Abstract
With these growing and evolving years, antimicrobial resistance has become a great subject of interest. The idea of using natural productive ways can be an effective measure against antimicrobial resistance. The growing prevalence of antimicrobial resistance indicates that advanced natural approaches are a topic of concern for fighting the resistance. Many natural products including essential oils, flavonoids, alkaloids and botanicals have been demonstrated as effective bactericidal agents. In this review, we will discuss in detail about the relevance of such natural products to tackle the problem of antimicrobial resistance, antibiotic adjuvants that aim towards non-essential bacterial targets to reduce the prevalence of resistant bacterial infections, latest bioinformatics approach towards antibacterial drug discovery along with an understanding of biogenic nanoparticles in antimicrobial activity.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
One of the emerging reasons of mortality and morbidity is antimicrobial resistance (AMR) these days which is generally regarded as non-responsiveness of pathogens for antimicrobial treatment regime; however, mainly broad spectrum bacterial resistance is a rising concern all over the world (Marianne et al. 2017). Hospitalar antibiotic resistance refers to the capacity to overcome previous antibiotic treatment doses given to susceptible individuals and is mostly focused on the minimum inhibitory concentration (MIC), whereas, environmental antibiotic resistance can be defined as a decreased susceptibility to an antibiotic compared with other strains of the same species. If bacteria resist an antibiotic notwithstanding its concentration, it is considered environmentally resistant (Larsson and Flach 2021). AMR has been observed to give rise to an increasing number of deaths (Fig. 1), extended infirmary stay and increased health care costs. In 2019, Murray et al. have documented that an estimated 4.95 million individuals died of bacterial AMR, with 1.27 million of those deaths associated with bacterial AMR. At the geographical level, the rate of all-age mortality due to resistance was estimated to be highest in western Sub-Saharan Africa (2.73 deaths per 100,000) and lowest in Australasia (6.5 deaths per 100,000). In 2019, the six most common bacteria causing mortality associated with resistance included Streptococcus pneumoniae, Staphylococcus aureus, Escherichia coli, Pseudomonas aeruginosa, Klebsiella pneumoniae and Acinetobacter baumannii causing 929,000 deaths (Murray et al. 2022). Natural antibiotics can be defined as the class of antibiotics produced by the microorganisms themselves, such as beta-lactam antibiotics comprising penicillin; semi-synthetic antibiotics are the ones produced from natural products, such as elavancin and dalbavancin and lastly synthetic antibiotics are the ones which are chemically synthesized in industries based on the structure of the natural products (Demain 2009). Studies have shown some of the natural products which have therapeutic properties and that can be used to combat AMR such as tannins, flavonoids, botanicals, essential oils, alkaloids and many more (WHO 2018). Leading causes of AMR includes overuse of the antimicrobials due to easy access, which leads to overuse (Michael et al. 2014); the second cause includes inappropriate prescribing which leads to patient exposure to complications of antimicrobials (Lushniak, 2014); and the third cause is extensive agricultural use; it is when humans consume food, they ingest antibiotics with it (Golkar et al. 2014). Some of the essential determinants of AMR are efflux pumps, β-lactamases and many more. Basically, the mechanism of AMR includes enzyme inactivation, modification of the antibiotic target site, overproduction of the target, replacement of the target site, efflux and reduced permeability (Manar Ali et al., 2020).
Natural products against spread of AMR
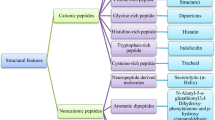
Infectious diseases are a leading cause of death around the world. Drug-resistant microorganisms are a major public health concern. To combat AMR and prevent the spread of infectious illnesses, new tactics are required (Shuman 2011). Antimicrobials are found in medicinal plants. For primary health, more than half of the world’s population is reliant on medicinal plants. These plants consist of a diverse chemical armory such as essential oils, phenolics, polyphenols, flavonoids, quinones, tannins, coumarins and alkaloids (Saleem et al. 2009).
Essential oils
These are combinations of volatile chemicals produced by aromatic plants’ secondary metabolism. Bioactive characteristics can be found in essential oils from Asteraceae plants (Yap et al. 2014). Essential oils contain chemicals that have medical and pharmacological characteristics. Various plant sources of Asteraceae species are used in the extraction of essential oils, and they possess inhibitory action against various bacterial species as demonstrated in Table 1 (Phan Canh et al. 2020).
Botanicals
Herbs show a spectrum of biological activity and can be used effectively for illness management. The existence of active compounds, such as flavonoids, quinones, phenols, alkaloid, essential oil, terpenoids, tannins, lignans and some secondary metabolites is related to antimicrobial activities in plants. The peptides that comprise their defence systems are other antibacterial agents of plants. Botanicals can provide potential sources of new antibiotics for medical and scientific communities (Veličković et al. 2003). It was demonstrated that the minimum inhibitory concentrations (MIC) of beta-lactam antibiotics for the methicillin-resistant Staphylococcus aureus was reduced by corilagin, a compound in Arctostaphylos. Corilagin possessed MIC value equivalent to 128 μg/mL. However, at 16 μg/mL, MIC of oxacillin against methicillin-resistant Staphylococcus aureus strains, gave fractional inhibitory concentration (FIC) index value of 0.13 (Shimizu et al. 2001).
Phenols and polyphenols
These are the large class of biologically active secondary metabolites of plants including flavonoids, anthocyanins, tannins, stilbenes, coumarins, phenolic acids, lignans and lignins (Pereira et al. 2009). Multiple antibacterial processes have been described in phenolic compounds such as morin, kaempferol and galangin. These compounds engage with bacterial proteins and components of the cell wall, can degrade cytoplasmic membranes, decrease membrane fluidity and hinder the synthesis of nucleic acids and cell wall, or energy metabolization (Daglia 2012). Moreover, morin is an inhibitor of sortases A and B (membrane cysteine transpeptidases found in bacteria) at sub-MIC levels. There are evidences of this action at the cellular level which indicate that morin reduced the binding of the whole S. aureus cells to fibrinogen, one of the host ligands to which pathogens adhere upon infections (Cushnie and Lamb, 2011).
Flavonoids
Flavonoids are characterized by a phenylbenzopyran chemical structure. Three different modes of antibacterial activity of flavonoids include nucleic acid synthesis inhibition, damage via a piercing mechanism to the cytoplasmic membrane and reduction in membrane fluidity and lastly through the inhibition of energy metabolism (Avila et al. 2008). They also exhibit anti-biofilm activities such as Red Wine which was found to be effective in the suppression of S. aureus-biofilm formation (Cho et al. 2015). Xantohumol was discovered to prevent S. aureus adhesion and biofilm formation from prenylated Humulus lupulus chalconoid. Bacteria were also inactivated in previously formed biofilms, most likely by compromising the stability of the cytoplasmic membrane after lipid metabolism has been inhibited (Rozalski et al. 2013).
Quinones
Quinones are compounds comprising two isomers of cyclohexadienedione and consisting of a fully conjugated cyclic dion structure, for example benzoquinone. Henna (Lawsonia inermis), which is related to the presence of quinine, possesses antibacterial activity against Pseudomonas aeruginosa (Habbal et al. 2011). Hypericin, an anthraquinone, was shown to have general antimicrobial effects against methicillin-resistant and methicillin-sensitive staphylococcus as well (Dadgar et al. 2006).
Tannins
The polymeric phenolic substances, found almost in all plants, possess anti-bacterial activity both against gram-positive and gram-negative bacteria. They can permeate lipid bilayers and interact with them (Taylor et al. 2005). They may also induce membrane fusion, which leads to the leakage and aggregation of intramembrane components (Ikigai et al. 1993). Roccaro et al. demonstrated to the impact of regulation of bacterial drug resistance by catechin gallatas. Epigallocatechin gallate (EGCg) has been demonstrated to have a number of anti-bacterial activity that reduces bacterial growth and invasion and works synergistically with various antibiotics (SudanoRoccaro et al., 2004). Alnus japonica extract was the most active among plant extracts screened with quercetin and tannic acid being the key anti-S. aureus-biofilm components. It reduced the formation of biofilms via affecting genes in the development of biofilms (Lee et al. 2013).
Coumarins
The coumarins and its derivative products 5, 6-benzo-2-pyrone, which exist naturally, show a number of biological activities from photosensitization, vasodilation or analgesic qualities via good anti-inflammatory and anti-bacterial activity. In 2014, Lee et al. examined the anti-biofilm abilities of various coumarins such as esculetin, scopoletin and umbelliferone. They have indicated that coumarin and umbelliferone have had antibiotic-film forming activities without reducing planktonic cell development against enterohaemorrhagic E. coli O157:H7 (Lee et al. 2014).
Alkaloids
Alkaloids are organic heterocyclic nitrogen compound containing nitrogen and are derived from amino acid. They usually form water-soluble salts. The alkaloids are capable of intercalating with DNA. The cell division and cell death are hindered. Berberine’s mode of action is based on its capacity to interpose with DNA and disturb the membrane structure by raising bacterial membrane permeability (Peng et al. 2015). Anti-microbial action against Staphylococcus aureus, Streptococcus mutans, Microsporum gypseum, Microsporum canis and Trichophyton rubrum is detected in Hasubanalactam alkaloid isolated from Stefanian tubers (Semwal and Rawat 2009) (Fig. 2).
Targeting major microbial drug resistance determinants
Efflux pumps
Efflux pump was the first mechanism which was determined for resistance to tetracycline in Escherichia coli (Van Bambeke et al. 2000). Efflux pumps are mainly transport proteins which are present in gram-negative bacteria and gram-positive bacteria. Efflux pumps are engaged in banishing out harmful substances from the interior of the cell to the outer surrounding. These pumps work by using proton motive force as a propellant. Unicellular microorganisms contain five types of efflux transporters such as major facilitator, multidrug and toxic efflux, resistance nodulation division, small multidrug resistance and, ATP binding cassette. Currently, to enhance and reinforce the working of anti-microbials, the usage of efflux pump inhibitor (EPI) has been found effective (McMurry et al. 1980). EPI helps in inhibiting efflux pumps, and EPIs are of two types based on their mechanism of action; one is energy dissipation mechanism which enhances the activity of antibiotics and inhibits efflux pump function, as these are dependent on the proton gradient. The second type is inhibition by direct binding in this mechanism. EPI binds to the efflux pumps which decreases its strength to pair with substrates. Some of the EPIs also inhibit biofilm formation so EPI can be used to intensify antibiotic activities (Sharma et al. 2009).
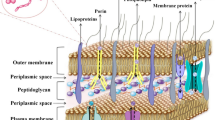
β-Lactamases
β-Lactamases are also one of the determinants of AMR. The very first enzyme was discovered in E. coli. Gram-positive bacteria produce β-Lactamases extracellularly, and gram-negative bacteria produce it in the periplasmic space. β-Lactamases work by inhibiting β-lactam antibiotics through hydrolysing the β-lactam ring and binding covalently to their carbonyl moiety (Sandanayaka and Prashad 2002). To control this, a development has been done which is β-lactamase inhibitors. These tiny molecule inhibitors of β-lactamases, such as clavulanic acid, sulbactam, and tazobactam, have been made in combination with β-lactams for treatment purposes (Bush 2017). Three of them have a similar structure as of penicillin. Clavulanate combinations have shown remarkable activity against many bacteria such as Streptococci, Staphylococcus aureus, and Bacteroides fragilis.
Biofilm formation
Biofilm formation is a complex process, in which microbes stick and breed to form extracellular polymers of polysaccharides, proteins, enzymes, and nucleic acids that in return helps in linking and matrix formation (Macià et al., 2014). Biofilm formation and beta-lactamases synergistically cause dissemination of multidrug resistant strains of gram-negative bacteria. A cross-sectional study was performed by Dumaru et al. 2019 to recognize the production of biofilms by gram-negative bacteria, and hence evaluate their anti-biogram, as well as to assess the production of extended-spectrum beta-lactamases and metallo-beta-lactamases. The results revealed that there was a significant association between metallo-beta-lactamases and biofilm formation (Dumaru et al. 2019). They are known to be responsible for recurrent infections, more number of deaths, and some serious issues (Sanchez et al. 2013). To get rid of biofilms and to reduce the spread of AMR, the study of natural products is increasing day by day around the world (Lewis and Ausubel 2006).
QS
Quorum sensing (QS), which is fundamentally detecting thickness, keeps up physiological behaviours in microscopic organisms. This component is displayed in both gram-positive and gram-negative microscopic organisms. These systems are divided into three types. One of the QS systems with acyl-homoserine lactone (AHL) as the self-inducible molecule exists in gram-negative bacteria. The oligopeptides are QS systems that are self-inducing molecules and exist in gram-positive bacteria. The other types are QS systems that use furan borate diesters as self-inducing molecules and exist in gram-negative and gram-positive bacteria (Monnet and Gardan 2015).
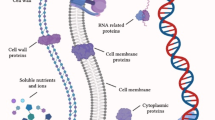
MGE
Mobile genetic elements (MGEs) are the types of moving genetic material that has the ability to change a place in chromosomes by moving between DNA molecules such as transposons, integrins and insertion sequences. They interact with each other to create a complex network with the potential to recruit and disseminate genes throughout a bacterial population. Enzymes that cut and paste or copy genetic material inside the bacterial genome, as well as between bacterial cells via horizontal gene transfer through conjugation, transduction or transformation cause DNA sequences to be transferred from chromosomal to episomal DNA and vice versa. Hence, these elements could prompt horizontal genetic exchange and spread of resistant genes and might play an important role in the spread of AMR (Partridge et al. 2018; Frost et al. 2005).
Role of bioinformatics and omics-based approach in discovery of microbial natural products in antibiotic resistance
Microbial natural products (MNPs) have long been a significant source for drug development. MNPs have played a crucial role in the development of therapeutically effective medications for infectious diseases, antioxidants, cancer, diabetes and other conditions. In recent decades, plant-derived compounds have been replaced as a source of medicinal drugs, and MNPs have emerged as a viable source for the discovery of new medications (Knight et al. 2003). Unique niche conditions comprising extreme climate, unusual landscapes and geographic location could result in different adaptations for various living species, and furthermore, these reservoirs could be used to isolate a diverse range of microorganisms with therapeutic value (Merino et al. 2019). Advances in metagenomics have resulted in a growing body of knowledge about the numerous and complex microbial communities that live in various habitats including lakes, rivers, sediments and extreme environmental conditions such as hydrothermal vents, caves and ice cores. For instance, due to adaptations to their environment, some bacteria such as hyperacidophilic Picrophilus oshimae and Picrophilus torridus have the potential to survive in hot springs and can withstand very high temperatures, whereas on the other hand, Proteobacteria, Bacteroidetes and Actinobacteria can survive at ice surface and withstand very cold temperatures (Anesio et al. 2017). Because these bio-diverse populations have divergent and varying chemical structures of medicinal significance, they could be used as an antibiotic mining source (Grossart et al. 2020). Bioinformatics helps in lead discovery by utilising high-throughput structure determination tools that are useful for screening complicated bacterium proteins (Bansal 2005). This is one of the most important tools for combating bacterial resistance, since it allows for the identification of potential relatives, alignment of sequences, and modelling of three-dimensional structures. The various bioinformatics strategies to limit antibiotic resistance are mentioned as below (Iskandar et al. 2021).
Genome mining
Genome mining has been explored as a potential alternative for conventional drug discovery screening (Zerikly and Challis 2009). The genome mining method aids in the identification and investigation of biosynthetic gene clusters spanning from genes to molecules. Genome mining has proven to be a time-saving, user-friendly and cost-effective technique. The mining technique will enable target-directed discovery of novel bioactive metabolites by predicting the chemical structure and class using bioinformatics in combination with bioassay-guided isolation, silent gene activation and heterologous expression procedures for novel bioactive metabolites (Mohana et al. 2018).
WGS
Genome sequencing can be defined as a method to determine the entire genetic makeup of an organism. It is a very flexible method; however, it is quite expensive and could be showing less efficiency in predicting some conditions, and suitable data analysis platforms are required before introducing whole-genome sequencing (WGS) on a large scale. Moreover, many of the components necessary for clinical usage are unavailable in current WGS analysis automation systems (Köser et al. 2014). In human and agricultural research, WGS of pathogens has become a more approachable and economical technique for genotyping (Collineau et al. 2019). The entire bacteria genome can be analysed using WGS. It is being used for various pathogens including Mycobacterium tuberculosis, Salmonella campylobacter and Neisseria gonorrhoeae (Hendriksen et al. 2019). Genome sequencing is a good way to advance scientific research, especially in biomolecular modelling and designing of drugs, with a major focus on antibiotic resistance. Sequencing of deoxyribonucleic acid (DNA) is an excellent platform for protein modelling and drug development (Blundell et al. 2006). Recent advancements in sample preparation have allowed WGS directly from single bacterial colonies grown under typical diagnostic settings, eliminating the requirement for subculture to obtain enough DNA for sequencing (Wyres et al. 2020). WGS can help in the identification of novel antibiotic-producing microorganisms, and hence can be useful to study the emergence of antibiotic resistance, for surveillance, to develop diagnostic tests, monitor multidrug evolution and transmission, to discover novel anti-bacterial drugs, therapeutics and assessment of their preventability and to monitor the emergence of bacterial infectious agents in different healthcare settings (Borelli et al. 2021). It rapidly identifies the resistant mechanisms like in the case of 454 pyrosequencing, which identified F0 subunit of the ATP synthase as the target of bedaquiline, which went on to become the first member of a new class of anti-tuberculosis drugs to be approved (Zumla et al. 2013). WGS-based early identification of resistance pathways has ramifications for clinical trial design as well. For instance, it can be used to screen a phylogenetically diverse group of pathogens for the presence and variability of the drug’s target during the early stages of drug development. It will help in demonstrating that this target and its variations are genuine and significant for all species and progenitors of the pathogen, lowering the risk of overlooking any resistant strains. Moreover, the mechanism of action of a drug can be easily identified using WGS. Bottromycins, for instance, are anti-bacterial peptides that possess inhibitory actions against a variety of gram-positive bacteria and mycoplasma. Subsequently, a research indicated that the binding A-site on the 50S ribosome of these peptides inhibits protein synthesis, suggesting that they could be a new potential class of antibiotics for vancomycin-resistant Enterococci (VRE) and methicillin-resistant Staphylococcus aureus (Punina et al. 2015). Such research will demonstrate that this target and its variations are genuine and significant for all species and lineages of the pathogenic genus, lowering the risk of missing any resistant strains. More frequent dosing or higher doses could be used in clinical trials to overcome this level of resistance if resistance mechanisms are identified that only result in marginally elevated MICs relative to wild type MIC distributions. WGS is increasingly being utilised to identify exogenous reinfection from relapse of the initial infection, which is critical when evaluating the efficacy of the drug or regimens under study and has also played a role in determining the rate of resistance (Köser et al. 2014). Anti-microbial susceptibility testing using whole genome sequencing (WGS-AST) has the potential to anticipate every known resistance phenotype for a strain quickly, consistently, and accurately, while also providing rich surveillance data (Van Belkum and Dunne 2013). The procedure behind WGS-AST begins from selective culturing of the bacterium from a clinical sample, followed by direct shotgun sequencing of clinical samples to fragment DNA and assembling the reads into genomic scaffolds in silico (Su et al. 2019), as demonstrated in the figure.
Genome shuffling
Genome shuffling has been proposed as an alternative technique that might be used in complex phenotypes that were previously complicated. Genome shuffling, along with modern genetic techniques, has allowed for targeted genetic manipulation of phenotypes (Stephanopoulos 2002). The technique has proven to be effective in improving strains as well as providing insights into cellular details of the desired phenotype. Many compounds have been generated employing DNA shuffling and targeted evolution, which has proven to be a useful technique for whole-cell and metabolic engineering (Petri and Schmidt-Dannert 2004). DNA shuffling enables for faster-directed evolution and pathways engineering, which are essential for improving industrially significant microorganism strains (Zhang et al. 2002). Avilamycin, produced from Streptomyces viridochromogenes, is an approved anti-microbial agent, which inhibits the growth of multidrug-resistant gram-positive bacteria. Xa et al. demonstrated that a powerful strategy for molecular breeding of high-yield industrial strains is genome shuffling combined with ribosome engineering to improve production of avilamycin in S. viridochromogenes. The results indicated that avilamycin production of S. viridochromogenes was increased from 0.24 to 1.4 g/L after employing the method of genome shuffling with streptomycin resistance screening (Lv et al. 2013) (Fig. 3).
Combinatorial biosynthesis
Induced evolution for designer antibiotics by genetics and/or medicinal chemistry is known as combinatorial biosynthesis. The major enzymes involved in the process of chemical entity engineering by combinatorial approach include polyketide synthase (PKS), non-ribosomal peptide synthase (NRPS), thioesterase (TE) and acyl transferase (AT) (Nguyen et al. 2006).
Metabolomics
Metabolomics, as well as the closely related fields of metabolomics and metabolite profiling, is a promising field that involves the quantitative identification of several small molecule metabolites in biological systems (Gowda et al. 2008). In bacterial physiology, metabolism of cellular lipids, nucleotides, amino sugars and energy are all common pathways. The metabolome of bacterial cells does have the potential to lead to novel anti-bacterial treatment strategies (Zampieri et al. 2017). Metabolites in biological samples are considered as biomarkers of the disease and biomarkers of efficacy. By properly quantifying the array of biochemical alterations and mapping these variations to metabolic pathways, metabolomics has had a significant impact on drug discovery and development procedures (Wang et al. 2006). In comparison to genomes, transcriptomics and proteomics, this method gives data that is less detailed, precise, relevant and quantitative (Weinshilboum 2003). Natural product libraries, which enable quick characterisation of known molecules, are critical in the metabolomics technique for natural product discovery. Data libraries such as traditional Chinese medicine integrative database (TCMID), Chinese ethnic minority traditional drug database (CEMTDD), SuperToxic and NPACT have been developed. By analysing drug response and adverse reactions, as well as the variance involved in natural product biodiscovery, metabolomics has the potential to greatly aid the study of systems biology for drug revelation (Xie et al. 2015).
Advances in machine/deep learning on antibiotic resistance research
The DeepARG models were created to analyse next-generation sequencing data, such as metagenomes, computationally. The deepARG models’ primary feature is their low false negative rate during prediction. The gene-like sequence concept is also intended to discover novel antibiotic resistance genes based on sequence homology. The computational technique of comparing metagenomic DNA sequences against existing internet databases is currently used to identify antibiotic resistance genes from livestock manure, wastewater treatment plants, compost and soil. Using softwares like BLAST, Bowtie, or DIAMOND, raw reads or predicted open reading frames from compiled contigs are aligned to the database of preference, and then the groups of antibiotic resistance genes present are predicted or assigned (Arango-Argoty et al. 2018). The Comprehensive Antibiotic Resistance Database or CARD is a regulated database that provides reference DNA and protein sequences, detection models and bioinformatics techniques on the molecular basis of AMR (McArthur et al. 2013). CARD brings together molecular biology, biochemistry and bioinformatics in an ontological framework to create a database that is both practical and feasible for doctors, scientists, industry and government agencies. CARD’s main goal is to unify and standardise AMR molecular sequence understanding through expert human curation, resulting in a credible and trustworthy central database of sequences and mutations known to cause AMR (Alcock et al. 2020). HMD-ARG utilises sequence encoding to evaluate if an input sequence is an ARG, and which antibiotic class it is resistant to, mechanism behind resistance, and if it is intrinsic or acquired. The HMD-ARG database is the world’s largest and outperforms existing approaches in terms of recall, high accuracy and precision (Li et al. 2021).
Conclusion
Thus, it can be concluded that idea of botanical alliance can be an effective measure against AMR. Many natural products have been demonstrated to boost immunity which can combat AMR such as botanicals, which provide potential sources of new antibiotics for scientific communities and tannins, which reduce bacterial invasion and growth. Also we can combat AMR by directly targeting its determinants such as efflux pumps which are the main transport proteins, biofilm formation and quorum sensing which basically maintains physiological behaviours in bacteria. Moreover, bioinformatics is the most important tools for combating bacterial resistance, since it allows for the identification of potential relatives, alignment of sequences and modelling of three-dimensional structures. By focusing on these parameters, we can reduce the number of deaths which are due to AMR these days.
Future prospective
Promising strategies for future resistance to resistance are divided into five categories; each of them requires additional social investment in basic and applied research and policy activities. These interventions are generally designed to prevent infections initially, foster new economic models that stimulate investment in anti-infective treatments, delay the spread of drug resistance to extend the shelf life of antibiotics and discover new ways to directly attack microorganisms; one that does not generate resistance or change the host-microbe interacts to change the way the disease occurs without directly attacking the microbe, preventing infection eliminates the need for antibiotics. Traditional infection prevention work must be supported by new technologies that can more effectively disinfect environmental surfaces, people and food. In this hour we are in need of technologies that can achieve intensive care without the need to implant foreign objects such as plastics or metals (for example, better delivery of drugs through the intestine, skin or respiratory mucosa to replace intravenous treatment and technical regeneration of tissues, thereby avoiding need to be used for prosthetic implants). Improvements in population health and health care systems can reduce the number of admissions in hospitals and specialized nursing facilities, thereby reducing infections. Finally, the new vaccine is expected to prevent antibiotic-resistant infections.
Data availability
Not applicable.
References
Alcock BP, Raphenya AR, Lau TT, Tsang KK, Bouchard M, Edalatmand A, Huynh W, Nguyen AL, Cheng AA, Liu S, Min SY (2020) CARD 2020: antibiotic resistome surveillance with the comprehensive antibiotic resistance database. Nucleic Acids Res D1:D517–D525. https://doi.org/10.1093/nar/gkz935
Anesio AM, Lutz S, Chrismas NA, Benning LG (2017) The microbiome of glaciers and ice sheets. Npj Biofilms and Microbiomes 3(1):1–1. https://doi.org/10.1038/s41522-017-0019-0
Arango-Argoty G, Garner E, Pruden A, Heath LS, Vikesland P, Zhang L (2018) DeepARG: a deep learning approach for predicting antibiotic resistance genes from metagenomic data. Microbiome 6(1):1–5. https://doi.org/10.1186/s40168-018-0401-z
Avila HP, Smânia Ede F, Monache FD, Smânia A Jr (2008) Structure-activity relationship of antibacterial chalcones. Bioorg Med Chem 22:9790–9794. https://doi.org/10.1016/j.bmc.2008.09.064
Bansal AK (2005) Bioinformatics in microbial biotechnology–a mini review. Microb Cell Fact 4:19. https://doi.org/10.1186/1475-2859-4-19
Blundell TL, Sibanda BL, Montalvão RW, Brewerton S, Chelliah V, Worth CL, Harmer NJ, Davies O, Burke D (2006) Structural biology and bioinformatics in drug design: opportunities and challenges for target identification and lead discovery. Philos Trans R Soc Lond B Biol Sci 361(1467):413–423. https://doi.org/10.1098/rstb.2005.1800
Borelli TC, Lovate GL, Scaranello AFT, Ribeiro LF, Zaramela L, Pereira-Dos-Santos FM, Silva-Rocha R, Guazzaroni ME (2021) Combining functional genomics and whole-genome sequencing to detect antibiotic resistance genes in bacterial strains co-occurring simultaneously in a Brazilian hospital. Antibiotics (Basel) 10(4):419. https://doi.org/10.3390/antibiotics10040419
Bousaada O, Ammar S, Saidana D, Chriaa J, Chraif I, Daami M, Helal AN, Mighri Z (2008) Chemical composition and antimicrobial activity of volatile compounds from capitula and aerial parts of Rhaponticum acaule DC. growing wild in Tunisia. Microbiol Res 163:87e95
Bush K (2017) Game changers: new β-lactamase inhibitor combinations targeting antibiotic resistance in gram-negative bacteria. ACS Infect Dis 4(2):84–87. https://doi.org/10.1021/acsinfecdis.7b00243
Cavar S, Maksimovic M, Vidic D, Paric A (2012) Chemical composition and antioxidant and antimicrobial activity of essential oil of Artemisia annua L. from Bosnia. Ind Crops Prod 37:479e85
Cetin B, Ozer H, Cakir A, Mete E, Oztürk E, Polat T, Kandemir A (2009) Chemical composition of hydrodistillated essential oil of Artemisia incana (L) Druce and antimicrobial activity against foodborne microorganisms. Chem Biodivers 6:2302e10
Cho HS, Lee JH, Cho MH, Lee J (2015) Red wines and flavonoids diminish Staphylococcus aureus virulence with anti-biofilm and anti-hemolytic activities. Biofouling. 31(1):1–11. https://doi.org/10.1080/08927014.2014.991319
Collineau L, Boerlin P, Carson CA, Chapman B, Fazil A, Hetman B, McEwen SA, Parmley EJ, Reid-Smith RJ, Taboada EN, Smith BA (2019) Integrating whole-genome sequencing data into quantitative risk assessment of foodborne AMR: a review of opportunities and challenges. Front Microbiol 10:1107. https://doi.org/10.3389/fmicb.2019.01107
Cushnie TP, Lamb AJ (2011) Recent advances in understanding the antibacterial properties of flavonoids. Int J Antimicrob Agents 38(2):99–107. https://doi.org/10.1016/j.ijantimicag.2011.02.014
Dadgar T, Asmar M, Saifi A, Mazandarani M, Bayat H, Moradi A, Bazueri M, Ghaemi EA (2006) Antibacterial activity of certain Iranian medicinal plants against methicillin-resistant and sensitive Staphylococcus aureus. Asian J Plant Sci 5https://doi.org/10.3923/ajps.2006.861.866
Daglia M (2012) Polyphenols as antimicrobial agents. CurrOpinBiotechnol 23(2):174–181. https://doi.org/10.1016/j.copbio.2011.08.007
Demain AL (2009) Antibiotics: natural products essential to human health. Med Res Rev 2009(6):821–842. https://doi.org/10.1002/med.20154
Dumaru R, Baral R, Shrestha LB (2019) Study of biofilm formation and antibiotic resistance pattern of gram-negative Bacilli among the clinical isolates at BPKIHS, Dharan. BMC Res Notes 12(1):1–6. https://doi.org/10.1186/s13104-019-4084-8
Formisano C, Mignola E, Rigano D, Senatore F, Arnold DA, Bruno M, Rosselli S (2009) Constituents of leaves and flowers essential oils of Helichrysum pallaswii (Spreng.) Ledeb. growing wild in Lebanon. J Med Food 12:203e7
Frost LS, Leplae R, Summers AO, Toussaint A (2005) Mobile genetic elements: the agents of open source evolution. Nat Rev Microbiol 3(9):722–732. https://doi.org/10.1038/nrmicro1235
Golkar Z, Bagasra O, Pace DG (2014) Bacteriophage therapy: a potential solution for the antibiotic resistance crisis. J Infect Dev Ctries 8(2):129–136. https://doi.org/10.3855/jidc.3573
Gowda GA, Zhang S, Gu H, Asiago V, Shanaiah N, Raftery D (2008) Metabolomics-based methods for early disease diagnostics. Expert Rev Mol Diagn 8(5):617–633. https://doi.org/10.1586/14737159.8.5.617
Grossart HP, Massana R, McMahon KD, Walsh DA. (2020) Linking metagenomics to aquatic microbial ecology and biogeochemical cycles. Limnol Oceanogr
Habbal O, Hasson SS, El-Hag AH, Al-Mahrooqi Z, Al-Hashmi N, Al-Bimani Z, Al-Balushi MS, Al-Jabri AA (2011) Antibacterial activity of Lawsoniainermis Linn (Henna) against Pseudomonas aeruginosa. Asian Pac J Trop Biomed 1(3):173–176. https://doi.org/10.1016/S2221-1691(11)60021-X
Havlik J, Budesinsky M, Kloucek P, Kokoska L, Valterova I, Vasickova S, Zeleny V (2009) Norsesquiterpene hydrocarbons, chemical composition and antimicrobial activity of Rhaponticumcarthamoides root essential oil. Phytochemistry 70:414e8
Hendriksen RS, Bortolaia V, Tate H, Tyson GH, Aarestrup FM, McDermott PF (2019) Using genomics to track global AMR. Front Public Health 7:242. https://doi.org/10.3389/fpubh.2019.00242
Ikigai H, Nakae T, Hara Y, Shimamura T (1993) Bactericidal catechins damage the lipid bilayer. BiochimBiophys Acta 1147(1):132–136. https://doi.org/10.1016/0005-2736(93)90323-r
Iskandar K, Molinier L, Hallit S et al (2021) Surveillance of AMR in low- and middle-income countries: a scattered picture. Antimicrob Resist Infect Control 10:63. https://doi.org/10.1186/s13756-021-00931-w
Kamatou GPP, Viljoen AMA (2010) review of the application and pharmacological properties of a-bisabolol and abisabolol-rich oils. J Am Oil Chem Soc 87:1e7
Knight V, Sanglier JJ, DiTullio D, Braccili S, Bonner P, Waters J, Hughes D, Zhang L (2003) Diversifying microbial natural products for drug discovery. ApplMicrobiolBiotechnol 62(5–6):446–458. https://doi.org/10.1007/s00253-003-1381-9
Köser CU, Ellington MJ, Peacock SJ (2014) Whole-genome sequencing to control AMR. Trends Genet 30(9):401–407. https://doi.org/10.1016/j.tig.2014.07.003
Larsson DG, Flach CF (2021) Antibiotic resistance in the environment. Nat Rev Microbiol 4:1–3. https://doi.org/10.1038/s41579-021-00649-x
Lee JH, Kim YG, Cho HS, Ryu SY, Cho MH, Lee J (2014) Coumarins reduce biofilm formation and the virulence of Escherichia coli O157:H7. Phytomedicine 21(8–9):1037–1042. https://doi.org/10.1016/j.phymed.2014.04.008
Lee JH, Park JH, Cho HS, Joo SW, Cho MH, Lee J (2013) (2013) Anti-biofilm activities of quercetin and tannic acid against Staphylococcus aureus. Biofouling. https://doi.org/10.1080/08927014.2013.788692
Lewis K, Ausubel FM (2006) Prospects for plant-derived antibacterials. Nat Biotechnol 24(12):1504–1507. https://doi.org/10.1038/nbt1206-1504
Li Y, Xu Z, Han W, Cao H, Umarov R, Yan A, Fan M, Chen H, Duarte C, Li L, Ho PL (2021) HMD-ARG: Identifying antibiotic resistance genes through machine learning. Microbiome 9:40. https://doi.org/10.1186/s40168-021-01002-3
Lorenzi V, Muselli A, Bernardini AF, Berti L, Pages JM, Amaral L, Bolla JM (2009) Geraniol restores antibiotic activities against multidrug-resistant isolates from Gram-negative species. Antimicrob Agents Chemother 53:2209e11
Lushniak BD (2014) Antibiotic resistance: a public health crisis. Public Health Rep 129(4):314–316. https://doi.org/10.1177/003335491412900402
Lv XA, Jin YY, Li YD, Zhang H, Liang XL (2013) Genome shuffling of Streptomyces viridochromogenes for improved production of avilamycin. ApplMicrobiolBiotechnol 97(2):641–648. https://doi.org/10.1007/s00253-012-4322-7
Macià MD, Rojo-Molinero E, Oliver A (2014) Antimicrobial susceptibility testing in biofilm-growing bacteria. Clin MicrobiolInfect. https://doi.org/10.1111/1469-0691.12651
Manar Ali A, Muzaheed, Amal Jamil F, Mohammed A, Wael M, Merin G, Sadananda A, Sanjay R, Darshan Devang D, Chitra J, Sajith V, Aftab Ahmed K, Jilani S, Poojdev J, (2020) AMR, mechanisms and its clinical significance, Dis-a-Mon 66,(6), 100971,ISSN 0011–5029, https://doi.org/10.1016/j.disamonth.2020.100971
Marianne F, Krishan K, Anthony B, (2017) Antibiotic resistance, Journal of Infection and Public Health,Volume 10, (Issue 4) ,Pages 369-378,ISSN 1876-0341https://doi.org/10.1016/j.jiph.2016.08.007
McArthur AG, Waglechner N, Nizam F, Yan A, Azad MA, Baylay AJ, Bhullar K, Canova MJ, De Pascale G, Ejim L, Kalan L (2013) The comprehensive antibiotic resistance database. Antimicrob Agents Chemother 57(7):3348–3357. https://doi.org/10.1128/AAC.00419-13
McMurry L, Petrucci RE Jr, Levy SB (1980) Active efflux of tetracycline encoded by four genetically different tetracycline resistance determinants in Escherichia coli. Proc Natl Acad Sci USA 77(7):3974–3977. https://doi.org/10.1073/pnas.77.7.3974
Merino N, Aronson HS, Bojanova DP, Feyhl-Buska J, Wong ML, Zhang S, Giovannelli D (2019) Living at the extremes: extremophiles and the limits of life in a planetary context. Front Microbiol 10:780. https://doi.org/10.3389/fmicb.2019.00780
Michael CA, Dominey-Howes D, Labbate M (2014) The AMR crisis: causes, consequences, and management. Front Public Health 2:145. https://doi.org/10.3389/fpubh.2014.00145
Mohana NC, Rao HY, Rakshith D, Mithun PR, Nuthan BR, Satish S (2018) Omics based approach for biodiscovery of microbial natural products in antibiotic resistance era. J Genet Eng Biotechnol
Monnet V, Gardan R (2015) Quorum-sensing regulators in gram-positive bacteria: ‘cherchez le peptide.’ Mol Microbiol 97(2):181–184. https://doi.org/10.1111/mmi.13060
Murray CJ, Ikuta KS, Sharara F, Swetschinski L, Aguilar GR, Gray A, Han C, Bisignano C, Rao P, Wool E, Johnson SC (2022) Global burden of bacterial antimicrobial resistance in 2019: a systematic analysis. Lancet 399:629–655. https://doi.org/10.1016/S0140-6736(21)02724-0
Nguyen KT, Ritz D, Gu JQ, Alexander D, Chu M, Miao V, Brian P, Baltz RH (2006) Combinatorial biosynthesis of novel antibiotics related to daptomycin. Proc Natl Acad Sci U S A 103(46):17462–17467. https://doi.org/10.1073/pnas.0608589103
Partridge SR, Kwong SM, Firth N, Jensen SO (2018) Mobile genetic elements associated with AMR. Clin Microbiol Rev 31(4):e00088-e117. https://doi.org/10.1128/CMR.00088-17
Peng L, Kang S, Yin Z, Jia R, Song X, Li L, Li Z, Zou Y, Liang X, Li L, He C, Ye G, Yin L, Shi F, Lv C, Jing B (2015) Antibacterial activity and mechanism of berberine against Streptococcus agalactiae. Int J Clin Exp Pathol 8(5):5217–5223
Pereira DM, Valentão P, Pereira JA, Andrade PB (2009) Phenolics: from chemistry to biology. Molecules 14(6):2202–2211. https://doi.org/10.3390/molecules14062202
Petri R, Schmidt-Dannert C (2004) Dealing with complexity: evolutionary engineering and genome shuffling. Curr Opin Biotechnol 15(4):298–304. https://doi.org/10.1016/j.copbio.2004.05.005
Phan Canh T, Le-Thi-Thanh T, Hoang-Tran-Viet H, Tuanh N (2020) DPPH-scavenging and antimicrobial activities of Asteraceae medicinal plants on uropathogenic bacteria. Evid-Based Complement Alternat Med 1–9. https://doi.org/10.1155/2020/7807026
Punina NV, Makridakis NM, Remnev MA, Topunov AF (2015) Whole-genome sequencing targets drug-resistant bacterial infections. Hum Genomics 9:19. https://doi.org/10.1186/s40246-015-0037-z
Radulovic NS, Dekic MS, Randelovic PJ, Stojanovic NM, Zarubica AR, Stojanovic-Radic ZZ (2012) Toxic essential oils: anxiolytic, antinociceptive and antimicrobial properties of the yarrow Achillea umbellate Sibth et Sm (Asteraceae) volatiles. Food Chem Toxicol 50:2016e26
Rozalski M, Micota B, Sadowska B, Stochmal A, Jedrejek D, Wieckowska-Szakiel M, &Rozalska B (2013) Antiadherent and antibiofilm activity of Humulus lupulus L. derived products: new pharmacological properties. BioMed Res Int 101089 https://doi.org/10.1155/2013/101089
Saleem M, Nazir M, Ali MS, Hussain H, Lee YS, Riaz N, Jabbar A (2009) Antimicrobial natural products: an update on future antibiotic drug candidates. Nat Prod Rep 27(2):238–254. https://doi.org/10.1039/b916096e
Sanchez CJ, Mende K, Beckius ML et al (2013) Biofilm formation by clinical isolates and the implications in chronic infections. BMC Infect Dis 13:47. https://doi.org/10.1186/1471-2334-13-47
Sandanayaka VP, Prashad AS (2002) Resistance to beta-lactam antibiotics: structure and mechanism based design of beta-lactamase inhibitors. Curr Med Chem 9(12):1145–1165. https://doi.org/10.2174/0929867023370031
Semwal DK, Rawat U (2009) Antimicrobial hasubanalactam alkaloid from Stephania glabra. Planta Med 75(4):378–380. https://doi.org/10.1055/s-0028-1112223
Sharma A, Gupta VK, Pathania R (2009) Efflux pump inhibitors for bacterial pathogens: from bench to bedside. Indian J Med Res 149(2):129–145. https://doi.org/10.4103/ijmr.IJMR_2079_17
Shimizu M, Shiota S, Mizushima T, Ito H, Hatano T, Yoshida T, Tsuchiya T (2001) Marked potentiation of activity of beta-lactams against methicillin-resistant Staphylococcus aureus by corilagin. Antimicrob Agents Chemother 45(11):3198–3201. https://doi.org/10.1128/AAC.45.11.3198-3201.2001
Shuman EK (2011) Global climate change and infectious diseases. Int J Occup Environ Med 2(1):11–19
Stephanopoulos G (2002) Metabolic engineering by genome shuffling. Nat Biotechnol 20(7):666–668. https://doi.org/10.1038/nbt0702-666
Su M, Satola SW, Read TD (2019) Genome-based prediction of bacterial antibiotic resistance. J Clin Microbiol 57(3):e01405-e1418. https://doi.org/10.1128/JCM.01405-18
SudanoRoccaro A, Blanco AR, Giuliano F, Rusciano D, Enea V (2004) Epigallocatechin-gallate enhances the activity of tetracycline in staphylococci by inhibiting its efflux from bacterial cells. Antimicrob Agents Chemother 48(6):1968–1973. https://doi.org/10.1128/AAC.48.6.1968-1973
Tabanca N, Demirci F, Demirci B, Wedge DE, Baser KHC (2007) Composition, enantiomeric distribution and antimicrobial activity of Tanacetum argenteum subsp. flabellifolium essential oil. J Pharm Biomed Anal 45:714e9
Taylor PW, Hamilton-Miller JM, Stapleton PD (2005) Antimicrobial properties of green tea catechins. Food Sci Technol Bull. https://doi.org/10.1616/1476-2137.14184
Teixeira MC, Leme EE, Delarmelina C, Almeida A, Figueira GM, Sartoratto A (2007) Activity of essential oils from Brazilian medicinal plants on Escherichia coli. J Ethnopharmacol 111:197e201
Van Bambeke F, Balzi E, Tulkens PM (2000) Antibiotic efflux pumps. Biochempharmacol 60(4):457–470. https://doi.org/10.1016/s0006-2952(00)00291-4
van Belkum A, Dunne WM Jr (2013) Next-generation antimicrobial susceptibility testing. J Clin Microbiol. https://doi.org/10.1128/JCM.00313-13
Veličković DT, Ranđelović NV, Ristić MS, Veličković AS, Šmelcerović AA (2003) Chemical constituents and antimicrobial activity of the ethanol extracts obtained from the flower, leaf and stem of Salvia officinalis L. J Serb Chem Soc 68(1):17–24
Wang L, McLeod HL, Weinshilboum RM (2006) Genomics and drug response. N Engl J Med 364(12):1144–1153. https://doi.org/10.1056/NEJMra1010600
Weinshilboum R (2003) Inheritance and drug response. N Engl J Med 348(6):529–537. https://doi.org/10.1056/NEJMra020021 (PMID: 12571261)
World Health Organization Antibiotic Resistance. [(accessed on 15 June 2018)];(2018) Available online: http://www.who.int/en/news-room/fact-sheets/detail/antibiotic-resistance
Wyres KL, Lam MMC, Holt KE (2020) Population genomics of Klebsiella pneumoniae. Nat Rev Microbiol 18(6):344–359. https://doi.org/10.1038/s41579-019-0315-1
Xie T, Song S, Li S, Ouyang L, Xia L, Huang J (2015) Review of natural product databases. Cell Prolif 48(4):398–404. https://doi.org/10.1111/cpr.12190
Yap PS, Yiap BC, Ping HC, Lim SH (2014) Essential oils, a new horizon in combating bacterial antibiotic resistance. Open Microbiol J 8:6–14. https://doi.org/10.2174/1874285801408010006
Yavli N, Yasar A, Gulec C, Usta A, Kolavli S, Coskuncelebi K, Karaoglu S (2005) Composition and antimicrobial activity of essential oils from Centaurea sessilis and Centaurea armena. Phytochemistry 66:1741e5
Zampieri M, Enke T, Chubukov V, Ricci V, Piddock L, Sauer U (2017) Metabolic constraints on the evolution of antibiotic resistance. Mol Syst Biol 13(3):917. https://doi.org/10.15252/msb.20167028
Zebra-Kucukbay F, Kuyumcu E, Bilenler T, Yildiz B (2011) Chemical composition and antimicrobial activity of the essential oil of Achillea cretica L. (Asteraceae) from Turkey. Nat Prod Res 25:1e6
Zerikly M, Challis GL (2009) Strategies for the discovery of new natural products by genome mining. ChemBioChem 10(4):625–633. https://doi.org/10.1002/cbic.200800389
Zhang YX, Perry K, Vinci VA, Powell K, Stemmer WP, del Cardayré SB (2002) Genome shuffling leads to rapid phenotypic improvement in bacteria. Nature 415(6872):644–646. https://doi.org/10.1038/415644a
Zumla A et al (2013) Advances in the development of new tuberculosis drugs and treatment regimens. Nat Rev Drug Discov 12:388–404
Author information
Authors and Affiliations
Contributions
RK, MK and TB: conceived the study and wrote the first draft of the paper; AS, AAHA and SB: data compilation; AAH and CVDLA: figure work; LA and TB: proof read.
Corresponding author
Ethics declarations
Ethical approval
Not applicable.
Consent to participate
Not applicable.
Consent for publication
All the authors have approved the manuscript for publication.
Competing interests
The authors declare no competing interests.
Additional information
Responsible Editor: Philippe Garrigues
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Kaur, R., Kanotra, M., Sood, A. et al. Emergence of nutriments as a nascent complementary therapy against antimicrobial resistance. Environ Sci Pollut Res 29, 49568–49582 (2022). https://doi.org/10.1007/s11356-022-20775-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11356-022-20775-0