Abstract
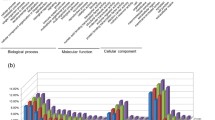
Panax ginseng C. A. Meyer is an important traditional herb in eastern Asia. It contains ginsenosides, which are primary bioactive compounds with medicinal properties. Although ginseng has been cultivated since at least the Ming dynasty to increase production, cultivated ginseng has lower quantities of ginsenosides and lower disease resistance than ginseng grown under natural conditions. We extracted root RNA from six varieties of fifth-year P. ginseng cultivars representing four different growth conditions, and performed Illumina paired-end sequencing. In total, 163,165,706 raw reads were obtained and used to generate a de novo transcriptome that consisted of 151,763 contigs (76,336 unigenes), of which 100,648 contigs (66.3%) were successfully annotated. Differential expression analysis revealed that most differentially expressed genes (DEGs) were upregulated (246 out of 258, 95.3%) in ginseng grown under natural conditions compared with that grown under artificial conditions. These DEGs were enriched in gene ontology (GO) terms including response to stimuli and localization. In particular, some key ginsenoside biosynthesis-related genes, including HMG-CoA synthase (HMGS), mevalonate kinase (MVK), and squalene epoxidase (SE), were upregulated in wild-grown ginseng. Moreover, a high proportion of disease resistance-related genes were upregulated in wild-grown ginseng. This study is the first transcriptome analysis to compare wild-grown and cultivated ginseng, and identifies genes that may produce higher ginsenoside content and better disease resistance in the wild; these genes may have the potential to improve cultivated ginseng grown in artificial environments.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Baeg IH, So SH. The world ginseng market and the ginseng (Korea). J Ginseng Res, 2013, 37: 1–7
Goldstein B. Ginseng: its history, dispersion, and folk tradition. Am J Chin Med (Gard City N Y), 1975, 3: 223–234
Yun TK. Panax ginseng—a non-organ-specific cancer preventive? Lancet Oncol, 2001, 2: 49–55
Rausch WD, Liu S, Gille G, Radad K. Neuroprotective effects of ginsenosides. Acta Neurobiol Exp (Wars), 2006, 66: 369–375
Choi JS, Chun KS, Kundu J, Kundu JK. Biochemical basis of cancer chemoprevention and/or chemotherapy with ginsenosides (Review). Int J Mol Med, 2013, 32: 1227–1238
Lu JM, Yao Q, Chen C. Ginseng compounds: an update on their molecular mechanisms and medical applications. Curr Vasc Pharmacol, 2009, 7: 293–302
Karmazyn M, Moey M, Gan XT. Therapeutic potential of ginseng in the management of cardiovascular disorders. Drugs, 2011, 71: 1989–2008
Gu J, Li W, Xiao D, Wei S, Cui W, Chen W, Hu Y, Bi X, Kim Y, Li J, Du H, Zhang M, Chen L. Compound K, a final intestinal metabolite of ginsenosides, enhances insulin secretion in MIN6 pancreatic beta-cells by upregulation of GLUT2. Fitoterapia, 2013, 87: 84–88
Xu Y, Lin L, Tang L, Zheng M, Ma Y, Huang L, Meng W, Wang W. Notoginsenoside R1 attenuates hypoxia and hypercapnia-induced vasoconstriction in isolated rat pulmonary arterial rings by reducing the expression of ERK. Am J Chin Med, 2014, 42: 799–816
Lee KS, Kim GH, Kim HH, Chang YI, Lee GH. Volatile compounds of Panax ginseng C.A. Meyer cultured with different cultivation methods. J Food Sci, 2012, 77: C805–810
Kwon KR, Park WP, Kang WM, Jeon EY, Jang JH. Identification and analysis of differentially expressed genes in mountain cultivated ginseng and mountain wild ginseng. J Acupunct Meridian Stud, 2011, 4: 123–128
Xu G, Fan X, Miller AJ. Plant nitrogen assimilation and use efficiency. Annu Rev Plant Biol, 2012, 63: 153–182
Sun C, Li Y, Wu Q, Luo H, Sun Y, Song J, Lui EM, Chen S. De novo sequencing and analysis of the American ginseng root transcriptome using a GSFLX Titanium platform to discover putative genes involved in ginsenoside biosynthesis. BMC Genomics, 2010, 11: 262
Chen S, Luo H, Li Y, Sun Y, Wu Q, Niu Y, Song J, Lv A, Zhu Y, Sun C, Steinmetz A, Qian Z. 454 EST analysis detects genes putatively involved in ginsenoside biosynthesis in Panax ginseng. Plant Cell Rep, 2011, 30: 1593–1601
Luo H, Sun C, Sun Y, Wu Q, Li Y, Song J, Niu Y, Cheng X, Xu H, Li C, Liu J, Steinmetz A, Chen S. Analysis of the transcriptome of Panax notoginseng root uncovers putative triterpene saponin-biosynthetic genes and genetic markers. BMC Genomics, 2011, 12 Suppl 5: S5
Li C, Zhu Y, Guo X, Sun C, Luo H, Song J, Li Y, Wang L, Qian J, Chen S. Transcriptome analysis reveals ginsenosides biosynthetic genes, microRNAs and simple sequence repeats in Panax ginseng C. A. Meyer. BMC Genomics, 2013, 14: 245
Wu D, Austin RS, Zhou S, Brown D. The root transcriptome for North American ginseng assembled and profiled across seasonal development. BMC Genomics, 2013, 14: 564
Subramaniyam S, Mathiyalagan R, Natarajan S, Kim YJ, Jang MG, Park JH, Yang DC. Transcript expression profiling for adventitious roots of Panax ginseng Meyer. Gene, 2014, 546: 89–96
Jayakodi M, Lee SC, Park HS, Jang W, Lee YS, Choi BS, Nah GJ, Kim DS, Natesan S, Sun C, Yang TJ. Transcriptome profiling and comparative analysis of Panax ginseng adventitious roots. J Ginseng Res, 2014, 38: 278–288
Jayakodi M, Lee SC, Lee YS, Park HS, Kim NH, Jang W, Lee HO, Joh HJ, Yang TJ. Comprehensive analysis of Panax ginseng root transcriptomes. BMC Plant Biol, 2015, 15: 138
Liu MH, Yang BR, Cheung WF, Yang KY, Zhou HF, Kwok JS, Liu GC, Li XF, Zhong S, Lee SM, Tsui SK. Transcriptome analysis of leaves, roots and flowers of Panax notoginseng identifies genes involved in ginsenoside and alkaloid biosynthesis. BMC Genomics, 2015, 16: 265
Grabherr MG, Haas BJ, Yassour M, Levin JZ, Thompson DA, Amit I, Adiconis X, Fan L, Raychowdhury R, Zeng Q, Chen Z, Mauceli E, Hacohen N, Gnirke A, Rhind N, di Palma F, Birren BW, Nusbaum C, Lindblad-Toh K, Friedman N, Regev A. Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat Biotechnol, 2011, 29: 644–652
Huang Y, Niu B, Gao Y, Fu L, Li W. CD-HIT Suite: a web server for clustering and comparing biological sequences. Bioinformatics, 2010, 26: 680–682
Langmead B, Salzberg SL. Fast gapped-read alignment with Bowtie 2. Nat Methods, 2012, 9: 357–359
Moriya Y, Itoh M, Okuda S, Yoshizawa AC, Kanehisa M. KAAS: an automatic genome annotation and pathway reconstruction server. Nucleic Acids Res, 2007, 35: W182–185
Haas BJ, Papanicolaou A, Yassour M, Grabherr M, Blood PD, Bowden J, Couger MB, Eccles D, Li B, Lieber M, Macmanes MD, Ott M, Orvis J, Pochet N, Strozzi F, Weeks N, Westerman R, William T, Dewey CN, Henschel R, Leduc RD, Friedman N, Regev A. De novo transcript sequence reconstruction from RNA-seq using the Trinity platform for reference generation and analysis. Nat Protoc, 2013, 8: 1494–1512
Eddy SR. Accelerated Profile HMM Searches. PLoS Comput Biol, 2011, 7: e1002195
Du Z, Zhou X, Ling Y, Zhang Z, Su Z. agriGO: a GO analysis toolkit for the agricultural community. Nucleic Acids Res, 2010, 38: W64–70
Sanseverino W, Hermoso A, D'Alessandro R, Vlasova A, Andolfo G, Frusciante L, Lowy E, Roma G, Ercolano MR. PRGdb 2.0: towards a community-based database model for the analysis of R-genes in plants. Nucleic Acids Res, 2013, 41: D1167–1171
Han JY, Kim HJ, Kwon YS, Choi YE. The Cyt P450 enzyme CYP716A47 catalyzes the formation of protopanaxadiol from dammarenediol-II during ginsenoside biosynthesis in Panax ginseng. Plant Cell Physiol, 2011, 52: 2062–2073
Davidson NM, Oshlack A. Corset: enabling differential gene expression analysis for de novo assembled transcriptomes. Genome Biol, 2014, 15: 410
Gonzalez E, Joly S. Impact of RNA-seq attributes on false positive rates in differential expression analysis of de novo assembled transcriptomes. BMC Res Notes, 2013, 6: 503
Roberts A, Pachter L. Streaming fragment assignment for real-time analysis of sequencing experiments. Nat Methods, 2013, 10: 71–73
Anders S, Huber W. Differential expression analysis for sequence count data. Genome Biol, 2010, 11: R106
Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods, 2001, 25: 402–408
Zhang CY, Dong L, Chen SL, Xie CX, Chang DL. UPLC fingerprint for quality assessment of ginsenosides of ginseng radix et rhizoma (in chinese). Yao Xue Xue Bao, 2010, 45: 1296–1300
Barkan A, Small I. Pentatricopeptide repeat proteins in plants. Annu Rev Plant Biol, 2014, 65: 415–442
Kobe B, Kajava AV. The leucine-rich repeat as a protein recognition motif. Curr Opin Struct Biol, 2001, 11: 725–732
Li D, Roberts R. WD-repeat proteins: structure characteristics, biological function, and their involvement in human diseases. Cell Mol Life Sci, 2001, 58: 2085–2097
Jung SC, Kim W, Park SC, Jeong J, Park MK, Lim S, Lee Y, Im WT, Lee JH, Choi G, Kim SC. Two Ginseng UDP-Glycosyltransferases Synthesize Ginsenoside Rg3 and Rd. Plant Cell Physiol, 2014, 55: 2177–2188
Ellis J, Dodds P, Pryor T. Structure, function and evolution of plant disease resistance genes. Curr Opin Plant Biol, 2000, 3: 278–284
Sanseverino W, Ercolano MR. In silico approach to predict candidate R proteins and to define their domain architecture. BMC Res Notes, 2012, 5: 678
Belkhadir Y, Subramaniam R, Dangl JL. Plant disease resistance protein signaling: NBS-LRR proteins and their partners. Curr Opin Plant Biol, 2004, 7: 391–399
Author information
Authors and Affiliations
Corresponding authors
Additional information
Contributed equally to this work
This article is published with open access at springerlink.bibliotecabuap.elogim.com
Deng XingWang is a university endowed professor of plant biology at Peking University. He graduated from Peking University with B.S. (1982) and M.S. (1982) degrees, and then graduated from University of California at Berkeley in 1989 with Ph.D. degree in plant biology. Before moving back to China, he was a faculty of Yale since 1992, and promoted to Daniel C. Eaton Professor of Yale University from 2003 till 2014. He worked on signaling process in plant photomorphogenesis, noncoding RNAs, heterosis and molecular design breeding in plant. He has been awarded the Kumho Science International Award by the International Society for Plant Molecular Biology (ISPMB) in 2003 and elected Member of US National Academy of Sciences in 2013.
Electronic supplementary material
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (https://creativecommons.org/licenses/by/4.0), which permits use, duplication, adaptation, distribution, and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Zhen, G., Zhang, L., Du, Y. et al. De novo assembly and comparative analysis of root transcriptomes from different varieties of Panax ginseng C. A. Meyer grown in different environments. Sci. China Life Sci. 58, 1099–1110 (2015). https://doi.org/10.1007/s11427-015-4961-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11427-015-4961-x