Abstract
Plants can stimulate the microbes to degrade ubiquitous petroleum hydrocarbons (PHCs), which has prompted a novel view on rhizoremediation. In the present study, the degradation rate of PHCs was investigated and 16S rRNA gene analysis was performed to investigate the PHC-degrading bacteria in petroleum-contaminated soil with different plants. Mirabilis jalapa (M. jalapa) has a higher PHC degradation rate than Lolium perenne (L. perenne) under petroleum contamination. The bacterial diversity in rhizospheric soil was decreased but the relative abundance of Actinobacteriota, Proteobacteria, and Candidatus Saccharibacteria were significant increased on 45 days petroleum-contaminated rhizospheric soil. In addition, the relative expression of PHC degradation–related genes, the content of malic acid and citric acid of the root exudates in the two plants was significantly increased in response to petroleum stress. The content of citric acid increased 11.9 times in M. jalapa and 3.4 times in L. perenne, respectively, in response to petroleum stress. These results indicate that M. jalapa changes the hydrocarbon-degrading microbial community to enhance the degradation of PHCs by root exudates and phytostimulation.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
The exploration, transportation, and processing of oil lead to serious soil contamination and environmental problems over the last decades, especially the accumulation of petroleum and petroleum byproducts in soils (Brakorenko and Korotchenko 2016; Mahmud et al. 2022; Mohanta, et al. 2024). Petroleum poses a potential threat to the soil structure, groundwater quality, ecosystems, and health of humans (Ramadass et al. 2015; Varjani 2017). In particular, polycyclic aromatic hydrocarbons (PAHs) and some components of petroleum, including benzene, toluene, ethylbenzene, and xylene (BTEX), are relatively recalcitrant, highly toxic, carcinogenic, and teratogenic (Zhang et al. 2011; Costa et al. 2012; Meckenstock et al. 2016). Petroleum hydrocarbons (PHCs) are known to be toxic to many organisms and are also considered common persistent organic pollutants in the environment. PHCs alter the soil structure, decline soil porosity, increase soil pH, and also change the soil’s electrochemical properties (Orozco et al. 2021; Pan et al. 2021). PHCs also inhibit soil enzyme activity, the normal respiration of roots, and the absorption of nutrients, which adversely affect soil fertility and plant growth (Mohanta et al. 2024).
Phytoremediation has become a main method for soil remediation of crude oil–contaminated soil. Phytoremediation uses living organisms such as plant, bacteria, or both, to enhance and stimulate the remediation of crude oil–contaminated soil. Unlike the phytoremediation of heavy metals through phytoextraction or accumulation of above-ground contaminant, rhizospherical degradation is the primary mechanism driving the phytoremediation of crude oil-contaminated soils (Liu et al. 2015; Agnello et al. 2016; Correa-García et al. 2018; Lü et al. 2024). Rhizosphere microorganisms, especially bacteria, are generally considered to be the main participants in the degradation of organic contaminants during rhizoremediation (El Amrani et al. 2015; Bhuyan et al. 2023).
Rhizospherical degradation of PHC-contaminated soil is the process in which PHCs are decomposed and metabolized by plant rhizosphere microorganisms (Hussain et al. 2018). Burkholderia, Pseudomonas, Mycobacterium, and Rhodococcus are believed to contribute to remediation in petroleum-contaminated areas, owing to their stronger aerobic metabolism of organic pollutants (Wei et al. 2017; Xia et al. 2020). In addition, plant roots can enhance the microbial activity 10–100 times in the rhizosphere soil through the release of exudates containing compounds such as amino acids, organic acids, and carbohydrates (Weller and Thomashow 2007). Root exudation stimulate microbial growth, which affects PHC-degrading bacteria diversity, particularly those exudations that induce PAH-degrading metabolic reactions (Liu et al. 2015; Rohrbacher and Starnaud 2016). Artificial root exudate mixtures have enhanced the degradation rates of three- to five-ring PAHs by increasing bacterial densities and shifting metabolic profiles (Gao et al. 2010; Guo et al. 2017a). However, the research focus on the specific types of root exudates for PHCs dissipation is still limited.
In the past decades, numerous plant species have been tested to determine their abilities to remove PHCs from contaminated soil. Alfalfa (Medicago sativa), ryegrass (Lolium multiflorum), tall fescue (Festuca arundinacea), vetiver grass (Vetiveria zizanioides L.), birdsfoot trefoil (Lotus corniculatusvar), and sorghum (Sorghum bicolor) (Kaimi et al. 2007; Liu et al. 2015; Rajaei and Seyedi 2018; Umeh et al. 2018; Yousaf et al. 2010) have been used for the removal of PHCs from soil. These plants are fast-growing species that develop a strong root system favorable for establishing rhizosphere-associated microorganisms to decontaminate polluted sites. M. jalapa was a petroleum tolerant pants which can effectively enhance the dissipation of soil pollutants (Cen et al. 2016; Peng et al. 2009). Our previous studies have demonstrated that M. jalapa has a higher capacity to resist and remediate the crude oil–contaminated soil (Ma et al. 2018). In order to further illustrate the mechanism of M. jalapa to facilitate phytoremediation of petroleum-contaminated soil, the rhizosphere microbial community abundance differences between M. jalapa and L. perenne were compared, as well as the root exudates were analyzed in the present study.
Materials and methods
Soil preparation
The petroleum used in the present study was obtained from Fushun oil field (Fushun, Liaoning Province, China, 123.97E, 41.97N). Clean soil collected from the greenhouse in Shenyang Agricultural University (Liaoning province, China, 123.38E, 41.80N) was air-dried and sieved through a 4-mm sieve to ensure soil homogeneity. The physicochemical properties of clean soil are as follows: pH 5.98, 2.87% organic material content, 263.6 mg/kg total nitrogen content, 91.3 mg/kg total phosphorus content, and a total potassium content of 611.0 mg/kg. For the highest petroleum-concentrated soil, 20 g of crude petroleum was dissolved in 50 mL of diethyl ether. Then the solution was mixed fully with 1 kg clean soil and air-dried for 24 h. The clean soil was fully mixed with the highest petroleum-concentrated soil in different ratios to form the discrete concentration of petroleum (0 g, 5 g, 10 g, 15 g and 20 g petroleum per kilogram soil).
Experimental design
For pot experiments, 10 cm × 12 cm (diameter × height) containers with 1 kg of soil were used. All experiments were with three replicates. Each pot contained one of the petroleum-contaminated soil with 5, 10, 15, and 20 g/kg, respectively. The pot containing 1 kg of clean soil was used as control. All of the pots were placed in a growth chamber (25 °C, 16-h light and 8-h dark), and watered to maintain a soil moisture rate of above 60%. M. jalapa and L. perenne seedlings (the three-leaf stage) were transplanted into pots containing 1 kg of soil, respectively. To ensure that the two plants with similar root biomass in each pot, each pot transplanted one M. jalapa or ten L. perenne, respectively.
Soil samples were collected at 15, 30, and 45 days after transplantation. Rhizosphere soil (adhere to root surface 1 ~ 2 mm) was collected carefully as follows. The entire plant root system was dug out and patted to loosen all the loose soil clumps. Then the soil attached to the root surface was collected with a sterile brush. Soil from plant-free pot was collected at a 5-cm depth, served as the plant-free rhizosphere soil samples. Soil sample was sieved to remove the root system, plant residues, and other impurities. Each soil sample was stored at − 80 °C till analyzed. For PHC analysis, rhizosphere soil was collected from five different points in each pot.
PHC and PAH determination
PHCs in soil were determined gravimetrically as described in Peng et al. (2009). Briefly, 5.0 g petroleum-contaminated soils had been pre-sieved through an 80 mesh and transferred into a 50-mL glass centrifuge tube for PHC extraction. Then 25 mL chloroform was added and covered with the lid. After ultrasonically extracting for 1 h, the tubes were centrifuged for 10 min at 3000 r/min. The extracts were transferred into a clean conical flask and dried to a constant weight. After the complete removal of the solvent by evaporation, the amount of residual PHCs was determined gravimetrically. Finally, the degradation rates of PHCs were calculated as the degradation rate (%) = (C0-C1)/C0 × 100%. C0 was initial petroleum hydrocarbon content, and C1 was residual petroleum hydrocarbon content after degradation.
Polycyclic aromatic hydrocarbons (PAHs) were determined by gas chromatography equipped with a mass spectrometry detector (GC–MS) as published method (Kao et al. 2015). Briefly, 2.0 g pre-sieved by 80 mesh petroleum-contaminated soils was mixed with 10 mL of methylene chloride/acetone (vol/vol = 1:1). After ultrasonically extracting for 20 min, 300 g of anhydrous sodium sulfate was added for dehydration. Then the supernatant was pre-concentrated to 1 mL by a rotary evaporator and solvent exchanged to hexane. The hexane extract was fractionated and cleaned up by DB-5MS column (30 m × 0.25 mm I.D. × 0.25 μm film thickness). The column was eluted with methylene chloride/hexane (vol/vol = 3:7) to obtain PAH. The PAH fraction was finally concentrated to 1 mL under a gentle stream of nitrogen. The concentrations of PAH in the extracts were determined by gas chromatograph-mass spectrometry (Shimadzu 17001029S).
DNA extraction and sequencing
To further analyze the higher PHC degradation by M. jalapa planting soils, the diversity of the microbial community in rhizosphere soil of M. jalapa was investigated. The DNA was extracted from 45 days petroleum-contaminated rhizosphere soils using an Ezup Column Soil DNA Purification Kit (B518263, Sangon Biotech, Shanghai, China) following the manufacturer’s instructions. The hypervariable region 4 (V4) of the 16S rRNA gene was amplified as described previously (Caporaso et al. 2012) with 520F and 802R as forward and reverse primers, respectively (Supplementary Table S1). Amplicons were sequenced using Illumina MiSeq by Shanghai Personalbio Biotechnology, Co., Ltd. (Shanghai, China). The raw sequences were processed as described previously (Wu et al. 2021). The Operational Taxonomic Unit (OTU) matrix was established through a multivariate analysis to further explore the bacterial community variations between the different treatments. Taxonomic abundance was presented in the phylum, class, order, family, and genus. The abundance-based coverage estimator (ACE) and Chao estimator were calculated, and then, the rarefaction curves were plotted using Qiime (version1.7.0, http://qiime.org/). The Shannon and Simpson indexes were calculated to assess microbial community diversity.
Quantification of genes related to PHC degradation
RNA was extracted from 45 days petroleum-contaminated rhizosphere soils using the EZNA Soil RNA Kit (R6825-00, Omega Bio-Tek, USA) following the manufacturer’s instructions. The quantitative PCR (qPCR) was carried out using a FastKing RT Kit with gDNase (KR116, Tiangen, Beijing, China) and SuperReal PreMix Plus (SYBR Green) (FP205, Tiangen, Beijing, China) according to the manufacturer’s protocols. Primer sequences of microbial genes related to PHC degradation, alkB and nahAc, are presented in supplementary Table S1.
Root exudate analysis
After 45 days of petroleum treatment at 10 g/kg, the plants were selected for the exudate collection. Root exudate collection was according to the method described by Huang et al. (2016).
Statistical analysis
Significant differences in alpha diversity in petroleum-contaminated soil that influence by plants were obtained by the Wilcoxon rank sum test. One-way analysis of variance for independent samples was performed using the R software package, version 3.0.2. Differences were considered statistically significant at p < 0.05. All the values were presented as means ± standard deviation (S.D.) of three replicates.
Results
Phytoremediation efficiency of PHCs
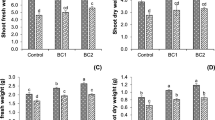
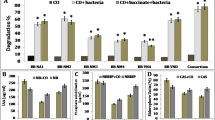
Petroleum stress inhibited the growth of M. jalapa and L. perenne. With the increase of petroleum concentration, the growth inhibition is more serious in both plants (Supplementary Figure S1). Both M. jalapa and L. perenne can promote the degradation rates of PHCs in petroleum-contaminated soil. The degradation rate of PHCs increased during the plant growth (Fig. 1). Petroleum-contaminated soil with M. jalapa has a higher degradation rate of PHCs than the contaminated soil with L. perenne (Fig. 1). The degradation rate of PHCs by M. jalapa was up to 89.1% under the 10 g/kg of petroleum-contaminated soil at 45 days (Fig. 1B). After 45 days, the degradation rate of two-ring PAH (naphthalene) in M. jalapa and L. perenne planted petroleum-contaminated soil was 71.5% and 48.2%, respectively, while the degradation rates of naphthalene in plant-free petroleum-contaminated soil was only 18.5% (Fig. 2). The degradation rates of three-ring PAH (acenaphthylene, acenaphthene, fluorene, and anthracene) in plant-free petroleum-contaminated soil were 30.9%, 41.5%, 22.4%, and 57.7%, respectively, while the degradation rates of these three-ring PAH in petroleum-contaminated soil with M. jalapa were 73.4%, 81.0%, 72.3%, and 88.1%, respectively (Fig. 2). And the degradation rates of these three-ring PAH in petroleum-contaminated soil with L. perenne were 59.0%, 73.5%, 53.0%, and 87.8%, respectively (Fig. 2). These results indicated that both M. jalapa and L. perenne could significantly promote the degradation of two-ring and three-ring PAHs. The degradation of four-ring PAHs (pyrene, fluoranthene, and benzanthracene) was increased significantly by M. jalapa in petroleum-contaminated soil, but only benzanthracene was significantly degraded by L. perenne (Fig. 2). The degradation rate of five-ring PAHs (benzofluoranthrene) was not affected by plants (Fig. 2). These results indicated that plants significantly enhanced the degradation of PAHs, and M. jalapa had a higher degradation rate of PAHs than L. perenne under 10 g/kg petroleum-contaminated soil.
Quantification of degradation genes for PHCs
In order to clarify whether plants increase the degradation of PHC in petroleum-contaminated soil by increasing the expression of gene related to PHC degradation, the relative expressions of two genes, alkB and nahAc, were investigated. Both alkB and nahAc were significantly upregulated under M. jalapa and L. perenne planted rhizosphere soil for 45 days (Fig. 3). However, the expression of alkB and nahAc in rhizosphere soil showed no significant difference between M. jalapa and L. perenne (Fig. 3). These results indicated that both M. jalapa and L. perenne enhanced the expression of microbial genes, alkB and nahAc, to promote PHC degradation.
Diversity of the microbial community
Based on the results of sequencing, the alpha-diversity indexes (Chao, ACE, Simpson and Shannon) were calculated to evaluate the diversity. The microbial alpha-diversity indexes (Chao, ACE, Simpson and Shannon) of rhizospheric soils were significantly decreased under petroleum stress for 45 days, no matter planted or not (Table 1). These results indicated that petroleum reduced microbial diversity (Simpson and Shannon) and abundance (Chao, ACE) in rhizospheric soils.
OTU matrix was used for multivariate analysis to further explore the bacterial community variations. The high coverage estimator (coverage index > 97%) indicates that the bacterial OTUs in each soil sample were well captured. After 45 days, 39 bacterial phyla were identified in the petroleum-contaminated rhizospheric soils. The ten bacterial phyla (with relative abundance > 1%) accounted for 94% of the all identified bacterial phyla (Fig. 4). Petroleum contamination increased the relative abundance of Proteobacteria and Candidatus Saccharibacteria. In rhizospheric soil of M. jalapa, Proteobacteria and Candidatus Saccharibacteria increased from 31.17 to 49.57%, from 3.30 to 17.70%, respectively. In rhizospheric soil of L. perenne, Proteobacteria and Candidatus Saccharibacteria increased from 29.07 to 45.60%, from 3.30 to 17.70%, respectively (Fig. 4). However, Chloroflexi, Gemmatimonadetes, Acidobacteria, and Bacteroidetes decreased in abundance under petroleum contamination, no matter planted or not (Fig. 4).
The relative abundance of the top 50 classified bacterial genera in rhizospheric soil was compared. The top 50 genera were distributed into four major clusters (Supplementary Figure S2). The bacterial genera increasing in abundance under petroleum-contaminated soil which planted M. jalapa and L. perenne belong to cluster I. In cluster II, most of the bacterial genera increasing in abundance under petroleum-contaminated soil when M. jalapa was planted (Supplementary Figure S2).
Analysis of organic acids and related factors
To clarify whether the different root exudates cause the microbial distribution difference in rhizospheric soils, the low-molecular-weight organic acids in the root exudates of the rhizospheric soils of M. jalapa and L. perenne were investigated. Under petroleum contamination, the malic acid and citric acid were significantly increased both in M. jalapa and in L. perenne root exudates, especially the citric acid in M. jalapa (Fig. 5). Petroleum contamination increased the content of malic acid in root exudates of L. perenne. However, more citric acid was observed in root exudates of M. jalapa in response to petroleum contamination (Fig. 5). The oxalic acid of root exudates in M. jalapa was not affect by petroleum stress. However, the oxalic acid of root exudates in L. perenne was significantly decreased under petroleum stress (Fig. 5). The differences in Candidatus Saccharibacteria, Proteobacteria, and Actinomycetes also correlated with the differences in organic acids, root lengths, and degradation rates in L. perenne (Fig. 6B). Generally, Candidatus Saccharibacteria and Proteobacteria exhibited the strongest correlations with root lengths, oxalic acid, malic acid, and citric acid. We also correlated the differences in Candidatus Saccharibacteria, Proteobacteria, and Actinomycetes with the differences in organic acids and degradation rates in M. jalapa (Fig. 6A). These results indicated that citric acid and malic acid exhibited the strongest correlations with Actinobacteria, Proteobacteria, and Candidatus Saccharibacteria in M. jalapa.
Discussion
Several species have been utilized in previous studies to remove organic pollutants from contaminated soil with great success, such as Fire Phoenix (Dai et al. 2020), Festuca arundinacea L. (Liu et al. 2015), Hibiscus cannabinus (Saratale et al. 2019), and L. perenne (a plant commonly used in phytoremediation studies), which has exhibited a prominent degrading effect on PHC-contaminated soil (Guo et al. 2017b; Iqbal et al. 2019; Meng et al. 2011). Our previous studies showed that the growth of M. jalapa in soil containing 10 g/kg of petroleum was only slightly repressed (Ma et al. 2018). The growth of M. jalapa in 10 g/kg of petroleum-contaminated soil for 45 days was not significant as compared with the control. However, the soil containing 15 g and 20 g petroleum significant decreased the plant height of M. jalapa and L. perenne for 45 days (Supplementary Figure S1). After cultivation in petroleum-polluted soil for 45 days, the degradation rate determined from the PHC concentrations in the rhizosphere soil of M. jalapa was significantly higher than that of L. perenne (Fig. 1). These results indicated that M. jalapa has stronger PHC removal capacity than L. perenne.
In the present study, petroleum contamination significantly decreased microbial diversity of rhizospheric soils (Table 1). However, Proteobacteria and Candidatus Saccharibacteria were the predominant bacteria which significantly increased in the petroleum-contaminated rhizospheric soils, especially the M. jalapa planted hizospheric soil (Fig. 4). Proteobacteria are a group of gram-negative bacteria that have been widely reported to be able to degrade PHCs (Hou et al. 2015). Most studies have demonstrated that higher PHC degradation mainly depend on the dominance of practical bacteria rather than the soil microbial diversity (Cébron et al. 2011; Guo et al. 2017a; Phillips et al. 2012; Wu et al. 2016). In essence, it is more important to selectively stimulate the specific petroleum-degrading bacteria than to stimulate the diversity of the entire microbial community (Bell et al. 2013). Alcanivorax, Hyphomicrobium, Proshecobacter, Opitutus, and Shewanella were dominant bacteria at genera level under petroleum contamination soil planting M. jalapa, while these bacteria were less dominant in the control group without petroleum contamination and the contaminated soil planting L. perenne (Supplementary Figure S2). These results indicated that a unique PHC-degrading bacterial community were induced in petroleum-polluted soil planting M. jalapa. Alcanivorax can degrade alkanes and can also utilize PHC components as a carbon source in polluted surroundings (Singh and Sedhuraman 2015; Zadjelovic et al. 2020). In this study, Mycobacterium and Phenylobacterium were the top 50 classified bacterial genera of the rhizospheric soil that belong to cluster I (Supplementary Figure S2). Both bacteria have been considered promoting PAH degradation (Lu et al. 2019). Flavobacterium (belong to cluster III) is related to denitrification and nitrogen fixation, and plays important roles in the cycling of carbon, nitrogen, and other nutrients in the soil (Ueki et al. 2010; Zhang et al. 2020). Pseudomonas (belong to cluster I) has been shown to promote the growth of roots and their aerial parts, protect plants from phenanthrene toxicity, and manipulate the antioxidant stress defense system of host plants to help the plants resist oxidative stresses caused by organic pollutants (Liu et al. 2015). Root exudate can induce the Actinobacteriota to increase the degradation of phenanthrene (Panchenko et al. 2022). Actinobacteria is also the principal phyla in bioremediation of hydrocarbon-polluted soils (Ros et al. 2010). Plants promote the enrichment of these microorganisms in petroleum-contaminated soil and accelerates the degradation of PHC pollutants in the soil. Therefore, it is feasible to apply a cooperative framework based on the dominant degrading bacteria and host plants to remediate soil polluted by oil at moderate concentrations.
The relative expression of alkB and nahAc increased significantly in the transplanted soil of M. jalapa and L. perenne (p < 0.05) (Fig. 3). This is consistent with the previous studies that soil amended with petroleum hydrocarbons had a higher abundance of catabolic genotypes and metabolic genes than uncontaminated soils (Varjani 2017). Naphthalene dioxygenases encoded by the nah gene can also metabolize broad substrates such as PAHs with few rings (Guo et al. 2017b). In an experiment on the degradation of hydrocarbons by L. perenne, Pseudomonas ITRI53 demonstrated a higher alkB gene expression, which indicates that there was a strong positive correlation between the abundance of the metabolizing genes and hydrocarbon degradation (Thomas et al. 2019). In addition, some flora orchestrated different gene expressions to cope with changes in the external environment. For example, Burkholderia is equipped with glutathione S-transferase genes to degrade and detoxify complex organic compounds (Mitter et al. 2013). Unlike the metabolic gene expression of bacteria in planted soil, the gene expression of bacteria in bare soil was relatively low. These results indicated that plants promoted the activity of degrading gene–carrying bacteria in the soil.
Among the many root secretions, low-molecular-weight organic acids play a vital role in the nutrient and energy supply of microorganisms (Jones 1998). Plant roots influence the rhizosphere microenvironment by the release of root exudates (Segura and Ramos 2013). Organic acids play an important role in providing substrates for microbial metabolism to affect phytoremediation (Eze and Amuji 2024; He et al. 2022; Franchi et al. 2022). For M. jalapa, little information is available on the response of plant organic acid exudation to PHC stress. The alterations of organic acid and microorganisms in M. jalapa response to petroleum stress were investigated in the present study. Citric acid may play a critical role in upgrading the survival of PHC-degrading bacteria and enhancing the PHC-degrading activity, because M. jalapa has the highest citric acid exudation under petroleum-contaminated for 45 days (Fig. 5) with the maximum total degradation of PHCs (89.1%) (Fig. 1B). The relative abundance of Proteobacteria and Candidatus Saccharibacteria was significantly increased in M. jalapa response to petroleum contamination (Fig. 4). These results are in line with the previous study that hydrocarbon-contaminated soil was dominated by Proteobacteria (Devi et al. 2022; Phulpoto et al. 2024).
The major limitation of phytoremediation is the sphere of action which restricted to the plant itself or to the rhizosphere. Typically, phytoremediation treatments typically proceed at a slower rate than other bioremediation techniques. Meanwhile, tolerance to PHCs and remediation efficiency continue to be limitations in plant remediation applications. Considering this limitation, further blending of root exudates and plant roots could increase the microbial activity in PHC-stressed soil. In addition, the selection of dominant flora under phytoremediation for bioaugmentation may represent a new avenue for bioremediation research. However, it is fundamental to further examine the root exudation execution of distinctive organic ligands to assess their roles in the tolerance of M. jalapa.
Conclusions
Petroleum stress alter the hydrocarbon-degrading microbial community. The relative abundance of Actinobacteriota, Proteobacteria, and Candidatus Saccharibacteria was significantly increased on 45 days petroleum-contaminated rhizospheric soil. The growth of functional bacteria was significantly correlated with malic and citric acid in the roots of both M. jalapa and L. perenne. Based on the findings of the present study, we propose that transplanted M. jalapa and supplemented with citric acid as a complex surfactant could lead to the higher degradation of PHCs under petroleum stress.
Data Availability
The data will be available on request.
References
Agnello AC, Bagard M, van Hullebusch ED, Esposito G, Huguenot D (2016) Comparative bioremediation of heavy metals and petroleum hydrocarbons co-contaminated soil by natural attenuation, phytoremediation, bioaugmentation and bioaugmentation-assisted phytoremediation. Sci Total Environ 563–564:693–703. https://doi.org/10.1016/j.scitotenv.2015.10.061
Bell TH, Yergeau E, Juck DF, Whyte LG, Greer CW (2013) Alteration of microbial community structure affects diesel biodegradation in an Arctic soil. FEMS Microbiol Ecol 85(1):51–61. https://doi.org/10.1111/1574-6941.12102
Bhuyan B, Kotoky R, Pandey P (2023) Impacts of rhizoremediation and biostimulation on soil microbial community, for enhanced degradation of petroleum hydrocarbons in crude oil-contaminated agricultural soils. Environ Sci Pollut Res 30:94649–94668. https://doi.org/10.1007/s11356-023-29033-3
Brakorenko NN, Korotchenko TV (2016) Impact of petroleum products on soil composition and physical-chemical properties. IOP Conf Ser: Earth Environ Sci 33:012028. https://doi.org/10.1088/1755-1315/33/1/012028
Caporaso JG, Lauber CL, Walters WA, Berglyons D, Huntley J, Fierer N, Owens SM, Betley JR, Fraser L, Bauer M (2012) Ultra-high-throughput microbial community analysis on the Illumina HiSeq and MiSeq platforms. ISME J 6(8):1621–1624. https://doi.org/10.1038/ismej.2012.8
Cébron A, Louvel B, Faure P, France-Lanord C, Chen Y, Murrell JC, Leyval C (2011) Root exudates modify bacterial diversity of phenanthrene degraders in PAH-polluted soil but not phenanthrene degradation rates. Environ Microbiol 13(3):722–736. https://doi.org/10.1111/j.1462-2920.2010.02376.x
Cen Y, Li Y, Jiao H, Wang X, Bai Z (2016) Growth on microbial community in bioremediation of petroleum-contaminated saline-alkali soil. Agric Sci Technol 17(5):1223–1230. https://doi.org/10.16175/j.cnki.1009-4229.2016.05.041
Correa-García S, Pande P, Séguin A, St-Arnaud M, Yergeau E (2018) Rhizoremediation of petroleum hydrocarbons: a model system for plant microbiome manipulation. Microb Biotechnol 11(5):819–832. https://doi.org/10.1111/1751-7915.13303
Costa AS, Romao LPC, Araujo BR, Lucas SCO, Maciel STA, Wisniewski A, Alexandre MR (2012) Environmental strategies to remove volatile aromatic fractions (BTEX) from petroleum industry wastewater using biomass. Biores Technol 105:31–39. https://doi.org/10.1016/j.biortech.2011.11.096
Dai Y, Liu R, Zhou Y, Li N, Hou L, Ma Q, Gao B (2020) Fire Phoenix facilitates phytoremediation of PAH-Cd co-contaminated soil through promotion of beneficial rhizosphere bacterial communities. Environ Int 136:105421. https://doi.org/10.1016/j.envint.2019.105421
Devi SP, Jani K, Sharma A, Jha DK (2022) Bacterial communities and their bioremediation capabilities in oil-contaminated agricultural soils. Environ Monit Assess 194:9. https://doi.org/10.1007/s10661-021-09669-9
El Amrani A, Dumas AS, Wick LY, Yergeau E, Berthomé R (2015) “Omics” Insights into PAH degradation toward improved green remediation biotechnologies. Environ Sci Technol 49(19):11281–11291. https://doi.org/10.1021/acs.est.5b01740
Eze MO, Amuji CF (2024) Elucidating the significant roles of root exudates in organic pollutant biotransformation within the rhizosphere. Sci Rep 14:2359. https://doi.org/10.1038/s41598-024-53027-x
Franchi E, Cardaci A, Pietrini I, Fusini D, Conte A, De Folly DA, Grifoni M, Pedron F, Barbafieri M, Petruzzelli G, Vocciante M (2022) Nature-based solutions for restoring an agricultural area contaminated by an oil spill. Plants 11(17):2250. https://doi.org/10.3390/plants11172250
Gao Y, Ren L, Ling W, Kang F, Zhu X, Sun B (2010) Effects of low-molecular-weight organic acids on sorption-desorption of phenanthrene in soils. Soil Sci Soc Am J 74(1):51–59. https://doi.org/10.2136/sssaj2009.0105
Guo M, Gong Z, Miao R, Rookes J, Cahill D, Zhuang J (2017a) Microbial mechanisms controlling the rhizosphere effect of ryegrass on degradation of polycyclic aromatic hydrocarbons in an aged-contaminated agricultural soil. Soil Biol Biochem 113:130–142. https://doi.org/10.1016/j.soilbio.2017.06.006
Guo M, Gong Z, Miao R, Su D, Li X, Jia C, Zhuang J (2017b) The influence of root exudates of maize and soybean on polycyclic aromatic hydrocarbons degradation and soil bacterial community structure. Ecol Eng 99:22–30. https://doi.org/10.1016/j.ecoleng.2016.11.018
He M, Li Z, Chen C, Mei P (2022) Impact of soil types and root exudates on cadmium and petroleum hydrocarbon phytoremediation by Sorghum sudanense, Festuca arundinace, and Lolium perenne. Front Ecol Evol 10:1036765. https://doi.org/10.3389/fevo.2022.1036765
Hou J, Liu W, Wang B, Wang Q, Luo Y, Franks AE (2015) PGPR enhanced phytoremediation of petroleum contaminated soil and rhizosphere microbial community response. Chemosphere 138:592–598. https://doi.org/10.1016/j.chemosphere.2015.07.025
Huang G, Guo G, Yao S, Zhang N, Hu H (2016) Organic acids, amino acids compositions in the root exudates and Cu-accumulation in castor (Ricinus communis L.) Under Cu stress. Int J Phytoremediation 18(1):33–40. https://doi.org/10.1080/15226514.2015.1058333
Hussain I, Puschenreiter M, Gerhard S, Schöftner P, Yousaf S, Wang A, Syed JH, Reichenauer TG (2018) Rhizoremediation of petroleum hydrocarbon-contaminated soils: improvement opportunities and field applications. Environ Exp Bot 147:202–219. https://doi.org/10.1016/j.envexpbot.2017.12.016
Iqbal A, Mukherjee M, Rashid J, Khan SA, Ali MA, Arshad M (2019) Development of plant-microbe phytoremediation system for petroleum hydrocarbon degradation: an insight from alkb gene expression and phytotoxicity analysis. Sci Total Environ 671:696–704. https://doi.org/10.1016/j.scitotenv.2019.03.331
Jones DL (1998) Organic acids in the rhizosphere - a critical review. In Plant and Soil 205(1):25–44. https://doi.org/10.1023/A:1004356007312
Kaimi E, Mukaidani T, Tamaki M (2007) Screening of twelve plant species for phytoremediation of petroleum hydrocarbon-contaminated soil. Plant Prod Sci 10(2):211–218. https://doi.org/10.1626/pps.10.211
Kao NH, Su MC, Fan JR, Chung YY (2015) Identification and quantification of biomarkers and polycyclic aromatic hydrocarbons (PAHs) in an aged mixed contaminated site: from source to soil. Environ Sci Pollut Res 22:7529–7546. https://doi.org/10.1007/s11356-015-4237-9
Liu W, Hou J, Wang Q, Yang H, Luo Y, Christie P (2015) Collection and analysis of root exudates of Festuca arundinacea L. and their role in facilitating the phytoremediation of petroleum-contaminated soil. Plant Soil 389(1–2):109–119. https://doi.org/10.1007/s11104-014-2345-9
Lu L, Zhang J, Peng C (2019) Shift of soil polycyclic aromatic hydrocarbons (PAHs) dissipation pattern and microbial community composition due to rhamnolipid supplementation. Water Air Soil Pollut 230(5):107. https://doi.org/10.1007/s11270-019-4118-9
Lü H, Tang GX, Huang YH, Mo CH, Zhao HM, Xiang L, Li YW, Li H, Cai QY, Li QX (2024) Response and adaptation of rhizosphere microbiome to organic pollutants with enriching pollutant-degraders and genes for bioremediation: a critical review. Sci Total Environ 912:169425. https://doi.org/10.1016/j.scitotenv.2023.169425
Ma H, Wang A, Zhang M, Li H, Du S, Bai L, Chen S, Zhong M (2018) Compared the physiological response of two petroleum tolerant-contrasting plants to petroleum stress. Int J Phytorem 20(10):1043–1048. https://doi.org/10.1080/15226514.2018.1460303
Mahmud T, Sabo IA, Lambu ZN, Danlami D, Shehu AA (2022) Hydrocarbon degradation potentials of fungi: a review. J Environ Bioremediation Toxicol 5(1):50–56. https://doi.org/10.54987/jebat.v5i1.681
Meckenstock RU, Boll M, Mouttaki H, Koelschbach JS, Tarouco PC, Weyrauch P, Dong X, Himmelberg AM (2016) Anaerobic degradation of benzene and polycyclic aromatic hydrocarbons. J Mol Microbiol Biotechnol 26:92–118. https://doi.org/10.1159/000441358
Meng L, Qiao M, Arp HPH (2011) Phytoremediation efficiency of a PAH-contaminated industrial soil using ryegrass, white clover, and celery as mono- and mixed cultures. J Soils Sediments 11(3):482–490. https://doi.org/10.1007/s11368-010-0319-y
Mitter B, Petric A, Shin MW, Chain PSG, Hauberg-Lotte L, Reinhold-Hurek B, Nowak J, Sessitsch A (2013) Comparative genome analysis of Burkholderia phytofirmans PsJN reveals a wide spectrum of endophytic lifestyles based on interaction strategies with host plants. Front Plant Sci 4:120. https://doi.org/10.3389/fpls.2013.00120
Mohanta S, Pradhan B, Behera ID (2024) Impact and remediation of petroleum hydrocarbon pollutants on agricultural land: a review. Geomicrobiol J 41(4):345–359. https://doi.org/10.1080/01490451.2023.2243925
Orozco AF, Ciampi P, Katona T, Censini M, Papini MP, Deidda GP, Cassiani G (2021) Delineation of hydrocarbon contaminants with multi-frequency complex conductivity imaging. Sci Total Environ 768:144997. https://doi.org/10.1016/j.scitotenv.2021.144997
Pan Y, Zhang Q, Yu Y, Tong Y, Wu W, Zhou Y, Hou W, Yang J (2021) Three-dimensional migration and resistivity characteristics of crude oil in heterogeneous soil layers. Environ Pollut 268:115309. https://doi.org/10.1016/j.envpol.2020.115309
Panchenko LV, Kuzyanov DA, Pleshakova YV, Pozdnyakova NN, Muratova AY, Turkovskaya OV (2022) Effect of plant root exudate constituents on the degradation of phenanthrene by the rhizobacterium Mycolicibacterium gilvum (Mycobacteriaceae, Actinobacteria). Biology Bulletin 49:1958–1964. https://doi.org/10.1134/S1062359022100284
Peng S, Zhou Q, Cai Z, Zhang Z (2009) Phytoremediation of petroleum contaminated soils by Mirabilis Jalapa L. in a greenhouse plot experiment. J Hazard Mater 168(2–3):1490–1496. https://doi.org/10.1016/j.jhazmat.2009.03.036
Phillips LA, Greer CW, Farrell RE, Germida JJ (2012) Plant root exudates impact the hydrocarbon degradation potential of a weathered-hydrocarbon contaminated soil. Appl Soil Ecol 52(1):56–64. https://doi.org/10.1016/j.apsoil.2011.10.009
Phulpoto IA, Qi Z, Qazi MA, Yu Z (2024) Biosurfactants-based mixed polycyclic aromatic hydrocarbon degradation: from microbial community structure toward non-targeted metabolomic profile determination. Environ Int 184:108448. https://doi.org/10.1016/j.envint.2024.108448
Rajaei S, Seyedi SM (2018) Phytoremediation of petroleum-contaminated soils by Vetiveria zizanioides (L.) Nash. Clean – Soil Air Water 46(8):1800244. https://doi.org/10.1002/clen.201800244
Ramadass K, Mallavarapu M, Kadiyala V, Naidu R (2015) Ecological implications of motor oil pollution: earthworm survival and soil health. Soil Biol Biochem 85. https://doi.org/10.1016/j.soilbio.2015.02.026
Rohrbacher F, Starnaud M (2016) Root exudation: the ecological driver of hydrocarbon rhizoremediation. Agronomy 6(1):19. https://doi.org/10.3390/agronomy6010019
Ros M, Rodríguez I, García C, Hernández T (2010) Microbial communities involved in the bioremediation of an aged recalcitrant hydrocarbon polluted soil by using organic amendments. Biores Technol 101(18):6916–6923. https://doi.org/10.1016/j.biortech.2010.03.126
Saratale RG, Saratale GD, Cho SK, Kim DS, Ghodake GS, Kadam A, Kumar G, Bharagava RN, Banu R, Shin HS (2019) Pretreatment of kenaf (Hibiscus cannabinus L.) biomass feedstock for polyhydroxybutyrate (PHB) production and characterization. Bioresour Technol 282(February):75–80. https://doi.org/10.1016/j.biortech.2019.02.083
Segura A, Ramos JL (2013) Plant–bacteria interactions in the removal of pollutants. Curr Opin Biotechnol 24:467–473. https://doi.org/10.1016/j.copbio.2012.09.011
Singh MJ, Sedhuraman P (2015) Biosurfactant, polythene, plastic, and diesel biodegradation activity of endophytic Nocardiopsis sp. mrinalini9 isolated from Hibiscus rosasinensis leaves. Bioresour Bioprocess 2(1). https://doi.org/10.1186/s40643-014-0034-4
Thomas F, Corre E, Cébron A (2019) Stable isotope probing and metagenomics highlight the effect of plants on uncultured phenanthrene-degrading bacterial consortium in polluted soil. ISME J 13(7):1814–1830. https://doi.org/10.1038/s41396-019-0394-z
Ueki A, Kodama Y, Kaku N, Shiromura T, Satoh A, Watanabe K, Ueki K (2010) Rhizomicrobium palustre gen. nov., sp. nov., a facultatively anaerobic, fermentative stalked bacterium in the class Alpha proteobacteria isolated from rice plant roots. J Gen Appl Microbiol 56(3):193–203. https://doi.org/10.2323/jgam.56.193
Umeh AC, Vázquez-Cuevas GM, Semple KT (2018) Mineralisation of 14C-phenanthrene in PAH-diesel contaminated soil: impact of Sorghum bicolor and Medicago sativa mono- or mixed culture. Appl Soil Ecol 125:46–55. https://doi.org/10.1016/j.apsoil.2017.12.010
Varjani SJ (2017) Microbial degradation of petroleum hydrocarbons. Biores Technol 223:277–286. https://doi.org/10.1016/j.biortech.2016.10.037
Wei X, Lyu S, Yu Y, Wang Z, Liu H, Pan D, Chen J (2017) Phylloremediation of air pollutants: exploiting the potential of plant leaves and leaf-associated microbes. Front Plant Sci 8(2):1–23. https://doi.org/10.3389/fpls.2017.01318
Weller DM, Thomashow LS (2007) Current challenges in introducing beneficial microorganisms into the rhizosphere. Mol Ecol Rhizosphere Microorganisms: Biotechnol Release GMOs 293:1–18. https://doi.org/10.1002/9783527615810.ch1
Wu M, Dick WA, Li W, Wang X, Yang Q, Wang T, Xu L, Zhang M, Chen L (2016) Bioaugmentation and biostimulation of hydrocarbon degradation and the microbial community in a petroleum-contaminated soil. Int Biodeterior Biodegradation 107:158–164. https://doi.org/10.1016/j.ibiod.2015.11.019
Wu N, Li Z, Meng S, Wu F (2021) Soil properties and microbial community in the rhizosphere of Populus alba var. pyramidalis along a chronosequence. Microbiol Res 250:126812. https://doi.org/10.1016/j.micres.2021.126812
Xia M, Chakraborty R, Terry N, Singh RP, Fu D (2020) Promotion of saltgrass growth in a saline petroleum hydrocarbons contaminated soil using a plant growth promoting bacterial consortium. Int Biodeterior Biodegradation 146:104808. https://doi.org/10.1016/j.ibiod.2019.104808
Yousaf S, Andria V, Reichenauer TG, Smalla K, Sessitsch A (2010) Phylogenetic and functional diversity of alkane degrading bacteria associated with Italian ryegrass (Lolium multiflorum) and Birdsfoot trefoil (Lotus corniculatus) in a petroleum oil-contaminated environment. J Hazard Mater 184(1–3):523–532. https://doi.org/10.1016/j.jhazmat.2010.08.067
Zadjelovic V, Chhun A, Quareshy M, Silvano E, Hernandez-Fernaud JR, Aguilo-Ferretjans MM, Bosch R, Dorador C, Gibson MI, Christie-Oleza JA (2020) Beyond oil degradation: enzymatic potential of Alcanivorax to degrade natural and synthetic polyesters. Environ Microbiol 22(4):1356–1369. https://doi.org/10.1111/1462-2920.14947
Zhang ZZ, Hou Z, Yang C, Ma C, Tao F, Xu P (2011) Degradation of n-alkanes and polycyclic aromatic hydrocarbons in petroleum by a newly isolated Pseudomonas aeruginosa DQ8. Biores Technol 102(5):4111–4116. https://doi.org/10.1016/J.BIORTECH.2010.12.064
Zhang L, Zhong M, Li X, Lu W, Li J (2020) River bacterial community structure and co-occurrence patterns under the influence of different domestic sewage types. J Environ Manag 266:110590. https://doi.org/10.1016/j.jenvman.2020.110590
Funding
This research was supported by the National Natural Science Foundation of China (Grant No. 31070448).
Author information
Authors and Affiliations
Contributions
M.Z.: conceptualization; writing—original draft; supervision; resources. C.Y. and L.S.: investigation, data curation, methodology. Z.S. and J.X.: investigation, visualization, software. J.Z., Q.L., and Y.H.: methodology, software, validation. H.M., H.C., and J.C.: methodology, writing—review and editing. S.C.: conceptualization, data curation, software, writing—review and editing.
Corresponding author
Ethics declarations
Ethical approval
Not applicable.
Consent to participate
Not applicable.
Consent to publish
Not applicable.
Competing interests
The authors declare no competing interests.
Additional information
Responsible Editor: Diane Purchase
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhong, M., Yang, C., Su, L. et al. Interactions between plants and bacterial communities for phytoremediation of petroleum-contaminated soil. Environ Sci Pollut Res 31, 37564–37573 (2024). https://doi.org/10.1007/s11356-024-33667-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11356-024-33667-2