascua (Demers et al. (2017)—are necessary to fully characterize this species complex; however, we strongly support the recognition of these taxa at the specific level, particularly because of the inferred aecial host of P. pascua—Aecidium stillingiae (syn. A. crotonopsidis; Figs. 1 and 4). Arthur (1934) reports that the aecial state of P. graminicola, Aecidium stillingiae, occurs across several euphorbs including taxa in the genera Croton, Ditrysinia, Sebastiana, and Stillingia. Yet, at present, there is no evidence in the scientific literature supporting the telia-aecial association of P. graminicola and A. stillingiae. In their taxonomic study of Panicum rust, Demers et al. (2017) included Puccinia crotonopsidis (= A. crotonopsidis, = A. stillingiae) BPI 006810 on Michaux’s croton (Crataegus michauxii G.L. Webster) as part of their separate phylogenetic analyses of the ITS-28S and intergenic spacer region. Results of these analyses were identical to that revealed in the present DNA analyses using the ITS2 and ITS2-28S sequence data: P. crotonopsidis was distantly related, phylogenetically, to P. graminicola and, hence, was not the aecial state of P. graminicola (Figs. 1 and 4). In addition to P. crotonopsidis, we included material from A. stillingiae PUR 11681 and PUR 11682 collected on queen’s delight (S. sylvatica) in Oklahoma in the early 1910s. However, unlike P. crotonopsidis, the two taxa of A. stillingiae were recovered within the P. graminicola complex (clade IV) and grouped with P. pascua (Figs. 1 and 4). Thus, this result represents the first evidence of host alternation for the newly described switchgrass rust fungus, P. pascua. Moreover, this further supports the recognition of taxa in the P. graminicola complex at the species level and points to the likely importance of euphorbs in the life histories and evolution of these species. Future taxonomic studies of switchgrass rust fungi should be expanded to include multi-gene phylogenetic analysis and fungus material with Aecidium-type aecia on taxa in the euphorbiaceous genus Stillingia to delineate species boundaries and telial-aecial host associations within the P. graminicola complex.
The research presented here clarifies the species identity and diversity of switchgrass rust fungi previously reported by Kenaley et al. (2016) and lends support to the recent taxonomic revision of switchgrass rust fungi by Demers et al. (2017). Thus, to date, three Puccinia spp.—P. graminicola, P. pammelii, and P. novopanici—are known to cause leaf rust diseases on common and economically important switchgrass cultivars in agricultural settings within the North Central and Eastern U.S. However, future studies elucidating the geographic distributions, biology, intraspecific diversity, and life cycles of switchgrass rust fungi are necessary in order to effectively implement regional disease management strategies and switchgrass breeding programs for rust resistance.
References
Aime MC (2006) Toward resolving family-level relationships in rust fungi (Uredinales). Mycoscience 47:112–122. https://doi.org/10.1007/s10267-006-0281-0
Aime MC, Bell C, Wilson AW (2018) Deconstructing the evolutionary complexity of rust fungi (Pucciniales) and their hosts. Stud Mycol 89:143–152. https://doi.org/10.1016/j.simyco.2018.02.002
Akaike H (1974) A new look at the statistical model identification. IEEE Trans Automat Contr AC-19:716–723. https://doi.org/10.1109/tac.1974.1100705
Alaei H, De Backer M, Nuytinck J, Maes M, Höfte M, Heungens K (2009) Phylogenetic relationships of Puccinia horiana and other rust pathogens of Chrysanthemum × morifolium based on rDNA ITS sequence analysis. Mycol Res 113:668–683. https://doi.org/10.1016/j.mycres.2009.02.003
Anonymous (1970) Index of plant diseases in the United States. United States Department of Agriculture, Agricultural handbook no. 165. U.S. Government Printing Office, Washington, D.C. 531
Arias Aguirre A, Studer B, Frei U, Lübberstedt T (2012) Prospects for hybrid breeding in bioenergy grasses. Bioenergy Res 5:10–19. https://doi.org/10.1007/s12155-011-9166-y
Arthur JC (1902) Cultures of Uredineae in 1900 and 1901. J Mycol 8:51–56. https://doi.org/10.2307/3752295
Arthur JC (1903) Cultures of Uredineae in 1902. Bot Gaz 35:10–23. https://doi.org/10.1086/328313
Arthur JC (1905) Cultures of Uredineae in 1904. J Mycol 11:50–67. https://doi.org/10.2307/3752724
Arthur JC (1906) Cultures of Uredineae in 1905. J Mycol 12:11–27. https://doi.org/10.2307/3752702
Arthur JC (1907) Cultures of Uredineae in 1906. J Mycol 13:189–205. https://doi.org/10.2307/3752588
Arthur JC (1908) Cultures of Uredineae in 1907. J Mycol 14:7–26. https://doi.org/10.2307/3752603
Arthur JC (1909) Cultures of Uredineae in 1908. Mycologia 1:225–256. https://doi.org/10.2307/3753158
Arthur JC (1910) Cultures of Uredineae in 1909. Mycologia 2:213–240. https://doi.org/10.2307/3753158
Arthur JC (1915) Cultures of Uredineae in 1912, 1913 and 1914. Mycologia 7:61–89. https://doi.org/10.2307/3753130
Arthur JC (1917) Cultures of Uredineae in 1916 and 1917. Mycologia 9:294–312. https://doi.org/10.2307/3753130
Arthur JC (1934) Manual of the rusts in United States and Canada. Purdue Research Foundation, Lafayette, IN. The Science Press Printing Co., Lancaster, PA. 438
Barnes CW, Szabo LJ (2007) Detection and identification of four common rust pathogens of cereals and grasses using real-time polymerase chain reaction. Phytopathology 97:717–727. https://doi.org/10.1094/phyto-97-6-0717
Bennett C, Aime MC, Newcombe G (2011) Molecular and pathogenic variation within Melampsora on Salix in western North America reveals numerous cryptic species. Mycologia 103:1004–1018. https://doi.org/10.3852/10-289
Burrill TJ (1884) New species of Uredineae (collection made by a. B. Seymour). Bot Gaz 9:187–191. https://doi.org/10.1086/325802
Casler MD, Vogel KP (2014) Selection for biomass yield in upland, lowland, and hybrid switchgrass. Crop Sci 54:626–636. https://doi.org/10.2135/cropsci2013.04.0239
Casler MD, Stendal CA, Kapich L, Vogel KP (2007) Genetic diversity, plant adaptation regions, and gene pools for switchgrass. Crop Sci 47:2261–2273. https://doi.org/10.2135/cropsci2006.12.0797
Casler MD, Tobias CM, Kaeppler SM, Buell CR, Wang Z-Y, Cao P, Schmutz J, Ronald P (2011) The switchgrass genome: tools and strategies. Plant Genome 4:273–282. https://doi.org/10.3835/plantgenome2011.10.0026
Casler MD, Vogel KP, Harrison M (2015) Switchgrass germplasm resources. Crop Sci 55:2463–2478. https://doi.org/10.2135/cropsci2015.02.0076
Cornelius DR, Johnston CO (1941) Differences in plant type and reaction to rust among several collections of Panicum virgatum L. J Am Soc Agron 33:115–124. https://doi.org/10.2134/agronj1941.00021962003300020003x
Cortese LM, Bonos SA (2013) Bioenergy traits of ten switchgrass populations grown in the northeastern/mid-Atlantic USA. Bioenergy Res 6:580–590. https://doi.org/10.1007/s12155-012-9271-6
Crouch JA, Beirn LA, Cortese LM, Bonos SA, Clarke BB (2009) Anthracnose disease of switchgrass caused by the novel fungal species Colletotrichum navitas. Mycol Res 113:1411–1421. https://doi.org/10.1016/j.mycres.2009.09.010
Cummins GB (1971) The rust fungi of cereals, grasses and bamboos, 2nd edn. Springer, New York, p 570
Cummins GB, Hiratsuka Y (2003) Illustrated genera of rust fungi, 3rd edn. The American Phytopathological Society, APS Press, St. Paul, Minnesota, U.S.A. 225
Darriba D, Taboada GL, Doallo R, Posada D (2012) jModelTest 2: more models, new heuristics and parallel computing. Nat Methods 9:772. https://doi.org/10.1038/nmeth.2109
Das MK, Fuentes RG, Taliaferro CM (2004) Genetic variability and trait relationships in switchgrass. Crop Sci 44:443–448. https://doi.org/10.2135/cropsci2004.4430
Davis SC, Anderson-Teixeira KJ, DeLucia EH (2009) Life-cycle analysis and the ecology of biofuels. Trends Plant Sci 14:140–146. https://doi.org/10.1016/j.tplants.2008.12.006
Demers JE, Liu M, Hambleton S, Castlebury LA (2017) Rust fungi of Panicum. Mycologia 109:1–17. https://doi.org/10.1080/00275514.2016.1262656
van der Merwe M, Ericson L, Walker J, Thrall PH, Burdon JJ (2007) Evolutionary relationships among species of Puccinia and Uromyces (Pucciniaceae, Uredinales) inferred from partial protein coding gene phylogenies. Mycol Res 111:163–175. https://doi.org/10.1016/j.mycres.2006.09.015
van der Merwe MM, Walker J, Ericson L, Burdon JJ (2008) Coevolution with higher taxonomic host groups within the Puccinia/Uromyces rust lineage obscured by host jumps. Mycol Res 112:1387–1408. https://doi.org/10.1016/j.mycres.2008.06.027
Dietel P (1895) New North American Uredineae. Erythea 3:57–82
Dietel P (1928) Unterklasse Hemibasidii (Ustilaginales and Uredinales). Reihe Uredinales. In: Engler A (ed) Die natürlichen Pflanzenfamilien, vol 6, pp 24–98
Farr DF, Rossman AY (2018) Fungal databases, systematic mycology and microbiology laboratory, ARS, USDA. http://nt.ars-grin.gov/fungaldatabases/. Accessed 06 April 2018
Frazier T, Shen Z, Zhao B, Bush E (2013) First report of Puccinia emaculata infection on switchgrass in Virginia. Plant Dis 97:424. https://doi.org/10.1094/pdis-09-12-0814-pdn
Gardes M, Brun TD (1993) ITS primers with enhanced specificity for basidiomycetes—application to identification of mycorrhizae and rusts. Mol Ecol 2:113–118. https://doi.org/10.1111/j.1365-294X.1993.tb00005.x
Gilley M, Tomaso-Peterson M, Orquera GK, Marek SM (2013) First report of rust caused by Puccinia emaculata on switchgrass in Mississippi. J Miss Acad Sci 58:197–200
Gravert C, Munkvold G (2002) Fungi and diseases associated with cultivated switchgrass in Iowa. J Iowa Acad Sci 109:30–34
Gravert CE, Tiffany LH, Munkvold GP (2000) Outbreak of smut caused by Tilletia maclaganii on cultivated switchgrass in Iowa. Plant Dis 84:596. https://doi.org/10.1094/pdis.2000.84.5.596a
Guindon S, Gascuel O (2003) A simple, fast, and accurate algorithm to estimate large phylogenies by maximum likelihood. Syst Biol 52:696–704. https://doi.org/10.1080/10635150390235520
Gustafson DM, Boe A, Jin Y (2003) Genetic variation for Puccinia emaculata infection in switchgrass. Crop Sci 43:755–759. https://doi.org/10.2135/cropsci2003.7550
Helaers R, Milinkovitch MC (2010) MetaPIGA v2.0: maximum likelihood large phylogeny estimation using the metapopulation genetic algorithm and other stochastic heuristics. BMC Bioinformatics 11:379. https://doi.org/10.1186/1471-2105-11-379
Hirsch RL, TeBeest DO, Bluhm BH, West CP (2010) First report of rust caused by Puccinia emaculata on switchgrass in Arkansas. Plant Dis 94:381. https://doi.org/10.2135/cropsci2003.7550
Jackson HS (1917) The Uredinales of Delaware. Proc Indiana Acad Sci 27:311–385
Kenaley SC, Smart LB, Hudler GW (2014) Genetic evidence for three discrete taxa of Melampsora (Pucciniales) affecting willows (Salix spp.) in New York State. Fungal Biol 118:704–720. https://doi.org/10.1016/j.funbio.2014.05.001
Kenaley SC, Hudler GW, Bergstrom GC (2016) Detection and phylogenetic relationships of Puccinia emaculata and Uromyces graminicola (Pucciniales) on switchgrass in New York State using rDNA sequence information. Fungal Biol 120:791–806. https://doi.org/10.1016/j.funbio.2016.01.016
Kiniry JR, Johnson M-VV, Bruckerhoff SB, Kaiser JU, Cordsiemon RL, Harmel RD (2012) Clash of the titans: comparing productivity via radiation use efficiency for two grass giants of the biofuel field. Bioenergy Res 5:41–48. https://doi.org/10.1007/s12155-011-9116-8
Kropp BR, Albee S, Flint KM, Zambino P, Szabo L, Thomson SV (1995) Early detection of systemic rust infections of dyers woad (Isatis tinctoria) using the polymerase chain reaction. Weed Sci 43:467–472. https://doi.org/10.1094/phyto-86-891
Larkin MA, Blackshields G, Brown NP, Chenna R, McGettigan PA, McWilliams H, Gibson TJ, Higgins DG (2007) Clustal V and Clustal X version 2.0. Bioinformatics 23:2947–2948. https://doi.org/10.1093/bioinformatics/btm404
Lenne JM (1990) World list of fungal diseases of tropical pasture species. Phytopathological Papers No. 31. CAB International, Oxon, UK. 162
Lipka AE, Lu F, Cherney JH, Buckler ES, Casler MD, Costich DE (2014) Accelerating the switchgrass (Panicum virgatum L.) breeding cycle using genomic selection approaches. PLoS One 9:e112227. https://doi.org/10.1371/journal.pone.0112227
Liu M, Hambleton S (2010) Taxonomic study of stripe rust, Puccinia striiformis sensu lato, based on molecular and morphological evidence. Fungal Biol 114:881–899. https://doi.org/10.1016/j.funbio.2010.08.005
Liu M, Hambleton S (2013) Laying the foundation for a taxonomic review of Puccinia coronata s.l. in a phylogenetic context. Mycol Prog 12:63–89. https://doi.org/10.1007/s11557-012-0814-1
Lowry DB, Behrman KD, Grabowski P, Morris GP, Kiniry JR, Juenger TE (2014) Adaptations between ecotypes and along environmental gradients in Panicum virgatum. Am Nat 183:682–692. https://doi.org/10.1086/675760
Maddison WP, Maddison DR (2015) Mesquite: a modular system for evolutionary analysis. Version 3.04. http://mesquiteproject.org. Accessed 06 April 2018
Maier W, Begerow D, Weiß M, Oberwinkler F (2003) Phylogeny of the rust fungi: an approach using large subunit ribosomal DNA sequences. Can J Bot 81:12–23. https://doi.org/10.1139/b02-113
McDonald BA, Linde C (2002) Pathogen population genetics, evolutionary potential, and durable resistance. Annu Rev Phytopathol 40:349–379. https://doi.org/10.1146/annurev.phyto.40.120501.101443
McLaughlin SB, Adams Kszos L (2005) Development of switchgrass (Panicum virgatum) as a bioenergy feedstock in the United States. Biomass Bioenergy 28:515–535. https://doi.org/10.1016/j.biombioe.2004.05.006
Minnis AM, McTaggart A, Rossman A, Aime MC (2012) Taxonomy of mayapple rust: the genus Allodus resurrected. Mycologia 104:942–950. https://doi.org/10.3852/11-350
Orquera-Tornakian GK, Garrido P, Kronmiller B, Hunger R, Tyler BM, Garzon CD, Marek SM (2017) Identification and characterization of simple sequence repeats (SSRs) for population studies of Puccinia novopanici. J Microbiol Methods 139:113–122. https://doi.org/10.1016/j.mimet.2017.04.011
Parish ES, Hilliard MR, Baskaran LM, Dale VH, Griffiths NA, Mulholland PJ, Sorokine A, Thomas, Downing ME, Middleton RS (2012) Multimetric spatial optimization of switchgrass plantings across a watershed. Biofuel Bioprod Bior 6:58–72
Ramachar P, Cummins G (1963) The species of Uromyces on the tribe Paniceae. Mycopathol Mycol Appl 19:49–61. https://doi.org/10.1007/bf02049014
Ramachar P, Cummins G (1965) The species of Puccinia on the tribe Paniceae. Mycopathol Mycol Appl 25:7–60. https://doi.org/10.1007/bf0204961
Rambaut A, Drummond J (2009) Tracer v1.5. (Distributed by the first author, Department of Biology, University of York, UK) http://tree.bio.ed.ac.uk/software/tracer/. Accessed 8 April 2018
Ronquist F, Teslenko M, van der Mark MP, Ayres DL, Darling A, Höhna S, Larget B, Suchard MA, Huelsenbeck JP (2012) MrBayes 3.2: efficient Bayesian phylogenetic inference and model choice across a large model space. Syst Biol 61:539–542. https://doi.org/10.1093/sysbio/sys029
Roy BA, Vogler DR, Bruns TD, Szaro TM (1998) Cryptic species in the Puccinia monoica complex. Mycologia 90:846–853. https://doi.org/10.2307/3761326
Sanford GR, Oates LG, Jasrotia P, Thelen KD, Robertson GP, Jackson RD (2016) Comparative productivity of alternative cellulosic bioenergy cropping systems in the North Central USA. Agric Ecosyst Environ 216:344–355. https://doi.org/10.1016/j.agee.2015.10.018
Schmer MR, Vogel KP, Mitchell RB, Perrin RK (2008) Net energy of cellulosic ethanol from switchgrass. Proc Natl Acad Sci U S A 105:464–469. https://doi.org/10.1073/pnas.0704767105
Serba DD, Uppalapati SR, Mukherjee S, Krom N, Tang Y, Mysore KS, Saha MC (2015) Transcriptome profiling of rust resistance in switchgrass using RNA-seq analysis. Plant Genome 8. https://doi.org/10.3835/plantgenome2014.10.0075
Stewart A, Cromey M (2011) Identifying disease threats and management practices for bio-energy crops. Curr Opin Environ Sustain 3:75–80. https://doi.org/10.1016/j.cosust.2010.10.008
Strange RN, Scott PR (2005) Plant disease: a threat to global food security. Annu Rev Phytopathol 43:83–116. https://doi.org/10.1146/annurev.phyto.43.113004.133839
Stuart W (1901) Some additions to the flora of Indiana. Proc Indiana Acad Sci 11:282–284
Swofford DL 2003 Phylogenetic analysis using parsimony (*and other methods). Version 4. Sinauer 758 Associates, Sunderland, Massachusetts, U.S.A.
Sykes VR, Allen FL, Mielenz JR, Stewart CN, Windham MT, Hamilton CY, Rodriguez M, Yee KL (2016) Reduction of ethanol yield from switchgrass infected with rust caused by Puccinia emaculata. Bioenergy Res 9:239–247. https://doi.org/10.1007/s12155-015-9680-4
Szabo LJ (2006) Deciphering species complexes: Puccinia andropogonis and Puccinia coronata, examples of differing modes of speciation. Mycoscience 47:130–136. https://doi.org/10.1007/S10267-006-0287-7
Tiffany LH, Shearer JF, Knaphus G (1990) Plant parasitic fungi of four tallgrass prairies of northern Iowa: distribution and prevalence. J Iowa Acad Sci 97:157–166
Tracy SM, Earle FS (1899) New fungi from Mississippi. Bull Torrey Bot Club 26:493–495. https://doi.org/10.2307/2478018
Uppalapati SR, Serba DD, Ishiga Y, Szabo LJ, Mittal S, Bhandari HS, Bouton JH, Mysore KS, Saha MC (2013) Characterization of the rust fungus, Puccinia emaculata, and evaluation of genetic variability for rust resistance in switchgrass populations. Bioenergy Res 6:458–468. https://doi.org/10.1007/s12155-012-9263-6
USDA-NRCS (2018) The PLANTS Database. National Plant Data Team, Greensboro, NC http://plants.usda.gov. Accessed on 06 April 2018
Vialle AN, Feau N, Frey P, Bernier L, Hamelin RC (2013) Phylogenetic species recognition reveals host-specific lineages among poplar rust fungi. Mol Phylogenet Evol 66:628–644. https://doi.org/10.1016/j.ympev.2012.10.021
Vilgalys R, Hester M (1990) Rapid genetic identification and mapping of enzymatically amplified ribosomal DNA from several Cryptococcus species. J Bacteriol 172:4238–4246
Wingfield BD, Ericson L, Szaro T, Burdon JJ (2004) Phylogenetic patterns in the Uredinales. Australas Plant Pathol 33:327–335. https://doi.org/10.1071/ap04020
Yun HY, Minnis AM, Kim YH, Castlebury L, Aime MC (2011) The rust genus Frommeëlla revisited: a later synonym of Phragmidium after all. Mycologia 103:1451–1463. https://doi.org/10.3852/11-120
Zale J, Freshour L, Agarwal S, Sorochan J, Ownley BH, Gwinn KD, Castlebury LA (2008) First report of rust on switchgrass (Panicum virgatum) caused by Puccinia emaculata in Tennessee. Plant Dis 92:1710. https://doi.org/10.1094/pdis-92-12-1710b
Zambino PJ, Szabo LJ (1993) Phylogenetic relationships of selected cereal and grass rusts based on rDNA sequence analysis. Mycologia 85:401–414. https://doi.org/10.2307/3760702
Zhu Q, Bennetzen JL, Smith SM (2013) Isolation and diversity analysis of resistance gene homologues from switchgrass. G3-Genes Genom Genet 3:1031–1042. https://doi.org/10.1534/g3.112.005447
Funding
This study was funded in part by the Cornell University Hatch Project NYC 153743 from the United States Department of Agriculture-National Institute for Food and Agriculture (USDA-NIFA). The findings, conclusions, and/or recommendations expressed in this publication are those of the authors and do not necessarily reflect the view of the USDA-NIFA.
Author information
Authors and Affiliations
Corresponding author
Additional information
Section Editor: Meike Piepenbring
Rights and permissions
About this article
Cite this article
Kenaley, S.C., Quan, M., Aime, M.C. et al. New insight into the species diversity and life cycles of rust fungi (Pucciniales) affecting bioenergy switchgrass (Panicum virgatum) in the Eastern and Central United States. Mycol Progress 17, 1251–1267 (2018). https://doi.org/10.1007/s11557-018-1434-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11557-018-1434-1