Abstract
The ability of bacteria to efficiently remove phenolic pollutants depends on their genetic makeup and environmental conditions. This study examined a novel strain, Pseudomonas aeruginosa STV1713, for degrading higher concentrations of phenol and 2,4-dichlorophenol. After optimization, a combination of degradation parameters, such as pH (7.0), temperature (32.5 °C), and ammonium nitrate concentration (0.7 g/L), was found to reduce degradation time while promoting cell growth. Under these optimal conditions, the bacterium effectively degraded up to 2000 mg/L of phenol and 1400 mg/L of 2,4-dichlorophenol, while maximum tolerance was observed till 2100 mg/L and 1500 mg/L, respectively. Metabolic profiling identified crucial metabolites in the ortho-degradation pathway during pollutant removal. Additionally, transcriptome analysis revealed that P. aeruginosa STV1713 utilizes different branches of the beta ketoadipate pathway for phenol and 2,4-DCP removal. Moreover, under high pollutant stress, the bacterium survived through differential gene expression in ribosome biogenesis, chemotaxis, membrane transport, and other pathways.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Phenolic contaminants are widely distributed in the environment, especially in water and soil, causing various health hazards to living organisms while being a major reason for groundwater pollution. (Chen et al. 2019). Continuous exposure of living cells to these pollutants produces various toxic metabolites via transformation processes through the cytochrome P450 enzyme system. These toxic metabolites cause DNA damage or hamper the production of crucial metabolic enzymes, leading to mutations or malignancy (Barik et al. 2021). Besides their health impacts, phenol and 2,4-dichlorophenol possess strong antimicrobial characteristics, which make them highly resistant to microbial degradation, unlike other aromatic hydrocarbon compounds (Pinheiro et al. 2018). It disrupts the lipid bilayer structure of the cell membrane, leading to increased membrane permeability and leakage of cellular contents. This disturbance in membrane integrity can result in the loss of ions and nutrients, causing damage to the overall cellular homeostasis. In addition to membrane and cell wall damage, phenol exposure can lead to intracellular damage. It can affect enzymes, proteins, and nucleic acids, disrupting various cellular functions (Lucchini et al. 1990). Furthermore, the chemical stability of the C–Cl bond poses challenges for most environmental microorganisms to degrade polychlorinated compounds (Oluwasanu 2018).
Several Pseudomonas species have been reported for their efficacy in phenol removal; however, only a few bacteria demonstrated degradation capabilities above 1500 mg/L (Asimakoula et al. 2023). Regarding 2,4-DCP degradation, only a few Pseudomonas species were documented, but none of which exhibited pollutant tolerance above 1000 mg/L (Kargi and Eker 2005). The low tolerance levels of these bacteria were attributed to the strong antimicrobial effects of these pollutants (Hasan and Jabeen 2015). Moreover, many external factors such as pH, temperature, nitrogen source, other carbon sources, initial pollutant concentration, and the use of bacterial consortia were found to influence microbial phenol degradation. Many researchers have analyzed the effects of these parameters on phenolic pollutant degradation (Aisami et al. 2020). However, the key metabolic enzymes play a major role in pollutant removal. Therefore, optimizing the activities of major enzymes can accelerate both pollutant removal and cell growth by hastening metabolism. Response Surface Methodology (RSM) refers to a collection of mathematical and statistical techniques that help to design, refine, and enhance a process where multiple variables impact a response of importance, intending to optimize the response (Subramaniam et al. 2020).
The fundamental mechanism of bacterial phenol degradation, including ortho cleavage and meta-degradation pathways, has been extensively studied and thoroughly explored (Lee et al. 2022). In both pathways, the initial step involves the conversion of phenolic pollutants into catechol by the phenol hydroxylase enzyme (van Schie and Young 2000). Furthermore, in the ortho-cleavage pathway, catechol is then cleaved into cis-cis muconic acid by the enzyme catechol 1,2-dioxygenase. Conversely, in the meta-pathway, the enzyme catechol 2,3-dioxygenase transforms catechol into 2-hydroxymuconic semialdehyde. Following these steps, a cascade of reactions occurs, and the end products of both pathways enter the Krebs cycle, eventually breaking down pollutants into simpler compounds that bacteria can use for growth and energy production (Krastanov et al. 2013). Therefore, metabolic profiling is the easiest way to determine the key metabolic pathways in a given biological system during pollutant degradation (Barik et al. 2021). Apart from these pathways, bacteria may sometimes employ novel mechanisms for pollutant removal as a part of their adaptation processes.
Understanding the genetic mechanisms behind degradation processes is important for developing efficient bioremediation strategies. Transcriptome analysis is a powerful tool used to study gene expression patterns at the mRNA level. It can help to identify essential genes involved in phenol degradation pathways and understand their regulation (Shakya et al. 2019). Moreover, transcriptome analysis can be used to identify novel genes involved in phenolic pollutant degradation, which can lead to the discovery of new enzymes, metabolic pathways, and survival mechanisms that can be exploited for bioremediation purposes (Gu et al. 2018).
In our study, we aimed to isolate and characterize a bacterium capable of tolerating and degrading higher concentrations of phenol and 2,4-DCP from oil-contaminated soil. Optimization of degradation parameters, transcriptome analysis, and metabolic profiling were conducted on the best isolate to study the molecular mechanisms underlying pollutant degradation and survival mechanisms under high pollutant stress.
Materials and methods
Isolation and screening
Soil samples for isolating phenol degrading bacterium were collected (3–5 cm below the surface level) from an oil-contaminated site near a petrol pump at Kattangal, Kerala, India. The composition of the Mineral Salt Medium (BH-MSM) used in the current study was adopted from previous literature (Subramaniam et al. 2019). Previous studies have shown that the bacteria capable of degrading 2,4-DCP were also highly proficient in degrading phenol (Farag et al. 2021). Consequently, we employed only 2,4-DCP to isolate and screen the most tolerant bacteria in the current study.
The soil sample (5 g) was added to autoclaved BH-MSM (100 mL) containing 100 mg/L of 2,4-DCP and incubated at 30 °C for 72 h with continuous shaking (125 rpm). Then, 1 mL of the culture was serially transferred to the fresh 2,4-DCP-BH-MSM (200 mg/L) and incubated under the same conditions. Next, 1 mL of the enrichment suspension was streaked onto BHA plates with 300 mg/L of 2,4-DCP and incubated for two days at 30 °C. Four colonies (A1, A2, A3, and A4) formed on the BHA plates were screened (secondary screening) by growing them individually in BH-MSM with 400 mg/L of 2,4-DCP. The final enrichment culture was streaked onto BHA plates with the same pollutant concentration. After incubation, the single bacterial colony (A1) that appeared on the plate was purified for further degradation studies.
Morphological, biochemical, and molecular characterizations of the isolate
The best tolerant isolate, A1, was preliminarily identified by morphological observation (grams staining and hanging drop method) and biochemical characterization (indole, methyl-Red, Voges-Proskauer, oxidase, catalase, starch hydrolysis, urease, citrate utilization, growth on cetrimide agar, and Triple Sugar Iron (TSI) test) (Mahon and Lehman 2019). The 16 rRNA sequencing was performed to confirm the genetic identity of the bacterial species (Chromous biotech Pvt. Ltd., Bangalore).
Optimization of degradation conditions
Effect of process variables on phenol hydroxylase activity
Metabolic enzymes play a pivotal role in every degradation process. Phenol hydroxylase, also known as phenol monooxygenase, is considered the primary enzyme for degrading phenolic pollutants, as it catalyzes the initial hydroxylation reaction. Therefore, optimizing the activity of phenol hydroxylase can significantly impact the entire degradation process. Consequently, the first stage of RSM analysis involved checking the effects of cultivation variables on the phenol hydroxylase activity (supplementary Table 1). The variables and their lowest and highest values were selected based on the preliminary studies.
Briefly, the bacterium inoculated in BH-MSM separately containing 1000 mg/L phenol and 2,4-DCP were incubated under varying pH, temperature, and ammonium nitrate (NH4NO3) conditions while keeping two parameters constant each time. Cells were collected by centrifugation at 2508 × g for 10 min. The resulting pellets were washed twice using 0.33 M Tris–HCl buffer (pH 7.6) and sonicated (Sonics Vibra-cell™) on ice for 4 min (30 s on and 30 s off cycle). After sonication, cell extracts were centrifuged at 15775 × g for 20 min at 4 °C, and cell-free extracts were immediately used to test the activity of the phenol hydroxylase enzyme, as described previously (Cafaro et al. 2004). One unit of phenol hydroxylase activity was defined as the amount of enzyme required for catalyzing the oxidation of 1 µmol NADH per min. Lowry’s method was used to determine the total protein concentration in the cell-free extract, with bovine serum albumin (BSA) as the standard (Lowry et al. 1951). Phenol and 2,4-DCP degradation percentages were estimated using the 4-aminoantipyrine technique, as described previously (Subramaniam et al. 2020). Cell growth was measured by taking optical density (OD) values at 600 nm in a UV spectrophotometer (PerkinElmer Lambda 365 UV/Vis spectrophotometer).
RSM-experimental design and data analysis
Response Surface Methodology is a collection of mathematical and statistical approaches for determining the relationship between the response and the independent variable (Nor Suhaila et al. 2013). The current study employed a central composite design (CCD) (Zhou et al. 2011) with three variables, NH4NO3 concentration (A), temperature (B), and pH (C), as they were expected to impact pollutant degradation time and cell growth substantially. For each model (phenol and 2,4-DCP), 20 experimental combinations were constructed, with six replications at the center point, and data were analyzed with Design Expert Software (version 13.0.5.0—Stat-Ease, Inc., Minneapolis, USA). ANOVA was used to evaluate relationships between process factors and responses. The R2 coefficient was used to determine the fitness of the coefficients, while the F-test was employed to determine the statistical significance of the models (Singh and Gupta 2016). The lowest and highest values of the variables were selected based on preliminary laboratory experiments.
Mathematical relationships of responses (Y) to independent variables A, B, and C (NH4NO3 concentration, temperature, and pH, respectively) were determined by the following quadratic polynomial equation:

where E is the response (degradation time and cell growth); A, B, and C are significant independent variables; ɼ1, ɼ2, ɼ3 are linear regression coefficients; ɼ11, ɼ22, and ɼ33 are quadratic regression coefficients; ɼ12, ɼ23, and ɼ13 are interactive regression coefficients; and ɼ0 is a constant.
The results were evaluated by growing the bacterium in BH-MSM containing phenol/2,4-DCP with an initial concentration of 1 g/L under various pH, temperature, and NH4NO3 combinations suggested by the RSM software (supplementary Table 2). All experiments were performed in triplicates.
Validation of optimal conditions
After RSM analysis, validation experiments were conducted three times using the recommended optimal conditions to validate their accuracy and reliability. The bacterium was separately inoculated into BH-MSM containing 1 g/L phenol and 2,4-DCP and incubated under the optimal conditions suggested by the RSM software. Results were compared with the predicted experimental values. Percentage error values were calculated to verify the reliability of models (Mohd Ariff and Abdullah 2019). In addition, all remaining experiments in this study were carried out under optimized conditions.
Degradation studies
Degradation studies were conducted with an initial concentration of 400 mg/L (the pollutant concentration at which the bacterium was isolated) phenol and 2,4-DCP. All experiments were conducted under optimized conditions. Phenolic-BH-MSM medium without inoculum was used as the control. Additionally, degradation studies involving mixed pollutants were performed to confirm the stability and activity of the bacterium. BH-MSM containing phenol and 2,4-DCP in equal concentrations was used for the study. The experiments were continued until the bacterium reached its maximum ability to degrade the pollutants. Samples aseptically collected at regular intervals (4–8 h) were analyzed for pollutant removal and cell growth analysis (Subramaniam et al. 2020). Each experiment was performed in triplicate.
Metabolic profiling
The isolate A1 was separately inoculated into BH-MSM containing 1000 mg/L of phenol and 2,4-DCP and incubated under the optimum conditions. Samples collected at various intervals (e.g., every 8 h) were subjected to centrifugation at 15775 × g for 10 min at 4 °C. The resulting cell pellets were disrupted using an ultrasonicator for 4 min (with 30 s on and 30 s off cycles). After disruption, the extracts were centrifuged at 15775 × g at 4 °C for 30 min. Supernatants were filtered to completely remove cell debris using a 0.45 µm membrane filter (MF-Millipore™ Membrane Filter, Merck, India). Subsequently, the supernatants were acidified to pH 2 with 0.5 M H2SO4, and metabolites were extracted using the solvent extraction method with ethyl acetate. The ethyl acetate-extracted metabolites were dried in a vacuum desiccator and then dissolved in dichloromethane for GC/MS (Gas Chromatography-Mass Spectrometry) analysis. Samples were injected into the GC/MS with a split ratio of 5:1. The MS (mass spectrum) system was equipped with an EI source, and the masses of the samples were analyzed within a range of 30–250 m/z (Gas Chromatograph, Agilent Technologies 6890N, GC system coupled with JEOL Mass Spectroscopy; CareKeralam, Kochi) (Panigrahy et al. 2020).
Transcriptome analysis
A1 cells, separately grown in phenol (1000 mg/L) and 2,4-DCP (1000 mg/L), were collected [(in the mid-log phase) phenol grown sample, 16 h; 2,4-DCP grown sample, 17 h; and control, 14 h] by centrifugation (2508 × g for 10 min at 4 °C), and the total RNA of each sample was extracted using the GeneJET RNA Purification Kit (Thermo Fisher Scientific). RNA concentration and quality were evaluated, and high-throughput RNA sequencing was conducted, as previously described (Gu et al. 2017). Protein sequences were annotated against the NCBI nr, Uniprot, Swissprot, and Uniref100 protein databases using the BLASTp module of Diamond. Gene ontology analysis (http://www.geneontology.org/) and KEGG analysis (http://www.genome.jp/kegg/) were performed on differentially expressed genes (Clevergene Pvt. Ltd., Bangalore) to identify metabolic pathways and cellular survival mechanisms under high phenol and 2,4-DCP concentrations.
Statistical analysis
All the experiments were done in triplicates, and the experimental data were analyzed using Design Expert Software (version 13.0.5.0—Stat-Ease, Inc. Minneapolis, USA), Origin 2018 (V9.5.1), and Microsoft Excel 2016.
Results
Isolation and screening of phenol-degrading bacteria
Among four phenotypically different colonies (A1, A2, A3, and A4) observed during the initial screening stage, only one colony, A1, was found to have tolerance at 400 mg/L 2,4-DCP concentration during the secondary screening. It was selected for further degradation studies and was purified and stored as glycerol stocks at − 20 °C for future use.
Morphological, biochemical, and molecular characterizations of the isolate
The isolate, A1, was a rod-shaped, gram-negative, and motile organism. Biochemical characterization revealed that the isolate could produce enzymes such as catalase and oxidase but could not hydrolyze starch, indicating the absence of amylase. Similarly, it tested negative for urease, indole, Voges-Proskauer, and methyl red tests. Also, the TSI test indicated that the organism could not utilize sugars such as glucose, sucrose, and lactose. In contrast, it showed positive results for citrate utilization. The isolate had proliferous growth on cetrimide agar medium (Supplementary Fig. 1). After the 16 s rRNA gene analysis, the organism was identified as a novel strain of P. aeruginosa, and the 16S rRNA gene sequence information was deposited to the GenBank database with the strain name P. aeruginosa STV1713 (Accession no. OP753705.1) (Fig. 1).
Optimization of degradation conditions
Effect of process variables on phenol hydroxylase activity
An initial screening was conducted to assess the impact of various factors, including temperature, pH, nitrogen sources, initial pollutant concentration, and glucose concentration on phenol and 2,4-DCP degradations. The most influential factors, NH4NO3 (A), temperature (B), and pH (C), were subsequently optimized for phenol hydroxylase activity, cell growth, and degradation time (keeping two parameters constant each time). Effects of individual degradation parameters on cell growth, phenol hydroxylase activity, and degradation time of phenol and 2,4-DCP are summarized in supplementary Table 1. Phenol hydroxylase activity increased with an increasing NH4NO3 concentration, ranging from 0.4 to 0.8 g/L. It was also significantly influenced by the cultivation temperature and pH, with the highest activities observed at 33 °C and the pH 7.0. The maximum phenol hydroxylase activity was closely correlated with the maximum cell concentration obtained during cultivation, and the phenol degradation time also reduced proportionally with an increase in phenol hydroxylase activity, suggesting that this enzyme played a crucial role in the degradations of phenol and 2,4-DCP by P. aeruginosa STV1713. Consequently, the combinational effects of these parameters on pollutant degradation time and cell growth were analyzed by the RSM-CCD method.
RSM-experimental design and data analysis
A total of 40 experiments (20 each for phenol and 2,4-DCP) with various combinations of NH4NO3 (Factor A, 0.4 to 1 g/L), temperature (Factor B, 25 to 40 °C), and pH (Factor C, pH 6 to 8) were conducted. The results were analyzed using the CCD of RSM. The specific responses measured were the degradation time of pollutants (phenol and 2,4-DCP) and maximum cell growth. Predicted and experimental results are provided in supplementary Table 2.
During the optimization factors for phenol degradation, the highest and lowest cell concentrations were observed during runs 8 (0.832) and 17 (0.0853), respectively. Similarly, the fastest phenol degradation time (25.5 h) was achieved in run 8, whereas the slowest degradation time (41 h) was recorded in run 17. In the context of 2,4-DCP degradation, the maximum and minimum cell concentrations were noted in runs 2 (0.6642) and 17 (0.0986), respectively. Similarly, the most rapid degradation time for 2,4-DCP (62 h) was attained in run 2, while the longest phenol degradation time (88 h) was observed in run 17. The findings of this study also revealed a direct correlation between lower cell production and extended pollutant degradation time. Conversely, the highest cell growth was associated with the shortest pollutant degradation time (supplementary Table 2).
After multiple regression analysis on the experimental data, the following second-order polynomial equations (Eqs. 2 and 3, in actual units) were obtained, and they could connect the pollutant degradation time and cell growth to the parameters evaluated.
where A, B, and C are the independent variables; Y1 and Y2 are the responses (cell growth and degradation time, respectively).
Data from the analysis of variance (ANOVA) for the response surface reduced quadratic model are shown in supplementary Table 3. Data obtained on predicted and observed values revealed that the empirical models fit the actual data with the R2 (determination of coefficients) values of 0.9877 (cell growth during phenol degradation), 0.9805 (phenol degradation time), 0.9892 (cell growth during 2,4-DCP degradation), and 0.9777 (2,4DCP degradation time). The R2 values were close to 1, indicating that the empirical models fit the actual data well. The adjusted R2 values (0.9766 for cell growth and 0.9629 for phenol degradation time; 0.9794 for cell growth and 0.9576 for 2,4-DCP degradation time) revealed that changes in model parameters accounted for more than 95% of the variation in the responses. Additionally, predicted R2 values for all models were in reasonable agreement with their respective adjusted R2. The F value for each model, 88.97 (cell growth during phenol degradation), 55.81 (maximum phenol degradation), 101.54 (cell growth during 2,4-DCP degradation), and 48.64 (maximum 2,4DCP degradation), indicated that models were significant. The Lack of Fit values for each model, viz. 0.0516 (cell growth during phenol degradation), 0.1891 (phenol degradation time), 0.0827 (cell growth during 2,4-DCP degradation), and 0.6582 (2,4-DCP degradation time), indicated that the Lack of Fit values were not significant in comparison to the pure error. A non-significant Lack of Fit is considered good as it indicates the fitness of the model (Favier et al. 2021). The Adequate Precision metric assessed the signal-to-noise ratio. A ratio greater than 4 was preferred (Singh and Gupta 2016). Meanwhile, coefficient of variation (CV) values of 11%, 3.48%, 8.9%, and 2.3% indicated that the experiments were reliable and precise. The p-value, found to be below 0.0001 in all models, played a pivotal role in establishing the significance of each coefficient.
The bacterial cell growth and degradation time of pollutants concerning different NH4NO3 concentrations, temperature, and pH are shown in (Fig. 2A, B). During both phenol and 2,4-DCP degradations, cell growth was analyzed against NH4NO3 concentration and temperature when pH was kept at 7. Maximum cell concentrations were obtained at a temperature of 32.5 °C and an NH4NO3 concentration of 0.7 g/L. Similarly, when temperature and pH were tested against cell growth at an NH4NO3 concentration of 0.7 g/L, the maximum values were observed at pH 0.7 and the temperature 32.5 °C. Furthermore, when NH4NO3 concentration and pH were tested at the middle range of temperature 32.5 °C, maximum biomass concentration was obtained at the pH 0.7 and NH4NO3 concentration of 0.7 g/L. In the case of 2,4-DCP degradation, the same trend was observed. Finally, the results suggested that a combination of pH 7.0, a temperature of 32.5 °C, and an NH4NO3 concentration of 0.7 g/L has highly favored maximum cell growth during both pollutant degradation (phenol, 0.832; 2,4-DCP, 0.6642) while reducing the degradation time (phenol, 25.5; 2,4-DCP, 62 h) due to increased microbial activity.
Validation of optimal conditions
Degradation experiments using phenol and 2,4-DCP (initial concentration of 1 g/L) were repeated three times under the suggested optimal conditions (pH, 7.0; temperature, 32.5 °C; NH4NO3, 0.7 g/L) to verify their accuracy and reliability. Based on these conditions, the predicted maximum biomass concentrations during phenol and 2,4-DCP degradations were 0.7652 and 0.6176, respectively. Similarly, under the optimized conditions, predicted degradation durations for phenol and 2,4-DCP were 27.55 h and 63.17 h, respectively (supplementary Table 2).
Upon validation of the optimized conditions through further experiments, maximum cell growth during phenol and 2,4-DCP degradations were determined to be 0.764 and 0.6034, respectively. The corresponding degradation durations were 27 h and 62.4 h, respectively. As depicted in Table 1, percentage error values below 5% confirmed the reliability of the optimized parameters.
Similarly, the phenol hydroxylase activity in the samples grown under optimized conditions showed maximum activity (phenol, 138.54 µm/mL/min and 2,4-DCP 116.65 μm/mL/min) compared to the initial optimization.
Degradation studies
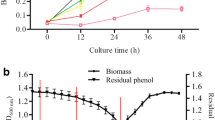
Degradation studies started with an initial pollutant concentration of 400 mg/L, the same concentration at which the bacterium was isolated, using phenol and 2,4-DCP as the sole carbon sources under optimum conditions (pH, 7.0; temperature, 32.5 °C; NH4NO3, 0.7 g/L). The degradations of phenol and 2,4-DCP were studied separately. Overnight cultures of A1 cells grown separately in 400 mg/L phenol and 400 mg/L 2,4-DCP were used as inocula. The maximum tolerance levels for phenol and 2,4-DCP in P. aeruginosa STV1713 were observed until 2100 mg/L and 1500 mg/L, respectively. However, 100% degradation ability was shown for phenol up to 2000 mg/L within 60 h and 2,4-DCP up to 1400 mg/L within 108 h (Fig. 3a, b). Similarly, degradation studies involving mixed pollutants started at an initial pollutant concentration of 400 mg/L (200 mg/L each phenol and 2,4-DCP). Remarkably, the cells showed maximum tolerance till 2000 mg/L, while maximum degradation capabilities were observed till 1900 mg/L (Fig. 3c). These findings strongly suggest that the bacterium exhibited efficient degradation of higher concentrations of mixed pollutants, similar to its performance with individual pollutants. Furthermore, the study highlights that the presence of the pollutant mixture did not hinder the degradation potential of P. aeruginosa STV1713. However, there was a gradual reduction in the cell viability as pollutant concentration increased above 1000 mg/L of phenol and 2,4-DCP. The reduction in cell viability could be related to the harmful antimicrobial effects of phenolic pollutants on bacterial cells (Fig. 3d).
Graphs show the pollutant removal ability of P. aeruginosa STV1713 at various concentrations of phenol (a), 2,4-DCP (b), and mixed pollutants (c). The growth of P. aeruginosa STV1713 with respect to different initial phenol and 2.4-DCP concentrations (d). Error bars are added to the graphs to show the experimental variability or error observed among triplicate data points
Metabolic profiling
Metabolites formed during the degradation of phenol and 2,4-DCP by P. aeruginosa STV1713 were detected by GC/MS analysis. Metabolites were identified based on their mass spectrum characteristics and retention times. The analysis revealed three metabolites, viz. catechol (molecular weight (mw), 110.1; m/z, 110.04; retention time, 10.24 min), cis-cis muconic acid (mw, 142.11, m/z, 142.01; retention time, 13.25 min), and succinate (mw, 118.09; m/z, 118.03; retention time, 10.45 min), during phenol degradation (Fig. 4). Similarly, four intermediates, viz. 4-chlorophenol (mw, 128.56; m/z, 128; RT, 12.32), phenol (mw, 94.11; m/z, 94.04; RT, 6.24), cis-cis muconic acid (mw, 142.11; m/z, 142.01; RT, 13.16), and fumarate (mw, 116.07; m/z, 116.01; RT, 10.43) (Fig. 4) were identified during 2,4-DCP degradation. The presence of cis-cis muconic acid was verified in both samples, indicating the utilization of the ortho degradation pathway by the bacterium for the removal of pollutants. Furthermore, the detection of succinate and fumarate provided evidence that the intermediates of the ortho pathway ultimately reached the Krebs cycle. GC/MS analysis results were later compared with the outcomes of transcriptome-KEGG analysis for detailed metabolic pathway analysis (“Metabolic pathway analysis” section).
Transcriptome analysis
Transcriptomes of P. aeruginosa STV1713 cultivated in BH-MSM containing phenol (1000 mg/L; sample 1) or 2,4-dichlorophenol (1000 mg/L; sample 2), and BH-MSM without phenolic pollutants (control) were examined to understand its genetic mechanism of pollutant degradation. High-throughput RNA-sequencing yielded clean reads with total lengths of 27,364,942 from the control, 20,361,356 reads from sample 1, and 24,353,506 reads from sample 2. The Q20 of all samples was above 99%, indicating excellent transcriptome sequencing. The sequence data were uploaded to the NCBI GenBank database with the following unique identifiers: bio-project: PRJNA964760, bio-samples SAMN34490102 (Phenol), SAMN34490102 (2,4-DCP), and SAMN34489986 (control).
Data processing and de novo transcriptome assembly
Assembled transcriptomes were processed in rMLST to identify the species, and it was identified as P. aeruginosa STV1713 with 100% support. Among the three transcriptomes, 10,601 transcripts were annotated and classified into 50 clusters of orthologous protein groups (Fig. 5). The most abundant cluster was related to signaling and cellular processes, followed by a cluster involved in genetic information processing. P. aeruginosa STV1713 employed a complex regulatory system for phenol and 2,4-DCP degradation, as evidenced by the abundance of proteins associated with metabolic processes, catalytic activities, and cellular processes.
Differential gene expression analysis
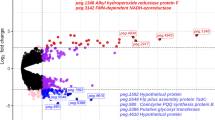
During DGE analysis, the data were normalized using the relative log expression method, and transcripts with an absolute log2 fold change ≥ 1 and a p-value ≤ 0.05 were deemed statistically significant (supplementary Fig. 2). In sample 1 (phenol), a significant differential gene expression was observed in 614 genes compared to the control group, with 479 genes upregulated and 135 genes downregulated. In 2,4-DCP grown sample, 630 genes exhibited significant differential expression, with 468 genes upregulated and 162 genes downregulated. Among the upregulated genes in the phenol-grown sample, 298 were associated with metabolism, including xenobiotic degradation and phenol degradation. Genes related to cellular processes (103) and genetic information processing (56) were also identified. Some upregulated genes (22) remained unclassified, indicating their unknown functions. In sample 2, 272 genes in the upregulated genes were involved in metabolism, including xenobiotic degradation and 2,4-DCP degradation. Additionally, 99 genes were associated with signaling and cellular processes, and 61 were related to membrane transport. The remaining 37 genes were categorized as unclassified. The functions of key genes and proteins detected during transcriptome analysis are listed in supplementary Table 4.
Metabolic pathway analysis
After transcriptome analysis, the predicted metabolic pathways for phenol and 2,4-DCP degradation in P. aeruginosa STV1713 are represented in Fig. 6.
In P. aeruginosa STV1713, the initial conversion of phenol into catechol was catalyzed by the enzyme phenol/toluene 2-monooxygenase via the hydroxylation reaction. Catechol was transformed into cis-cis-muconate by catechol 1,2-dioxygenase. Further conversion of cis-cis-muconate yielded muconolactone by muconolactone cycloisomerase. Muconolactone underwent reactions, including the formation of 3-oxoadipate enol lactone catalyzed by muconolactone delta-isomerase (not detected in transcriptome analysis). 3-Oxoadipate enol lactone was then converted to 3-oxoadipate by 3-oxoadipate enol lactone hydrolase. Furthermore, the enzyme 3-oxoadipate CoA-transferase catalyzed the conversion of 3-oxoadipate, resulting in the production of 3-oxoadipyl-CoA. Finally, 3-oxoadipyl-CoA was converted to TCA cycle intermediates like succinyl-CoA or acetyl CoA through the action of 3-oxoadipate CoA-transferase and 3-oxoadipyl-CoA thiolase.
Furthermore, the findings suggested that the bacterium employed an alternative pathway for phenol degradation, the protocatechuate pathway, as evidenced by the presence of major enzyme-coding genes among the DEGs (Fig. 6). Despite the downregulation of many genes related to this pathway, the substantial upregulation of the gene encoding the protocatechuate 3,4-dioxygenase enzyme suggests the activation of this pathway induced by phenol. Briefly, the pathway commenced with the conversion of phenol into 4-hydroxybenzoate by the enzyme protocatechuate 3,4-dioxygenase. Subsequently, 4-hydroxybenzoate was metabolized into protocatechuate through the action of 4-hydroxybenzoate 3-monooxygenase. Protocatechuate then underwent cleavage into beta-carboxy-cis,cis-muconate by protocatechuate 3,4-dioxygenase. This compound was further transformed into cis,cis-muconate via cis,cis-muconate cycloisomerase. Cis,cis-muconate entered the ortho-degradation pathway for further degradation and finally reached the Krebs cycle.
In P. aeruginosa STV1713, the initial step in 2,4-DCP degradation involved dehalogenation at the second position, forming 4-chlorophenol. Subsequently, the same enzyme acted upon 4-chlorophenol and converted it into phenol. Following that the enzyme phenol/toluene 2-monooxygenase facilitated the transformation of phenol into catechol. The presence of catechol 1,2-dioxygenase, muconolactone cycloisomerase, 3-oxoadipate CoA-transferase, 3-oxoadipate enol lactone hydrolase, and 3-oxoadipyl-CoA thiolase (according to KEGG annotation) indicated that the bacterium employed the ortho degradation pathway for pollutant removal.
In addition to the major genes responsible for 2,4-DCP degradation, the gene encoding the benzene/toluene/chlorobenzene dioxygenase ferredoxin component was upregulated in P. aeruginosa STV1713. This gene played a crucial role in the dioxygenation pathway of 2,4-DCP degradation by facilitating electron transfer and aiding in the initial breakdown of 2,4-DCP to 2,4-dichlorocatechol through the action of the benzene/toluene/chlorobenzene dioxygenase enzyme. Although other genes or enzymes associated with the dioxygenation pathway were not detected in the current study, the upregulation of the benzene/toluene/chlorobenzene dioxygenase ferredoxin component gene suggested that P. aeruginosa STV1713 might be employing alternative pathways, potentially involving dioxygenation mechanisms, for efficient 2,4-DCP degradation. Other common upregulated genes identified during the degradation of phenol and 2,4-DCP include genes coding putative transposase, acetyl esterase, and NADH-dependent peroxiredoxin subunit C. These proteins lack characterization within known metabolic pathways and hierarchies, and their specific roles in phenolic pollutant degradation remain unknown. However, their similar upregulation in both samples suggests their potential involvement in phenolic pollutant degradation, which requires further extensive research for confirmation.
The transcriptome analysis findings provided additional support for the initial identification of metabolic pathways established through GC/MS analysis. During the degradation of phenol and 2,4-DCP, GC/MS analysis successfully detected all the key intermediates of the ortho degradation pathway. Subsequently, transcriptome analysis elucidated the enzymes responsible for catalyzing the transformations of these intermediates.
Furthermore, a wide array of transcriptional regulators played crucial roles in the process of phenol and 2,4-DCP biodegradation. These included members from various families, such as LysR, AraC, TetR, GntR, and Fis family transcriptional regulators. The LysR family transcriptional regulators have been identified as activators responsible for enhancing the expression of genes associated with the ortho-cleavage pathway (β-ketoadipate pathway) including the catB, catC, and catA genes (Gerischer 2002). In this study, LysR family transcriptional regulators exhibited a similar up-regulation pattern. Preliminary data also indicated the involvement of AraC family transcriptional regulators in the control of catabolic operons, typically acting as transcription activators when chemical effector molecules were present (Gu et al. 2017). Additionally, the TetR family showed down-regulation in our study, aligning with the known role of TetR-type regulators as repressors for genes related to aromatic compound degradation (Eaton 1997). Members of the GntR family were previously assumed to act as repressors, controlling the degradation of aromatic compounds when the pathway substrate was absent (Morawski et al. 2000). Our study corroborated this notion by detecting a down-regulation of GntR family transcriptional regulators. Next, Fis family regulators were found to be up regulated in the present study. The role of Fis regulators, known as global regulators of transcription in Escherichia coli, in the context of phenol biodegradation remains to be further elucidated. Similarly, the LuxR-type regulators, typically functioning as transcriptional activators, are of increasing interest for their precise function in phenol metabolism.
Upregulation of other important genes in P. aeruginosa STV1713 under phenol and 2,4-DCP stresses
Significant upregulation in the transcription of genes encoding key ribosome regulatory proteins, such as rimP, rpsL, rp1O, and rp1A, was observed during phenol and 2,4-DCP degradations. This upregulation promoted the regulation of ribosome biosynthesis and helped to reduce the toxicity of phenol and 2,4-DCP by promoting the synthesis of key enzymes and proteins involved in their degradation (Gu et al. 2017). Similarly, under stress, we observed a significant upregulation of chemotaxis pathway genes in phenol and 2,4-DCP samples compared to the control. This included the sensor kinase gene cheA, the response regulator protein gene cheY, and the regulatory genes cheD, cheR, motA, and motB. Differential expression of flagellar biosynthesis genes such as fliH, fliP, flhA, flhB, flil, flgE, and fliI were also observed along with the chemotaxis genes. All these genes maintained elevated expression levels to enhance the strain’s mobility and substrate degradation capacity. The RNA-seq analysis revealed three distinct, well-annotated two-component regulatory systems (TCS) that displayed higher expression patterns. These trends in expression suggested their diverse functions for these pathways in response to phenol and 2,4-DCP stress. Specifically, the EnvZ/OmpR system, known for its role in pressure-responsive regulation in bacteria, exhibited significantly higher gene expression levels during pollutant degradation, implying its involvement in enhancing tolerance to phenol stress. Similarly, the LuX/FixJ system, responsible for responding to oxygen concentration, also exhibited a high expression pattern. Additionally, the upregulation of the NtrC/DctB system, which governs dicarboxylate acid transport regulation, suggests that it may contribute to the tolerance of the bacterium under high phenol and 2,4-DCP induced stress (Ma et al. 2020).
Furthermore, high expression of genes associated with membrane transport, such as benE, lptA, macB, and lptC, could be related to the physiological status and stress resistance of cells. benE is primarily associated with the uptake of benzoate or similar substances. Lipopolysaccharide (LPS) is a crucial component of the outer membrane in Gram-negative bacteria, and its regulation can be influenced by various environmental factors, including phenol and 2,4-DCP-induced stresses. lptA and lptC are two protein-coding genes that transfer and assemble LPS (Ma et al. 2020). They are typically expressed in a stress-free environment, but their high expression was observed under phenol and 2,4-DCP-induced stresses. Therefore, increased LPS production contributed to strengthening the integrity of the bacterial outer membrane against phenolic stress.
Discussion
Phenol and 2,4-DCP are two ubiquitous antimicrobial pollutants in the environment. Strong antimicrobial properties, structural characteristics, and toxicity of these compounds make them resistant to microbial degradations (Oluwasanu 2018; Pinheiro et al. 2018). However, some bacteria found in oil-contaminated soil were found to have capabilities for degrading phenolic pollutants (Hossain et al. 2022). Oil-contaminated soil is a rich source of phenol-degrading microbes due to the selective pressure exerted by petroleum-derived hydrocarbons, including phenolic compounds. These specialized microorganisms have evolved the genetic capacity to break down and metabolize complex hydrocarbons (Hossain et al. 2022). Therefore, the current study used oil-contaminated soil to isolate phenol and 2,4-DCP degrading bacteria. The bacterium was identified as a novel strain of P. aeruginosa, STV1713.
Similarly, several Pseudomonas bacterial species have been characterized to degrade phenolic pollutants effectively (Abarian et al. 2019; Mahgoub et al. 2023). Their enzymes have low specificity, and metabolic pathways demonstrate high convergence, enabling simultaneous utilization of similar substances without unwanted genetic coding for enzyme induction (Xu et al. 2021). However, only a few species (pure culture) displayed degradation capabilities at higher concentrations of phenol and 2,4-DCP (only a few Pseudomonas species demonstrated degradation abilities above 1500 mg/L phenol, while none of them could degrade 2,4-DCP above 1000 mg/L) (Kargi and Eker 2005; Hasan and Jabeen 2015). In the present study, P. aeruginosa STV1713 was able to degrade the highest concentrations of phenol up to 2000 mg/L and 2,4-DCP up to 1400 mg/L. Additionally, the bacterium showed good tolerance capacity against phenol up to 2100 mg/L and 2,4-DCP up to 1500 mg/L.
The pollutant removal capabilities of bacteria greatly depend on their enzyme activities. Also, various factors, such as pH, temperature, carbon source, and nitrogen source, can exert a huge influence on bacterial activity and pollutant removal (Lee et al. 2020). In the current study, parameters such as pH, temperature, and NH4NO3 were found to influence the activity of the phenol hydroxylase enzyme. Phenol hydroxylase is the first enzyme to catalyze the initial conversion of most of the phenolic compounds into catechol (Asimakoula et al. 2023). Therefore, these parameters were further optimized for cell growth and degradation time by the RSM-CCD method (Lee et al. 2020). A combination of pH 7.0, a temperature of 32.5 °C, and an NH4NO3 concentration of 0.7 g/L was found to promote maximum cell growth during phenol and 2,4-DCP degradation while reducing the degradation time. Also, maximum phenol hydroxylase activities for phenol and 2,4-DCP grown samples were observed under the optimum conditions. In line with the current study, Subramaniam et al. (2020) conducted the optimization of maximum phenol degradation by Arthrobacter sp. strain AQ15 under different conditions of pH, temperature, NaCl, and (NH4)2SO4. The results showed that maximum phenol degradation was favored at conditions of 0.15 g/L (NH4)2SO4, 0.13 g/L NaCl, pH 7.25, and incubation at 15 °C. When comparing these results, it was evident that each bacterium has specific growth requirements for their growth and survival; also, these growth requirements can greatly affect their functional capabilities.
In bacteria, phenol and 2,4-DCP, after an initial breakdown into catechol, are further degraded via two metabolic pathways: the ortho-cleavage pathway and the meta-cleavage pathway (R. Sasi and Suchithra 2023). Therefore, the detection of metabolic intermediates formed during degradation can be helpful for the rapid detection of the metabolic pathways involved (Panigrahy et al. 2020). In this study, GC/MS analysis of metabolic pathway intermediates revealed the crucial metabolites formed during phenol and 2,4-DCP degradations. Catechol, cis-cis muconic acid, and succinate were the major intermediates detected during phenol degradation, while 4-chlorophenol, phenol, cic-cis muconic acid, and fumaric acid were detected during 2,4-DCP degradation. The presence of cis-cis muconic acid in both samples indicates the utilization of the ortho degradation pathway by the bacterium through the activity of catechol 1,2-dioxygenase enzyme for pollutant removal (Asimakoula et al. 2023). Furthermore, the detection of succinate and fumarate provided evidence that the intermediates of the ortho pathway ultimately reached the Krebs cycle (Prochownik and Wang 2021). Similarly, Hasan and Jabeen (2015) used P. aeruginosa (IES-Ps-1) and Bacillus subtilis (IES-B) during the degradation of phenol. The GC/MS analysis during degradation studies revealed an increasing concentration of catechol after 24 h of degradation. Later, when more than 95% of phenol and 99% of catechol were degraded, traces of cis-muconate were observed.
Furthermore, understanding the intricate mechanisms of pollutant degradation is essential for developing efficient bioremediation strategies. Transcriptome analysis is a powerful tool that enables researchers to study the global gene expression profiles of organisms under specific conditions (Gu et al. 2021). It can provide insights into the molecular mechanisms of microbial degradation of phenolic pollutants. In the current study, P. aeruginosa STV1713 was subjected to transcriptome analyses during phenol and 2,4-DCP degradations. The findings revealed that the bacterium used multiple mechanisms simultaneously to remove the pollutants. Upregulation of major enzyme coding genes related to ortho-degradation (Gu et al. 2017) and protocatechuate pathways (Daisuke et al. 2009) were observed during phenol degradation. Furthermore, genes and proteins related to ortho-degradation involving the reduced dehalogenation mechanism and evidence for a dioxygenation mechanism were identified during 2,4-DCP degradation. Previous studies have shown that the dehalogenation mechanism is rare in aerobic bacteria and specific for highly adapted bacteria (Mohn and Kennedy 1992). Similarly, the upregulation of the gene that codes benzene/toluene/chlorobenzene dioxygenase ferredoxin component indicates that the bacterium might be employing a unique dioxygenation mechanism for 2,4-DCP removal.
Moreover, the tolerance mechanisms of P. aeruginosa STV1713 under high phenol and 2,4-DCP-induced stresses were analyzed during transcriptome analysis. It revealed the upregulation of major genes related to ribosome biogenesis, flagellar biosynthesis, chemotaxis proteins, membrane proteins, and bacterial two-component systems. Ribosomes function as the sites where protein synthesis occurs, and they typically modulate their activity in response to environmental stress by regulating the synthesis of ribosomal proteins (Zhu and Dai 2020). Similarly, motility and chemotaxis play multiple roles in the collective behaviors of bacteria, including stress response (Colin et al. 2021). The two-component system refers to a fundamental regulatory mechanism that allows bacteria to sense and respond to changes in their environment (Hirakawa et al. 2020). Additionally, the overexpression of membrane transport genes such as benE and LPS synthesis genes contribute to membrane stability in gram-negative bacteria under stress conditions (Li et al. 2019).
In line with the current investigation, a study by Gu et al. ( 2017) focused on the biodegradation of phenol by immobilized Acinetobacter sp. DW-1 cells and analyzed the proteome and transcriptome during degradation. The results showed that phenol was degraded primarily by the ortho pathway, induced by phenol hydroxylase and catechol 1,2-dioxygenase. The Fis family transcriptional regulator was identified as a likely activator of gene expression. Similarly, Gu et al. (2018) analyzed the transcriptome of Rhodococcus sp. CS-1 during phenol degradation. They found that Rhodococcus sp. CS-1 can degrade phenol through the catechol and protocatechuate branch of the β-ketoadipate pathway. This study identified some new biomarkers potentially involved in phenol biodegradation.
Table 2 compares the phenol and 2,4-DCP degradation capabilities of P. aeruginosa STV1713 with other reported phenolic pollutant degraders.
In summary, all experiments conducted in the present study demonstrate that the bacterium P. aeruginosa STV1713 is an ideal candidate for wastewater treatment processes due to its exceptional tolerance and degradation capabilities. Transcriptome analysis has revealed that the bacterium is genetically evolved to tolerate and degrade high-strength phenolic pollutants. However, further investigation is needed to understand the contributions of the unclassified genes that have been identified. Characterizing these genes may provide better insights into the bacterial degradation mechanisms under high phenol stress. Furthermore, time-dependent transcriptome analysis can offer deeper insights into the tolerance and degradation mechanisms of phenol and 2,4-DCP by bacteria.
Conclusion
In summary, this study demonstrated the ability of P. aeruginosa STV1713 to tolerate higher phenol and 2,4-DCP concentrations up to 2100 mg/L and 1500 mg/L, respectively. Statistical optimization based on phenol hydroxylase activity revealed that pH, temperature, and NH4NO3 concentration greatly influenced pollutant degradation time and cell growth. Transcriptome analysis showed that the mechanisms by which cells degraded higher concentrations of phenol and 2,4-DCP were complex processes involving the timely expression of various genes. KEGG pathway analysis revealed evidence of a dioxygenation mechanism in P. aeruginosa STV1713 for the removal of 2,4-DCP. Apart from this, phenol and 2,4-DCP metabolisms depended on different branches of the ortho-degradation pathway, which may be a novel phenolic pollutant degradation characteristic of microbes. Therefore, this finding may contribute to a better understanding of the self-regulation and degradation mechanisms of bacteria under phenol and 2,4-DCP-induced stresses. It may also provide future directions to enhance the efficiency of strains for application in wastewater treatment.
Data availability
The data supporting this study’s findings are available from the corresponding author, Suchithra Tharamel Vasu, upon reasonable request.
References
Abarian M, Hassanshahian M, Esbah A (2019) Degradation of phenol at high concentrations using immobilization of Pseudomonas putida P53 into sawdust entrapped in sodium-alginate beads. Water Sci Technol 79:1387–1396. https://doi.org/10.2166/wst.2019.134
Aisami A, Yasid N, Shukor Y (2020) Effect of temperature and ph on phenol biodegradation by a newly identified Serratia sp. AQ5–03. Open J Educ Res 1:28–43. https://doi.org/10.52417/ojbr.v1i1.57
Asimakoula S, Marinakos O, Tsagogiannis E, Koukkou A-I (2023) Phenol degradation by pseudarthrobacter phenanthrenivorans Sphe3. Microorganisms 11:524. https://doi.org/10.3390/microorganisms11020524
Barik M, Das CP, Kumar Verma A et al (2021) Metabolic profiling of phenol biodegradation by an indigenous Rhodococcus pyridinivorans strain PDB9T N-1 isolated from paper pulp wastewater. Int Biodeterior Biodegradation 158:105168. https://doi.org/10.1016/j.ibiod.2020.105168
Cafaro V, Izzo V, Scognamiglio R et al (2004) Phenol hydroxylase and toluene/o-xylene monooxygenase from pseudomonas stutzeri OX1: interplay between two enzymes. Appl Environ Microbiol 70:2211–2219. https://doi.org/10.1128/AEM.70.4.2211-2219.2004
Chen L, Li C, Xu B et al (2019) Microbial degradation of organic pollutants in groundwater related to underground coal gasification. Energy Sci Eng 7:2098–2111. https://doi.org/10.1002/ese3.415
Chris Felshia S, AshwinKarthick N, Thilagam R, Gnanamani A (2020) Elucidation of 2, 4-dichlorophenol degradation by Bacillus licheniformis strain SL10. Environ Technol 41:366–377. https://doi.org/10.1080/09593330.2018.1498923
Colin R, Ni B, Laganenka L, Sourjik V (2021) Multiple functions of flagellar motility and chemotaxis in bacterial physiology. FEMS Microbiol Rev 45:1–19. https://doi.org/10.1093/femsre/fuab038
Daisuke K, Toshihiro F, Tomokuni A et al (2009) Uncovering the protocatechuate 2,3-cleavage pathway genes. J Bacteriol 191:6758–6768. https://doi.org/10.1128/jb.00840-09
Dankaka SM, Muhammad JB, Usman S et al (2023) Phenol biodegradation by Acinetobacter baumanii and Citrobacter sedlakii isolated from petroleum products contaminated environment. Case Stud Chem Environ Eng 8:100468. https://doi.org/10.1016/j.cscee.2023.100468
Eaton RW (1997) p-Cymene catabolic pathway in Pseudomonas putida F1: cloning and characterization of DNA encoding conversion of p-cymene to p-cumate. J Bacteriol 179:3171–3180. https://doi.org/10.1128/jb.179.10.3171-3180.1997
Farag AM, Fawzy A, El-Naggar MY, Ghanem KM (2021) Biodegradation and enhancement of 2,4-dichlorophenol by marine halophilic Bacillus subtilis AAK. Egypt J Aquat Res 47:117–123. https://doi.org/10.1016/j.ejar.2021.04.005
Favier L, Ungureanu CV, Simion AI et al (2021) Enhancing the biodegradation efficiency of a emergent refractory water pollutant by a bacterial isolate through a statistical process optimization approach. Process Saf Environ Prot 148:1133–1145. https://doi.org/10.1016/j.psep.2021.02.012
Gerischer U (2002) Specific and global regulation of genes associated with the degradation of aromatic compounds in bacteria. J Mol Microbiol Biotechnol 4:111–121
Gu Q, Wu Q, Zhang J et al (2017) Acinetobacter sp. DW-1 immobilized on polyhedron hollow polypropylene balls and analysis of transcriptome and proteome of the bacterium during phenol biodegradation process. Sci Rep 7:1–18. https://doi.org/10.1038/s41598-017-04187-6
Gu Q, Wu Q, Zhang J et al (2018) Isolation and transcriptome analysis of phenol-degrading bacterium from carbon–sand filters in a full-scale drinking water treatment plant. Front Microbiol 9:21–62. https://doi.org/10.3389/fmicb.2018.02162
Gu Q, Chen M, Zhang J et al (2021) Genomic analysis and stability evaluation of the phenol-degrading bacterium Acinetobacter sp. DW-1 during water treatment. Front Microbiol 12:1–13. https://doi.org/10.3389/fmicb.2021.687511
Hasan SA, Jabeen S (2015) Degradation kinetics and pathway of phenol by Pseudomonas and Bacillus species. Biotechnol Biotechnol Equip 29:45–53. https://doi.org/10.1080/13102818.2014.991638
Hirakawa H, Kurushima J, Hashimoto Y, Tomita H (2020) Progress overview of bacterial two-component regulatory systems as potential targets for antimicrobial chemotherapy. Antibiotics 9:635. https://doi.org/10.3390/antibiotics9100635
Hossain MF, Akter MA, Sohan MSR et al (2022) Bioremediation potential of hydrocarbon degrading bacteria: isolation, characterization, and assessment. Saudi J Biol Sci 29:211–216. https://doi.org/10.1016/j.sjbs.2021.08.069
Kargi F, Eker S (2005) Kinetics of 2,4-dichlorophenol degradation by Pseudomonas putida CP1 in batch culture. Int Biodeterior Biodegradation 55:25–28. https://doi.org/10.1016/j.ibiod.2004.06.010
Krastanov A, Alexieva Z, Yemendzhiev H (2013) Microbial degradation of phenol and phenolic derivatives. Eng Life Sci 13:76–87. https://doi.org/10.1002/elsc.201100227
Lee GLY, Zakaria NN, Convey P et al (2020) Statistical optimisation of phenol degradation and pathway identification through whole genome sequencing of the cold-adapted Antarctic bacterium, Rhodococcus Sp. Strain AQ5-07. Int J Mol Sci 21:1–20. https://doi.org/10.3390/ijms21249363
Lee GL, Zakaria NN, Futamata H et al (2022) Metabolic pathway of phenol degradation of a cold-adapted Antarctic bacteria. Arthrobacter Sp Catalysts 12:1422. https://doi.org/10.3390/catal12111422
Li Y, Orlando BJ, Liao M (2019) Structural basis of lipopolysaccharide extraction by the LptB2FGC complex. Nature 567:486–490. https://doi.org/10.1038/s41586-019-1025-6
Lowry O, Rosebrough N, Farr AL, Randall R (1951) Protein measurement with the folin phenol reagent. J Biol Chem 193:265–275. https://doi.org/10.1016/S0021-9258(19)52451-6
Lucchini JJ, Corre J, Cremieux A (1990) Antibacterial activity of phenolic compounds and aromatic alcohols. Res Microbiol 141:499–510. https://doi.org/10.1016/0923-2508(90)90075-2
Ma Y, Li L, Awasthi MK et al (2020) Time-course transcriptome analysis reveals the mechanisms of Burkholderia sp. adaptation to high phenol concentrations. Appl Microbiol Biotechnol 104:5873–5887. https://doi.org/10.1007/s00253-020-10672-2
Mahgoub SA, Qattan SYA, Salem SS et al (2023) Characterization and biodegradation of phenol by Pseudomonas aeruginosa and Klebsiella variicola strains isolated from sewage sludge and their effect on soybean seeds germination. Molecules 28:1203. https://doi.org/10.3390/molecules28031203
Mahon CR, Lehman DC (2019) Textbook of diagnostic microbiology. Elsevier Health Sciences, Saunders
Mohd Ariff MA, Abdullah N (2019) Optimization of reflux extraction for cat’s whiskers leaves extract using response surface methodology. Chem Ind Chem Eng Q 26:24. https://doi.org/10.2298/CICEQ190228024A
Mohn WW, Kennedy KJ (1992) Reductive dehalogenation of chlorophenols by Desulfomonile tiedjei DCB-1. Appl Environ Microbiol 58:1367–1370. https://doi.org/10.1128/aem.58.4.1367-1370.1992
Morawski B, Segura A, Ornston LN (2000) Repression of Acinetobacter vanillate demethylase synthesis by VanR, a member of the GntR family of transcriptional regulators. FEMS Microbiol Lett 187:65–68. https://doi.org/10.1111/j.1574-6968.2000.tb09138.x
Nor Suhaila Y, Ramanan RN, Rosfarizan M et al (2013) Optimization of parameters for improvement of phenol degradation by Rhodococcus UKMP-5M using response surface methodology. Ann Microbiol 63:513–521. https://doi.org/10.1007/s13213-012-0496-6
Oluwasanu AA (2018) Fate and Toxicity of chlorinated phenols of environmental implications: a review. Med Anal Chem Int J 2:000126. https://doi.org/10.23880/macij-16000126
Panigrahy N, Barik M, Sahoo RK, Sahoo NK (2020) Metabolic profile analysis and kinetics of p-cresol biodegradation by an indigenous Pseudomonas citronellolis NS1 isolated from coke oven wastewater. Int Biodeterior Biodegrad 147:104837. https://doi.org/10.1016/j.ibiod.2019.104837
Pinheiro PF, Menini LAP, Bernardes PC et al (2018) Semisynthetic phenol derivatives obtained from natural phenols: antimicrobial activity and molecular properties. J Agric Food Chem 66:323–330. https://doi.org/10.1021/acs.jafc.7b04418
Prochownik EV, Wang H (2021) The metabolic fates of pyruvate in normal and neoplastic cells. Cells 10:762. https://doi.org/10.3390/cells10040762
Sasi R, Suchithra TV (2023) Transcriptome analysis of a novel and highly resistant hydrocarbon-degrading bacteria during the removal of phenol and 2,4-dichlorophenol. Biomass Convers Biorefinery 11:1–21. https://doi.org/10.1007/s13399-023-04509-x
Shakya M, Lo C-C, Chain PSG (2019) Advances and challenges in metatranscriptomic analysis. Front Genet 10:453–469. https://doi.org/10.1007/978-981-15-9758-9_21
Singh P, Gupta S (2016) Optimization of biodegradation process of river water containing phenol using response surface methodology by Psuedomonas putida (P. putida ). Int J Curr Res 8:44217–44222
Subramaniam K, Shaharuddin NA, Tengku-Mazuki TA et al (2020) Statistical optimisation for enhancement of phenol biodegradation by the Antarctic soil bacterium Arthrobacter sp. strain AQ5–15 using response surface methodology py. J Environ Biol 41:1560–1569. https://doi.org/10.22438/JEB/41/6/MRN-1496
Subramaniam K, Athirrah T, Mazuki T et al (2019) Isolation and optimisation of phenol degradation by antarctic isolate using one factor at time. Malaysian J Biochem Mol Biol 1:79–86
van Schie PM, Young LY (2000) Biodegradation of phenol: mechanisms and applications. Bioremediat J 4:1–18. https://doi.org/10.1080/10588330008951128
Xu N, Qiu C, Yang Q et al (2021) Analysis of phenol biodegradation in antibiotic and heavy metal resistant Acinetobacter lwoffii NL1. Front Microbiol 12:725–755. https://doi.org/10.3389/fmicb.2021.725755
Zhou J, Yu X, Ding C et al (2011) Optimization of phenol degradation by Candida tropicalis Z-04 using Plackett-Burman design and response surface methodology. J Environ Sci 23:22–30. https://doi.org/10.1016/S1001-0742(10)60369-5
Zhu M, Dai X (2020) Bacterial stress defense: the crucial role of ribosome speed. Cell Mol Life Sci 77:853–858. https://doi.org/10.1007/s00018-019-03304-0
Funding
Kerala State Council for Science Technology and Environment (Sasthra Bhavan, Pattom, Thiruvananthapuram, Kerala, India-695004) (Research Fellowship-Sanction order No.29/FSHP/2016/KSCSTE dated 24.03.2017).
Author information
Authors and Affiliations
Contributions
All authors, RS and STV, contributed to the study's conception and design. RS performed material preparation, data collection, and experimental analyses. RS wrote the first draft of the manuscript, and all authors commented on previous versions. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
Not applicable
Consent for publication
Not applicable
Competing interests
The authors declare no competing interests.
Additional information
Responsible Editor: Robert Duran
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Sasi, R., Tharamel Vasu, S. Revealing the degradation mechanisms of the hyper-tolerant bacterium Pseudomonas aeruginosa STV1713 under high phenol and 2,4-DCP-induced stress conditions through RNA-seq analysis. Environ Sci Pollut Res 31, 5625–5640 (2024). https://doi.org/10.1007/s11356-023-31500-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11356-023-31500-w