Abstract
We have proposed an algorithm to guide radiofrequency catheter ablation procedures. This algorithm employs the single equivalent moving dipole (SEMD) to model cardiac electrical activity. The aim of this study is to investigate the optimal time instant during the cardiac cycle as well as the number of beats needed to accurately estimate the location of a pacing site. We have evaluated this algorithm by pacing the ventricular epicardial surface and inversely estimating the locations of pacing electrodes from the recorded body surface potentials. Two pacing electrode arrays were sutured on the right and left ventricular epicardial surfaces in swine. The hearts were paced by the electrodes sequentially at multiple rates (120–220 bpm), and body surface ECG signals from 64 leads were recorded for the SEMD estimation. We evaluated the combined error of the estimated interelectrode distance and SEMD direction at each time instant during the cardiac cycle, and found the error was minimum when the normalized root mean square (RMS n ) value of body surface ECG signals reached 15 % of its maximum value. The beat-to-beat variation of the SEMD locations was significantly reduced (p < 0.001) when estimated at 15 % RMS n compared to the earliest activation time (EAT). In addition, the 5–95 % interval of the estimated interelectrode distance error decreased exponentially as the number of beats used to estimate a median beat increased. When the number of beats was 4 or larger, the 5–95 % interval was smaller than 3.5 mm (the diameter of a commonly used catheter). In conclusion, the optimal time for the SEMD estimation is at 15 % of RMS n , and at that time instant a median beat estimated from 4 beats is associated with a beat-to-beat variability of the SEMD location that is appropriate for catheter ablation procedures.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Ventricular tachycardia (VT) is an abnormally rapid heart rhythm originating from one of the ventricles and may develop to ventricular fibrillation (VF), an even faster and totally uncoordinated heart rhythm, causing sudden cardiac death. Radiofrequency catheter ablation (RCA) is an established treatment of VT, during which the operator attempts to deliver curative radiofrequency energy, for example to the exit of the reentry circuit. However, the success of the procedure greatly depends on the accurate identification of the target site for ablation. There have been techniques to identify the target site and guide the ablation catheter to the site using conventional endocardial mapping [18] and electro-anatomical mapping systems [17, 25]. These methods determine the activation pattern and locate the target site based on the pattern.
We have proposed an algorithm that aims to guide RCA procedures to the target ventricular site [2, 3]. In this method, the heart’s electrical activity is modeled by a single equivalent moving dipole (SEMD), and the location of the SEMD is inversely estimated from the measured body surface potentials. An important aspect of this method involves determining the optimal time instant during the cardiac cycle to estimate the target site location for ablation. Early during the cardiac cycle, the excitation wave-front is localized and can be accurately approximated with a SEMD; however, a small excitation wave-front gives rise to small body surface potentials which result in low signal-to-noise ratio with significant implications on the accuracy of the SEMD estimation. In prior studies, we employed the earliest activation time (EAT) in the body surface ECG as the optimal time instant to estimate the target site [1–3, 6–9, 21, 22]. However, one of the drawbacks of EAT is its relatively large beat-to-beat variation, especially in the diseased heart.
In this study, we aim to determine the time instant during the cardiac cycle that would best approximate the excitation wave-front emanating from an epicardial pacing electrode to a SEMD, and to assess the minimum number of beats required optimally estimate the location of the SEMD. For that matter, we evaluate a new “optimal” instant during the cardiac cycle to estimate the SEMD that is determined by the root mean square (RMS) of the body surface ECG signals. We hypothesized that the RMS “signal” is more robust to ECG variability than the EAT. Furthermore, we examine what is the minimum number of beats needed to estimate the SEMD with sufficient accuracy in order to guide radiofrequency catheter ablation procedures.
2 Methods
2.1 Experimental protocol
The experimental protocol was approved by the MIT Committee on Animal Care. We performed 13 swine studies. Weight, radius and length of the swine were 39.2 ± 4.3 kg, 14.8 ± 1.2 cm and 59.4 ± 1.6 cm, respectively.
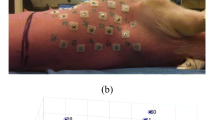
The surgical preparation has been described previously [2, 3]. Briefly, a linear electrode array was sutured on the epicardial surface of the right ventricle longitudinally from apex to base direction, and another electrode array was also sutured on the left ventricle in the same way. There were nine electrodes on each electrode array. Electrodes on the right ventricular array were named A1–A9 from apex to base. Electrodes on the left ventricular array were named B1–B9, following the same configuration. The interelectrode distances of A1A2 and B1B2 were 0.2 cm, and all other adjacent electrode distances were 1 cm. The chest was closed in layers, and the air was evacuated from the chest. We paced the heart from each epicardial electrode at a wide range of rates, suitable to simulate ventricular tachycardia: 120, 140, 160, 180, 200 and 220 beats/min.
Sixty-four recording electrodes were placed on the torso to measure body surface potentials. The ECG acquisition system included high-fidelity BioSemi amplifiers (Active Two, BioSemi, Netherlands), a very flexible, 24-bit, true DC, battery-powered front-end, with fiber optic data transfer system which allowed the recording of 64 body surface potentials that were digitized at 8192 Hz (Labview, National Instrument Corp, Austin, TX). Data were analyzed off-line using custom-built software.
2.2 SEMD algorithm
The algorithm for estimating the SEMD has been previously described. Briefly, in off-line analysis, the R-wave of each QRS complex was identified, and an iterative, adaptive QRS template matching scheme was used to refine initial fiducial point estimates [19]. Next, a median beat was computed for each ECG lead by aligning each beat within the lead data segment according to the R-wave and identifying the median value on a point-by-point basis within the beat.
To further examine the robustness of our SEMD parameter identification technique under clinically relevant conditions, we studied the beat-to-beat variation of the spatial SEMD parameters at identical instants of the cardiac cycle. By applying our algorithm to a sequence of successive beats, we can estimate the uncertainty [3, 22] that might arise from time-varying physiological effects. One then may compute the statistical properties of the parameter estimates for each sample-point across the sequence of time-aligned beats.
2.3 Method to determine the optimal time instant during the cardiac cycle
Estimation of the SEMD employs a three-step approach: (1) processing body surface potentials during ventricular tachycardia (a few beats) to obtain the SEMD parameters for a multiplicity of points during the beginning of ventricular depolarization, and from analysis of the SEMD one identifies the SEMD location corresponding to the site of origin of the arrhythmia when the bioelectrical source is mostly localized; (2) using the same algorithm to identify in real time (while the arrhythmia is ongoing) the location of the tip of the ablation catheter from analysis of potentials resulting from dipolar current pulses delivered at the tip; (3) advancing the ablation catheter until the SEMD corresponding to the tip of the catheter is superposed with the SEMD corresponding to the site of origin of the arrhythmia.
Therefore, the accuracy of the relative distance and direction of the two estimated locations (in steps (1) and (2) above) is more important than the accuracy of the two estimated locations themselves. In this study, we employed multiple pacing electrodes (as described above), and calculated their interelectrode distance. We determined the “optimal” time instant of the SEMD location corresponding to each pacing site as the instant that minimizes the error in estimating the interelectrode distance and direction. We defined the error of the interelectrode distance (E d ) and direction (E θ ) as follows.
where d e and d r are the estimated and real interelectrode distances, and \(\hat{e}\) and \(\hat{r}\) are the estimated and real unit vectors determined by two pacing electrodes, respectively.
Propagation of cardiac electrical activity is indicated by the RMS value of body surface potentials. We define the normalized RMS (RMS n ) relative to the maximum RMS, as follows.
where ϕ i (t): the potential values of the ith electrode at time t; N: number of electrodes; and n: normalized. As the depolarization wavefront propagates in the ventricles, RMS n increases from 0 to 100 % and decreases again. We evaluated the estimation errors from 1 to 100 % RMS n values, with 1 % increment. We defined E d,max and E θ,max as the maximum absolute E d and E θ values of RMS n (from 1 to 100 %) and then defined the normalized errors E d,n , E θ,n and E t,n , as follows. We determined the optimal instant for the pacing site estimation as the value of RMS n (Eq. 4), which minimizes E t,n :
2.4 Statistical analysis
We calculated the optimal timing to estimate the SEMD using the RMS n and EAT, with respect to the R-wave (time zero) of a predetermined channel, and calculated the standard deviation (std) of the estimated SEMD locations as a measure of beat-to-beat SEMD location variation across all animals, heart rates and pacing electrodes. Specifically, we calculated the std of the SEMD x-, y- and z-locations (stdx, stdy and stdz) at the optimal RMS n and EAT, and calculated the overall std as
We then applied ANOVA to determine whether RMS n timing is more stable than EAT (p < 0.05 indicates statistical significance).
3 Results
3.1 Determining the optimal time instant during the cardiac cycle
We calculated the normalized errors for all animals, heart rates and pacing electrodes, and displayed the median values in Fig. 1. We observed that the normalized total error (E t,n ) exhibits a minimum at ~12 to 23 % of RMS n , and therefore we chose the RMS n value of 15 % as the optimal time instant for estimating the SEMD parameters corresponding to the depolarization wavefront resulting from the pacing pulse.
An advantage of the RMS n over EAT is that the beat-to-beat variation of the RMS n timing is more stable (Fig. 2a, b). We observe that for each electrode (across all animals and heart rates), the std of the estimated beat-to-beat variation of the timing of the 15 % RMS n is much smaller than the corresponding std of the EAT. The overall std of the 15 % RMS n and EAT is 0.35 ± 0.28 ms and 1.70 ± 1.48 ms, respectively, in the right ventricle (p < 0.0001) and 0.39 ± 0.32 ms and 1.59 ± 1.13 ms, respectively, in the left ventricle (p < 0.0005).
Statistical representation across all beats, heart rates and animals, of the standard deviation of the timings of 15 % RMS n and EAT (a and b), and the standard deviation of the corresponding SEMD locations (c and d). For each recording, the timings of 15 % RMS n and EAT were calculated for every beat, and then the standard deviations were calculated across beats, heart rates and animals (a and b). In addition, the std of the SEMD locations was estimated (as described in the text) at the corresponding above-mentioned 15 % RMS n and EAT timings (c and d). Results from pacing at the right ventricle (A1–A9) are presented in a and c and the left ventricle (B1–B9) in b and d. The five nodes at each bar graph correspond to the values of 5, 25, 50, 75 and 95 %. The std difference of EAT and 15 % RMS n is statistically significant for both ventricles (p < 0.0001), and the mean std of locations estimated at EAT and 15 % RMS n is also significantly different for both ventricles (p < 0.0001 for the right ventricle and p < 0.0005 for the left ventricle)
In addition, we observe that the higher stability of the timing of the 15 % RMS n resulted in smaller dispersion of the estimated SEMD locations (Fig. 2c, d). The overall std for 15 % RMS n and EAT are 0.10 ± 0.08 and 0.15 ± 0.16 cm in the right ventricle and 0.10 ± 0.15 and 0.14 ± 0.18 cm in the left ventricle, and the differences are statistically significant (p < 0.001) for both ventricles.
Figure 3a, b shows the interelectrode distance calculated at the time instant of 15 % RMS n using the beat-to-beat analysis. When all hearts and all heart rates are considered together, the estimated A1A2 interelectrode distance is 0.34 ± 0.42 cm, while the interelectrode distance of the remaining electrodes in the right ventricle is 0.89 ± 0.66 cm. Similarly, in the left ventricle, the estimated B1B2 interelectrode distance is 0.34 ± 0.44 cm, while the remaining interelectrode distance is 0.92 ± 0.69 cm. The overall interelectrode distance error is 0.36 ± 0.46 cm in the right ventricle and 0.41 ± 0.47 cm in the left ventricle.
Statistical representation of the distance between adjacent pacing electrodes estimated at the time corresponding to 15 % of maximum RMS n , using the beat-to-beat analysis. Results from pacing at the right ventricle (A1–A9) and at the left ventricle (B1–B9) are presented in a and b, respectively. At each interelectrode pair, the distances at different heart rates are displayed separately. The five nodes at each bar graph correspond to the values of 5, 25, 50, 75 and 95 %. The real distances between A1 and A2 (A1A2) and B1 and B2 (B1B2) are 0.2 cm, and all others are 1.0 cm
Figure 4a, b shows the interelectrode distance calculated at the time instant of 15 % RMS n using the median beat. When all hearts and all heart rates are considered together, the estimated A1A2 interelectrode distance is 0.38 ± 0.43 cm, while the interelectrode distance of the remaining electrodes in the right ventricle is 0.90 ± 0.61 cm. Similarly, in the left ventricle, the estimated B1B2 interelectrode distance is 0.39 ± 0.42 cm, while the remaining interelectrode distance is 0.95 ± 0.62 cm. The overall interelectrode distance error is 0.34 ± 0.42 cm in the right ventricle and 0.40 ± 0.39 cm in the left ventricle.
Statistical representation of the distance between adjacent pacing electrodes estimated at the time corresponding to 15 % of maximum RMS n , using the median beat analysis. Results from pacing at the right ventricle (A1–A9) and at the left ventricle (B1–B9) are presented in a and b, respectively. At each interelectrode pair, the distances at different heart rates are displayed separately. The five nodes at each bar graph correspond to the values of 5, 25, 50, 75 and 95 %. The real distances between A1 and A2 (A1A2) and B1 and B2 (B1B2) are 0.2 cm, and all others are 1.0 cm
The significance of these figures lies not only in the fact that the spread of the spatial SEMD parameters is small when we want to ablate, but also in the fact that it demonstrates that analysis of only a few beats of a monomorphic rhythm like monomorphic ventricular tachycardia could provide meaningful results about the locus of the tachycardia.
3.2 Determining the optimal number of individual beats needed to estimate a median beat
Thereafter, we evaluated the effect of the number of beats needed to estimate a median beat that would reduce the beat-to-beat variation of the estimated interelectrode distance. Body surface potentials were recorded during pacing from each of the epicardial electrodes on the right- and left-ventricle, such that at each stimulation frequency ECG signals were recorded for ~20 s. After that, we estimated a median beat by randomly selecting a specific number of beats from the recorded signals. We repeated this process 10 times for each specific number of beats and estimated the error in the estimated interelectrode distance.
Figure 5 shows the error of the estimated interelectrode distance using the EAT [1–3, 6–9, 21, 22] and the 15 % RMS n as a function of the number of beats used to compute the median beat; results from all hearts, all heart rates and all pacing electrodes were included, and the median values of all those cases are displayed. Figure 5 shows that while the median (50 %) values do not change, the 5 and 95 % values converge toward the 50 % value. Using the 15 % RMS n , the 5–95 % interval decreases exponentially from 0.45 to 0.11 cm as the number of individual beats used to form the median beat increases from 1 to 40. When the SEMD is calculated at EAT, the 5–95 % intervals are ~0.06 cm larger than the intervals calculated at 15 % RMS n , which suggests that using the RMS n provides better accuracy of the SEMD parameter estimation.
Effect of the number of individual beats used to estimate the median beat, on the error of the interelectrode distance. The x-axis represents the number of beats used in the median beat calculation, and the y-axis presents a statistical representation of the interelectrode distance error values of 5, 50 and 95 % (left y-axis in mm, right y-axis in French) from all hearts, all heart rates and all pacing electrodes. SEMD was estimated at both EAT (square) and 15 % RMS n (circle)
4 Discussion
The most common etiology of VT in the presence of myocardial infarction, is the formation of reentrant circuit. Delivery of radiofrequency current to the exit site of the reentrant circuit remains the most successful approach to eliminate the reentrant VT [23, 26]. However, the most challenging aspect of this treatment option is the identification of the exit site [13, 24]. The selection of the appropriate ablation site is achieved by painstaking mapping of the electrical activity on the endocardial surface of the heart, and often, this recording must be done while the arrhythmia is ongoing.
To overcome these challenges, we have developed a method that models cardiac electrical activity with a SEMD, and we have performed a detailed evaluation in a swine animal model of the ability of this method to accurately identify the location of the SEMD [1–3, 6–9, 21, 22]. The choice of fitting the SEMD to potentials at EAT, as the time instant during the cardiac cycle that would be suitable for ablation, is based on the idea that the depolarization wave-front during ventricular tachycardia will be close to the exit and thus well localized, but also large enough to give rise to potentials that exceed the noise level. The EAT has been determined by finding the time instant of the fastest potential change of each body surface electrode and then choosing the earliest one among them. However, there are two concerns with determining the EAT. First, the automatically determined EAT is vulnerable to being distorted by noise, overlap of previous and current beats, etc., and thus the time instant of the fastest potential change may depend on the specific algorithm used. Second, it may not be accurate to calculate the EAT from body surface potentials. Therefore, we searched for the optimal time instant during the cardiac cycle to estimate the pacing site location, using the RMS n values of body surface potentials.
In this study, by employing multiple epicardial pacing electrodes and estimating the distance between the pacing electrodes, we determined the best time instant for the pacing site estimation as the one that minimizes the combined error of the estimated interelectrode distance as well as the direction between adjacent electrodes. In addition, an optimal number of beats needed to estimate a median beat was determined as the number that gives acceptable precision in the error of the estimated interelectrode distance. Specifically, we examined the interelectrode estimation errors from the time instant of the pacing time to the time instant of the maximum RMS n of body surface potentials, and found that the estimation error is minimum at 15 % RMS n . Furthermore, the automatic determination of the time instant of 15 % RMS n has been computationally efficient and robust.
Ilg et al. [11] used post-ablation cardiac magnetic resonance images to measure the ablation lesion size resulting from radiofrequency catheter ablation of ventricular arrhythmias in 35 patients, where either conventional or open irrigated-tip catheters were used, and reported the distribution of endocardial lesion area to be 3.5 ± 3.0 cm2 (mean ± std). Stevenson et al. [23] measured the epicardial radiofrequency ablation lesion size in the normal ventricular myocardium of 16 dogs and reported the length and width of the lesions to be 8.7 ± 3.3 mm and 6.5 ± 2.5 mm, respectively, with a conventional catheter and 9.3 ± 2.6 mm and 8.1 ± 1.9 mm, respectively, with a saline irrigation catheter. In endocardial ventricular radiofrequency ablation of 16 sheep performed with an open irrigated-tip catheter [24], the lesion length and width size were found to be 7.7 ± 2.1 mm and 5.3 ± 1.7 mm, respectively. To determine whether our method can achieve resolution appropriate for catheter ablation procedures, we sought to determine the minimum number of beats required to calculate the median beat used in the estimation of the SEMD, by minimizing the error in estimating the interelectrode distance. Using the beat-to-beat analysis, the 5–95 % variation of the interelectrode distance errors has been ~0.45 cm; use of the median beat reduces this variation and improves precision of estimation, and this reduction depends on the number of beats used to calculate the median beat (Fig. 5). The 5–95 % interval is smaller than a 3.5-mm-diameter catheter (typical size of an ablation catheter tip) at four beats (Fig. 5).
The CARTO™ (BiosenseWebster, Johnson and Johnson, Diamond Bar, CA, USA) system has been an established method that provides an electro-anatomical map of the heart under the assumption that the activation pattern and chamber geometry are constant from beat to beat. However, multiple morphologies of VT are common in patients with scar-related VT. Thus, monitoring of the activation sequence of the ongoing arrhythmia [13, 25] has resulted in a success rate of terminating the target VT of 82 %, while when targeting all inducible monomorphic VTs the complete procedural success was 64 %. An alternative to CARTO system has been the Ensite (St. Jude Medical, St. Paul, MN, USA) non-contact mapping system that utilizes a multi-electrode catheter which allows the simultaneous recording of multiple areas of endocardial activation [10]. The multi-electrode catheter is an inflatable balloon with 64 electrodes on its surface; hence, relatively high-density mapping can be performed from even a single beat. However, the movement of the heart during systole is not taken into account, which might limit the clinical applicability of this system in patients with large cardiac chambers, or it could require balloon repositioning to the areas of interest [14]. New cardiac imaging techniques [27], by either MRI [4, 12, 20], CT [15, 16, 28] or ultrasound [5] that have been introduced to overcome the above-mentioned limitations, require the acquisition of 3D anatomical information, and therefore the corresponding inverse solutions become particularly time consuming.
In conclusion, in this manuscript we evaluate an algorithm aiming to provide a reliable, accurate and fast means in guiding the ablation catheter to the site of the origin of the arrhythmia. Our approach is based on the principle that for many arrhythmias, the electrical activity within the heart is highly localized for a portion of the cardiac cycle. If one can localize the site of origin of the arrhythmia during the cardiac cycle, then one can ablate it. We have found that the optimal instant for the SEMD estimation is when the RMS n of body surface ECG signals reaches 15 % of its maximum. The beat-to-beat variation of the SEMD locations corresponding to the pacing site, estimated at 15 % RMS n is statistically significantly smaller than at EAT; this implies that during the guidance of the catheter to the target site, the beat-to-beat estimation of the distance of the catheter tip to the target site is more consistent. Furthermore, when a median beat is estimated from 4 beats only, the variability of the SEMD locations is smaller than the diameter of commonly used catheter ablation tips. In future studies, we need to further evaluate the proposed algorithm in a real catheter guidance procedure; for example, guide the catheter tip to a pacing electrode on the endocardial surface.
References
Armoundas AA, Feldman AB, Sherman DA, Cohen RJ (2001) Applicability of the single equivalent point dipole model to represent a spatially distributed bio-electrical source. Med Biol Eng Comput 39:562–570
Armoundas AA, Feldman AB, Mukkamala R, Cohen RJ (2003) A single equivalent moving dipole model: an efficient approach for localizing sites of origin of ventricular electrical activation. Ann Biomed Eng 31:564–576
Armoundas AA, Feldman AB, Mukkamala R, He B, Mullen TJ, Belk PA, Lee YZ, Cohen RJ (2003) Statistical accuracy of a moving equivalent dipole method to identify sites of origin of cardiac electrical activation. IEEE Trans Biomed Eng 50:1360–1370
Berger T, Fischer G, Pfeifer B, Modre R, Hanser F, Trieb T, Roithinger FX, Stuehlinger M, Pachinger O, Tilg B, Hintringer F (2006) Single-beat noninvasive imaging of cardiac electrophysiology of ventricular pre-excitation. J Am Coll Cardiol 48:2045–2052
Cheng LK, Sands GB, French RL, Withy SJ, Wong SP, Legget ME, Smith WM, Pullan AJ (2005) Rapid construction of a patient-specific torso model from 3D ultrasound for non-invasive imaging of cardiac electrophysiology. Med Biol Eng Comput 43:325–330
Fukuoka Y, Armoundas AA, Oostendorp TF, Cohen RJ (2000) Accuracy of a single equivalent moving dipole model in a realistic anatomic geometry torso model. Comput Cardiol 27:439–442
Fukuoka Y, Oostendorp TF, Cohen RJ, Armoundas AA (2001) A simulation study on a catheter navigation method for guiding the ablative therapy of cardiac arrhythmias. In 2001-Conference-Proceedings-of-the-23rd-Annual-International-Conference-of-the-IEEE-Engineering-in-Medicine-and-Biology-Society, Instanbul, pp 402–404
Fukuoka Y, Oostendorp TF, Sherman DA, Armoundas AA (2006) Applicability of the single equivalent moving dipole model in an infinite homogeneous medium to identify cardiac electrical sources: a computer simulation study in a realistic anatomic geometry torso model. IEEE Trans Biomed Eng 53:2436–2444
Fukuoka Y, Oostendorp TF, Armoundas AA (2009) Method for guiding the ablation catheter to the ablation site: a simulation and experimental study. Med Biol Eng Comput 47:267–278
Gornick CC, Adler SW, Pederson B, Hauck J, Budd J, Schweitzer J (1999) Validation of a new noncontact catheter system for electro anatomic mapping of left ventricular endocardium. Circulation 99:829–835
Ilg K, Baman TS, Gupta SK, Swanson S, Good E, Chugh A, Jongnarangsin K, Pelosi F Jr, Crawford T, Oral H, Morady F, Bogun F (2010) Assessment of radiofrequency ablation lesions by CMR imaging after ablation of idiopathic ventricular arrhythmias. JACC Cardiovasc Imaging 3:278–285
Modre R, Tilg B, Fischer G, Hanser F, Messnarz B, Seger M, Schocke MF, Berger T, Hintringer F, Roithinger FX (2003) Atrial noninvasive activation mapping of paced rhythm data. J Cardiovasc Electrophysiol 14:712–719
Nademanee K, Kosar EM (1998) A nonfluoroscopic catheter-based mapping technique to ablate focal ventricular tachycardia. Pacing Clin Electrophysiol 21:1442–1447
Okishige K, Kawabata M, Umayahara S, Yamashiro K, Gotoh M, Isobe M, Strickberger SA (2003) Radiofrequency catheter ablation of various kinds of arrhythmias guided by virtual electrograms using a noncontact, computerized mapping system. Circ J 67:455–460
Ramanathan C, Ghanem RN, Jia P, Ryu K, Rudy Y (2004) Noninvasive electrocardiographic imaging for cardiac electrophysiology and arrhythmia. Nat Med 10:422–428
Ramanathan C, Jia P, Ghanem R, Ryu K, Rudy Y (2006) Activation and repolarization of the normal human heart under complete physiological conditions. Proc Natl Acad Sci USA 103:6309–6314
Schilling RJ, Peters NS, Davies DW (1999) Feasibility of a noncontact catheter for endocardial mapping of human ventricular tachycardia. Circulation 99:2543–2552
Schmitt C, Zrenner B, Schneider M, Karch M, Ndrepepa G, Deisenhofer I, Weyerbrock S, Schreieck J, Schomig A (1999) Clinical experience with a novel multielectrode basket catheter in right atrial tachycardias. Circulation 99:2414–2422
Smith JM, Clancy EA, Valeri CR, Ruskin JN, Cohen RJ (1988) Electrical alternans and cardiac electrical instability. Circulation 77:110–121
Smith NP, Buist ML, Pullan AJ (2003) Altered T wave dynamics in a contracting cardiac model. J Cardiovasc Electrophysiol 14:S203–S209
Sohn K, Lv W, Lee K, Galea AM, Hirschman GB, Hayward AM, Cohen RJ, Armoundas AA (2013) The single equivalent moving dipole model does not require spatial anatomical information to determine cardiac sources of activation. IEEE J Biomed Health Inform 18:222–230
Sohn K, Lv W, Lee K, Galea A, Hirschman G, Barrett C, Cohen RJ, Armoundas AA (2014) A method to noninvasively identify cardiac bioelectrical sources. Pacing Clin Electrophysiol 37:1038–1050
Stevenson WG, Delacretaz E (2000) Radiofrequency catheter ablation of ventricular tachycardia. Heart 84:553–559
Stevenson WG, Sager PT, Natterson PD, Saxon LA, Middlekauff HR, Wiener I (1995) Relation of pace mapping QRS configuration and conduction delay to ventricular tachycardia reentry circuits in human infarct scars. J Am Coll Cardiol 26:481–488
Stevenson WG, Delacretaz E, Friedman PL, Ellison KE (1998) Identification and ablation of macro reentrant ventricular tachycardia with the CARTO electro anatomical mapping system. Pacing Clin Electrophysiol 21:1448–1456
Stevenson WG, Friedman PL, Kocovic D, Sager PT, Saxon LA, Pavri B (1998) Radiofrequency catheter ablation of ventricular tachycardia after myocardial infarction. Circulation 98:308–314
Wang Y, Cuculich PS, Zhang J, Desouza KA, Vijayakumar R, Chen J, Faddis MN, Lindsay BD, Smith TW, Rudy Y (2011) Noninvasive electro anatomic mapping of human ventricular arrhythmias with electrocardiographic imaging. Sci Trans Med 3:98ra84
Zhang X, Ramachandra I, Liu Z, Muneer B, Pogwizd SM, He B (2005) Noninvasive three-dimensional electrocardiographic imaging of ventricular activation sequence. Am J Physiol Heart Circ Physiol 289:H2724–H2732
Acknowledgments
The work was supported by NIH grant 1RO1HL103961.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Sohn, K., Armoundas, A.A. On the efficiency and accuracy of the single equivalent moving dipole method to identify sites of cardiac electrical activation. Med Biol Eng Comput 54, 1611–1619 (2016). https://doi.org/10.1007/s11517-015-1437-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11517-015-1437-x