Abstract
2,4,6-trinitrotoluene (TNT) is a nitroaromatic compound that causes soil and groundwater pollution during manufacture, transportation, and use, posing significant environmental and safety hazards. In this study, a TNT-degrading strain, Bacillus cereus strain T4, was screened and isolated from TNT-contaminated soil to explore its degradation characteristics and proteomic response to TNT. The results showed that after inoculation with the bacteria for 4 h, the TNT degradation rate reached 100% and was transformed into 2-amino-4,6-dinitrotoluene (2-ADNT), 4-amino-2,6-dinitrotoluene (4-ADNT), 2,4-diamino-6-nitrotoluene (2,4-DANT), and 2,6-diamino-4-nitrotoluene (2,6-DANT), accompanied by the accumulation of nitrite and ammonium ions. Through proteomic sequencing, we identified 999 differentially expressed proteins (482 upregulated, 517 downregulated), mainly enriched in the pentose phosphate, glycolysis/gluconeogenesis, and amino acid metabolism pathways. In addition, the significant upregulation of nitroreductase and N-ethylmaleimide reductase was closely related to TNT denitration and confirmed that the strain T4 converted TNT into intermediate metabolites such as 2-ADNT and 4-ADNT. Therefore, Bacillus cereus strain T4 has the potential to degrade TNT and has a high tolerance to intermediate products, which may effectively degrade nitroaromatic pollutants such as TNT in situ remediation in combination with other bacterial communities.
Graphical Abstract

Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
2,4,6-Trinitrotoluene (TNT), which is stable and cost-effective, has become one of the most widely used explosives. Approximately 1,000 tons of TNT are produced globally, and nearly two million liters of TNT-contaminated wastewater and other nitroaromatic compounds are released daily. These wastewater seriously pollute surface water and groundwater under the action of surface runoff and infiltration, and pose a threat to the health and ecosystem of surrounding residents (Su et al. 2023). The US Environmental Protection Agency has set the environmental discharge standard for TNT at 0.06 mg/L and the drinking water discharge standard at 2.0 μg/L. However, TNT contamination in soil and water surrounding military factories and ammunition disposal sites exceeds these standards significantly (Lewis et al. 2004). Long-term exposure to TNT can cause skin irritation, liver dysfunction, anemia and induce carcinogenic and mutagenic effects in humans (Leffler et al. 2014; Hsu et al. 2019; Ni et al. 2022; Lin et al. 2023). TNT pollution can also disrupt the structure of soil microbial communities (Yang et al. 2022) and induce damage to plant root systems (Johnston et al. 2015; Das et al. 2017; Yang et al. 2021). The issue of environmental TNT pollution has garnered considerable attention.
Microbial remediation technology has been widely used in the remediation of TNT-contaminated soil and water due to its lack of secondary pollution. Several highly efficient TNT-degrading microorganisms have been reported recently. For example, Pseudomonas sp. TNT3, isolated and screened from an Antarctic environmental sample, could transform 100 mg/L of TNT within 48 h, demonstrating higher biotransformation capacity than the well-known TNT-degrading strain Pseudomonas putida KT2440 (Cabrera et al. 2020). Two mildly halophilic alkaline bacterial strains from TNT-contaminated soil have been isolated (Ali-Begloui et al. 2020), which could rapidly remove up to 200 mg/L of TNT within 24 h under aerobic conditions. However, existing strains face problems such as poor environmental adaptability, low TNT degradation rates, and incomplete TNT degradation during in situ soil remediation processes. Enzymatic degradation of TNT can effectively avoid the toxicity of TNT degradation products on microbial metabolism. This method has the advantages of simple operation, low cost, and strong environmental adaptability. Previous studies have shown that the degradation of TNT mainly involves the reduction of nitro groups. The nitro group of TNT is reduced to the nitroso group, hydroxy amino group, and amino group under the action of nitro reductase or xanthate family enzymes (OYEs), forming amino aromatic derivatives (Williams et al. 2004; Alothman et al. 2020). However, the enzyme production capacity and enzymatic properties of existing TNT-degrading bacteria are still unclear.
Therefore, this study aims to isolate and domesticate highly efficient TNT-degrading strains from TNT-contaminated soil in a Chinese ammunition disposal plant and analyze their TNT degradation ability and degradation pathway. We used proteomic technology to study the protein composition of TNT-degrading bacteria, identify critical proteins involved in TNT degradation, and reveal the key pathways of microbial TNT degradation. In addition, we screened out the key protease in the process of TNT degradation and interpreted the molecular mechanism of TNT biodegradation through proteome changes. The results of this study provide theoretical references for the subsequent construction of TNT-degrading enzyme expression vectors.
Materials and methods
Isolation and Identification of TNT-degrading Strains
Soil samples were collected from a munitions destruction plant in northern China. 5 g of soil was added to 95 mL of Luria–Bertani liquid medium (LB) (consisting of 10 g/L tryptone, 5 g/L yeast extract, 10 g/L NaCl) containing 100 mg/L of TNT and shaken at 37 ℃, and 150 rpm for 24 h (The average pollution concentration of TNT in some ammunition demolition sites are 100 mg/L)(Zhang et al. 2023). 5 mL of the culture was then transferred to 95 mL of Nitrogen-free medium (consisting of glucose 10 g/L, KH2PO4 0.2 g/L, MgSO4·7H2O 0.2 g/L, NaCl 0.2 g/L, CaSO4·2H2O 0.2 g, pH = 7) containing 100 mg/L of TNT and passaged for three consecutive times. The culture was then streaked on a Nitrogen-free solid medium containing 50 mg/L of TNT to isolate and purify single colonies. Genomic DNA was extracted using a bacterial DNA extraction kit, and the bacterial 16S rDNA sequence was amplified using universal primers (27F: 5'-AGAGTTTGATCCTGGCTCAG-3; 1492R: 5'-GGTTACCTTGTTACGACTT-3). Beijing Qink Bio-tech Co., Ltd sequenced the PCR products. The sequencing data were subjected to a Blast search in the NCBI database, and a phylogenetic tree was constructed using MEGA7.0 (Kalyan et al. 2022).
Phenotypic analysis of strain
The strain was inoculated in LB medium and cultured at 37 ℃ and 150 rpm for 4 h. A small amount of bacterial liquid was taken and air-dried on a microscope slide for observation of cell morphology under a microscope. Gram staining was also performed, and the bacterial cells were observed under an oil-immersion lens. The strains were streaked on Nitrogen-free solid medium containing different concentrations of TNT (0, 25, 50, and 100 mg/L) and cultured at 37 ℃ for 12 h. The colony morphology was observed, and a small number of bacterial cells were spread on a cover slip and fixed with electron microscopy fixative (4% glutaraldehyde, phosphate). The samples were dehydrated with a gradient of ethanol solutions (30, 50, 70, 80, 90, and 95%) for 15 min at each concentration and observed under a scanning electron microscope (Liu et al. 2013).
Growth and TNT Degradation Analysis of Strain
The strain was inoculated in the Nitrogen-free medium containing different concentrations of TNT (0, 25, 50, and 100 mg/L) and shaken at 37 ℃ and 150 rpm for 12 h. 1 mL of the culture was collected at regular intervals, and the bacterial biomass was estimated by measuring the optical density at 600 nm. The culture was centrifuged at 12,000 rpm for 5 min, and collect the supernatant. TNT content in the culture medium was determined spectrophotometrically by N-cetyl pyridine-sodium sulfite (Cabrera et al. 2020).
Single-factor experiment and response surface optimization
Single-factor experiment
A single-factor experiment was conducted to investigate the effects of different factors on the strain's biomass and TNT degradation rate. The specific methods are as follows: the strain was inoculated into the Nitrogen-free medium containing 100 mg/L TNT at different pH (4, 5, 6, 6.5, 7, 7.5, 8, 9, 10), and the initial OD600 was adjusted to 0.05. The culture was shaken at 37 °C and 150 rpm for 4 h, and then 1 mL of the culture was taken into a centrifuge tube to measure the biomass and TNT degradation rate. The strain was inoculated into Nitrogen-free medium containing 100 mg/L TNT at pH 7.0, and the initial OD600 was adjusted to 0.05. The culture was shaken at different temperatures (10, 20, 25, 30, 35, 40, and 45 °C) and 150 rpm for 4 h, and then 1 mL of the culture was taken into a centrifuge tube to measure the biomass and TNT degradation rate. The strain was inoculated into Nitrogen-free medium containing 100 mg/L TNT at pH 7.0, and the initial OD600 was adjusted to (0.05, 0.10, 0.15, 0.20, 0.25, 0.30, 0.35). The culture was shaken at 37 °C, and at 150 rpm for 4 h, 1 mL of the culture was taken into a centrifuge tube to measure the biomass and TNT degradation rate.
Response surface optimization
Based on the results of the single-factor experiment, the optimal degradation conditions for the strain were further optimized using response surface methodology (Li et al. 2019). DesignExpert 8.0.6 software was used for experimental design and statistical analysis, and a Box-Behnken design with three factors and three levels was employed. Based on the experimental design, the binomial equation obtained was shown in the supplementary materials.
TNT degradation dynamic analysis
The strain was inoculated separately into Nitrogen-free medium containing 0 and 100 mg/L TNT and was cultured at 37 °C with shaking at 150 rpm for 12 h. The medium containing 100 mg/L TNT without inoculated strain was used as a control. At every 2 h interval, 2 mL of culture was collected and centrifuged at 12,000 rpm for 5 min. Nitrite content was determined using a nitrite content detection kit (Solarbio, Beijing) for soil and water. Ammonium nitrogen content was determined using a Soil ammonium nitrogen content detection kit (Solarbio, Beijing). The amount of TNT and its intermediates were determined by high-performance liquid chromatography (HPLC) according to US Environmental Protection Agency (USEPA) method 8330A (Chien et al. 2014; Cabrera et al. 2020). 500 μL of the supernatant was extracted with an equal volume of acetonitrile. After filtering with a 0.22 μm microporous membrane, the extract was transferred to a chromatographic vial. The liquid chromatography parameters were as follows: mobile phase A was water, and mobile phase B was methanol; the flow rate was 0.7 mL/min, and the sample injection volume was 10 μL. Detection was performed at 33 °C and 254 nm.
TMT labeled quantitative proteomics analysis
Bacterial culture and sample preparation: The strains were inoculated separately into Nitrogen-free medium containing 0 and 100 mg/L TNT and were cultured at 37 °C with shaking at 150 rpm for 4 h. 10 mL of culture was collected and centrifuged at 8,000 rpm for 5 min. The pellets were washed three times with phosphate buffer and then resuspended in phosphate buffer. The resuspended cells were transferred to 2 mL grinding tubes and centrifuged again. The supernatant was discarded, and the pellets were immediately frozen in liquid nitrogen and stored at -80 °C. Each group of samples was divided into four replicates.
Protein extraction, digestion, and TMT labeling: After grinding the samples into fine powder in liquid nitrogen, the powder was added to lysis buffer, vortexed for 10 s, and placed on ice for 10 min. This process was repeated 3–4 times. After lysis, the cell lysate was centrifuged at 4 °C and 12,000 × g for 30 min. The supernatant was collected, and the total protein concentration was determined using the BCA protein assay kit (Bio-Rad, USA). The protein quality was detected by SDS-PAGE gel electrophoresis. The protein solution was then digested with trypsin (Promega, Madison, WI) and freeze-dried. TMT labeling was performed using the TMT labeling kit (ThermoScientific)(Wang et al. 2022).
LC–MS/MS Analysis: Each collected peptide was reconstituted, separated, and subjected to LC–MS/MS analysis. Reverse phase chromatography conditions were used where the sample was loaded onto an Acclaim PepMap RSLC analytical column, 75 μm × 50 cm (RP-C18, Thermo Fisher), at a 300 nL/min flow rate. The mobile phase A consisted of H2O-FA (99.9:0.1, v/v), while the mobile phase B consisted of ACN-H2O-FA (80:19.9:0.1, v/v/v). The gradient elution conditions were as follows: 0–50 min, 2–28% B; 50–60 min, 28–42% B; 60–65 min, 42–90% B; 65–75 min, 90% B. Mass spectrometry conditions were set with a resolution of 60,000 for the first MS scan, automatic gain control of 1 × 106, and a maximum injection time of 50 ms. Full scan MS/MS spectra were acquired using a data-dependent acquisition mode in positive ion mode, with high energy collision-induced dissociation (HCD) at a collision energy of 36. MS/MS resolution was set at 30,000 with automatic gain control of 1 × 105, and the maximum injection time of ions was 80 ms. Dynamic exclusion time was set at 30 s.
Bioinformatics Analysis: The OmicShare bioinformatics learning platform (www.omicshare.com/tools) was used to analyze the protein data matrix. Principal component analysis (PCA) was performed to detect the degree of variation within and between sample groups. All identified proteins were annotated by searching the GO and KEGG databases. Significant enrichment of GO functions and KEGG pathways were determined using differentially expressed proteins (fold change ≥ 1.2). Detailed proteomic experimental methods are provided in the supplementary materials.
Statistical analysis
Column and line graphs were created using Origin 2022, Statistical Analysis was performed using IBM SPSS Statistics 22.0 (IBM, USA), and image layout and design were done using Adobe Illustrator 2022 (Adobe, USA).
Results
Isolation of strains from the soil for TNT biodegradation
A strain capable of efficiently degrading TNT was isolated and purified by streaking the culture on Nitrogen-free solid medium containing 100 mg/L TNT and incubating at 37 ℃ for 3 days. The strain T4 is a Gram-positive, aerobic, short rod-shaped strain (Fig. 1a). 16S rDNA sequencing results showed that strain T4 has the closest phylogenetic relationship with the Bacillus genus, with a similarity of 99.9% to Bacillus cereus (Fig. 1b). Under different concentrations of TNT (0–100 mg/L) and incubation at 37 ℃ for 12 h, the strain appeared unaffected based on strain morphology observations (Fig. 1c). Based on the bacterial morphology, and 16S rDNA phylogenetic Analysis, strain T4 was preliminarily identified as Bacillus cereus. The 16S rRNA data have been submitted to NCBI and can be accessed in the GenBank under the accession number OR564170.
Bacterial growth and TNT degradation analysis
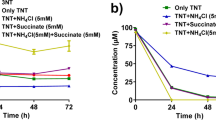
As the TNT concentration increased from 0 to 100 mg/L, the growth rate of strain T4 gradually decreased. When the TNT pollution concentration was 25–100 mg/L, the OD600 of strain T4 after 4 h of growth was 0.18–0.42, representing a decrease of 36.7%-70% compared with the control group (Fig. 2a). The results of TNT degradation efficiency showed that after 4 h of growth, the residual TNT in the culture medium was significantly reduced, and the TNT degradation rate of strain T4 reached 92.56%-100% (Fig. 2b-d).
Growth of strain T4 and its degradation efficiency of TNT. Note: (a) Growth curves of strain T4 under different concentrations of TNT treatment after 4 h of cultivation; (b) Growth and TNT degradation of strain T4 under 25 mg/L TNT treatment; (c) Growth and TNT degradation of strain T4 under 50 mg/L TNT treatment; (d) Growth and TNT degradation of strain T4 under 100 mg/L TNT treatment. Different letters indicate significant differences at the 0.05 level (n = 3)
Response surface model optimization of strain T4 for TNT degradation
Single-factor experiments showed that the biomass of strain T4 and TNT degradation rate increased first and then decreased with pH ranging from 4 to 10. When the pH was 6–7, the OD600 of strain T4 reached 0.1–0.39 after 4 h of growth, and the TNT degradation rate reached 2.6–98.6% (Fig. 3a). As the cultivation temperature increased from 10 to 45 ℃, the biomass of strain T4 and TNT degradation rate increased first and then decreased. When the temperature was 35 ℃, the OD600 of strain T4 reached 0.25 after 4 h of growth, and the TNT degradation rate reached 98.7% (Fig. 3b). With the increase of inoculation amount (OD600: 0.05–0.35), the biomass of strain T4 gradually increased. When the inoculation amount reached 0.1–0.35, the TNT degradation rate of strain T4 reached 99.7–100% after 4 h of cultivation (Fig. 3c).
Effects of pH, temperature, and initial OD600 on the growth and TNT degradation of strain T4. Note: (a) The effect of pH on the growth and TNT degradation of strain T4 (initial OD600 of 0.05, temperature of 37 ℃, incubation for 4 h); (b) The effect of temperature on the growth and TNT degradation of strain T4 (initial OD600 of 0.05, initial pH of 7.0, incubation for 4 h); (c) The effect of initial OD600 on the growth and TNT degradation of strain T4 ( temperature of 37 ℃, initial pH of 7.0, incubation for 4 h). Different letters indicate significant differences at the 0.05 level (n = 3)
Based on the results of the single-factor experiments, we determined the appropriate ranges of the three main factors that affect TNT degradation: initial pH of 6–8, incubation temperature of 20–40 ℃, and initial OD600 of 0.05–0.15. We conducted experiments according to the Box-Behnken design and measured the TNT content in the culture after 4 h of incubation. We calculated the TNT degradation rate, used the Design-Expert software to fit a quadratic equation and response surface to the final results (Table S1), and performed statistical analysis on the obtained data (Table S2). The small p value of the quadratic polynomial model indicates that the predictive model is significant. The set parameters for the optimization conditions can well reflect the effects of the three factors on the TNT degradation process. The adjusted coefficient of determination (Adj R2) and coefficient of determination (R2) were 0.8083 and 0.9161, respectively, indicating that the model can predict the optimal values of the three factors well.
The three-factor interaction effects on the TNT degradation process were visualized using surface and contour plots created using Design-Expert software (Fig. S1a-c, Fig. S2). The results showed that the interaction between pH and temperature significantly impacted the TNT degradation rate.
As shown in Attachment S1(d-f), the typical probability plot of residuals displayed a roughly linear pattern, indicating that the residuals were normally distributed. The scatter plot of residuals vs TNT degradation rate showed a random distribution. The linear fit plot demonstrated a strong correlation between the predicted and experimental values of the TNT degradation rate. These results indicate that the response surface model fits the data well.
Based on the software prediction results, the optimal cultivation conditions for strain T4 to degrade TNT were determined to be: initial pH of approximately 7, cultivation temperature of 37 ℃, and initial OD600 of 0.13.
TNT degradation and intermediate product analysis by strain T4
According to the optimized cultivation parameters, the initial pH was adjusted to 7.0, the cultivation temperature was 37 ℃, the inoculum was 0.13, and the cultivation was carried out for 12 h. The bacteria in the control group showed exponential growth from 0 to 4 h, and the OD600 reached 1.1 after 12 h. With the extension of cultivation time, the bacterial density in the TNT treatment group gradually increased, and the OD600 reached 0.7 after 12 h (Fig. 4a). Liquid chromatography analysis showed that the TNT degradation rate reached 100% after 4 h of cultivation with inoculated strain T4 (Fig. S3h). Dynamic Analysis of TNT degradation showed that the intermediate metabolites of TNT biodegradation, 2-ADNT, 4-ADNT, 2,6-DANT, and 2,4-DANT gradually appeared in the liquid chromatography spectrum (Fig. S3 g-l). After 4 h of cultivation, the levels of 2,4-DANT and 2,6-DANT continued to increase, and the content of 2,6-DANT reached 25.74 mg/L (Fig. 4b). After 12 h of cultivation, 2-ADNT was not detected in the chromatogram, and 4-ADNT showed a trend of first increasing and then decreasing. Some of the 2,4-DANT and 2,6-DANT may be produced by converting 2-ADNT and 4-ADNT.
Compared with the control group, after inoculation with strain T4, nitrite and ammonium nitrogen content in the culture medium gradually increased with the extension of cultivation time (0–12 h), indicating that the TNT degradation products participated in bacterial nitrogen metabolism (Fig. 4c,d).
TMT proteomic identification of critical proteins involved in TNT degradation by strain T4
As shown in Figure S4, TMT proteomic Analysis identified 4082 credible proteins. The positions of T0 and T1 on the score plot are relatively far apart, with significant separation in the first and second principal components, with differences of 78.38% and 6.96%, indicating significant differences in credible protein expression levels between the control group T0 without TNT and the experimental group T1 with 100 mg/L TNT. Compared with the T0 group, proteins with Fold change ≥ 2 or Fold change ≤ 1/2 and p-value < 0.05 were considered differentially expressed proteins. 999 differentially expressed proteins were identified, including 482 upregulated and 517 downregulated proteins.
GO enrichment pathways showed that under TNT exposure, the functional proteins of strain T4 were annotated in the GO database as cell component (CC) > molecular function (MF) > biological process (BP). At the BP classification level, the protein expression of UDP-N-acetylglucosamine biosynthetic process, pentose-phosphate shunt, and heme biosynthetic process involved in strain T4 induced by TNT exposure was upregulated. At the CC classification level, the relative abundance of proteins related to the cytoplasm, plasma membrane, and tricarboxylic acid cycle enzyme complex in strain T4 induced by TNT exposure was significantly upregulated. At the MF classification level, upregulated proteins in strain T4 were involved in FMN binding, NADP binding, and electron transfer activity. Down-regulated proteins were involved in metal ion binding, oxidoreductase activity, structural proteins molecule activity, and metalloendopeptidase activity (Fig. S5).
KEGG enrichment pathways showed that differential proteins were mainly enriched in metabolic pathways, including carbohydrate metabolism pathways (55 upregulated, 42 down-regulated), amino acid metabolism pathways (29 upregulated, 48 down-regulated), cofactor and vitamin metabolism pathways (28 upregulated, 14 down-regulated), and energy metabolism pathways (20 upregulated, 17 down-regulated). Among them, the significant up-regulation of proteins in strain T4 involved in the phosphogluconate pathway, glycolysis/gluconeogenesis pathway, and nitrotoluene degradation pathway related to TNT degradation may be one of the critical pathways for the degradation of TNT by this strain(Fig. 5).
KEGG pathway analysis of differentially expressed proteins. Note: (a) Distribution of differentially expressed proteins at KEGG Level 2; (b) KEGG enrichment bubble chart. Detailed information on KEGG enrichment pathways is provided in Table S4
Nitrotoluene degradation pathway and nitrogen metabolism network analysis
Differentially expressed proteins in the nitrotoluene degradation pathway are shown in Table S5. Proteins related to oxygen-insensitive nitroreductase (NADPH), nitroreductase, N-ethylmaleimide reductase, olefin reductase, N-ethylmaleimide reductase, and proteins with nitroreductase domain showed significant upregulation. Nitrate/nitrite transporters, nitrate reductase, and nitric oxide dioxygenase were significantly downregulated in the nitrogen metabolism pathway. In contrast, the expression of formyl glutamate synthase and hydroxylamine reductase, which are related to ammonia production, was down-regulated. However, glutamine synthetase, which synthesizes glutamine from ammonia, was significantly upregulated, and glutamate dehydrogenase showed upregulation and downregulation (Fig. 6).
Nitrogen metabolism network analysis of strain T4 under TNT exposure. Note: Detailed protein expression information related to nitrogen metabolism pathways can be found in Table S6
Discussion
In this study, we isolated a highly efficient TNT-degrading strain T4, from soil contaminated with TNT for an extended period. The ability of this microorganism to degrade TNT is significantly better than that of previously reported microbial strains. In the early stages of culture, strain T4 consumed much energy for TNT degradation and primary metabolism. After 4 h, the TNT degradation rate was up to 100% (Fig. S3), and strain T4 began to grow normally, entering the logarithmic phase. We identified the optimal degradation conditions for strain T4 through response surface optimization, which increased its TNT degradation rate by 33.22%. It may be because strain T4 could better express proteins related to TNT degradation under optimal culture conditions. The dynamic analysis of TNT degradation showed that after 4 h of culture in Nitrogen-free medium containing 100 mg/L TNT, strain T4 had a 100% TNT degradation rate. Then TNT was converted to 2-ADNT, 4-ADNT, 2,4-DANT, and 2,6-DANT, followed by nitrite accumulation (Fig. 4b,c). Nitrite accumulation is closely related to TNT conversion, confirming the production of intermediate products such as 2-ADNT (Ziganshin et al. 2007; Mercimek et al. 2015; Khilyas et al. 2017). Studies have shown that intermediate metabolites of TNT also have substantial toxicity to organisms (Sims and Steevens 2008; Khan et al. 2013). In this study, we found that in the presence of intermediate products such as 2-ADNT, the growth of the strain was not significantly inhibited, indicating that strain T4 also has a strong tolerance to intermediate products during TNT degradation. However, compared with the control group without TNT, the ammonium content decreased, which is significantly different from previous reports (Lamba et al. 2021). We believe this may be related to the differential expression of essential proteins in the nitrogen metabolism pathway. In addition, Yang (Yang et al. 2019) found that TNT and its intermediates entered cells through passive transport, and the membrane affinity of monoamine metabolites 2-ADNT and 4-ADNT is more significant than that of diamino metabolites 2, 4-DANT and 2,6-DANT, which was consistent with the findings in Fig. 4b.
Aerobic bacteria typically initiate the biotransformation of TNT by reducing one of its nitro groups (Lin et al. 2013). We identified intermediates of TNT metabolism using HPLC analysis, such as 2-ADNT (Fig. S3). Studies have shown that the catalysis of critical enzymes is essential for TNT biotransformation by bacterial strains. Many proteases are involved in TNT biotransformation, mainly including nitroreductases, the flavin monooxygenase family, and other enzymes such as nitrate reductases, ferredoxin-NADP+ reductases, and sulfite reductase (Watrous et al. 2003; Caballero et al. 2005; van Dillewijn et al. 2008). It has been reported that some strains possess dual pathways: nitro reduction (reducing the nitro group in TNT to hydroxyl or amino) and denitration (reducing the aromatic ring of TNT to release nitrite and form Mason's complex)(Smets et al. 2007). To reveal the degradation mechanism of strain T4 towards TNT and identify the vital degradative proteins, this study employed TNT proteomics technology to analyze the response of strain T4 proteome to TNT exposure. In the nitrotoluene degradation pathway, the N-ethylmaleimide reductase in nitroreductase and flavin monooxygenase family was significantly upregulated, indicating that strain T4 may possess dual pathways. In addition, nitrite accumulation confirmed the increase in nitroreductase expression in strain T4. The down-regulation of nitrate/nitrite transporters and nitrate reductase expression in the nitrogen metabolism pathway may be related to nitrite accumulation during TNT nitroreduction. The down-regulation of hydroxylamine reductase and up-regulation of glutamine synthetase led to a decrease in ammonia content, consistent with the results in Fig. 4d, and may be due to the decreased activity of hydroxylamine reductase induced by nitrite accumulation (Hucklesby and Hageman 1976).
Moreover, glutamine synthetase assimilates ammonia into glutamic acid, a key enzyme in nitrogen metabolism. Thus, the increase in glutamine synthetase expression promotes ammonia assimilation (Bernard and Habash 2009). Studies have shown that the pentose phosphate pathway, glycolysis/gluconeogenesis, and biosynthesis of macromolecules are closely related, including the production of NADPH, the biosynthesis of nucleotides, and the synthesis of amino acids (Christodoulou et al. 2018; Zhang et al. 2020). We found that under TNT exposure, key enzymes such as glycolysis/gluconeogenesis, 6-phosphofructokinase in the pentose phosphate pathway, glyceraldehyde-3-phosphate dehydrogenase, and 6-phosphogluconate dehydrogenase were significantly upregulated in the carbohydrate metabolism pathway of strain T4. A large amount of energy was used to synthesize macromolecules such as carbohydrates, indicating that the synthetic metabolism was more significant than the catabolic metabolism (Fig. 5). It may be that after the complete degradation of TNT, strain T4 began to consume energy for their growth and reproduction, which is also confirmed by the exponential growth of strain T4 after 4 h (Fig. 4a).
Based on the Analysis, the degradation pathway of TNT by strain T4 is illustrated in Fig. 7. The first step of TNT biodegradation is to convert it into 4-hydroxyamino-2,6-dinitrotoluene (4-HDNAT) or 2-hydroxyamino-4,6-dinitrotoluene (2-HDNT), which is mediated by nitroreductase and N-ethylmaleimide reductase. After these reactions, 4-ADNT or 2-ADNT is formed by oxygen-insensitive nitroreductase. Under the catalysis of different proteases, intermediate products such as 2,4-DANT, and 2,6-DANT are further produced. After multiple reaction steps, TAT is deaminated and transformed into toluene, which is opened, and finally, succinyl-CoA and Acetyl-CoA are generated, which enter the tricarboxylic acid cycle to provide a material basis for bacterial energy metabolism (Williams et al. 2004; Lin et al. 2013; Serrano-González et al. 2018; Lamba et al. 2021). Strain T4 has the potential to degrade TNT and exhibits strong tolerance to intermediate products such as 2-ADNT. Therefore, in situ, remediation of nitroaromatic pollutants such as TNT can be efficiently achieved by coacting strain T4 with other microbial communities.
Conclusion
This study isolated a highly efficient TNT-degrading strain T4, from TNT-contaminated soil and identified it as Bacillus cereus. Under optimal culture conditions (initial pH 7.0, cultivation temperature 37 ℃, and initial OD600 of 0.13), the degradation rate of TNT reached 100% after 4 h of cultivation. TMT proteomics analysis showed that the significant enrichment of proteins related to carbohydrate metabolism, energy metabolism, and other pathways was closely related to the tolerance and biodegradation of TNT by strain T4. In addition, nitroreductase and N-ethylmaleimide reductase, which can convert TNT to intermediate products such as 2-ADNT and 4-ADNT, were significantly upregulated in the TNT degradation pathway, consistent with the HPLC analysis results, further confirming the high efficiency of TNT degradation by strain T4.
Data availability
The data covered in this manuscript are available from the corresponding author on reasonable request.
References
Ali-Begloui M, Salehghamari E, Sadrai S et al (2020) Biotransformation of Trinitrotoluene (TNT) by Newly Isolated Slight Halophilic Bacteria. Microbiology 89:616–625. https://doi.org/10.1134/S0026261720050033
Alothman ZA, Bahkali AH, Elgorban AM et al (2020) Bioremediation of Explosive TNT by Trichoderma viride. Molecules 25:1393. https://doi.org/10.3390/molecules25061393
Bernard SM, Habash DZ (2009) The importance of cytosolic glutamine synthetase in nitrogen assimilation and recycling. New Phytol 182:608–620. https://doi.org/10.1111/j.1469-8137.2009.02823.x
Caballero A, Lazaro JJ, Ramos JL, Esteve-Nunez A (2005) PnrA, a new nitroreductase-family enzyme in the TNT-degrading strain Pseudomonas putida JLR11. Environ Microbiol 7:1211–1219. https://doi.org/10.1111/j.1462-2920.2005.00801.x
Cabrera MA, Márquez SL, Quezada CP et al (2020) Biotransformation of 2,4,6-Trinitrotoluene by Pseudomonas sp. TNT3 isolated from Deception Island, Antarctica. Environ Pollut 262:113922. https://doi.org/10.1016/j.envpol.2020.113922
Chien C-C, Kao C-M, Chen D-Y et al (2014) Biotransformation of trinitrotoluene (TNT) by Pseudomonas spp. isolated from a TNT-contaminated environment: Biotransformation of TNT by Pseudomonas spp. Environ Toxicol Chem 33:1059–1063. https://doi.org/10.1002/etc.2553
Christodoulou D, Link H, Fuhrer T et al (2018) Reserve Flux Capacity in the Pentose Phosphate Pathway Enables Escherichia coli’s Rapid Response to Oxidative Stress. Cell Syst 6:569-578.e7. https://doi.org/10.1016/j.cels.2018.04.009
Das P, Sarkar D, Datta R (2017) Proteomic profiling of vetiver grass ( Chrysopogon zizanioides ) under 2,4,6-trinitrotoluene (TNT) stress. GeoHealth 1:66–74. https://doi.org/10.1002/2017GH000063
Hsu D-W, Wang T-I, Huang D-J et al (2019) Copper promotes E. coli laccase-mediated TNT biotransformation and alters the toxicity of TNT metabolites toward Tigriopus japonicus. Ecotoxicol Environ Saf 173:452–460. https://doi.org/10.1016/j.ecoenv.2019.02.056
Hucklesby DP, Hageman RH (1976) Hydroxylamine Reductase Enzymes from Maize Scutellum and Their Relationship to Nitrite Reductase. Plant Physiol 57:693–698. https://doi.org/10.1104/pp.57.5.693
Johnston EJ, Rylott EL, Beynon E et al (2015) Monodehydroascorbate reductase mediates TNT toxicity in plants. Science 349:1072–1075. https://doi.org/10.1126/science.aab3472
Kalyan VSRK, Meena S, Karthikeyan S, Jawahar D (2022) Isolation, screening, characterization, and optimization of bacteria isolated from calcareous soils for siderophore production. Arch Microbiol 204:721. https://doi.org/10.1007/s00203-022-03322-w
Khan MI, Lee J, Park J (2013) A toxicological review on potential microbial degradation intermediates of 2,4,6-trinitrotoluene, and its implications in bioremediation. KSCE J Civ Eng 17:1223–1231. https://doi.org/10.1007/s12205-013-0305-1
Khilyas IV, Lochnit G, Ilinskaya ON (2017) Proteomic Analysis of 2,4,6-Trinitrotoluene Degrading Yeast Yarrowia lipolytica. Front Microbiol 8:2600. https://doi.org/10.3389/fmicb.2017.02600
Lamba J, Anand S, Dutta J et al (2021) Study on aerobic degradation of 2,4,6-trinitrotoluene (TNT) using Pseudarthrobacter chlorophenolicus collected from the contaminated site. Environ Monit Assess 193:80. https://doi.org/10.1007/s10661-021-08869-7
Leffler P, Brännäs E, Ragnvaldsson D et al (2014) Toxicity and Accumulation of Trinitrotoluene (TNT) and its Metabolites in Atlantic Salmon Alevins Exposed to an Industrially Polluted Water. J Toxicol Environ Health A 77:1183–1191. https://doi.org/10.1080/15287394.2014.920756
Lewis TA, Newcombe DA, Crawford RL (2004) Bioremediation of soils contaminated with explosives. J Environ Manage 70:291–307. https://doi.org/10.1016/j.jenvman.2003.12.005
Li H, van den Driesche S, Bunge F et al (2019) Optimization of on-chip bacterial culture conditions using the Box-Behnken design response surface methodology for faster drug susceptibility screening. Talanta 194:627–633. https://doi.org/10.1016/j.talanta.2018.10.048
Lin H, Yu C-P, Chen Z (2013) Aerobic and anaerobic biodegradation of TNT by newly isolated Bacillus mycoides. Ecol Eng 52:270–277. https://doi.org/10.1016/j.ecoleng.2012.11.004
Lin D, Chen Y, Liang L et al (2023) Effects of exposure to the explosive and environmental pollutant 2,4,6-trinitrotoluene on ovarian follicle development in rats. Environ Sci Pollut Res 30:96412–96423. https://doi.org/10.1007/s11356-023-29161-w
Liu Q, Chen J, Munyaneza JE et al (2013) Scanning electron microscopy and in vitro cultivation of endophytic bacteria from potato tubers afflicted with zebra chip disease. Can J Plant Pathol 35:192–199. https://doi.org/10.1080/07060661.2013.785446
Mercimek HA, Dincer S, Guzeldag G et al (2015) Degradation of 2,4,6-trinitrotoluene by P. aeruginosa and characterization of some metabolites. Braz J Microbiol 46:103–111. https://doi.org/10.1590/S1517-838246120140026
Ni S, Zhang H, Sun L et al (2022) Transgenerational reproductive toxicity of 2,4,6-trinitrotoluene (TNT) and its metabolite 4-ADNT in Caenorhabditis elegans. Environ Toxicol Pharmacol 92:103865. https://doi.org/10.1016/j.etap.2022.103865
Serrano-González MY, Chandra R, Castillo-Zacarias C et al (2018) Biotransformation and degradation of 2,4,6-trinitrotoluene by microbial metabolism and their interaction. Def Technol 14:151–164. https://doi.org/10.1016/j.dt.2018.01.004
Sims JG, Steevens JA (2008) The role of metabolism in the toxicity of 2,4,6-trinitrotoluene and its degradation products to the aquatic amphipod Hyalella azteca. Ecotoxicol Environ Saf 70:38–46. https://doi.org/10.1016/j.ecoenv.2007.08.019
Smets BF, Yin H, Esteve-Nuñez A (2007) TNT biotransformation: when chemistry confronts mineralization. Appl Microbiol Biotechnol 76:267–277. https://doi.org/10.1007/s00253-007-1008-7
Su K, Wang Y, Liu Y et al (2023) Screening of Bacillus altitudinis D47 from TNT red water-contaminated soil for highly dinitrotoluene sulfonate efficient biodegradation. J Clean Prod 407:137103. https://doi.org/10.1016/j.jclepro.2023.137103
van Dillewijn P, Wittich R-M, Caballero A, Ramos J-L (2008) Subfunctionality of Hydride Transferases of the Old Yellow Enzyme Family of Flavoproteins of Pseudomonas putida. Appl Environ Microbiol 74:6703–6708. https://doi.org/10.1128/AEM.00386-08
Wang Y-Y, Yan J-K, Ding Y, Ma H (2022) Effects of ultrasound on the thawing of quick-frozen small yellow croaker (Larimichthys polyactis) based on TMT-labeled quantitative proteomic. Food Chem 366:130600. https://doi.org/10.1016/j.foodchem.2021.130600
Watrous MM, Clark S, Kutty R et al (2003) 2,4,6-Trinitrotoluene Reduction by an Fe-Only Hydrogenase in Clostridium acetobutylicum. Appl Environ Microbiol 69:1542–1547. https://doi.org/10.1128/AEM.69.3.1542-1547.2003
Williams RE, Rathbone DA, Scrutton NS, Bruce NC (2004) Biotransformation of Explosives by the Old Yellow Enzyme Family of Flavoproteins. Appl Environ Microbiol 70:3566–3574. https://doi.org/10.1128/AEM.70.6.3566-3574.2004
Yang H, Li H, Liu L et al (2019) Molecular Simulation Studies on the Interactions of 2,4,6-Trinitrotoluene and Its Metabolites with Lipid Membranes. J Phys Chem B 123:6481–6491. https://doi.org/10.1021/acs.jpcb.9b03033
Yang X, Zhang Y, Lai J et al (2021) Analysis of the biodegradation and phytotoxicity mechanism of TNT, RDX, HMX in alfalfa (Medicago sativa). Chemosphere 281:130842. https://doi.org/10.1016/j.chemosphere.2021.130842
Yang X, Lai J, Zhang Y, Luo X (2022) Reshaping the microenvironment and bacterial community of TNT- and RDX-contaminated soil by combined remediation with vetiver grass (Vetiveria ziznioides) and effective microorganism (EM) flora. Sci Total Environ 815:152856. https://doi.org/10.1016/j.scitotenv.2021.152856
Zhang W, Chen X, Sun W et al (2020) Escherichia coli Increases its ATP Concentration in Weakly Acidic Environments Principally through the Glycolytic Pathway. Genes 11:991. https://doi.org/10.3390/genes11090991
Zhang H, Zhu Y, Wang S et al (2023) Spatial-vertical variations of energetic compounds and microbial community response in soils from an ammunition demolition site in China. Sci Total Environ 875:162553. https://doi.org/10.1016/j.scitotenv.2023.162553
Ziganshin AM, Gerlach R, Borch T et al (2007) Production of Eight Different Hydride Complexes and Nitrite Release from 2,4,6-Trinitrotoluene by Yarrowia lipolytica. Appl Environ Microbiol 73:7898–7905. https://doi.org/10.1128/AEM.01296-07
Acknowledgements
This study was financially supported by the Outstanding Talent Program of the Academy of Military Sciences (ZQ2021-01) and the National Natural Science Foundation of China (22176222, 22106182).
Author information
Authors and Affiliations
Contributions
Maoling Yin: Investigation, Data curation, Writing-original draft; Jinglong Lai: Writing-original draft, Writing-review & editing; Xu Yang: Investigation; Yongbing Zhu: Investigation; Bin Dong: Investigation; Yu Zhang: Writing-review & editing; Sanping Zhao: Conceptualization, Investigation, Supervision, Project administration.
Corresponding author
Ethics declarations
Ethics approval
Not applicable.
Consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare no competing interests.
Additional information
Responsible Editor: Gerald Thouand
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Yin, Ml., Zhao, Sp., Lai, Jl. et al. Oxygen-insensitive nitroreductase bacteria-mediated degradation of TNT and proteomic analysis. Environ Sci Pollut Res 30, 116227–116238 (2023). https://doi.org/10.1007/s11356-023-30568-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11356-023-30568-8