Abstract
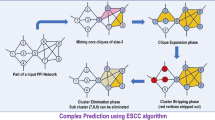
Proteins usually bind together to form complexes, which play an important role in cellular activities. Many graph clustering methods have been proposed to identify protein complexes by finding dense regions in protein-protein interaction networks. We present a novel framework (CPL) that detects protein complexes by propagating labels through interactions in a network, in which labels denote complex identifiers. With proper propagation in CPL, proteins in the same complex will be assigned with the same labels. CPL does not make any strong assumptions about the topological structures of the complexes, as in previous methods. The CPL algorithm is tested on several publicly available yeast protein-protein interaction networks and compared with several state-of-the-art methods. The results suggest that CPL performs better than the existing methods. An analysis of the functional homogeneity based on a gene ontology analysis shows that the detected complexes of CPL are highly biologically relevant.
Article PDF
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
References
Spirin V, Mirny L A. Protein complexes and functional modules in molecular networks. Proceedings of the National Academy of Sciences, 2003, 100(21): 12123–12128.
Chen B, Fan W, Liu J et al. Identifying protein complexes and functional modules | From static PPI networks to dynamic PPI networks. Briefings in Bioinformatics, 2014, 15(2): 177–194.
Geva G, Sharan R. Identification of protein complexes from co-immuno-precipitation data. Bioinformatics, 2011, 27(1): 111–117.
Ji J, Zhang A, Liu C et al. Survey: Functional module detection from protein-protein interaction networks. IEEE Knowledge and Data Engineering, 2014, 26(2): 261–277.
Li X, Wu M, Kwoh C K et al. Computational approaches for detecting protein complexes from protein interaction networks: A survey. BMC Genomics, 2010, 11(Suppl 1): S3.
Wang J, Li M, Deng Y et al. Recent advances in clustering methods for protein interaction networks. BMC Genomics, 2010, 11(Suppl 3): S10.
Nepusz T, Yu H, Paccanaro A. Detecting overlapping protein complexes in protein-protein interaction networks. Nature Methods, 2012, 9(5): 471–472.
Becker E, Robisson B, Chapple C E et al. Multifunctional proteins revealed by overlapping clustering in protein interaction network. Bioinformatics, 2012, 28(1): 84–90.
Chen B, Shi J, Zhang S et al. Identifying protein complexes in protein-protein interaction networks by using clique seeds and graph entropy. Proteomics, 2013, 13(2): 269–277.
Habibi M, Eslahchi C, Wong L. Protein complex prediction based on k-connected subgraphs in protein interaction network. BMC Systems Biology, 2010, 4(1): 129.
Zhang C, Liu S, Zhou Y. Fast and accurate method for identifying high-quality protein-interaction modules by clique merging and its application to yeast. Journal of Proteome Research, 2006, 5(4): 801–807.
Altaf-Ul-Amin M, Shinbo Y, Mihara K et al. Development and implementation of an algorithm for detection of protein complexes in large interaction networks. BMC Bioinformatics, 2006, 7(1): 207.
Bader G D, Hogue C W V. An automated method for finding molecular complexes in large protein interaction networks. BMC Bioinformatics, 2003, 4(1): 2.
Wu M, Li X, Kwoh C K et al. A core-attachment based method to detect protein complexes in PPI networks. BMC Bioinformatics, 2009, 10(1): 169.
Adamcsek B, Palla G, Farkas I J et al. CFinder: Locating cliques and overlapping modules in biological networks. Bioinformatics, 2006, 22(8): 1021–1023.
Palla G, Derényi I, Farkas I, Vicsek T. Uncovering the overlapping community structure of complex networks in nature and society. Nature, 2005, 435(7043): 814–818.
Wang S, Wu F. Detecting overlapping protein complexes in PPI networks based on robustness. Proteome Science, 2013, 11(Suppl 1): S18.
Enright A, Van Dongen S, Ouzounis C. An e±cient algorithm for large-scale detection of protein families. Nucleic Acids Research, 2002, 30(7): 1575–1584.
Pizzuti C, Rombo S. A co-clustering approach for mining largeprotein-protein interaction networks. IEEE/ACM Transactions on Computational Biology and Bioinformatics, 2012, 9(3): 717–730.
Anirban M, Sumanta R, Moumita D. Detecting protein complexes in a PPI network: A gene ontology based multi-objective evolutionary approach. Molecular BioSystems, 2012, 8(11): 3036–3048.
Eileen M H. Detection of overlapping protein complexes using a protein ranking algorithm. In Proc. the 9th Int. Conference on Innovations in Information Technology, March 2013, pp.233–236.
Zaki N, Berengueres J, Efimov D. Detection of protein complexes using a protein ranking algorithm. Proteins: Structure, Function, and Bioinformatics, 2012, 80(10): 2459–2468.
Wang Y, Qian X. Functional module identification in protein interaction networks by interaction patterns. Bioinformatics, 2014, 30(1): 81–93.
Raghavan U N, Albert R, Kumara S. Near linear time algorithm to detect community structures in large-scale networks. Physical Review E, 2007, 76(3): 036106.
Stark C, Breitkreutz B J, Reguly T et al. BioGRID: A general repository for interaction datasets. Nucleic Acids Research, 2006, 34(Suppl 1): D535–D539.
Salwinski L, Miller C S, Smith A J et al. The database of interacting proteins: 2004 update. Nucleic Acids Research, 2004, 32(Database Issue): D449-D451.
Mewes H W, Amid C, Arnold R et al. MIPS: Analysis and annotation of proteins from whole genomes. Nucleic Acids Research, 2004, 32(Database Issue): D41-D44.
Pu S, Wong J, Turner B et al. Up-to-date catalogues of yeast protein complexes. Nucleic Acids Research, 2009, 37(3): 825–831.
Wu Z H, Lin Y F, Gregory S et al. Balanced multi-label propagation for overlapping community detection in social networks. Journal of Computer Science and Technology, 2012, 27(3): 468–479.
Hong E L, Balakrishnan R, Dong Q et al. Gene ontology annotations at SGD: New data sources and annotation methods. Nucleic Acids Research, 2008, 36(Suppl 1): D577–D581.
Boyle E I, Weng S, Gollub J et al. GO::TermFinder — Open source software for accessing gene ontology information and finding significantly enriched gene ontology terms associated with a list of genes. Bioinformatics, 2004, 20(18): 3710–3715.
Berman H M, Westbrook J, Feng Z et al. The protein data bank. Nucleic Acids Research, 2000, 28(1): 235–242.
Naveed H, Han J J. Structure-based protein-protein interaction networks and drug design. Quantitative Biology, 2013, 1(3): 183–191.
Zhang Q C, Petrey D, Deng L et al. Structure-based prediction of protein-protein interactions on a genome-wide scale. Nature, 2012, 490(7421): 556–560.
Author information
Authors and Affiliations
Corresponding author
Additional information
The work was supported by the National Natural Science Foundation of China under Grant Nos. 61271346, 61172098, and 91335112, the Specialized Research Fund for the Doctoral Program of Higher Education of China under Grant No. 20112302110040, and the Fundamental Research Funds for the Central Universities of China under Grant No. HIT.KISTP.201418.
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(PDF 91 kb)
Rights and permissions
About this article
Cite this article
Dai, QG., Guo, MZ., Liu, XY. et al. CPL: Detecting Protein Complexes by Propagating Labels on Protein-Protein Interaction Network. J. Comput. Sci. Technol. 29, 1083–1093 (2014). https://doi.org/10.1007/s11390-014-1492-z
Received:
Revised:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11390-014-1492-z