Abstract
Phytophthora blight caused by Phytophthora capsici limits the production of cucurbits and peppers in the United States and is a growing threat to sustainable vegetable production in New England. Little is known about the genetic diversity of P. capsici in New England, and a total of 210 P. capsici isolates from 18 sites were genotyped using 46 single nucleotide polymorphism markers, revealing 85 unique and 34 repeated multi-locus genotypes. Both mating types were recovered from 7 of the 18 locations. Isolates with identical genotypes (clonal lineages) ranged from 2 to 16. Three clonal lineages were recovered from multiple sites within the same year, although none were recovered across multiple years. Bayesian clustering revealed individuals with a complex genetic cluster composition. This, coupled with a high outcrossing rate (mean t = 0.87) and no clear clustering in principal coordinates analysis, suggests outcrossing among the populations. Phylogenetic and genetic distance analysis indicate differentiation based on farm location and movement among farms may be infrequent. There was no obvious differentiation based on cucurbit, tomato or pepper hosts.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Phytophthora blight caused by Phytophthora capsici is recognized as the most destructive disease of cucurbits (Cucurbita spp.) and peppers (Capsicum spp.) in the United States (Babadoost 2004; Hausbeck and Lamour 2004; McGrath 2001; Wick 1994; Zitter 1989). First described on pepper in the United States in 1922, P. capsici has a cosmopolitan distribution across the globe (Lamour 2013; Leonian 1922; Park et al. 2008). In the northeast US, the pathogen is responsible for significant loss on peppers, eggplant, tomatoes, pumpkins, melons and squash. Currently, the pathogen is a growing threat to sustainable vegetable production in New England and surrounding states (UMass Extension 2016).
Phytophthora capsici is soil-borne and, once introduced to a new location, can survive many years as dormant thick-walled oospores, even after crop rotation to non-hosts (Bowers et al. 1990; Lamour and Hausbeck 2003). In general, P. capsici requires the interaction of A1 and A2 mating types to produce oospores (Hausbeck and Lamour 2004), although there is evidence of apomixis (Hurtado-Gonzales and Lamour 2009) and homothallic oospore production (Islam et al. 2005). Both mating types are often present in the same field, and in near-equal ratios, at farms in Massachusetts and the surrounding region (New York, Pennsylvania and Connecticut) (Pan 1997; Hausbeck and Lamour 2004; Gobena et al. 2012a; Dunn et al. 2010). In addition to oospores, some strains of P. capsici can produce asexual survival spores (chlamydospores) that have been reported from Illinois and could enable the persistence of highly virulent clonal lineages (Islam et al. 2005).
Population studies of P. capsici in the middle and eastern US indicate that clonal lineages generally do not overwinter (Dunn et al. 2010; Gobena et al. 2012a; Granke et al. 2012; Lamour and Hausbeck 2002; Quesada-Ocampo et al. 2011). The situation in North America is significantly different from populations in the Latin American countries of Peru and Argentina and in central China, where clonal lineages are widely dispersed and survive multiple years (Hulvey et al. 2011; Gobena et al. 2012b; Hurtado-Gonzalez et al. 2008). Our objective was to assess the genetic structure of P. capsici populations in New England (primarily Massachusetts) to better understand pathogen survival and spread.
Materials and methods
Isolate collection, DNA extraction and mating type determination
Infected plant material (root, crown, stem and fruit) with typical symptoms of Phytophthora blight were collected from farms in the New England region from 1997 to 2014. Each farm was assigned a unique identifier as follows: AQ: Aqua Vita Road, BG: Bagdon Farm, CU2: CU-2, FPP: Foppoma, GD: Gardner, HF: Harvest Farm, KG: Kitchen Garden, MED: Medeiros, MRF: Marini Farm, MTF: Matusko’s Farm, NBO: Next Barn Over, PTR: Plumtree Road, RLF: Riverland Farm, S: S1-S9, SS: Stone Soup, UMF: UMass Farm, UNK: Unkown, VRF: Verrill Farm. The collected material was transported to the laboratory for isolation. Small sections (≈1 cm2) of infected tissue were plated onto PARP-V8 agar media (160 mL V8 juice, 3 g CaCO3, 25 ppm pimaricin, 100 ppm ampicillin, 25 ppm rifampicin, 25 ppm pentachloronitrobenzene and 840 mL deionized water) and the plates checked daily for mycelial growth. Mycelium was sub-cultured as hyphal-tips onto new plates and a unique identifier assigned.
For DNA extraction, plates with mycelia of 7 days growth were flooded with lysis buffer and mycelium was scraped from plates and transferred into 1.5-mL tubes. Genomic DNA was extracted using a modified chloroform/isoamyl fungal protocol (Hulvey et al. 2011). DNA quality was assessed with a Quawell Q3000 micro-volume spectrophotometer (Quawell Technology, San Jose, USA). Mating type was determined by growing isolates on PARP-V8 agar with known A1 and A2 isolates and observing the margin between the growing mycelium for the presence/absence of amphigynous oospores under a light microscope.
Targeted sequencing and genotyping
Genomic DNA samples with a concentration of at least 5 ng/μL and a 260/280 ratio of ~1.8 were submitted to Floodlight Genomics (Floodlight Genomics, Knoxville, USA) for targeted sequencing for 45 single nucleotide polymorphism (SNP) loci based on P. capsici LT1534 genome v.11.0 available from the Joint Genome Institute (http://genome.jgi-psf.org/) which were previously used in other P. capsici population studies (Castro-Rocha et al. 2016; Hu et al. 2013). A subset of the isolates was duplicated (blindly) and all samples were processed twice. Floodlight Genomics PCR amplifies targets (80–100 bp) in a multiplexed PCR reaction and then generates sample-specific sequences using a next-generation sequencing device (e.g., Ion Proton or Illumina MiSeq). The sample amplifications and targeted sequencing were carried out at no charge as part of the Floodlight Genomics Educational and Research Outreach Program.
The raw sample-specific sequence data were made available via an FTP site and processed using CLC Genomics Workbench v.8.0 (CLC bio, Aarhus, Denmark) to map reads to a reduced representation reference genome composed solely of the genetic target sequences. The mapping parameters were set at 90% length and 90% similarity. After mapping, the genotypes were assigned for target sites with at least 20× coverage. Sites with <15% alternate allele were considered homozygous and >15% and <85% were considered heterozygous.
Data analysis
Genetic data were analyzed using GENALEX 6.5 to identify identical multi-locus genotypes (Peakall and Smouse 2006). Isolates with identical genotypes are considered members of the same mitotically-derived asexual clonal lineage (CL) and were assigned identifiers (CL-1, CL-2, etc.) and duplicate genotypes were removed before conducting further analysis. Genotypes were grouped into populations based on the farm from which the samples were collected. Bayesian algorithms were used to determine the most probable number of genetic clusters (K) among the populations using Structure 2.3.4, and to determine population outcrossing rate (t) and inbreeding coefficient (F) with the program BORICE using the default parameters and assuming isolates from the same farm belonged to the same population (Pritchard et al. 2000; Koelling et al. 2012). Structure analysis was conducted with the parameters set to no prior population information using an admixture model with allele frequencies correlated. Isolate collection site and hosts data were included in the structure analysis. The alpha value was set to 1 and simulations from K = 1 to K = 50 for 20 iterations with 200,000 burn-in cycles and 200,000 Markov Chain Monte Carlo runs were computed. Structure results were tested with Structure Harvester to select the most probable K (Earl and vonHoldt 2012). A condensed neighbor-joining tree of the multi-locus genotypes was determined using Mega 6.06 and using 1000 Bootstrap repetitions with the maximum composite likelihood method at default parameters and a cut-off value equal to 50% (Saitou and Nei 1987; Tamura et al. 2004, 2013). Neighbor net and parsimony splits phylogenetic networks were constructed from the multi-locus genotypes with SplitsTree4 at default parameters (Huson and Bryant 2006). GENALEX 6.5 was used to determine pairwise population fixation index (F ST) values, Nei’s genetic distances, Shannon diversity estimate and principal coordinate analysis (PCoA) based on site of collection. GENALEX was also used to calculate the allele frequencies for each marker when all unique multi-locus genotypes were combined and to test for deviations from Hardy–Weinberg equilibrium at 0.05, 0.01 and 0.001 levels of significance.
Results
A total of 210 P. capsici isolates were recovered from 18 sites (16 in Massachusetts and 2 on Long Island, New York) and genotyped for 45 polymorphic SNP loci (Tables 1, 2; Supplementary Fig. 1). All samples were genotyped in duplicate and showed identical multi-locus genotypes. A total of 86 genotypes were unique and 34 were repeated (clonal lineages) (Table 3). The number of duplicate genotypes ranged from 2 to 16 and most members of a clonal lineage were confined to single locations (Table 3). Both the A1 and A2 mating types were recovered from 7 of the 18 locations (Table 3). When combining all isolates, the allele frequencies for the clone-corrected set of data averaged 0.23, and 60% of the markers did not differ from the expectations of Hardy–Weinberg equilibrium (Table 2). Bayesian clustering analysis determined K = 38 (Delta K = 1214.23) as the most probable number of genetic clusters among the unique genotypes (Supplementary Fig. 2). The Bayesian clustering bar graph shows most populations have individuals with unique genetic clusters (Fig. 1). However, within those same populations appear individuals with a complex genetic cluster composition, typical for populations with outcrossing and intermixing. A high population outcrossing rate (mean t = 0.87) with a correspondingly low inbreeding coefficient (mean F = 0.07) also indicates that there is likely outcrossing among the populations. Phylogenetic analysis, either tree or networks (Fig. 2; Supplementary Figs. 3, 4, 5), grouped most isolates of the same farm closely together, indicating some degree of population differentiation, but this may be due to the relatively small number of isolates analyzed from individual locations. Clustering, from a host perspective, shows three groups for squash, three groups for pepper, two groups for pumpkin, one group for tomato and one group for melon' however, clustering of the same host at different locations do not group together. Pairwise population F ST values and Nei genetic distances indicate different degrees of differentiation between the populations (Supplementary Tables 1, 2). The values obtained from both metrics correspond to the phylogenetic analysis, which suggests that sufficient differentiation has occurred among the populations to distinguish them during phylogenetic reconstruction. Shannon’s diversity estimate indicates a lower diversity among populations (sH = 1.126) than within populations (sH = 1.548), which suggests inbreeding within farm populations is higher than outcrossing among farms (Supplementary Table 3). The PCoA supports the notion that, even though the populations have enough differentiation to be independently clustered one from another, some individuals appear intermediate between the clusters of multiple populations suggesting outcrossing among farms (Supplementary Fig. 5).
Bayesian clustering of the unique multi-locus genotypes obtained from P. capsici isolates recovered from infected farms (1 Aqua Vita Road, 2 Bagdon Farm, 3 CU-2, 4 Foppoma, 5 Gardner, 6 Harvest Farm, 7 Kitchen Garden, 8 Medeiros, 9 Marini Farm, 10 Matusko’s Farm, 11 Next Barn Over, 12 Plumtree Road, 13 Riverland Farm, 14 S1–S9, 15 Stone Soup, 16 UMass Farm, 17 Unkown, 18 Verrill Farm)
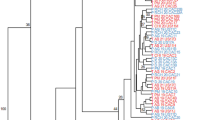
Phylogenetic analysis of unique multi-locus genotypes obtained from P. capsici isolates. Evolutionary history inferred using the neighbor-joining method computed with MEGA6 (optimal tree sum of branch length = 9.16365606). (AQ Aqua Vita Road, BG Bagdon Farm, CU2 CU-2, FPP Foppoma, GD Gardner, HF Harvest Farm, KG Kitchen Garden, MED Medeiros, MRF Marini Farm, MTF Matusko’s Farm, NBO Next Barn Over, PTR Plumtree Road, RLF Riverland Farm, S S1–S9, SS Stone Soup, UMF UMass Farm, UNK Unkown, VRF Verrill Farm)
Discussion
Overall, our results indicate that the survival and spread of P. capsici in Massachusetts is like the Midwest and Northeastern US, where sexual reproduction is crucial for overwintering and the dispersal of asexual sporangia and swimming zoospores is limited to the same or nearby farms (Dunn et al. 2010; Gobena et al. 2012a; Granke et al. 2012; Lamour and Hausbeck 2002; Quesada-Ocampo et al. 2011). Evidence supporting sexual reproduction includes the finding of both the A1 and A2 mating types at multiple locations, the overall high proportion of unique multi-locus genotypes and the limited survival and spread of clonal lineages. The importance of sexual oospores is not surprising as there are no hosts, such as rocoto (C. pubescens) in Peru, to serve as a “green bridge” through the fallow winter months (Hurtado-Gonzalez et al. 2008; Hulvey et al. 2011). This does not bode well for sustainable vegetable production in Massachusetts, because once P. capsici has been introduced to a new site, the thick-walled oospores can persist for many years and planting susceptible hosts incurs the risk of total crop failure (Lamour and Hausbeck 2000).
Once the growing season is underway and oospores germinate and colonize host tissue, the rise in frequency of clonal lineages is common, as P. capsici produces massive numbers of deciduous sporangia on the surface of infected vegetable fruit (Granke et al. 2012). Although we identified multiple clonal lineages, they were generally limited to a single farm location and were not found across multiple years. This same trend has been recorded in New York and Michigan (Dunn et al. 2010; Gobena et al. 2012a; Lamour and Hausbeck 2001, 2003; Quesada-Ocampo et al. 2011), and suggests that P. capsici is not actively dispersed over long distances. The three clonal lineages found at multiple sites were recovered from farms no more than ~5 miles (8 km) apart.
An important question is how vegetable blight is spreading to new locations. One factor may be equipment sharing. Due to the small scale of most farming operations in New England, equipment is shared to reduce operating costs (Buckler 2012). In addition, a single operator may plow multiple fields, used for diverse crops, in the spring and fall. From 2002 to 2007, the average area of farmland in Massachusetts decreased from 85 to 67 acres (34 to 27 ha), with the decrease attributed, in part, to a rising number of small farms in the state (USDA 2007). During that same 5-year period, market value of vegetable products increased from $36.4 to $58.9 million, driven by increasing interest in local and organic vegetables (USDA 2007). With an increasing number of small farms using shared equipment, the potential to spread soilborne pathogens like P. capsici is high. Irrigation water from above-ground sources can also be contaminated with P. capsici inoculum and must be considered (Gevens et al. 2007; Wang et al. 2009).
Another factor, difficult to assess due to the explosive epidemiology of P. capsici, is the potential for inoculum to be seedborne. Leonian states in the 1922 species description that P. capsici on pepper can be seedborne, and farmers in Tennessee (2004) and most recently, Maryland (2017), report to K. Lamour (quite emphatically) that P. capsici entered their farm through a single lot of pepper (TN, 2004) or melon (MA, 2017) seed. In both scenarios, the blight started at specific (previously un-infested) locations in the field where a new variety was planted. In the case of pepper in Tennessee, the farmer identified the single pepper plant that showed initial wilt symptoms (located on well-drained, elevated ground) and described the subsequent movement down the row and, eventually, throughout his entire field, which included multiple varieties of pepper, tomato and squash hosts. Like the findings here, the isolates recovered from this epidemic showed no genetic structuring based on host (K. Lamour, unreported data).
The obvious goal in New England is to halt the movement of not just P. capsici, but of both mating types of P. capsici into a new location. Based on two recent findings, preventing the movement of both mating types in P. capsici may be more challenging than previously recognized. First, a multi-year study in New York characterizing the genetic diversity and inbreeding of a controlled bi-parental field population found a specific mating type region (MTR) of the P. capsici genome (previously identified via genetic mapping) that maintains a high level of heterozygosity for A2 mating types; despite obvious inbreeding across the rest of the genome (Carlson et al. 2017; Lamour et al. 2012). Second, recent studies of vegetable blight in Taiwan and taro leaf blight (P. colocasiae) describe a novel dimension of plasticity for the Phytophthora genome: intra-genomic variations in ploidy (Barchenger et al. 2017; Shrestha et al. 2017). It appears that the P. capsici genome expands and contracts on a local (chromosome) scale during asexual clonal reproduction. Historical Taiwanese isolates were primarily of the A1 mating type and diploid across the 18 linkage groups of the P. capsici genome (Barchenger et al. 2017). In recent years, a shift occurred across the island and the 2016 populations were almost exclusively the A2 mating type. The change in mating type is likely related to increased ploidy across the MTR, and ploidy varied from diploid to triploid to tetraploid (and higher in P. colocasiae) within individual genomes.
The importance of these recent findings to New England (and elsewhere) is that P. capsici, due to expansion and contraction of the genome across different chromosomes (particularly the MTR) during clonal reproduction, may be able to generate an opposite mating type when only a single mating type has been introduced to a new site. More work in this area is needed, as an epidemic with both mating types will produce long-lived, thick-walled oospore and, in most cases, lead to the loss of the use of a field for vegetable production, indefinitely.
References
Babadoost M (2004) Phytophthora blight: a serious threat to cucurbit industries. APSnet Features. Online at: http://www.apsnet.org/publications/apsnetfeatures/Pages/PhytophthoraBlight.aspx. Accessed 12 March 2017
Barchenger, DW, Lamour K, Sheu Z-M, Shrestha S, Kumar S, Lin S-W, Burlakoti RR, Bosland PW (2017) Intra- and intergenomic variation of ploidy and clonality characterize Phytophthora capsici on Capsicum sp. in Taiwan. Mycol Prog 16:893–904. https://doi.org/10.1007/s11557-017-1323-z
Bowers JH, Papavizas GC, Johnston SA (1990) Effect of soil temperature and soil-water matric potential on the survival of Phytophthora capsici in natural soil. Plant Dis 74:771–777. https://doi.org/10.1094/PD-74-0771
Buckler B (2012) Farm equipment needs and cooperative solutions for southern New England. Northern Rhode Island Conservation District. http://www.nricd.org/EquipmentBankStudyReport.pdf. Accessed 12 March 2017
Carlson MO, Gazave E, Gore MA, Smart CD (2017) Temporal genetic dynamics of an experimental, biparental field populations of Phytophthora capsici. Front Genet 8. https://doi.org/10.3389/fgene.2017.00026
Castro-Rocha A, Shrestha S, Lyon B, Grimaldo-Pantoja GL, Flores-Marges JP, Valero-Galván J, Aguirre-Ramírez M, Osuna-Ávila P, Gómez-Dorantes N, Ávila-Quezada G, Luna-Ruíz JJ, Rodríguez-Alvarado G, Fernández-Pavía SP, Lamour KH (2016) An initial assessment of genetic diversity for Phytophthora capsici in northern and central Mexico. Mycol Prog 15:15. https://doi.org/10.1007/s11557-016-1157-0
Dunn AR, Milgroom MG, Meitz JC, McLeod A, Fry WE, McGrath MT, Dillard HR, Smart CD (2010) Population structure and resistance to mefenoxam of Phytophthora capsici in New York state. Plant Dis 94:1461–1468. https://doi.org/10.1094/PDIS-03-10-0221
Earl DA, vonHoldt BM (2012) STRUCTURE HARVESTER: a website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv Genet Resour 4:359–361. https://doi.org/10.1007/s12686-011-9548-7
Gevens AJ, Donahoo RS, Lamour KH, Hausbeck MK (2007) Characterization of Phytophthora capsici from Michigan surface irrigation water. Phytopathology 97:421–428. https://doi.org/10.1094/PHYTO-97-4-0421
Gobena D, McGrath MT, Lamour KH (2012a) Survival and spread of Phytophthora capsici on Long Island, New York. Mycol Prog 11:761–768. https://doi.org/10.1007/s11557-011-0787-5
Gobena D, Roig J, Galmarini C, Hulvey J, Lamour K (2012b) Genetic diversity of Phytophthora capsici isolates from pepper and pumpkin in Argentina. Mycologia 104:102–107. https://doi.org/10.3852/11-147
Granke LL, Quesada-Ocampo LM, Lamour KH, Hausbeck MK (2012) Advances in research on Phytophthora capsici on vegetable crops in the United States. Plant Dis 95:1588–1600. https://doi.org/10.1094/PDIS-02-12-0211-FE
Hausbeck MK, Lamour KH (2004) Phytophthora capsici on vegetable crops: research progress and management challenges. Plant Dis 88:1992–1303. https://doi.org/10.1094/PDIS.2004.88.12.1292
Hu J, Diao Y, Zhou Y, Lin D, Bi Y, Pang Z, Trout Fryxell R, Liu X, Lamour K (2013) Loss of heterozygosity drives clonal diversity of Phytophthora capsici in China. PLoS ONE 8:e82691. https://doi.org/10.1371/journal.pone.0082691
Hulvey J, Hurtado-Gonzalez O, Aragón-Caballero L, Gobena D, Storey D, Finley L, Lamour K (2011) Genetic diversity of the pepper pathogen Phytophthora capsici on farms in the Amazonian high jungle of Peru. Am J Plant Sci 2:461–466. https://doi.org/10.4236/ajps.2011.23054
Hurtado-Gonzales O, Lamour KH (2009) Evidence for inbreeding and apomixis in close crosses of Phytophthora capsici. Plant Pathol 58:715–722. https://doi.org/10.1111/j.1365-3059.2009.02059.x
Hurtado-Gonzalez O, Aragon-Caballero L, Apaza-Tapia W, Donahoo R, Lamour K (2008) Survival and spread of Phytophthora capsici in coastal Peru. Phytopathology 98:688–694. https://doi.org/10.1094/PHYTO-98-6-0688
Huson DH, Bryant D (2006) Application of phylogenetic networks in evolutionary studies. Mol Biol Evol 23:254–267. https://doi.org/10.1093/molbev/msj030
Islam SZ, Babadoost M, Lambert KN, Ndeme A (2005) Characterization of Phytophthora capsici isolates from processing pumpkin in Illinois. Plant Dis 89:191–197. https://doi.org/10.1094/PD-89-0191
Koelling VA, Monnahan PJ, Kelly JK (2012) A Bayesian method for the joint estimation of outcrossing rate and inbreeding depression. Heredity 109:393–400. https://doi.org/10.1038/hdy.2012.58
Lamour KH (2013) Phytophthora: a global perspective. CABI, Wallingford
Lamour KH, Hausbeck MK (2000) Mefenoxam insensitivity and the sexual stage of Phytophthora capsici in Michigan cucurbit fields. Phytopathology 90:396–400. https://doi.org/10.1094/PHYTO.2000.90.4.396
Lamour KH, Hausbeck MK (2001) Investigating the spatiotemporal genetic structure of Phytophthora capsici in Michigan. Phytopathology 91:973–980. https://doi.org/10.1094/PHYTO.2001.91.10.973
Lamour KH, Hausbeck MK (2002) The spatiotemporal genetic structure of Phytophthora capsici in Michigan and implications for disease management. Phytopathology 92:681–684. https://doi.org/10.1094/PHYTO.2002.92.6.681
Lamour KH, Hausbeck MK (2003) Effect of crop rotation on the survival of Phytophthora capsici in Michigan. Plant Dis 87:841–845. https://doi.org/10.1094/PDIS.2003.87.7.841
Lamour KH, Mudge J, Gobena D, Hurtado-Gonzáles OP, Schmutz J, Kuo A, Miller NA, Rice BJ, Raffaele S, Cano LM, Bharti AK, Donahoo RS, Finley S, Huitema E, Hulvey J, Platt D, Salamov A, Savidor A, Sharma R, Stam R, Storey D, Thines M, Win J, Haas BJ, Dinwiddie DL, Jenkins J, Knight JR, Affourtit JP, Han CS, Chertkov O, Lindquist EA, Detter C, Grigoriev IV, Kamoun S, Kingsmore SF (2012) Genome sequencing and mapping reveal loss of heterozygosity as a mechanism for rapid adaptation in the vegetable pathogen Phytophthora capsici. Mol Plant-Microbe Interact 25:1350–1360
Leonian LH (1922) Stem and fruit blight of peppers caused by Phytophthora capsici sp. nov. Phytopathology 12:401–408
McGrath MT (2001) Phytophthora blight of cucurbits. Cornell University Extension Factsheet. Online at: http://vegetablemdonline.ppath.cornell.edu/factsheets/Cucurbit_Phytoph2.htm. Accessed 12 March 2017
Pan Z (1997) Investigation of genetic variation in Phytophthora capsici. PhD Dissertation, University of Massachusetts, Amherst
Park J, Park B, Veeraraghaven N, Jung K, Lee YH, Blair JE, Geiser DM, Isard S, Mansfield MA, Nikolaeva E, Park SY, Russo J, Kim SH, Greene M, Ivors KL, Balci Y, Peiman M, Erwin DC, Coffey MD, Rossman A, Farr D, Cline E, Grunwald NJ, Luster DG, Schrandt J, Martin F, Ribeiro OK, Makalowska I, Kang S (2008) Phytophthora database: a forensic database supporting the identification and monitoring of Phytophthora. Plant Dis 92:966–972. https://doi.org/10.1094/PDIS-92-6-0966
Peakall R, Smouse PE (2006) GENALEX 6: genetic analysis in excel. Population genetic software for teaching and research. Mol Ecol Notes 6:288–295. https://doi.org/10.1111/j.1471-8286.2005.01155.x
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Quesada-Ocampo LM, Granke LL, Hausbeck MK (2011) Temporal genetic structure of Phytophthora capsici populations from a creek used for irrigation in Michigan. Plant Dis 95:1358–1369. https://doi.org/10.1094/PDIS-03-11-0191
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Shrestha S, Miyasaka SC, Shintaku M, Kelly H, Lamour K (2017) Phytophthora colocasiae from Vietnam, China, Hawaii and Nepal: intra- and inter-genomic variations in ploidy and a long-lived, diploid Hawaiian lineage. Mycol Prog. https://doi.org/10.1007/s11557-017-1323-z
Tamura K, Nei M, Kumar S (2004) Prospects for inferring very large phylogenies by using the neighbor-joining method. Proc Natl Acad Sci U S A 101:11030–11035. https://doi.org/10.1073/pnas.0404206101
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S (2013) MEGA6: molecular evolutionary genetics analysis version 6.0. Mol Biol Evol 30:2725–2729. https://doi.org/10.1093/molbev/mst197
UMass Extension (2016) Phytophthora blight. The center for agriculture, food and the environment. http://ag.umass.edu/vegetable/fact-sheets/phytophthora-blight. Accessed 12 March 2017
USDA Agricultural Census. 2007. Massachusetts Census Publications. http://www.agcensus.usda.gov/Publications/2007/Full_Report/Census_by_State/Massachusetts/. Accessed 12 March 2017
Wang Z, Langston DB, Csinos AS, Gitaitis RD, Walcott RR, Ji P (2009) Development of an improved isolation approach and simple sequence repeat markers to characterize Phytophthora capsici populations in irrigation ponds in southern Georgia. Appl Environ Microbiol 75:5467–5473. https://doi.org/10.1128/AEM.00620-09
Wick RL (1994) Diseases of vegetable crops: Phytophthora blight of cucurbits, peppers and tomato. Grower 9:4
Zitter TA (1989) Phytophthora blight of cucurbits, pepper, tomato and eggplant. Cornell University Extension Factsheet. Cornell University Extension Factsheet. http://vegetablemdonline.ppath.cornell.edu/factsheets/Cucurbit_Phytoph.htm. Accessed 12 March 2017
Acknowledgement
Permission to harvest infected plant material was obtained from all landowners.
Author information
Authors and Affiliations
Corresponding author
Additional information
Section Editor: Marco Thines
Electronic supplementary material
ESM 1
Supplementary Fig. 1 Location of Massachusetts farms from which P. capsici infected plant material was collected. Supplementary Fig. 2 Delta K graph for each K number of clusters for complete multi-locus data set of P. capsici isolates recovered from Massachusetts and Long Island, New York. Supplementary Fig. 3 Parsimony splits phylogenetic network computed with SplitsTree4 from infected farms P. capsici recovered isolates. (AQ: Aqua Vita Road, BG: Bagdon Farm, CU2: CU-2, FPP: Foppoma, GD: Gardner, HF: Harvest Farm, KG: Kitchen Garden, MED: Medeiros, MRF: Marini Farm, MTF: Matusko’s Farm, NBO: Next Barn Over, PTR: Plumtree Road, RLF: Riverland Farm, S: S1-S9, SS: Stone Soup, UMF: UMass Farm, UNK: Unkown, VRF: Verrill Farm). Supplementary Fig. 4 Neighbor net phylogenetic network computed with SplitsTree4 for unique P. capsici isolates. (AQ: Aqua Vita Road, BG: Bagdon Farm, CU2: CU-2, FPP: Foppoma, GD: Gardner, HF: Harvest Farm, KG: Kitchen Garden, MED: Medeiros, MRF: Marini Farm, MTF: Matusko’s Farm, NBO: Next Barn Over, PTR: Plumtree Road, RLF: Riverland Farm, S: S1-S9, SS: Stone Soup, UMF: UMass Farm, UNK: Unkown, VRF: Verrill Farm). Supplementary Fig. 5 Principal coordinates analysis of the unique multi-locus genotypes obtained from New England P. capsici isolates. (AQ: Aqua Vita Road, BG: Bagdon Farm, CU2: CU-2, FPP: Foppoma, GD: Gardner, HF: Harvest Farm, KG: Kitchen Garden, MED: Medeiros, MRF: Marini Farm, MTF: Matusko’s Farm, NBO: Next Barn Over, PTR: Plumtree Road, RLF: Riverland Farm, S: S1-S9, SS: Stone Soup, UMF: UMass Farm, UNK: Unkown, VRF: Verrill Farm). (PPTX 950 kb)
ESM 2
(DOCX 20 kb)
Rights and permissions
About this article
Cite this article
Castro-Rocha, A., Hulvey, J.P., Wick, R. et al. Genetic diversity of Phytophthora capsici recovered from Massachusetts between 1997 and 2014. Mycol Progress 16, 999–1006 (2017). https://doi.org/10.1007/s11557-017-1334-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11557-017-1334-9