Abstract
Background
Sorafenib represents one of the therapeutic strongholds for advanced hepatocellular carcinoma (HCC), but unfortunately, predictive factors are lacking. We previously reported that the VEGF single nucleotide polymorphisms (SNPs) rs2010963 and rs4604006 might correlate with clinical outcomes in sorafenib-treated HCC patients.
Objective
The objective of the ALICE-2 study is to define a prognostic angiogenesis profile to better identify HCC patients who are more likely to benefit from sorafenib treatment.
Patients and methods
From 2008 to 2015, all consecutive HCC patients receiving sorafenib according to the Italian label were tested for specific HIF-1α, VEGF, and VEGFR SNPs. Results from angiogenesis genotyping were then correlated with clinical outcome parameters.
Results
Globally, a total of 210 patients were enrolled. At multivariate analysis rs12434438 of HIF1α, rs2010963 of VEGF-A, and rs4604006 of VEGF-C were confirmed as independent predictive factors. At the combined analysis of significant SNPs, the presence of two favourable alleles of rs2010963 and rs4604006 of VEGF compared to only one or to none favourable alleles, was able to identify three separate patients populations with different time-to-progression (TTP) (10.8 vs. 5.6 vs. 3.7 months, respectively; p < 0.0001) and overall survival (OS) (19.0 vs. 13.5 vs. 7.5 months, respectively; p < 0.0001). Furthermore, the presence of the GG genotype of rs12434438 (HIF-1α) seemed able to select a population with a particularly poor outcome, independently from the clinical effect of the two VEGF SNPs (TTP: 2.6 months, HR: 0.54, p = 0.0374; OS: 6.6 months, p = 0.0061, HR: 0.43).
Conclusions
Our findings show that polymorphism analysis of HIF-1α, VEGF, and VEGFR genes may represent a prognostic panel to better identify HCC patients who are more likely to benefit from sorafenib treatment.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Over the last few years, the growing knowledge of molecular mechanisms regulating the onset and neoplastic progression of hepatocellular carcinoma (HCC) has been applied to the development of anticancer treatment strategies directed against new and more specific biological targets [1, 2]. Following the positive results of the SHARP [3] and the Asia Pacific trials [4], sorafenib, a multitarget tyrosine-kinase inhibitor (TKI) [2], was approved by regulatory authorities for the treatment of advanced HCC patients worldwide. Sorafenib has since then been considered the standard of care for first-line therapy of HCC patients who are Barcelona Clinic Liver Cancer (BCLC) advanced or intermediate, and refractory to locoregional therapy.
Sorafenib targets the RAF serine/threonine kinases (A-RAF, B-RAF, C-RAF) and inhibits tyrosine kinase receptors, such as vascular endothelial growth factor receptor (VEGFR) 2, VEGFR 3, platelet-derived growth factor receptor-β (PDGFR-β), Flt3, and c-Kit [5, 6]. The inhibition of VEGFR and PDGFR, mainly responsible for the antiangiogenic effect, is a crucial driver of the therapeutic activity of sorafenib. Unfortunately, not all patients seem to benefit from this treatment and are thus exposed to unnecessary toxicities without deriving any clinical improvement. Recently, several additional systemic treatment options have been trialled and approved in the first- and later-line treatment of HCC [7,8,9,10,11,12,13,14,15,16,17,18] (Table 1). Against this background, clinical or biological markers able to identify responding/resistant patients are an unmet medical need.
To date, many studies have investigated the role of different predictive factors for sorafenib efficacy. Besides its off-target effects and the Barcelona Clinic Liver Cancer stage, the most interesting data are arising from the analysis of the tumour-driven angiogenesis process, which is one of the main biological features of HCC [19]. Angiogenesis is largely regulated by cellular signalling mediated by the vascular endothelial growth factor (VEGF) and its receptors, VEGFR1 and VEGFR2, which are also the major therapeutic targets of sorafenib [20, 21]. In this context, hypoxia inducible factors (HIFs) seem to play a leading role as well. The HIF-family includes different transcriptional regulators that are sensitive to cellular oxygen levels. HIF-1 is the major HIF isoform in humans, and it has the ability to promote the transcription of a wide range of target genes involved in neoangiogenesis, glucose metabolism, cell proliferation and apoptosis [22]. The molecular structure of HIF-1 comprises two subunits: HIF-1α, which is sensitive to low oxygen levels, and HIF-1β, which is constitutively expressed [23]. Preclinical data indicated that VEGF is one of the major target genes influenced by HIF-1α. These findings also showed a correlation between HIF-1α and HCC through the activation of tumour-induced angiogenesis [24].
We previously presented the results of the ALICE-1 study [25], which evaluated the role of VEGF and VEGFR polymorphisms in determining the clinical outcome of HCC patients receiving sorafenib. In this study, the C allele of rs2010963 (VEGF-A) and the T allele of rs4604006 (VEGF-C) showed significant correlation with time-to-progression (TTP) and overall survival (OS) among 148 HCC patients receiving sorafenib.
Based on these findings, we conducted the present study with the aim of defining a prognostic angiogenesis profile to better identify HCC patients who are more likely to benefit from sorafenib treatment. The aim of the ALICE 2 study was then to ascertain whether HIF-1α single nucleotide polymorphisms (SNPs) along with VEGF and VEGFR SNPs might have a role in influencing the global outcome of advanced HCC patients receiving sorafenib.
2 Methods
2.1 Aim of the Study and Outcome Measures
The aim of the present analysis was to evaluate the role of HIF-1α, VEGF, and VEGFR SNPs in predicting the clinical outcome of HCC patients treated with sorafenib. The primary endpoint was TTP (as defined in the Statistical Analysis section). The secondary endpoints were OS (as defined in the Statistical Analysis section) and objective response rate.
2.2 Patient Selection
ALICE-2 (Angiogenesis LIver CancEr 2) was a retrospective multicentre study conducted in eight centres in Italy. From 2008 to 2015 all consecutive patients with advanced HCC or intermediate stage HCC refractory to or unsuitable for locoregional therapies, either histologically proven or diagnosed according to the American Association for the Study of Liver Diseases 2005 (AASLD) guidelines and receiving sorafenib according to the Italian label were eligible for our analysis.
The study was performed in accordance with the study protocol, which was approved by the local ethics committee (Comitato Etico della Romagna—C.E.ROM.) along with all experimental procedures. Written informed consent was obtained for all patients enrolled into the analysis and methods were carried out in accordance with the Declaration of Helsinki.
Follow-up consisted of physical examination and complete blood count every 3 weeks and CT/MRI scan every 8 weeks or as clinically indicated. Tumour response was evaluated by clinicians according to the modified Response Evaluation Criteria in Solid Tumours (mRECIST). All patients received sorafenib at a standard dose (400 mg twice daily continuously). According to the European Medical Agency (EMA) sorafenib technical summary and to the manufacturer’s instructions, dose reductions were applied as clinically indicated. Two levels of dose reduction were considered, 400 mg once daily as the first level and a single 400-mg dose every other day as the second level. In case of toxicity (grade 2 with no improvement after symptomatic therapy and grade 3) sorafenib was discontinued until toxicity resolved to grade 0–1 then the treatment was resumed applying the first level of dose reduction. If toxicity persisted after the first dose reduction, the second level was applied. Permanent interruption was considered in grade 4 toxicities. Toxicity was evaluated with the National Cancer Institute Common Terminology Criteria for Adverse Events (NCI-CTCAE).
2.3 Genotyping Analysis
HIF-1α, VEGF and VEGFR genotyping was performed using DNA extracted from formalin-fixed paraffin-embedded liver tissue blocks (about 30 mg) of HCC or from blood samples. For tissue blocks, paraffin wax was removed with xylene and the samples were washed twice with 100% ethanol. DNA was isolated from the deparaffinised tissue using the RecoverAll™ Total Nucleic Acid Isolation Kit for FFPE Tissues (Applied Biosystems, Foster City, CA, USA), according to the manufacturer’s instructions. DNA from each sample was then eluted in 120 μl of eluting solution. For blood samples, genomic DNA was extracted from 2 ml of whole blood by FlexiGene DNA kit (Qiagen Inc., Valencia, CA, USA), following the manufacturer’s instructions. Concentration and purity index of each sample were evaluated by UV spectrophotometry as the ratio absorbance 260/280 nm; a purity index of 1.5–2.0 was considered optimal.
Single nucleotide polymorphisms within each gene were selected using the Pupasuite software (http://pupasuite.bioinfo.cipf.es/index.jsf—version 2.0.0. bioinfo 2008), the CIPF (Centro de Investigacion Principe Felipe) Single Nucleotide Polymorphism database (dbSNP) generated by the National Centre for Biotechnology Information (http://www.ncbi.nlm.nih.gov/SNP) and by review of the medical literature, using the following criteria:
-
1.
The polymorphism had some degree of likelihood to alter the structure or the expression of the gene in a biologically relevant manner (i.e. affecting its sequence, 3′-untranslated region (UTR), or promoter region);
-
2.
the minor allele frequency was above 10% (with the only exception of rs2305948, rs6877011, and rs307822);
-
3.
the genetic polymorphism was established and well documented.
Further considerations led the selection of SNPs for our study. A correlation between the presence of a specific allele on a polymorphic site and the expression of the respective protein has been previously documented for VEGF [26, 27].
SNPs regulatory sequences, including introns, 5′- and 3′-UTR, have been demonstrated to influence mRNA stability, elaboration efficiency, expression and location of the isoform. Moreover, regulatory sequences in mRNA 3′-UTR were shown to influence the messenger stability and/or its translational efficiency. In this way, we can suppose that SNPs in these sequences can influence genic expression. Globally, we hypothesized that selected SNPs can impact on protein expression and thus on the biological function.
Selected SNPs, chromosomal locations, positions, and biological effects are summarised in Table 2.
SNP genotyping was performed by TaqMan technology, using a SNP genotyping assay (Applied Biosystems). Polymerase chain reaction (PCR) was performed and genotypes were analysed on the 7300 Real-Time PCR System (Applied Biosystems) using ABI Prism 7300 Sequence Detection System software (version 1.3.1, Applied Biosystems). Each reaction contained 0.2 μl of total genomic DNA. Laboratory personnel blinded to patient status performed genotyping, and a random 10% of the samples were repeated to validate genotyping procedures. All SNPs genotyped had to present an overall call rate of ≥ 90% to be included in our analysis.
2.4 Data Management and Statistical Analysis
In order to detect a difference in the effect size (hazard ratio) with statistical significance in the proportion of patients without disease progression at 6 months according to genotyping and assuming a 6 months TTP of 45% in the sorafenib population and ≥ 70% as a target, at least 96 patients were necessary with α = 0.05 and β = 0.05. A p value < 0.05 was considered statistically significant. Statistical analysis was performed with MedCalc software version 13.1.2 for Windows. The association between categorical variables was estimated by χ2 test. Survival probability over time was estimated by the Kaplan–Meier method. Significant differences in probability of survival between the strata were evaluated by log-rank test. Cox’s multiple regression analysis was used to assess the role of polymorphisms as prognostic factor adjusted for those variables resulted significant at univariate analysis. The Bonferroni correction was used to adjust the values for multiple comparisons.
After a literature review, including in particular the registration trials [3, 4] we selected and tested the following variables: gender (male vs. female), median age (≥ 69 vs. < 69 years), BCLC (Barcelona Clinic Liver Cancer) stage (B vs. C), aetiology (HCV vs. HBV vs. alcoholic vs. cryptogenic/metabolic vs. multiple aetiology), co-morbidities (> 5%: cardiovascular vs. diabetes vs. other previous neoplasm), serum α-FP level (≥ 400 ng/ml vs. < 400 ng/ml), type of samples (tumour tissues vs. peripheral blood), sorafenib dose reduction applied (yes vs. no), prior loco-regional transarterial chemoembolisation (TACE) treatments (yes vs. no), major target lesion diameter (> 5 cm vs. ≤ 5 cm), portal invasion (yes vs. no), extrahepatic spread (yes vs. no), metastatic site (lymph nodes vs. lung vs. bone vs. peritoneum), Child–Pugh class (A5 vs. A6), aspartate transaminase (AST) serum level (< upper normal limit (UNL) vs. < 2 × UNL), alanine transaminase (ALT) serum level (< UNL vs. < 2 × UNL), overall toxicity profile (no toxicity vs. any grade); (no toxicity vs. grade 1–2 vs. grade > 2); skin toxicity (no toxicity vs. any grade), (no toxicity vs. early onset < 1 months vs. late onset > 1 months), Japan Red Cross (JRC) prognostic score (low risk vs. intermediate risk vs. high risk), hepatoma arterial embolisation prognostic (HAP) score (A vs. B vs. C vs. D).
For statistical analysis, OS and TTP were respectively defined as the interval between the date of beginning of sorafenib treatment to death or last follow-up visit, and to clinical progression or last follow-up visit if not progressed.
The DCR was defined as the percentage of patients who had a best response rating of complete response, partial response or stable disease (according to mRECIST).
All genetic polymorphisms were examined for deviation from Hardy–Weinberg equilibrium using the Powermarker v. 3.25 package (www.statgen.ncsu.edu/powermarker) [28]. Linkage Disequilibrium (LD) analysis was also performed using the Powermarker v. 3.25 package. LD was estimated using r2, with r2 = 1 indicating complete LD and r2 = 0 indicating absent LD.
3 Results
3.1 Patient Characteristics
From 2008 to 2015, 210 Caucasian patients diagnosed with HCC and receiving sorafenib were available for our analysis: 178 males (85%) and 32 females (15%); median age at diagnosis was 69 years (range 41–86). The median follow-up time was 11.3 months. All patients belonged to Child–Pugh class A. The main demographic and clinical characteristics of the study population are summarized in Table 3.
In the entire study population, median TTP was 5.2 months (progression events: 100%), and median OS was 12.6 months (death events: 100%). Regarding the type of progression, 197 (94%) patients showed an increase of target lesions parameters, and 124 (59%) showed an increase in the number of liver nodules. Of 109 patients without extrahepatic spread at treatment initiation, 29 (27%) developed extrahepatic disease.
3.2 Toxicities and Dose Reductions
The toxicity profile for sorafenib in the study population is shown in Table 4. First- and second-level dose reductions were applied in 57 (27%) and 23 (11%) patients, respectively. All dose reductions were applied for the management of toxicity. Patients who underwent dose reductions continued sorafenib at the lower dose until disease progression, and only one patient interrupted sorafenib treatment after 5 weeks for grade 4 liver toxicity. Median treatment duration, considering discontinuation periods, was 4.8 months.
3.3 SNPs, Hardy–Weinberg Equilibrium, and Linkage Disequilibrium
Globally, 25 SNPs met our selection criteria: eight SNPs of HIF-1α, six SNPs of VEGF-A, two SNPs of VEGF-C, two SNPs of VEGFR1, four SNPs of VEGFR2, and three SNPs of VEGFR3. Chromosomal locations and, when known, the biological effect of these SNPs are reported in Table 2. Genotyping was performed using DNA extracted from formalin-fixed paraffin-embedded liver tissue blocks of HCC in 67/210 patients (32%) or from blood samples in 143/210 patients (68%). All SNPs genotyped presented an overall call rate ≥ 90%. We evaluated concentration and purity index of each sample by UV spectrophotometry as the ratio absorbance 260/280 nm. All samples presented a purity index between 1.5 and 2.0. The frequencies of the tested genotypes were comparable to those reported in Caucasians, with no significant deviation from the Hardy–Weinberg equilibrium. Linkage disequilibrium was observed for the tumour genotypes rs12434438, rs11158358, and rs10873142 of HIF-1α (p < 0.0001); rs833061, rs699947, and rs2010963 of VEGF-A (p < 0.0001), rs4604006 and rs7664413 of VEGF-C (p < 0.0001).
3.4 Univariate Analysis
At univariate analysis, 11 SNPs showed to be statistically significant in the prediction of TTP and OS as detailed in Table 5. A statistically significant difference was found for TTP and OS according to the BCLC stage B vs. C. (TTP: 8.3 months stage B vs. 4.9 months stage C, p = 0.027; OS: 14.7 months stage B vs. 10.7 months stage C, p = 0.008). No differences were found for the other clinical variables tested, including JRC and HAP scores (Supplementary Table S1).
3.5 Multivariate Analysis
Cox multiple regression analysis was used to assess the role of variables that were found to be significant in the univariate analysis (Table 5). rs12434438 (HIF-1α) (Fig. 1a, b), rs2010963 (VEGF-A) (Fig. 1c, d), and rs4604006 (VEGF-C) (Fig. 1e, f) were independent prognostic factors, able to predict clinical outcome in terms of TTP (p = 0.0374, p = 0.0008, and p = 0.0095, respectively) and OS (p = 0.0061, p < 0.0001, p = 0.0074, respectively). BCLC classification maintained a relevant and independent prognostic role at multivariate analysis.
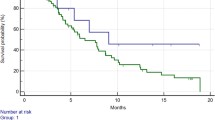
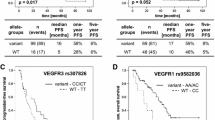
a Time to progression (TTP) according to HIF rs12434438 polymorphisms: AA/GA genotype (continuous line) 5.8 months vs. GG genotype (dotted line) 2.6 months; p < 0.0001. b Overall survival (OS) according to HIF rs12434438 polymorphism: AA/GA genotype (continuous line) 13.5 months vs. GG genotype (dotted line) 6.6 months; p < 0.0001. c TTP according to VEGF-A rs2010963 polymorphism: CC/CG genotype (continuous line) 7.6 months vs. GG genotype (dotted line) 3.9 months; p < 0.0001. d OS according to VEGF-A rs2010963 polymorphism: CC/CG genotype (continuous line) 15.1 months vs. GG genotype (dotted line) 8.4 months; p < 0.0001. e TTP according to VEGF-C rs4604006 polymorphism: TT/TC genotype (continuous line) 9.7 months vs.CC genotype (dotted line) 4.6 months; p < 0.0001. f OS according to VEGF-C rs4604006 polymorphism: TT/TC genotype (continuous line) 16.6 months vs. CC genotype (dotted line) 10.5 months; p < 0.0001
Furthermore, an exploratory combined analysis of polymorphisms statistically significant at multivariate analysis was conducted. Patients with expression of both favourable genotypes (FG) rs2010963 (VEGF-A) and rs4604006 (VEGF-C) showed a better TTP and OS (Fig. 2a, b), compared to patients expressing one or none of the favourable genotypes. At the combined analysis of HIF-1α and VEGF polymorphisms, patients with GG genotype of rs12434438 (HIF-1α) demonstrated a significantly worse TTP and OS then patients with genotype AA or AG (Fig. 3a, b). This observation was confirmed regardless of the number of favourable VEGF genotypes.
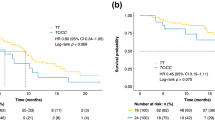
a Time to progression (TTP) according to rs2010963 VEGF-A and rs4604006 VEGF-C polymorphisms: two favourable alleles 10.8 months (dashed line) vs. one favourable allele 5.6 months (dash-dotted line) vs. 0 favourable alleles 3.7 months (dotted line); p < 0.0001. b Overall survival (OS) according to rs2010963 VEGF-A and rs4604006 VEGF-C polymorphisms: two favourable alleles 19.0 months (dashed line); one favourable allele 13.5 months (dash-dotted line); 0 favourable alleles 7.5 months (dotted line); p < 0.0001
a Time to progression (TTP) according to HIF rs12434438 GG vs. AA/AG combined with VEGF-A rs2010963 and VEGF-C rs4604006 polymorphisms. HIF rs12434438 GG 2.6 months. (continuous line) vs. rs12434438 AA/AG + : + two favourable VEGF-A/VEGF-C genotypes 10.8 months (dashed line); + one favourable VEGF-A/VEGF-C genotype 7.0 months (dash-dotted line); + 0 favourable VEGF-A/VEGF-C genotype 4.0 months (dotted line); p < 0.0001. b Overall survival (OS) according to HIF rs12434438 GG vs. AA/AG combined with VEGF-A rs2010963 and VEGF-C rs4604006 polymorphisms. HIF rs12434438 GG 6.6 months (continuous line) vs. rs12434438 AA/AG + : + two favourable VEGF-A/VEGF-C genotypes 19.0 months (dashed line); + one favourable VEGF-A/VEGF-C genotype 14.6 months (dash-dotted line); + 0 favourable VEGF-A/VEGF-C genotype 8.6 months (dotted line); p < 0.0001
3.6 Objective Response Analysis
A statistically significant correlation was found between the best objective response, evaluated by mRECIST criteria, and rs2010963 (VEGF-A) and rs4604006 (VEGF-C) genotypes. For rs2010963 (VEGF-A) CC + CG and GG genotypes CR + PR, SD and PD occur, respectively, in 11 (9%) versus six (7%), 71 (57%) versus 35 (41%) and 42 (34%) versus 45 (52%) patients (p = 0.027). The DCR was 66% versus 48%. For rs4604006 (VEGF-C) TT + CT and CC genotypes CR + PR, SD and PD occur, respectively, in nine (11%) versus eight (7%), 44 (59%) versus 62 (46%) and 23 (30%) versus 64 (47%) patients (p = 0.031). The DCR was 70% versus 53%. No statistically significant differences were found for rs12434438 (HIF-1α).
4 Discussion
Sorafenib represents the standard first-line systemic treatment for HCC patients. However, this molecule has a non-negligible toxicity profile, which may heavily impact on the quality of life, along with a limited survival benefit. Moreover, new systemic treatments became a part of the therapeutic scenario for HCC patients recently. It would be crucial to be able to distinguish non-responding patients to sorafenib in order to treat them with alternative systemic therapies or with best supportive care. Therefore, the identification of prognostic and predictive biomarkers of sorafenib represents a challenging area of research, and the results of the ALICE -2 study fit into this context. In particular, selected HIF-1α, VEGF, and VEGFR SNPs correlated with clinical outcomes of HCC patients receiving first-line sorafenib. rs12434438 (HIF-1α) allele A, rs2010963 (VEGF-A) allele C and rs4604006 (VEGF-C) allele T were significantly correlated with TTP and OS. Data about objective responses confirmed our previous findings from the ALICE-1 study. No differences were found for objective response and rs12434438 (HIF-1α), probably due to the genotype frequency found in the study population.
rs2010963 is located in the 5′-UTR region of the VEGF-A gene; whereas rs12434438 and rs4604006 are located in one of the intronic regions of the HIF-1α and VEGF-C genes, respectively. However, clear information on the biological effects of these genetic variants are largely lacking for most of the SNPs included in our analysis. Based on the available data, we can hypothesize that SNPs of the HIF-1α gene might imply a different HIF-1α sensitivity to hypoxic stimuli, leading to a heterogeneous transcription of target genes, including VEGF. Recent studies showed that HIF-1α SNPs might lead to variations in the oxygen-dependent degradation domain of HIF-1α, consequently influencing protein expression [29]. Likewise, we have evidence that different SNPs in the HIF-1α gene and VEGF might modify circulating VEGF levels, thus possibly affecting the response to antiangiogenetic treatment strategies [30, 31]. Nonetheless, results in this area of investigation are conflicting. On the one hand, some authors demonstrated a significant association for candidate polymorphisms in the promoter, 5′- and 3′-UTRs of the VEGF gene, but on the other hand, other studies were unable to identify this correlation for these and other VEGF SNPs [32,33,34,35,36]. Further studies are needed to validate these preliminary data and to fully unveil the molecular mechanisms underlying the biological implication of specific SNPs.
Although the impact on TTP might suggest a predictive effect of analysed SNPs our analysis seems to show mainly a prognostic role. Globally, our data show that polymorphism analysis of HIF-1α, VEGF and VEGFR genes might represent a valuable asset in order to better identify HCC patients who are more likely to benefit from sorafenib treatment. A relevant and independent prognostic role was maintained by the BCLC classification, thus confirming the reproducibility of analysis.
In conclusion, our study appears to be consistent with our previous findings (ALICE 1) and those from the SHARP trial, suggesting that variations in the angiogenesis pathway could have a relevant role in identifying different risk categories among HCC patients receiving sorafenib. Moreover, our data also suggest a role for HIF-1α. Globally, the ALICE 1 and the ALICE 2 studies could be then considered two complementary hypothesis-generating studies in this setting. The main limitations of our study are represented by the absence of a control arm and the retrospective nature of the research. The absence of a control arm does not allow us to distinguish between the prognostic and predictive role of the evaluated biomarkers, whereas the retrospective nature of the analysis prevents us to achieve definitive conclusions.
Further prospective trials would hopefully clarify the role of these potential biomarkers in the near future, thus allowing clinicians to better select the right treatment strategy for the right patient. The prospective, multicentre INNOVATE trial [37], currently being conducted by our research group, aims at determining whether eNOS, HIF-1, VEGF, Ang2 and VEGFR polymorphisms may play a role in predicting the objective response rate, PFS and OS of advanced HCC patients treated with sorafenib. The primary aim of the study is to validate the prognostic or predictive role of eNOS, HIF-1, VEGF, VEGFR and Ang2 polymorphisms in relation to PFS of HCC sorafenib-treated patients; the secondary aim is to verify the prognostic value of the same polymorphisms in relation to OS and the basal level of lactate dehydrogenase, blood pressure, MELD, Ang2 plasma level, VEGF and Ang2 in relation to PFS and OS in the same setting of patients. So, the results of our study will possibly give new insight into this area.
Change history
10 November 2020
The listing of the author names and affiliations, which previously read.
References
Faloppi L, Scartozzi M, Maccaroni E, Paolo MDP, Berardi R, Del Prete M, et al. Evolving strategies for the treatment of hepatocellular carcinoma: from clinical-guided to molecularly-tailored therapeutic options. Cancer Treat Rev. 2011;37(3):169–77.
Liu L, Cao Y, Chen C, Zhang X, McNabola A, Wilkie D, et al. Sorafenib blocks the RAF/MEK/ERK pathway, inhibits tumor angiogenesis, and induces tumor cell apoptosis in hepatocellular carcinoma model PLC/PRF/5. Cancer Res. 2006;66:11851–8.
Llovet JM, Ricci S, Mazzaferro V, Hilgard P, Gane E, Blanc JF, et al. Sorafenib in advanced hepatocellular carcinoma. N Engl J Med. 2008;359:378–90.
Cheng AL, Kang YK, Chen Z, Tsao CJ, Qin S, Kim JS, et al. Efficacy and safety of sorafenib in patients in the Asia-Pacific region with advanced hepatocellular carcinoma: a phase III randomized, double-blind, placebo-controlled trial. Lancet Oncol. 2009;10:25–34.
Galmiche A, Chauffert B, Barbare JC. New biological perspectives for the improvement of the efficacy of sorafenib in hepatocellular carcinoma. Cancer Lett. 2014;346:159–62.
Holderfield MM, Nagel TE, Stuart DD. Mechanism and consequences of RAF kinase activation by small-molecule inhibitors. Br J Cancer. 2014;111(4):640–5.
Kudo M, Finn RS, Qin S, Han KH, Ikeda K, Piscaglia F, et al. A Randomised Phase 3 trial of lenvatinib vs. sorafenib in first-line treatment of patients with unresectable hepatocellular carcinoma. Lancet. 2018;391:1163–73.
Bruix J, Qin S, Merle P, Granito A, Huang YH, Bodoky G, et al. Regorafenib for patients with hepatocellular carcinoma who progressed on sorafenib treatment (RESORCE): a randomised, double-blind, placebo-controlled, phase 3 trial. Lancet. 2017;389:56–66.
Abou-Alfa GK, Meyer T, Cheng AL, El-Khoueiry AB, Rimassa L, Ryoo BY, et al. Cabozantinib in patients with advanced and progressing hepatocellular carcinoma. N Engl J Med. 2018;379:54–63.
Zhu AX, Kang YK, Yen CJ, Finn RS, Galle PR, Llovet JM, et al. REACH-2: a randomized, double-blind, placebo-controlled phase 3 study of ramucirumab versus placebo as second-line treatment in patients with advanced hepatocellular carcinoma (HCC) and elevated baseline alpha-fetoprotein (AFP) following first-line sorafenib. J Clin Oncol. 2018;36:4003.
Bristol-Myers Squibb Announces Results from CheckMate -459 study evaluating opdivo (nivolumab) as a First-Line Treatment for Patients with Unresectable Hepatocellular Carcinoma. Bristol-Myers Squibb. 2019. https://bit.ly/2x70MSX. Accessed 24 Jun 2019.
Sangro B, Park JW, Dela Cruz CM, Anderson J, Lang L, Neelyet J, et al. A randomized, multicenter, Phase III study of nivolumab vs. sorafenib as first-line treatment in patients (pts) with advanced hepatocellular carcinoma (HCC): CheckMate-459. J Clin Oncol. 2016;34(Suppl. 15):TPS4147.
Finn RS, Ryoo BY, Merle P, Kudo M, Bouattour M, Lim HJ, et al. Results of KEYNOTE-240: phase 3 study of pembrolizumab (Pembro) vs. best supportive care (BSC) for second line therapy in advanced hepatocellular carcinoma (HCC). J Clin Oncol. 2019;37(Suppl. 15):4004.
Casadei Gardini A, Foca F, Scartozzi M, Silvestris N, Tamburini E, Faloppi L, et al. Metronomic capecitabine versus best supportive care as second-line treatment in hepatocellular carcinoma: a retrospective study. Sci Rep. 2017;7:42499.
Sangro B, Gomez-Martin C, de la Mata M, Iñarrairaegui M, Garralda E, Barrera P, et al. A clinical trial of CTLA-4 blockade with tremelimumab in patients with hepatocellular carcinoma and chronic hepatitis C. J Hepatol. 2013;59:81–8.
Duffy AG, Ulahannan SV, Makorova-Rusher O, Rahma O, Wedemeyer H, Pratt D, et al. Tremelimumab in combination with ablation in patients with advanced hepatocellular carcinoma. J Hepatol. 2017;66:545–51.
El-Khoueiry AB, Sangro B, Yau T, Crocenzi TS, Kudo M, Hsu C, et al. Nivolumab in patients with advanced hepatocellular carcinoma (CheckMate 040): an open-label, non-comparative, phase 1/2 dose escalation and expansion trial. Lancet. 2017;389(10088):2492–502.
Zhu AX, Finn RS, Edeline J, Cattan S, Ogasawara S, Palmer D, et al. Pembrolizumab in patients with advanced hepatocellular carcinoma previously treated with sorafenib (KEYNOTE-224): a non-randomised, open-label phase 2 trial. Lancet Oncol. 2018;19(7):940–52.
Marisi G, Cucchetti A, Ulivi P, Canale M, Cabibbo G, Solaini L, et al. Ten years of sorafenib in hepatocellular carcinoma: are there any predictive and/or prognostic markers? World J Gastroenterol. 2018;24(36):4152–63.
Ferrara N, Gerber HP, LeCouter J. The biology of VEGF and its receptors. Nat Med. 2003;9:669–76.
Peng S, Wang Y, Peng H, Chen D, Shen S, Peng B, et al. Autocrine vascular endothelial growth factor signaling promotes cell proliferation and modulates sorafenib treatment efficacy in hepatocellular carcinoma. Hepatology. 2014;60(4):1264–77.
Semenza GL, Wang GL. A nuclear factor induced by hypoxia via de novo protein synthesis binds to the human erythropoietin gene enhancer at a site required for transcriptional activation. Mol Cell Biol. 1992;12:5447–54.
Jiang B, Rue E, Wang GL, Roe R. Semenza GL Dimerization, DNA binding, and transactivation properties of hypoxia-inducible factor-1. J Biol Chem. 1997;30:17771–8.
Wang W, Xu GL, Jia WD, Wang ZH, Li JS, et al. Expression and correlation of hypoxia-inducible factor-1a, vascular endothelial growth factor and microvessel density in experimental rat hepatocarcinogenesis. J Int Med Res. 2009;2:417–25.
Scartozzi M, Faloppi L, Svegliati Baroni S, Loretelli C, Piscaglia F, Iavarone M, et al. VEGF and VEGFR genotyping in the prediction of clinical outcome for HCC patients receiving sorafenib: the ALICE-1 study. Int J Cancer. 2014;135:1247–56.
Formento JL, Etienne-Grimaldi MC, Francoual M, Pagès G, Onesto C, Formento P, et al. Influence of the VEGF-A 936C > T germinal polymorphism on tumoral VEGF expression in head and neck cancer. Pharmacogenomics. 2009;10:1277–83.
Chen MH, Tzeng CH, Chen PM, Lin JK, Lin TC, Chen WS, et al. VEGF-406T → polymorphism and its association with VEGF expression and outcome to FOLFOX-4 treatment in patients with colorectal carcinoma. Pharmacogenomics J. 2011;11(3):227–36.
North Carolina State University. 2019. www.statgen.ncsu.edu/powermarker.
Pages G, Puyssegur J. Transcriptional regulation of the vascular endothelial growth factor gene—a concert of activating factors. Cardiovasc Res. 2005;65:564–73.
Ruggiero D, Dalmasso C, Nutile T, Sorice R, Dionisi L, Aversano M, et al. Genetics of VEGF serum variation in human isolated populations of cilento: importance of VEGF polymorphisms. PLoS One. 2011;6(2):e16982.
Al-Habboubi HH, Mahdi N, Abu-Hijleh TM, Abu-Hijleh FM, Sater MS, Almawi WY. The relation of vascular endothelial growth factor (VEGF) gene polymorphisms on VEGF levels and the risk of vasoocclusive crisis in sickle cell disease. Eur J Haematol. 2012;89(5):403–9.
Zhai R, Gong MN, Zhou W, Thompson TB, Kraft P, Su L, et al. Genotypes and haplotypes of the VEGF gene are associated with higher mortality and lower VEGF plasma levels in patients with ARDS. Thorax. 2007;62:718–22.
Ferrante M, Pierik M, Henckaerts L, Joossens M, Claes K, Van Schuerbeek N, et al. The role of vascular endothelial growth factor (VEGF) in inflammatory bowel disease. Inflamm Bowel Dis. 2006;12:870–8.
Krippl P, Langsenlehner U, Renner W, Yazdani-Biuki B, Wolf G, Wascher TC, et al. A common 936 C/T gene polymorphism of vascular endothelial growth factor is associated with decreased breast cancer risk. Int J Cancer. 2003;106:468–71.
Awata T, Inoue K, Kurihara S, Ohkubo T, Watanabe M, Inukai K, et al. A common polymorphism in the 5′-untranslated region of the VEGF gene is associated with diabetic retinopathy in type 2 diabetes. Diabetes. 2002;51:1635–9.
Petrovic MG, Korosec P, Kosnik M, Osredkar J, Hawlina M, Peterlin B, et al. Local and genetic determinants of vascular endothelial growth factor expression in advanced proliferative diabetic retinopathy. Mol Vis. 2008;14:1382–7.
Gardini AC, Faloppi L, Aprile G, Brunetti O, Caparello C, Corbelli J, et al. Multicenter prospective study of angiogenesis polymorphism validation in HCC patients treated with sorafenib. An INNOVATE study protocol. Tumori. 2018;104(6):476–9.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Funding
No external funding was used in the preparation of this article.
Conflict of interest
Luca Faloppi, Marco Puzzoni, Andrea Casadei Gardini, Nicola Silvestris, Gianluca Masi, Giorgia Marisi, Caterina Vivaldi, Cosmo Damiano Gadaleta, Pina Ziranu, Maristella Bianconi, Cristian Loretelli, Laura Demurtas, Eleonora Lai, Riccardo Giampieri, Eva Galizia, Paola Ulivi, Nicola Battelli, Alfredo Falcone, Stefano Cascinu and Mario Scartozzi declare that they have no conflicts of interest that might be relevant to the contents of this article.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Faloppi, L., Puzzoni, M., Casadei Gardini, A. et al. Angiogenesis Genotyping and Clinical Outcomes in Patients with Advanced Hepatocellular Carcinoma Receiving Sorafenib: The ALICE-2 Study. Targ Oncol 15, 115–126 (2020). https://doi.org/10.1007/s11523-020-00698-x
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11523-020-00698-x