Abstract
In this work, we present the release of a novel easy to use software package called DGM or Directed-Graph-Mapping. DGM can automatically analyze any type of arrhythmia to find reentry or focal sources if the measurements are synchronized in time. Currently, DGM requires the local activation times (LAT) and the spatial coordinates of the measured electrodes. However, there is no requirement for any spatial organization of the electrodes, allowing to analyze clinical, experimental or computational data. DGM creates directed networks of the activation, which are analyzed with fast algorithms to search for reentry (cycles in the network) and focal sources (nodes with outgoing arrows). DGM has been mainly optimized to analyze atrial tachycardia, but we also discuss other applications of DGM demonstrating its wide applicability. The goal is to release a free software package which can allow researchers to save time in the analysis of cardiac data. An academic license is attached to the software, allowing only non-commercial use of the software. All updates of the software, user and installation guide will be published on a dedicated website www.dgmapping.com.

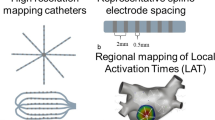
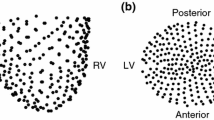
Direct-Graph-Mapping is a method to automatically analyze a given arrhythmia by converting measured data of the electrodes in a directed network. DGM requires the local activation times (LAT) and the spatial coordinates of the measured electrodes. There is no requirement for any spatial organization of the electrodes, allowing to analyze clinical, experimental or computational data (see left). An example could be the LATs and coordinates from a CARTO file. DGM creates a directed network of the activation by (1) determining the neighbors of each node, 2 (2) allowing a directed arrow between two neighbors if propagation of the electrical wave is possible, (3) repeating this process for all nodes, (4) if necessary, redistributing the nodes more uniformly and repeating step (1)-(3). Two possible steps are (5) to visualize the wavefront by creating an average graph or (6) find the cycles in the network which represent the reentry loops. Focal sources are nodes with only outgoing arrows.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
1.1 Different types of arrhythmias and sources
One possible treatment for curing an arrhythmia is via catheter ablation, which aims to restore the normal rhythm by destroying a part of the tissue. For this, it is important to determine the source and the location of the arrhythmia. It is important to destroy as little tissue as possible, since the larger the ablated area, the higher the risk of severe complications and reconnected tissue [1].
In general, there are two possible sources for an arrhythmia: reentry and triggered activity. The first driving source reentry is usually subdivided in two categories: anatomical reentry and functional reentry. Anatomical reentry occurs when the electrical wave starts to rotate around an obstacle, e.g., around a valve, a vein or scar tissue. Depending on the size of the smallest rotation, one can make a distinction between macro reentry (large obstacle, like a valve or a vein) or localized reentry (small obstacle, usually scar tissue, also called micro reentry). However, reentry is also possible without an obstacle, whereby the wave rotates around an excitable core, namely functional reentry. Functional reentry is also often called a spiral wave or a rotor. The second driving source is triggered activity or automaticity. The latter mechanism gives rise to focal activation, most likely accompanied by a concentric activation pattern.
We can divide most arrhythmia in two different types: the regular arrhythmia, called tachycardia, and the irregular arrhythmia, called fibrillation (except for Torsade de Pointes) as they make the atrial or ventricular walls fibrillate and therefore not contract regularly. When including the location of the arrhythmia, we distinguish 4 main types of arrhythmia: atrial tachycardia (AT) (e.g., atrial flutter), atrial fibrillation (AF), ventricular tachycardia (VT) and ventricular fibrillation (VF). A tachycardia can have two mechanisms, a regular reentry or a regular single focal source [2, 3]. For AT and VT, the reentry is usually reported to be anatomical, although it might be difficult to differentiate a small localized reentry from a functional reentry [4]. This is in contrast with fibrillation, which can be any combination of multiple irregular sources: reentries and focal sources which appear irregularly in the heart. For paroxysmal AF, Haissaguerre et al. [5] suggested it is triggered by irregular ectopic activity originating in the pulmonary veins region and demonstrated that pulmonary vein isolation is an effective way to manage paroxysmal AF. For persistent AF, the mechanism is still unclear, leading to a success rate in the range of only \(\sim \)50% [6,7,8]. Many different mechanisms have been proposed, which were recently summarized by Roney et al. [9]. All of these theories have been reported in experimental/clinical work, but were also reproduced in computer simulations. Among them are the multiple wavelet theory [10,11,12,13], the mother rotor fibrillation [14, 15], the multiple rotor theory where the rotors are stable [16,17,18,19] or meandering [19,20,21,22], the double layer hypothesis [23,24,25,26] and micro-anatomic intramural reentry [27, 28]. In addition for VF, the mechanism remains unclear, even though in the US alone VF (and VT) is responsible for about 700,000 death each year. Similarly as in AF, there exist multiple hypothesis. According to Krummen et al., VF can be divided in the following four stages: initiation, transition, maintenance and evolution [29]. VF is usually initiated by premature ventricular complexes [30] or nonsustained VT [29]. In the next three stages VF can be maintained by wandering wavelets of disorganized activity [31, 32], by mother rotor fibrillation [14], by rotors [31, 32], focal activity [31, 32]. Notice many similarities with AF, although the ventricles have the additional complexity of extra wall thickness.
1.2 Measuring an arrhythmia
In order to better understand the mechanism of an arrhythmia or to perform more precise ablation procedures, it is necessary to analyze the electrical activity in detail. In the clinic, this is done by inserting a catheter in the heart. If the arrhythmia is regular and tolerable by the patient, this can be done consecutively and many data points can be gathered. This is in contrast with an irregular arrhythmia, where it is necessary to simultaneously record the electrical activity, as the arrhythmia is continuously changing. As a consequence, irregular arrhythmia are often measured only partly via contact mapping, with a grid mapping catheter, or a circular catheter (like a lasso catheter). A way to have a more complete picture is via inserting a non-contact basket catheter inside the atrium or the ventricle. The downside is that usually not many electrodes are inserted providing a sparse view. In experimental studies in animals, much more is possible like inserting needle electrodes or performing optical measurements during Langendorf perfusion. Finally, computer simulations allow full control over the data, including millions of datapoints. After obtaining a dataset, the following methods exist to automatically detect functional, anatomical and focal activity.
1.3 Methods to analyze the data
First, in order to detect functional reentry or rotors, multiple methods were developed. The most popular method is phase mapping (PM), see Umapathy et al. for a review [33]. A first restriction is that PM requires a regular set of electrodes (like a 64-basket catheter, or a regular grid). PM assigns a phase to any signal per time frame, and looks for phase singularities, which is usually a set of 4 electrodes for which the phase differences add up to about 2π. Many different implementations were developed to assign the correct phase to the signal, using phase space [34], the Hilbert transform [35, 36], a sinusodal recomposition [37], or the filter method of Roney et al. [38]. Although PM is currently being used for clinical ablations of AF, AT and VT (FIRM procedure of Topera inc. [17]), many concerns have risen about the application of PM to guide ablations, mainly during AF. In Kuklik et al. [39], it was shown that at least eight electrodes are required to avoid frequent false positive PS detection due to chance. Also, it was shown that the number of detected rotors critically depends on the inter-electrode distance and parameters of the detection algorithm [39,40,41,42]. In addition, noise has a big effect on the detection of phase singularities [43, 44], and far field and interpolation effects can induce permanent false rotors [45]. Many other methods have been developed to find rotors. We mention the following: the CARTOFINDER (Biosense Webster) [46], a method using electrocardiographic imaging (ECGI) [12], a method based on displaying electrogram dispersion [47, 48], a method based on the optical flow of the wavefront dynamics [49], sequential ultra high density contact activation mapping [50], a machine learning method based on the 12 lead ECGs [51], the ASAP method for a multipole diagnostic catheter [52], the ICAN method for the circular lasso catheter [53], machine learning methods for mapping electrodes [54, 55], single-signal algorithms based on instantaneous amplitude modulation (iAM) and instantaneous frequency modulation (iFM) [56], method to identify repetitive-regular activities (RRas) [57], the RADAR (Real-Time Electrogram Analysis for Drivers of Atrial Fibrillation) system [58], Noncontact Charge Density Mapping [59], novel integrated mapping technique searching for regions with repetitive-regular (RR) activations [60], Stochastic trajectory analysis of ranked signals (STAR) mapping [61], the wavefront field method [62], the electrographic flow mapping [63], deep neural networks for rotor localization [64]. It is clear that an abundance of methods have been developed in the past years to find rotors. All of these methods have potential, but require further development or clinical testing and are often constructed for a particular type of catheter.
Second, although anatomical reentry arises often in AT and VT, there are almost no methods to analyze anatomical reentry automatically. Currently, if anatomical reentry is present during, e.g., AT, the electrophysiologist (EP) has to manually analyze (often guided) color maps from CARTO (Biosense Webster), RHYTHMIA (Boston Scientific) and EnSite NavX (Abbott). Only one automatic method was previously developed by Oesterlein et al. [65], but it only identifies the critical areas and it does not show the reentry path.
Thirdly, focal activity can be easily detected as the region of earliest activation in between two consecutive beats. However, for more complex cases where reentry and focal are present simultaneously, this method becomes more difficult. Again, FIRM can also detect focal sources [17], the method of Salmin et al. [66], the Focal Source and Trigger (FaST) computational algorithm [67] and the method of Oesterlein et al. [65].
1.4 DGM—a novel software package
Following the previous section, there is a clear need for new methods which can analyze any type of reentry mechanisms (functional and anatomical) in cardiac arrhythmia independent of type of measuring system. This is necessary for a better understanding of the mechanism of the arrhythmia as well as to guide the electrophysiologist (EP) during an ablation procedure. Also, it is important to make these methods public, allowing for a greater use and limiting the time researchers need to spend to each time re-implement existing methods [68]. Therefore, in this paper, we present an easy to use software package called Directed-Graph mapping or DGM, which can automatically analyze any type of arrhythmia to find reentry or focal sources. It is based on converting the excitation pattern of the electrical activity in a directed network and analyzing this directed network [44]. A platform independent (Windows, Mac, Linux) executable can be obtained after registration on www.dgmapping.com. DGM is available for investigators affiliated with a research institution or a hospital. It is important to note that the use of DGM is restricted under an academic license. DGM can be used for the analysis of clinical data, experimental data and in silico data. In time, we will also allow researchers to obtain the source code for restricted non-commercial use only. In the next sections we will discuss the different parts of the software.
2 General concept of DGM
In this section, we will provide more details on the basic concept of DGM. As explained in the previous section, DGM is based on transforming a given dataset of an arrhythmia into a directed network. It only requires a file including the spatial coordinates and the LATs of the electrodes. The LATs can be extracted from the measured signals. There is no need for any regularity of the spatial locations of the dataset, making it independent of the measuring system. Therefore, it can take as input any extracted data from a computational dataset, experimental dataset like needle data, socket data, ..., or from a clinical dataset, grid electrode data, basket catheter data, etc. The spatial locations of the electrodes will be the nodes of the network. In the next paragraph we will explain how we will create the directed edges of the network.
2.1 Creation of the network
In Fig. 1, the different steps DGM are shown [44]. First, for each electrical sample, the neighbors are determined using the spatial coordinates of the data points (1). A node is considered a neighbor of another node, if they are within a certain spherical distance. We currently do not take into account the geometry of the atria/ventricles to specify the neighbors. However, in a future release, we will enable a mode to read your own neighbors file, which can be created with your own algorithm. Second, for a given time t, each node’s first LAT value larger than t is selected. In case of a regular arrhythmia, it can be necessary to add the cycle length to the given LAT, as only one LAT is given per node. Therefore, one should provide the correct cycle length in the GUI. Then, for each node only a directed arrow is allowed to a neighbor if the spatial distance divided by the time difference in LATs is in between a certain allowed minimal and maximal conduction velocity, see Fig. 1(2). For standard parameter settings, the minimal conduction velocity is set to 0.2 mm/ms and the maximal conduction velocity is set to 2.0 mm/ms. Repeating this process for all the nodes, this creates a single directed network at a certain time t. In addition, a second network is created at a time t + δt, whereby δt is a parameter of the program, also referred to as the jump. This jump is set to 40 ms in standard settings. This second network is merged with the first network as follows. If an additional arrow is created which has the same originating LAT, it is added to the first network. This results the full directed network of excitation (3). Notice that this step is mandatory to create closed cycles in the network. As an intermediate step, the properties of this directed network are used to create more uniformly distributed nodes in case of regular datasets (4). Due to time constraints this step is currently not implemented for irregular datasets. If step 4 takes place, step (1)–(3) is repeated. Once the final network is created, DGM searches for directed cycles. In the default visualization mode, a band of loops is created for clear visualization of the reentrant pattern responsible for maintaining the arrhythmia (6). DGM also computes a wavefront representing the excitation pattern (5). In case of focal activity, the location of the focal source can be visualized by clicking on the corresponding button “focal”.
2.2 Detection of reentrant activity
To uncover the reentry loops in the arrhythmia, we search for closed cycles in the directed graph with a Breath First Search algorithm [44]. We only show the loops which have the best variance. This term was previously introduced in Van Nieuwenhuyse et al. [69] as phase variance. However, we choose to rename the feature as it has nothing to do with a phase and might lead to an incorrect interpretation. The concept can be explained as follows. We order the values of the LATs for each consecutive node of the cycle on the Y-axis of a graph and draw a straight line between the first and last node. The variance is the sum of the quadratic errors between the line and the actual LATs. We visualize the loops with the lowest variance. In case multiple cycles are detected, we calculate the geometrical mean of the cycle and in case the geometrical means of 2 cycles differ more than the value of the parameter Core separator, these reentries will be visualized separately. To enhance the intuitive visualization and corresponding interpretation, we have added a loopband feature, which emphasizes and colors the reentry loops (6).
2.3 Detection of focal activity or automaticity
In addition to reentry loops, DGM can also find focal sources. For this, LATs are put in different bins to receive a single LAT value. This reduces unwanted variation on the LATs. Based on these new LAT values, the directed network in (1)–(3) is reconstructed. We check if regions with only outgoing arrows are present, for which the middle of these regions corresponds to the source of the focal activity [44]. The arrows pointing outwards from these regions, correspond with the propagation of the focal sources.
2.4 Regular versus irregular arrhythmia
In case DGM analyzes a regular arrhythmia, only a single network requires analysis, which agrees with a single time frame. However, in case you analyze an irregular arrhythmia, DGM will sequentially repeat the previous described steps for the complete time interval in steps provided by the user. This will allow the user to analyze the arrhythmia sequentially. For regular arrhythmia, we have mainly analyzed AT, see Vandersickel et al. [44] and Van Nieuwenhuyse et al. [69] (see Section 5.1). For irregular arrhythmia, we have analyzed needle data of the CAVB dog model for the study of Torsade de Pointes [70] (see Section 5.3). Also, we have analyzed irregular in silico data of a meandering spiral in the right atrium [71] (see Section 5.2).
3 DGM software
3.1 Input files
In this section, we will discuss which input files DGM can handle (Fig. 2). In addition to a general input file format, DGM can also process the industry standards of two main mapping systems: CARTO (Biosense Webster) and RHYTHMIA (Boston Scientific). DGM will automatically pre-process these formats into the general input file format. For more information regarding the industry standard, we refer to Section 3.1.2. DGM can also handle files generated by openCARP [72], an open source software package for simulating a broad variety of different arrhythmia in 2D and 3D. Once the software is opened and the input file is loaded, a copy or modification of that file will be generated in the input folder. This way, the original file will solely be used for reading remains unaltered.
General flow of the software: Once a file is selected, DGM checks if you wish to render a company file such as CARTO or RHTYHMIA. In that case, the files are automatically pre-processed to a standard DGM format. This general input is used for the graphical user interface to visualize the electrodes and the mesh. Once you press calculate, the C back-end creates .csv and .txt files. These files are used as an input for the visualization in the GUI (basic or advanced view, depending on the user’s needs). In addition, the user can choose to change the files and visualize them in a software of choice, such as Paraview
3.1.1 General input file
DGM works with a standard input file format which contains (1) the spatial coordinates of the electrode recordings and (2) the annotated LATs of these electrode recordings. DGM can handle both regular arrhythmia (one LAT per location with a fixed cycle length) as irregular arrhythmia (multiple LATs per spatial location). As mentioned before, as long as the input file is in the right format, DGM is independent of the type of measuring system. In the supplementary material 8.1.1 the form of the obligatory LAT file is presented.
Optionally, the spatial coordinates of a mesh can be included in the folder. These coordinates will be used to create a triangulated mesh in the user interface. In the supplementary material 8.1.2, the format of mesh file is presented
3.1.2 Existing mapping systems
CARTO
DGM can handle the output format of the CARTO system of Biosense Webster. The latest system which was tested is CARTO 3 v7. However, older formats might have another header, in which case we will have to add another module to read this format. The mapping can be performed with single electrode ablation catheters or multi-electrode catheters like PentaRay. We have tested CARTO formats solely for the analysis of regular arrhythmias such as AT and VT. However, the CARTO system can also be used to measure the heart during sinus rhythm (SR), which can be considered a regular arrhythmia as well. In that case, DGM will currently not give any useful information for SR, although we plan to further develop DGM to extract additional structural information.
A full export of the study data of a CARTO case will generate at least two important files: a _car.txt file, which contains the LATs and the coordinates of the measured points, and a .mesh file which contains all the information about the mesh of the atrium or ventricle. Both files should be in a single folder. However, if the user decides not to include the mesh, DGM will generate a mesh based on some basic algorithms, but the results will be visually less appealing. In case the V isiTagExport/Sites.txt file is added to the folder, DGM will also be able to show ablation points. Additionally, in a case where the correct tags are labeled, also PPI-TCL points can be loaded. For this, the study’s .xml file should be put in the same folder as the _car.txt and mesh file. In case a PPI-TCL point is recorded and saved during the procedure, one should use the tags presented in Section 3.1.4.
RHYTHMIA
DGM can also process files generated by RHYTHMIA, the mapping software of Boston Scientific. Mapping can be performed with single electrode ablation catheters or multi-electrode catheters like Orion. All necessary information is merged within a single .mat file which contains the LAT and coordinates. The RHYTHMIA system is known for gathering a large amount of electrode recordings. DGM will use this abundance of electrode recordings to generate a mesh file as well as select a set of electrodes for the analysis of the reentry sources or focal activity.
Ensite
DGM is not yet adapted to automatically parse the mapping system of Abbott for which mapping can be performed with single electrode ablation catheters or multi-electrode catheters like HD grid. In case there is a need, we will implement this mapping system in the future.
3.1.3 OpenCARP
DGM can also automatically analyze a simulation generated by openCARP. Currently, DGM can interpret the .pts and .vtk (meshes) files required for the generation of a simulation, and the .igb file, which contains the output of the simulation. Again, these 3 files should be located in the same folder, together with a costume written openCARP.csv file in a comma-separated format. For clearness, we presented an example of this latter file in the supplementary material 8.1.3.
3.1.4 PPI-TCL, ablation points and corresponding tags
In case the study’s xml is uploaded for a CARTO file and the predefined tags as presented in Table 1 were used during the procedure to label certain electrodes, one can visualize not only the analysis of DGM but also PPI-TCL locations with corresponding colors. Scar regions, double potentials and fragmented signals can be annotated during the procedure on the CARTO module and visualized in DGM in case the tags are used correctly.
In case one uses RHYTHMIA, the spatial location of your ablation points are needed and a calculation editor such as Excel will be necessary to create the template PPI file. For simulations, one can use the same template format. The basic format should contain the XYZ coordinates and the color label of the PPI-TCL point. Default options are: “green”, “yellow”, “red”, “blue”, “black”, “white”, “orange”, “purple”, “pink” and “grey”. One can also use DGM to visualize the arrhythmia with additional nodes providing useful information or to tag certain regions.
The following table provides possible tags which can be used in the file.
In the supplementary material 8.1.4, the format is specified to show the PPI-TCL points in DGM.
3.2 Output generated by DGM
Once the input files are generated or extracted from a mapping system, a simulation or an experimental set-up, one can process the data with DGM by opening the GUI. Once loaded into the software, by pressing calculate, DGM will automatically search for rotational and/or focal activity present in the data.
For further processing of the data generated by DGM, we will give an overview of the output which is generated by DGM. First, in the main folder, a period.json file is generated which stores the period of the arrhythmia. In case the arrhythmia is irregular, this is also stored in this file. All the other output files are generated in a separate folder in the main data folder called “output”. These files are generated by the core algorithm, which are used for the visualization of the results. Of course, one can also use the files in the output folder for a different visualization tool. Note that most files have either the time of calculation and/or number of nodes in the filename. For example result_loops_t0.csv contains all the accepted loops found at 0 ms. The following output files are generated:
-
result_loops_t{time}.csv
$$ \begin{array}{@{}rcl@{}} &&{}\text{x\_core,~y\_core,~z\_core,~amount,~variance,~intervariability},\\ &&\text{length,~index0,~LAT0,~index1,~LAT1,} ... \end{array} $$Each row of this files provides detailed information about the different reentry cycles which are visualized in the GUI. The first 3 columns correspond with the location of core of the reentry loop. Amount is a measure for the number of different cycles which are detected by DGM and belong to the same reentry loop. V ariance is a measure for the uniformity of the reentry, see also Section 2.2. Intervariability is the absolute value of the maximal deviation of the LATs from the straight line used to calculate the variance. Length is the accumulative length of the reentry by taking the sum of the euclidean distances between the nodes. Next follows a series of indices and their corresponding LATs which belong to the chosen cycle. Each electrode or node has a unique index.
-
result_focal_t{time}.csv
$$ \begin{array}{@{}rcl@{}} &&{}\text{x\_core,~y\_core,~z\_core,~amount,~variance,~intervariability},\\ &&\text{length,~index0,~LAT0,~index1,~LAT1} \end{array} $$Each row determines represents an arrow which contributes to the visualization of the focal beats. For easy implementation, we copied the same format as the loops. However, only index0 and index1 are used as each row represents a directed arrow from index0 to index1.
-
front_t{time}_p{nodes}.csv
$$ \text{color}, \quad\text{scalar}, \quad\mathrm{x}, \quad\mathrm{y}, \quad\mathrm{z}, \quad\text{x\_dir}, \quad\text{y\_dir}, \quad\text{z\_dir} $$Each row in this file represents a single arrow of our wavefront. The wavefront is a collection of arrows which gives a visual representation of the electrical flow. Color represents a time associated to the arrow which is a value larger than the time t the directed network was created. Scalar is the normalized size of the arrow and should always be one. (x, y, z) is the 3D spatial location of the starting point of the arrow whilst (x_dir, y_dir, z_dir) is the normalized direction of the wave arrow. Drawing all these arrows creates a wavefront generated by DGM.
-
loopband_t{time}_p{nodes}.csv
$$ \text{color}, \quad\text{scalar}, \mathrm{x}, \mathrm{y}, \mathrm{z}, \text{x\_dir}, \text{y\_dir}, \text{z\_dir} $$Each row in this file represents an arrow in the loopband in the same format as the previously described wavefront. The loopband creates an easy-to-interpret visualization of a region around the detected loops.
-
mesh_t0_p0.csv
$$ \begin{array}{lll} x,& y,& z\\ x0,& y0,& z0\\ x1,& y1,& z1 \end{array} $$This file represents the mesh which was used for the visualization, which can also be used in other visualization tools (e.g., Paraview). It contains the spatial coordinates of the mesh nodes. The top line is also included in the mesh file.
3.3 Visualization
DGM has two layouts, the basic view and the advanced view. Depending on your default setting, the application will open in either the basic or the advanced view.
The basic view allows little room for adaptions and is ideal for a first use of DGM. The advanced layout allows the user to change many parameters of the program. In the supplementary material, all the possibilities of both layouts are explained in detail.
4 Some basic examples
In this section we will discuss all the possible combinations of input that the program can handle. This can be useful for the user to experiment with DGM and to create its own input files. These files can be downloaded from our site and should be bug free.
The first example presented is a regular simulation with reentry in 3D, corresponding to frame A in Fig. 3. To create this figure, the user should load LAT_p2000.txt. The mesh presented in panel A1 will be generated. After pressing Calculate, the outcome in A.2, A.3 and A.4 will be shown either by clicking the wavefront, loops or loopband button. In Supplementary Movie 1, an animated loopband is presented. As a second example, we show a regular simulation with focal activity in 2D starting from the lower left. Panel B.1 shows the original selected electrodes, while panel B.2 and B.3 present the results after calculation. In Supplementary Movie 2, we show the animated wavefront for a single focal beat. The third example is a irregular simulation with reentry in 2D. After processing the data, in panel C.1 one observes the electrodes including scar tissue. In panel C.2, we shown the wavefront, panel C.3 shows the reentry cycles which show anatomical reentry around the right scar tissue, and in panel C.4 we show the loopband. Additionally, one can download the corresponding ppi.txt file online and load it as a custom file. This file shows the simulated PPI-TCL measurements on the location indicated in panel C.5. Supplementary Movie 3 summarized the results of this case. In panel D, we present an irregular simulation with focal activity in 3D. The same case is also illustrated in Supplementary Movie 4.
In panel E, we show a Regular reentry recorded by CARTO (Fig. 4). Moreover, this is a double loop AT: one macro reentry around the mitral valve and one localized reentry at the anterior wall around some scar tissue. In panel F, we show a regular focal recorded by CARTO. To be more specific, this is actually a breakthrough from a right sided AT. As discussed before, the initial electrode positions of CARTO (E.1 and F.1) are redistributed (E.2 and F.2). Based on this new nodes, DGM calculated a wavefront (E.3 and F.3) and in case of reentry, displays the loopband (E.4). In case you have a focal arrhythmia, you will have to switch to the focal button (F.4). Also illustrated in panel E.5 are the PPI-TCL recordings from the .xml file and the ablation sites found in the V isiTagExport/Sites.txt file. We also present two additional supplementary movies, Supplementary Movie 5 and Supplementary Movie 6, which show the loopband of the reentry case E and the wavefront of the focal breakthrough F. For privacy reasons, we have created our own CARTO files, which are no longer linked to a certain patient.
Finally, we also show one example of a irregular reentry in 3D simulated by openCARP. To render these pictures, one has to load the openCARP .igb file. DGM will create the standard LAT_p.txt file. In panel G.1 the nodes are shown, in panel G.2 the loopband is presented, in panel G.3 the wavefront is shown, and in panel G.4 the cycles are plotted (Fig. 5).
5 Summary of the data which is already processed by DGM
In this section, we give an overview of the type of datasets which were already processed with DGM. We divide this section according to the type of arrhythmia.
5.1 Atrial tachycardia—AT
DGM has been mainly optimized to work with data from AT. In a first study [44], we compared the ablation target of 31 cases of AT which would be chosen based on DGM with the actual ablation target. For these cases, DGM had a 100% success rate. In a second study, 51 complex ATs were retrospectively analyzed with DGM, whereby entrainment mapping determined the actual reentry loop in case of reentry [69]. Experts diagnosed the correct AT mechanism and location with the latest mapping software of CARTO in 33 cases versus DGM in 38 cases. Even though a lot of thought went into these cases, we explicitly note that this was a single centered study. Hence we except that the automated performance will first drop and a modification of the now fixed parameters might be required to further develop and improve DGM.
So far all published data was on cases of Biosense Webster, although DGM can also work with the RHYTHMIA (Boston Scientific) system. Upon loading a .mat file, this data is processed and gives a result based on the analysis of a single center as well. However, no entrainment study on RHYTHMIA data is yet available to us, to know the ground truth. In the future, we aim to analyze and optimize DGM based on EM guided ablations of AT with the aid of RHYTHMIA electrode recordings.
5.2 Atrial fibrillation—AF
So far, we only analyzed simulated basket catheter data of a meandering spiral wave in the right atrium, causing the left and right atrium to fibrillate. It has been reported that upon analysis with PM, wrong rotors emerged due to the interpolation artefacts and the far field, obstructing correct interpretation of the single meandering rotor [45]. By changing the minimal conduction velocity in DGM, we were able to differentiate the false rotors from the true rotor [71], overcoming some of the artefacts of PM.
5.3 Torsade de Pointes—TdP
DGM was originally developed to analyze needle data of chronic atrioventricular block (CAVB) dog model, known for its high susceptibility for of Torsade de Pointes (TdP). TdP is a dangerous arrhythmia, as it can degenerate into VF and cause sudden cardiac death also in children en young adults [73]. Although TdP is an important arrhythmia, still many aspects of the mechanism remain unclear. Many studies agree that a TdP starts with a focal beat which is caused by a group a cells experiencing an early after depolarizations (EAD) at the same time [74,75,76,77,78,79,80,81]. However, contrary to the established mechanism of the trigger of TdP, the perpetuation of TdP remains controversial. Focal mechanisms [75, 79,80,81,82,83,84], non-stationary reentries [77, 78, 85] or both mechanisms [74, 76, 82] were claimed to underlie to basis of the perpetuation of a TdP. With an early version of DGM, we analyzed 54 episodes of TdP in 5 susceptible dogs and found that TdP can be driven by focal activity as well as by reentry depending on the duration of the episode. Short lasting episodes were maintained by focal sources, while longer lasting and non-terminating episodes were maintained by reentry [70]. With the latest version of DGM, we are currently further characterizing the types and numbers of reentry during the longer lasting episodes.
5.4 Simulations to compare DGM with PM
DGM could be a useful tool in comparison with PM, as we have previously shown that DGM is more robust compared to PM in case noise is added to the LATs [44]. In this paper, we simulated a single rotor which we measured with 256 evenly distributed electrodes. We added Gaussian noise to the LATs, which were derived as the steepest derivatives on the measured signals. We then compared the performance of DGM with PM, and found that for 20 ms of Gaussian noise, DGM was still 80% effective, while PM was unable to detect the single rotor correctly (0%). Instead of a single diagnosis, PM either did not found the center rotor or found the center rotor in addition with multiple other sites showing rotational activation.
The reason why PM fails in this in silico setting is twofold. First, PM easily detect false positives. The basic concept of PM is to search for a 2π enclosure around a line integral created by neighboring electrodes. Usually only four electrodes are taken into account. A single alteration of a LAT can therefore result in a misplaced detection. For the same reason, as only four neighbors are taken into account to find the true phase singularity, a mistake in one of these four neighbors can result in a false negative. As DGM is a global measure, it overcomes this shortcoming of the local PM.
However, a more profound analysis of DGM versus PM is necessary as we used a single implementation of PM based on the LATs. More on this in the Discussion section of this manuscript.
5.5 Future studies
We are currently also processing data of hemodynamical stable VTs, which were recorded by CARTO in a sequential way. We found that we can analyze these datasets in a very similar way as the ATs; however, we do need to adapt our parameters to optimize the analysis of DGM. Next, we are also analyzing extremely rare socket data of episodes of VF to better understand the mechanism of VF. We are currently comparing our results with previously performed analysis of PM. We aim to visualize the excitation fronts of the arrhythmia with the wavefront feature of DGM. We stress again that due to the generality of DGM, there is no limit to the type of dataset DGM can analyze, although each dataset will require specialized additional algorithms to optimize the analysis.
6 Discussion
6.1 DGM as a tool to process arrhythmia: moving forward to a clinical application
This manuscript presents the first version of DGM, a software package to automatically analyze any given dataset of an arrhythmia. The minimal input of this software is a single datafile containing the LATs and corresponding spatial distribution of the electrodes which were used to measure the arrhythmia under study. DGM can work with regular arrhythmia (a single LAT per coordinate) or irregular arrhythmia (multiple LATs per coordinate). DGM processes the data by creating a directed network of the excitation of the arrhythmia. In short, rotational activity is found by detected the cycles in the network and focal activity by detecting regions with only outgoing arrows. By making the program public for academic use, we hope to contribute to the community by making the analysis of arrhythmia fast and robust.
One of the major problems in science is reproducability of results. For DGM to be a reliable tool which could eventually be used for ablation procedures, it has to be validated on many different datasets, different systems and different hospitals. It is therefore of paramount importance that DGM will be further validated by different EPs in different hospitals. Moreover, based on new input, DGM needs to be further optimized. Our final goal would be to integrate DGM in an (multiple) existing mapping system(s) if it can be proven that DGM makes the procedures faster and more robust. AT is currently the most developed application of DGM, but we hope to show that DGM is also useful for other arrhythmias.
Another major problem is access to novel data. Due to privacy reasons, data sharing always needs consent of the patient, which can be difficult to obtain afterwards. We therefore recommend that EPs ask consent of the patient before the procedure, for data to be processed afterwards.
DGM is different than any other mapping method as is directly outputs the correct reentry circuit on the mapped region. For any other mapping system (moving LAT map in RHYTHMIA, the normal static LAT map, the activation mapping system of Biosense Webster with integrating vectors), the EP still have to interpret the maps manually to make a diagnosis. In comparison with PM, DGM can find anatomical reentry, while PM is more suitable to find rotors (functional reentry) instead of anatomical reentry. Moreover, we have shown that DGM is much more robust for finding rotors in noisy datasets, see Section 6.6.
6.2 Choosing the right parameters for optimal clinical outcome
DGM is currently manually and empirically optimized to find the best outcome based on datasets from a single center (AZ Sint Jan Bruges). We realize that smarter and more advanced tools are available to find an optimal parameter setting (such as genetic evolution and neural networks approaches). However, we need to collect huge amounts of data, from which we know the ground truth before we can automatically optimize DGM. For example, to automatically optimize AT we need a large cohort of data including a careful entrainment mapping study for each case.
6.3 LAT annotation: which annotation to use?
Currently, we completely rely on CARTO and RHYTHMIA annotated LAT for the analysis of DGM in clinical settings. However, we are currently working on implementing our own annotation for regular arrhythmia, which gives a similar outcome as using the company annotations. Although the LAT annotation of a mapping system might not be perfect, and methods exists to improve LAT annotation [86, 87], DGM is robust and can overcome some faulty LAT annotations due to its global nature. Indeed, if a few LATs are wrong, the network can form cycles around the false LAT. We have demonstrated this in earlier work [44].
For irregular arrhythmia, like AF, annotations can be tedious [86] to impossible. Therefore, one of our future goals is to make DGM independent of the LAT annotation and draw arrows based on the ECGM signals of two neighboring electrodes and their causal correlations.
6.4 Interpolation
DGM is currently not able to interpolate signals on unmapped regions. Therefore, DGM does not analyze regions where no electrode recordings are taken. Hence, it is key to have a uniform distribution of the spatial electrodes. The interpretation of the arrhythmia will not be accurate in case, e.g., solely the anterior wall is mapped.
6.5 Optimal amount of electrodes
Not only the uniformity but also the total amount of electrodes play an important role with respect to the scale of the arrhythmia. One can appreciate that if little electrode recordings are taken while also some electrode recordings are inaccurately annotated, the LAT-representation can differ significantly from the ground truth. We have noticed that this plays in important role in the common isthmus of a double loop circuit.
Therefore, in case of a regular AT, DGM needs around 500 to 1500 points which are uniformly distributed. However, the software can also handle ATs with a density of around 200 points, but this might lead to wrong diagnosis certainly in cases where wrongly annotated signals tend to play a keyrole in the interpretation of the AT. Changing a single electrode recording in a sparse network can have a big influence, while DGM would be able to discard this electrode if the network was more dense. If is of course logical that in case solely the anterior wall is mapped and the reentry or focal breakthrough/automaticity is located at the posterior wall, DGM will not be able to diagnose the AT. In the future, it might be a good idea to formalize the way the arrhythmia is measured (number of nodes/uniformity), to produce a warning when we expect DGM not to perform well.
On the other hand, in case you have an abundance amount of electrode recordings (such as over 3500), DGM will reduce this amount in a smart way. This is because the BFS algorithm slows down when more points are added. Also, we noticed that for regular ATs, having more than 20 points at the same 1 cm2 is not necessary as there is enough information in these 20 points.
In conclusion, we advise to have at least 500 uniformly distributed points in the atrium to record a regular AT. We do realize this is not always possible in unstable arrhythmias. Moreover, for VT, it might not be possible as the arrhythmia is not hemodynamically stable. To overcome this issues, we will have to look for additional algorithms, which can also analyze sparse data, or partial datasets.
6.6 DGM and PM
As PM remains one of the most popular methods to analyze cardiac data, we also plan to implement PM methods so the user can compare DGM with PM. This can save time for researchers which want to implement PM while also enabling a more standardized approach for PM guided research.
In addition, we have already implemented a hybrid approach combining DGM with PM (which we will be released in the future). In this approach the velocity restriction to construct the network is eliminated, while the locality of PM is globalized. It gives the possibility to give a likelihood on the detection by PM by means of the variance and intervariability, in contrast with the discrete yes/no outcome of PM. So in complex data, which is difficult to annotate, this hybrid method could be a solution. Indeed, this would allow us to use a Hilbert transform as presented in [37], overcoming the problem of annotating the LATs.
6.7 Comparison to the open software package openEP and COSMAS
Recently, openEP was launched [88], which is an open source software package for electrophysiology data analysis. OpenEP consists of two main modules: (1) it has a data parsing modules which allow proprietary data formats from major electroanatomic mapping platforms to be converted into the OpenEP data format and (2) a data analysis modules which covers 90% of the analysis techniques in use in contemporary electrophysiology research. The second module can create activation maps, voltage maps, compute the conduction velocities, display and annotate the electrograms, analyze the ablation lines, in addition to a few more features. However, openEP cannot analyze the arrhythmia by automatically displaying the circuit on the geometry and most of its features are also present in the current activation mapping systems. OpenEP also does not use any networks to analyze the mapping data. Therefore, openEP could be an additional useful tool in combination with DGM as they complement each other. In the future, it would therefore be useful to implement the data format of openEP to create additional workflow speed for the analysis of arrhythmia.
Another recent software package COSMOS was launched for the analysis of optical mapping recordings in cardiac preparations [89]. It generates activation and conduction velocity maps, and it visualizes the action potential and calcium transient duration. Also COSMOS does not analyze any reentry of focal activation automatically. Therefore, it would also be possible for DGM to analyze data generated by COSMOS automatically by adding an additional module which can read the output of this software package.
7 Conclusion
DGM is a versatile tool which can automatically analyze any type of arrhythmia to find reentry or focal sources, if the measurements are synchronized in time. Currently, DGM requires the spatial coordinates of the measurements and the corresponding LATs. However, we are currently implementing a feature so DGM will also be able to work with the full ECGM signal of the spatial coordinates. DGM is already well tested for cases of regular AT and can compete with the state of the art mapping systems [44]. However, we have demonstrated its wide applicability on many different arrhythmia in this manuscripts, beyond AT. We encourage researchers and EPs to register on wwww.dgmapping.com to receive the first version of DGM. Newer versions with additional features will be available in the future.
References
Gibson D N, Di Biase L, Mohanty P, Patel J D, Bai R, Sanchez J, Burkhardt J D, Heywood J T, Johnson A D, Rubenson D S et al (2011) Stiff left atrial syndrome after catheter ablation for atrial fibrillation: clinical characterization, prevalence, and predictors. Heart Rhythm 8(9):1364–1371
Pogwizd S M, Hoyt R H, Saffitz J E, Corr P B, Cox James Lewis, Cain M E (1992) Reentrant and focal mechanisms underlying ventricular tachycardia in the human heart. Circulation 86(6):1872–1887
Strisciuglio T, Vandersickel N, Lorenzo G, Van Nieuwenhuyse E, El Haddad M, De Pooter J, Kyriakopoulou M, Almorad A, Lycke M, Vandekerckhove Y et al (2020) Prospective evaluation of entrainment mapping as an adjunct to new-generation high-density activation mapping systems of left atrial tachycardias. Heart Rhythm 17(2):211–219
El Haddad M, Houben R, Tavernier R, Duytschaever M (2014) Stable reentrant circuit with spiral wave activation driving atrial tachycardia. Heart Rhythm 11(4):716–718
Haissaguerre M, Jaïs P, Shah D C, Takahashi A, Hocini M, Quiniou G, Garrigue S, Le Mouroux A, Le Métayer P, Clémenty J (1998) Spontaneous initiation of atrial fibrillation by ectopic beats originating in the pulmonary veins. N Engl J Med 339(10):659–666
Brooks AG, Stiles MK, Laborderie J, Lau D H, Kuklik P, Shipp NJ, Hsu L-F, Sanders P (2010) Outcomes of long-standing persistent atrial fibrillation ablation: a systematic review. Heart Rhythm 7 (6):835–846
Weerasooriya R, Khairy P, Litalien J, Macle L et al (2011) Catheter ablation for atrial fibrillation: are results maintained at 5 years of follow-up? JACC 57:160–166. 21. Altman RK, Proietti R, Barett CD, Perini AP et al. Management of refractory atrial fibrillation post surgical ablation. Ann Cardiothorac Surg 3(1):91–97, 2014
Weiss J N, Qu Z, Shivkumar K (2016) Ablating atrial fibrillation: a translational science perspective for clinicians. Heart Rhythm 13:1868–1877
Roney CH, Wit AL, Peters NS (2020) Challenges associated with interpreting mechanisms of af. Arrhythmia Electrophysiol Rev 8(4):273
Moe Gordon K, Rheinboldt Werner C, Abildskov J A (1964) A computer model of atrial fibrillation. Am Heart J 67(2):200–220
Allessie MA (1985) Experimental evaluation of moe’s multiple wavelet hypothesis of atrial fibrillation. In: Cardiac electrophysiology and arrhythmias, pp 265–275
Cuculich PS, Wang Y, Lindsay B D, Faddis M N, Schuessler RB, Damiano R J Jr, Li L, Rudy Y (2010) Noninvasive characterization of epicardial activation in humans with diverse atrial fibrillation patterns. Circulation 122(14):1364–1372
Child N, Clayton R H, Roney C R, Laughner J I, Shuros A, Neuzil P, Petru J, Jackson T, Porter B, Bostock J et al (2018) Unraveling the underlying arrhythmia mechanism in persistent atrial fibrillation: results from the starlight study. Circ Arrhythm Electrophysiol 11(6):e005897
Keldermann R H, ten Tusscher K H W J, Nash M P, Bradley C P, Hren R, Taggart P, Panfilov A V (2009) A computational study of mother rotor vf in the human ventricles. Am J Physiol Heart Circ Physiol 296(2):H370–H379
Jalife José, Berenfeld Omer, Mansour Moussa (2002) Mother rotors and fibrillatory conduction: a mechanism of atrial fibrillation. Cardiovasc Res 54(2):204–216
Narayan S M, Krummen D E, Shivkumar K, Clopton P, Rappel W -J, Miller J M (2012) Treatment of atrial fibrillation by the ablation of localized sources: confirm (conventional ablation for atrial fibrillation with or without focal impulse and rotor modulation) trial. J Am Coll Cardiol 60(7):628–636
Narayan SM, Baykaner T, Clopton P, Schricker A, Lalani GG, Krummen DE, Shivkumar K, Miller JM (2014) Ablation of rotor and focal sources reduces late recurrence of atrial fibrillation compared with trigger ablation alone: extended follow-up of the confirm trial (conventional ablation for atrial fibrillation with or without focal impulse and rotor modulation). J Am Coll Cardiol 63:1761–1768
Morgan R, Colman MA, Chubb H, Seemann G, Aslanidi OV (2016) Slow conduction in the border zones of patchy fibrosis stabilizes the drivers for atrial fibrillation: insights from multi-scale human atrial modeling. Front Physiol 7:474
Vigmond E, Pashaei A, Amraoui S, Cochet H, Hassaguerre M (2016) Percolation as a mechanism to explain atrial fractionated electrograms and reentry in a fibrosis model based on imaging data. Heart Rhythm. https://pubmed.ncbi.nlm.nih.gov/26976038/
Haissaguerre M, Hocini M, Denis A, Shah A J, Komatsu Y, Yamashita S, Daly M, Amraoui S, Zellerhoff S, Picat M -Q et al (2014) Driver domains in persistent atrial fibrillation. Circulation 130(7):530–538
Bayer J D, Roney C H, Pashaei A, Jaïs P, Vigmond E J (2016) Novel radiofrequency ablation strategies for terminating atrial fibrillation in the left atrium: a simulation study. Front Physiol 7:108
Zahid S, Cochet H, Boyle P M, Schwarz E L, Whyte K N, Vigmond E J, Dubois R, Hocini M, Haïssaguerre M, Jaïs P, Trayanova N A (2016) Patient-derived models link reentrant driver localization in atrial fibrillation to fibrosis spatial pattern. Cardiovasc Res. https://pubmed.ncbi.nlm.nih.gov/27056895/
Allessie MA, de Groot NMS, Houben RPM, Schotten U, Boersma E, Smeets JL, Crijns H (2010) Electropathological substrate of long-standing persistent atrial fibrillation in patients with structural heart diseaseclinical perspective. Circ: Arrhythm Electrophysiol 3(6):606–615
de Groot N, Van Der Does L, Yaksh A, Lanters E, Teuwen C, Knops P, van de Woestijne P, Bekkers J, Kik C, Bogers A et al (2016) Direct proof of endo-epicardial asynchrony of the atrial wall during atrial fibrillation in humans. Circ: Arrhythm Electrophysiol 9(5):e003648
de Groot NMS, Houben RPM, Smeets JL, Boersma E, Schotten U, Schalij MJ, Crijns H, Allessie MA (2010) Electropathological substrate of longstanding persistent atrial fibrillation in patients with structural heart diseaseclinical perspective. Circulation 122(17):1674–1682
Gharaviri A, Pezzuto S, Potse M, Verheule S, Conte G, Krause R, Schotten U, Auricchio A (2020) Left atrial appendage electrical isolation reduces atrial fibrillation recurrences: a simulation study. Circ: Arrhythm Electrophysiol CIRCEP–120
Hansen B J, Zhao J, Csepe T A, Moore B T, Li N, Jayne L A, Kalyanasundaram A, Lim P, Bratasz A, Powell K A et al (2015) Atrial fibrillation driven by micro-anatomic intramural re-entry revealed by simultaneous sub-epicardial and sub-endocardial optical mapping in explanted human hearts. Eur Heart J 36(35):2390–2401
Zhao J, Hansen BJ, Wang Y, Csepe TA, Sul LV, Tang A, Yuan Y, Li N, Bratasz A, Powell KA et al (2017) Three-dimensional integrated functional, structural, and computational mapping to define the structural “fingerprints” of heart-specific atrial fibrillation drivers in human heart ex vivo. J Am Heart Assoc 6(8):e005922
Krummen DE, Ho G, Villongco CT, Hayase J, Schricker AA (2016) Ventricular fibrillation: triggers, mechanisms and therapies. Future Cardiol 12(3):373–390
de Luna A B, Coumel P, Leclercq JF (1989) Ambulatory sudden cardiac death: mechanisms of production of fatal arrhythmia on the basis of data from 157 cases. Am Heart J 117(1):151–159
Nash MP, Bradley CP, Sutton PM, Clayton RH, Kallis P, Hayward MP, Paterson DJ, Taggart P (2006) Whole heart action potential duration restitution properties in cardiac patients: a combined clinical and modelling study. Exp Physiol 91(2):339–354
Nair K, Umapathy K, Farid T, Masse S, Mueller E, Sivanandan R V, Poku K, Rao V, Nair V, Butany J et al (2011) Intramural activation during early human ventricular fibrillationclinical perspective. Circ: Arrhythmia Electrophysiol 4(5):692–703
Umapathy K, Nair K, Masse S, Krishnan S, Rogers J, Nash MP, Nanthakumar K (2010) Phase mapping of cardiac fibrillation. Circ: Arrhythmia Electrophysiol 3(1):105–114
Gray RA, Pertsov AM, Jalife J (1998) Spatial and temporal organization during cardiac fibrillation. Nature 392(6671):75
Shors SM, Sahakian AV, Sih HJ, Swiryn S (1996) A method for determining high-resolution activation time delays in unipolar cardiac mapping. IEEE Trans Biomed Eng 43(12):1192– 1196
Bray M -A, Wikswo J P (2002) Use of topological charge to determine filament location and dynamics in a numerical model of scroll wave activity. IEEE Trans Biomed Eng 49(10):1086–1093
Kuklik P, Zeemering S, Maesen B, Maessen J, Crijns H, Verheule S, Ganesan AN, Schotten U (2014) Reconstruction of instantaneous phase of unipolar atrial contact electrogram using a concept of sinusoidal recomposition and hilbert transform. IEEE Trans Biomed Eng 62(1):296–302
Roney CH, Cantwell CD, Qureshi NA, Chowdhury RA, Dupont E, Lim P, Vigmond E, Tweedy JH, Ng FS, Peters NS (2017) Rotor tracking using phase of electrograms recorded during atrial fibrillation. Ann Biomed Eng 45(4):910–923
Kuklik P, Zeemering S, van Hunnik A, Maesen B, Pison L, Lau D H, Maessen J, Podziemski P, Meyer C, Schaffer B, Crijns H, Willems S, Schotten U (2017) Identification of rotors during human atrial fibrillation using contact mapping and phase singularity detection: technical considerations. IEEE Trans Bio-med Eng 64:310–318
Jacquemet V (2017) A statistical model of false negative and false positive detection of phase singularities. Chaos: An Interdisciplinary Journal of Nonlinear Science 27(10): 103124
Roney C H, Cantwell C D, Bayer J D, Qureshi N A, Lim P B, Tweedy J H, Kanagaratnam P, Peters N S, Vigmond E J, Ng F S (2017) Spatial resolution requirements for accurate identification of drivers of atrial fibrillation. Circ: Arrhythmia Electrophysiol 10(5):e004899
Laughner J, Shome S, Child N, Shuros A, Neuzil P, Gill J, Wright M (2016) Practical considerations of mapping persistent atrial fibrillation with whole-chamber basket catheters. JACC: Clin Electrophysiol 2(1):55–65
Jacquemet V (2018) Phase singularity detection through phase map interpolation: theory, advantages and limitations. Comput Biol Med 102:381–389
Vandersickel N, Van Nieuwenhuyse E, Van Cleemput N, Goedgebeur J, El Haddad M, De Neve J, Demolder A, Strisciuglio T, Duytschaever M, Panfilov A V (2019) Directed networks as a novel way to describe and analyze cardiac excitation: directed graph mapping. Front Physiol. https://pubmed.ncbi.nlm.nih.gov/31551814/
Martinez-Mateu L, Romero L, Ferrer-Albero A, Sebastian R, Rodríguez Matas JF, Jalife J, Berenfeld O, Saiz J (2018) Factors affecting basket catheter detection of real and phantom rotors in the atria: A computational study. PLoS Comput Biol 14(3):e1006017
Baykaner T, Zaman JAB (2019) Another method that shows organization in persistent af? That’sa raap. J Cardiovasc Electrophysiol 30(12):2713
Seitz J, Bars C, Théodore G, Beurtheret S, Lellouche N, Bremondy M, Ferracci A, Faure J, Penaranda G, Yamazaki M et al (2017) Af ablation guided by spatiotemporal electrogram dispersion without pulmonary vein isolation: a wholly patient-tailored approach. J Am Coll Cardiol 69(3):303–321
Qin M, Jiang W -F, Wu S -H, Xu K, Liu X (2020) Electrogram dispersion–guided driver ablation adjunctive to high-quality pulmonary vein isolation in atrial fibrillation of varying durations. J Cardiovasc Electrophysiol 31(1):48–60
Ríos-Muñoz GR, Arenal Á, Artés-Rodríguez A (2018) Real-time rotational activity detection in atrial fibrillation. Front Physiol 9:208
Laţcu D G, Enache B, Hasni K, Wedn AM, Zarqane N, Pathak A, Saoudi N Sequential ultra-high-density contact mapping of persistent atrial fibrillation: an efficient technique for driver identification. J Cardiovasc Electrophysiol. https://pubmed.ncbi.nlm.nih.gov/33155347/
Luongo G, Azzolin L, Rivolta MW, Almeida TP, Martínez JP, Soriano DC, Dössel O, Sassi R, Laguna P, Loewe A Machine learning to find areas of rotors sustaining atrial fibrillation from the ecg
Ganesan P, Cherry EM, Huang DT, Pertsov AM, Ghoraani B (2020) Atrial fibrillation source area probability mapping using electrogram patterns of multipole catheters. BioMedical Eng OnLine 19:1–23
Ganesan P, Salmin A, Cherry EM, Huang DT, Pertsov AM, Ghoraani B (2019) Iterative navigation of multipole diagnostic catheters to locate repeating-pattern atrial fibrillation drivers. J Cardiovasc Electrophysiol 30(5):758–768
McGillivray MF, Cheng W, Peters NS, Christensen K (2018) Machine learning methods for locating re-entrant drivers from electrograms in a model of atrial fibrillation. R Soc Open Science 5(4):172434
Zolotarev A M, Hansen B J, Ivanova E A, Helfrich K M, Li N, Janssen P M L, Mohler P J, Mokadam N A, Whitson B A, Fedorov M V et al (2020) Optical mapping-validated machine learning improves atrial fibrillation driver detection by multi-electrode mapping. Circ: Arrhythmia Electrophysiol 13(10):e008249
Quintanilla J G, Alfonso-Almazán J M, Pérez-Castellano N, Pandit S V, Jalife J, Pérez-Villacastín J, Filgueiras-Rama D (2019) Instantaneous amplitude and frequency modulations detect the footprint of rotational activity and reveal stable driver regions as targets for persistent atrial fibrillation ablation. Circ Res 125(6):609–627
Pappone C, Ciconte G, Vicedomini G, Mangual J O, Li W, Conti M, Giannelli L, Lipartiti F, McSpadden L, Ryu K et al (2018) Clinical outcome of electrophysiologically guided ablation for nonparoxysmal atrial fibrillation using a novel real-time 3-dimensional mapping technique: results from a prospective randomized trial. Circ: Arrhythmia Electrophysiol 11(3):e005904
Choudry S, Mansour M, Sundaram S, Nguyen DT, Dukkipati SR, Whang W, Kessman P, Reddy VY (2020) Radar: a multicenter food and drug administration investigational device exemption clinical trial of persistent atrial fibrillation. Circ: Arrhythmia Electrophysiol 13(1):e007825
Willems S, Verma A, Betts TR, Murray S, Neuzil P, Ince H, Steven D, Sultan A, Heck PM, Hall M C et al (2019) Targeting nonpulmonary vein sources in persistent atrial fibrillation identified by noncontact charge density mapping: uncover af trial. Circ: Arrhythmia Electrophysiol 12(7):e007233
Ciconte G, Vicedomini G, Li W, Mangual JO, McSpadden L, Ryu K, Saviano M, Vitale R, Conti M, Ćalović ž et al (2019) Non-paroxysmal atrial fibrillation mapping: characterization of the electrophysiological substrate using a novel integrated mapping technique. EP Europace 21(8):1193–1202
Honarbakhsh S, Hunter RJ, Ullah W, Keating E, Finlay M, Schilling RJ (2019) Ablation in persistent atrial fibrillation using stochastic trajectory analysis of ranked signals (star) mapping method. JACC: Clin Electrophysiol 5(7):817–829
Bhatia NK, Rogers AJ, Krummen DE, Hossainy S, Sauer W, Miller JM, Alhusseini MI, Peszek A, Armenia E, Baykaner T et al (2020) Termination of persistent atrial fibrillation by ablating sites that control large atrial areas. EP Europace 22(6):897–905
Swerdlow M, Tamboli M, Alhusseini MI, Moosvi N, Rogers AJ, Leef G, Wang PJ, Rillig A, Brachmann J, Sauer WH et al (2019) Comparing phase and electrographic flow mapping for persistent atrial fibrillation. Pacing Clin Electrophysiol 42(5):499–507
Lebert J, Ravi N, Fenton F, Christoph J (2021) Rotor localization and phase mapping of cardiac excitation waves using deep neural networks. arXiv:2109.10472
Oesterlein TG, Loewe A, Lenis G, Luik A, Schmitt C, Doessel O (2018) Automatic identification of reentry mechanisms and critical sites during atrial tachycardia by analyzing areas of activity. IEEE Trans Biomed Eng 65(10):2334–2344
Salmin AJ, Ganesan P, Shillieto KE, Cherry EM, Huang DT, Pertsov AM, Ghoraani B (2016) A novel catheter-guidance algorithm for localization of atrial fibrillation rotor and focal sources. In: 2016 38th Annual international conference of the IEEE engineering in medicine and biology society (EMBC). IEEE, pp 501–504
Chauhan V S, Verma A, Nayyar S, Timmerman N, Tomlinson G, Porta-Sanchez A, Gizurarson S, Haldar S, Suszko A, Ragot D et al (2020) Focal source and trigger mapping in atrial fibrillation: randomized controlled trial evaluating a novel adjunctive ablation strategy. Heart Rhythm 17(5):683–691
Anzt H, Bach F, Druskat S, Löffler F, Loewe A, Renard B Y, Seemann G, Struck A, Achhammer E, Aggarwal P et al (2020) An environment for sustainable research software in germany and beyond:, current state, open challenges, and call for action. arXiv:2005.01469
Van Nieuwenhuyse E, Strisciuglio T, Lorenzo G, El Haddad M, Goedgebeur J, Van Cleemput N, Ley C, Panfilov AV, de Pooter J, Vandekerckhove Y et al (2021) Evaluation of directed graph-mapping in complex atrial tachycardias. JACC: Clin Electrophysiol. https://pubmed.ncbi.nlm.nih.gov/33812833/
Vandersickel N, Bossu A, De Neve J, Dunnink A, Meijborg VMF, van der Heyden MAG, Beekman JDM, De Bakker JMT, Vos MA, Panfilov AV (2017) Short-lasting episodes of torsade de pointes in the chronic atrioventricular block dog model have a focal mechanism, while longer-lasting episodes are maintained by re-entry. JACC: Clin Electrophysiol 3(13):1565– 1576
Van Nieuwenhuyse E, Martinez-Mateu L, Saiz J, Panfilov AV, Vandersickel N (2021) Directed graph mapping exceeds phase mapping in discriminating true and false rotors detected with a basket catheter in a complex in-silico excitation pattern. Comput Biol Med 104381:133
Plank G, Loewe A, Neic A, Augustin C, Huang Y-L, Gsell MAF, Sanchez J, Karabelas E, Nothstein M, Prassl AJ, Seemann G, Vigmond EJ (2021) The openCARP simulation environment for cardiac electrophysiology, pages under review. bioRxiv preprint available
Vincent GM (1998) The molecular ggenetics of the long qt syndrome: genes causing fainting and sudden death. Ann Rev Med 49(1):263–274
Asano Y, Davidenko JM, Baxter WT, Gray RA, Jalife J (1997) Optical mapping of drug-induced polymorphic arrhythmias and torsade de pointes in the isolated rabbit heart. J Am Coll Cardiol 29(4):831–842
Choi B-R, Burton F, Salama G (2002) Cytosolic ca2+ triggers early afterdepolarizations and torsade de pointes in rabbit hearts with type 2 long qt syndrome. J Physiol 543(Pt 2):615–631
El Sherif N, Caref EB, Yin H, Restivo M (1996) The electrophysiological mechanism of ventricular arrhythmias in the long qt syndrome. Tridimensional mapping of activation and recovery patterns. Circ Res 79(3):474–492
El-Sherif N, Chinushi M, Caref EB, Restivo M (1997) Electrophysiological mechanism of the characteristic electrocardiographic morphology of torsade de pointes tachyarrhythmias in the long-qt syndrome: detailed analysis of ventricular tridimensional activation patterns. Circulation 96(12):4392–4399
Kozhevnikov DO, Yamamoto K, Robotis D, Restivo M, El-Sherif N (2002) Electrophysiological mechanism of enhanced susceptibility of hypertrophied heart to acquired torsade de pointes arrhythmias: tridimensional mapping of activation and recovery patterns. Circulation 105(9):1128–1134
Murakawa Y, Sezaki K, Yamashita T, Kanese Y, Omata M (1997) Three-dimensional activation sequence of cesium-induced ventricular arrhythmias. Am J Physiol 273(3 Pt 2):H1377– H1385
Schreiner KD, Voss F, Senges JC, Becker R, Kraft P, Bauer A, Kelemen K, Kuebler W, Vos MA, Schoels W (2004) Tridimensional activation patterns of acquired torsade-de-pointes tachycardias in dogs with chronic av-block. Basic Res Cardiol 99(4):288–298
Senges JC, Sterns LD, Freigang KD, Bauer A, Becker R, Kübler W, Schoels W (2000) Cesium chloride induced ventricular arrhythmias in dogs: three-dimensional activation patterns and their relation to the cesium dose applied. Basic Res Cardiol 95(2):152–162
Boulaksil M, Jungschleger JG, Antoons G, Houtman MJ C, de Boer TP, Wilders R, Beekman JD, Maessen J, van der Hulst FF, van der Heyden MAG, van Veen TAB, van Rijen HVM, de Bakker JMT, Vos MA (2011) Drug-induced torsade de pointes arrhythmias in the chronic av block dog are perpetuated by focal activity. Circ Arrhythm Electrophysiol 4(4):566–576
Kim TY, Kunitomo Y, Pfeiffer Z, Patel D, Hwang J, Harrison K, Patel B, Jeng P, Ziv O, Lu Y, Peng X, Qu Z, Koren G, Choi B-R (2015) Complex excitation dynamics underlie polymorphic ventricular tachycardia in a transgenic rabbit model of long qt syndrome type 1. Heart Rhythm 12 (1):220–228
Dunnink A, Stams TRG, Bossu A, Meijborg VMF, Beekman JDM, Wijers SC, de Bakker JMT, Vos MA (2016) Torsade de pointes arrhythmias arise at the site of maximal heterogeneity of repolarization in the chronic complete atrioventricular block dog. Europace
Akar FG, Yan G-X, Antzelevitch C, Rosenbaum DS (2002) Unique topographical distribution of m cells underlies reentrant mechanism of torsade de pointes in the long-qt syndrome. Circulation 105 (10):1247–1253
Cantwell CD, Roney CH, Ng FS, Siggers JH, Sherwin S J, Peters NS (2015) Techniques for automated local activation time annotation and conduction velocity estimation in cardiac mapping. Comput Biol Med 65:229–242
Barber F, Langfield P, Lozano M, García-fernández I, Duchateau J, Hocini M, Haïssaguerre M, Vigmond E, Sebastian R (2021) Estimation of personalized minimal purkinje systems from human electro-anatomical maps. IEEE Trans Med Imaging. https://pubmed.ncbi.nlm.nih.gov/33856987/
Williams SE, Roney CH, Connolly A, Sim I, Whitaker J, O’Hare D, Kotadia I, O’Neill L, Corrado C, Bishop M et al (2021) Openep: a cross-platform electroanatomic mapping data format and analysis platform for electrophysiology research. Front Physiol 12:160
Tomek J, Wang ZJ, Burton R-AB, Herring N, Bub G (2021) Cosmas: a lightweight toolbox for cardiac optical mapping analysis. Sci Rep 11(1):1–17
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Van Nieuwenhuyse, E., Hendrickx, S., Abeele, R.V.d. et al. DG-Mapping: a novel software package for the analysis of any type of reentry and focal activation of simulated, experimental or clinical data of cardiac arrhythmia. Med Biol Eng Comput 60, 1929–1945 (2022). https://doi.org/10.1007/s11517-022-02550-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11517-022-02550-y