Abstract
The environment plays an important role in the dissemination of clinically relevant antimicrobial-resistant bacteria and genes. In this study, we described genomic features of a plasmid-mediated colistin-resistant mcr-1-positive Escherichia coli strains (PK-3225) isolated from a dairy farm wastewater sample. After initial isolation and PCR detection of mcr-1-positive E. coli, whole-genome sequencing was performed using Illumina Hiseq 2500 followed by in silico analysis. Genetic context surrounding the mcr-1 gene was determined and SNP-based phylogenomic analysis was performed. Furthermore, plasmid analysis and conjugation assays were performed to determine transferability of mcr-1. E. coli PK-3225 belonged to ST10 and carried a broad resistome that included colistin (mcr-1), beta-lactam (blaTEM-IB), tetracycline (tetB), phenicol (catA1), macrolide (mdfA), trimethoprim (dfrA17), aminoglycosides (aadA5, aph(3”)-Ib, aph(6)-Id), and sulphonamide (sul2) resistance genes. The draft genome of E. coli calculated as 4.9 Mbp. Conjugation experiment showed successful transfer of the mcr-1 gene to E. coli recipient strain J53. In silico analysis showed that mcr-1 was located on IncI2 plasmid of > 59 kb in length, with the nikB-mcr-1-pap2 gene array, and lack ISApl1. The phylogenomic analysis revealed that the PK-3225 was closely related to human ST10 E. coli from Brazil and USA. To our knowledge, this is the first draft genome sequence of mcr-1 carrying E. coli isolated from the farm environment in Pakistan. Considering the high burden of colistin resistance in Pakistan, presence of pandemic high-risk E. coli clones in the environment requires strict surveillance.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Polymyxins are the small lipopolypeptide of the molecular weight 1200 Da with a polycationic ring attached to the hydrophobic fatty acids (Kola et al. 2016). Polymyxins including colistin are the well-organized drugs and have regained significant interest due to the increasing incidence of multidrug-resistant (MDR) gram-negative bacterial infections (Poirel et al. 2017). The mechanism of colistin is mediated by binding to the lipopolysaccharides of gram-negative bacteria and subsequently disintegrates the membrane. Colistin (Polymyxin E) is known as a last-resort antibiotic to treat MDR gram-negative bacterial infections in clinical use (Paterson et al. 2019). Colistin-resistant Enterobacteriaceae are globally recognized as an important threat for public health (Yao et al. 2016). The discovery of plasmid-mediated colistin resistance gene mcr-1 in China in 2015 (Liu et al. 2016) and worldwide dissemination of mcr-1 in bacteria through conjugation has been found a great threat to the public. To date, ten variants of the mcr-1 gene (mcr-1 to mcr-10) have been identified in different bacterial species isolated from humans, food, animals, farms, and the environment (Hussein et al. 2021). Colistin resistance gene mcr-1 has been found on plasmids of different incompatibility types IncX4, IncI2, IncY, and IncHI2 (Li et al. 2017). In Pakistan, mcr-1 has been identified in E. coli isolates from wildlife (Mohsin et al. 2016), human (Mohsin et al. 2017), healthy broiler, and sick broilers (Azam et al. 2020; Rafique et al. 2020). However, to the best of our knowledge, mcr-1 has not been reported from the farm environment.
Materials and methods
A total of 100 cattle samples were screened along with farm wastewater samples collected in 2019 from 4 corporate commercial dairy farms in Punjab, Pakistan. The loopful inoculum was streaked on MacConkey agar (Oxoid, UK) supplemented with colistin 2 μg/ml for 18–24 hours at 37°C. In this regard, one colistin-resistant E. coli isolate (PK-3225 strain) was recovered from a dairy farm wastewater. Antimicrobial susceptibility profile was determined by disk-diffusion and/or microdilution broth methods (EUCAST 2021; CLSI 2021; CLSI 2020). DNA of PK-3225 was extracted using DNeasy blood and tissue kit (Qiagen, Hilden, Germany). Conjugation experiment was performed to determine the transferability of colistin resistance using sodium azide–resistant E. coli J53 as recipient strain. The sequencing library was prepared using an Illumina Nextera XT DNA Library with 2 × 150-bp paired-end reads on an Illumina HiSeq 2500 instrument (Illumina, San Diego, CA, USA). De novo assembly was performed using SPAdes v3.9 (Wick et al. 2017) and annotation of the genome was performed using DFAST (https://dfast.nig.ac.jp/) (Tanizawa et al. 2019). The sequence was submitted to NCBI/ENA/DDBJ (Arnemann 2018). The tools of the Center for Genomic Epidemiology (http://www.genomicepidemiology.org) were used for the determination of multilocus sequence type (MLST) (Larsen et al. 2012), acquired antibiotic resistance genes (Zankari et al. 2012), plasmid replicon typing (Carattoli et al. 2014), and serotyping (Joensen et al. 2015) using MLST v2.0, ResFinder v4.1, VirulenceFinder v2.0, and SerotypeFinder v2.0, respectively. The complete sequence of the mcr-encoding plasmid was performed using Geneious R9 (Biomatters) and Bandage assembly graph visualizer (https://github.com/rrwick/Bandage), using an interactive anchoring of short-reads and contigs alignment combined approach, as previously described (Fuga et al. 2021). Plasmid comparative analysis was performed using the Proksee platform (https://beta.proksee.ca/) and BLASTN. The genomic sequences of 103 E. coli strains belonging to ST10 were downloaded from the Enterobase Escherichia/Shigella database (https://enterobase.warwick.ac.uk/) and submitted to antimicrobial resistance gene prediction by using ABRicate (https://github.com/tseemann/abricate) with Resfinder database. All sequences were aligned to the reference sequence using CSI- phylogeny v 1.4 and the phylogenetic tree was constructed based upon the SNP content. The tree was visualized using iTOL v6 (https://itol.embl.de) and resistome information based upon ABRicate analysis was added into phylogenetic tree.
Results and discussion
The antimicrobial susceptibility profile showed that the PK-3225 E. coli was resistant to ampicillin, cefotaxime, gentamicin, tetracycline, sulfamethoxazole/trimethoprim, tylosin, and colistin (MIC= 4 mg/L), remaining susceptible to moxifloxacin, enrofloxacin, and meropenem. Conjugation experiment showed that mcr-1 was transferable to sodium azide–resistant E. coli J53. De novo assembly result generated a total of 165 contigs, with the total length of 4957826 bp and the GC contents were 50.43%.
The PK-3225 carried antibiotic resistance genes against beta-lactams (blaTEM-IB), tetracycline (tetB), phenicol (catA1), colistin (mcr-1.1) macrolides (mdfA), trimethoprim (dfrA17), aminoglycosides [aadA5, aph(3”)-Ib, aph(6)-Id], and sulphonamide (sul2), as well as chromosomal mutations in gyrA (S83L, D85L) and parC (S80I). Plasmid incompatibility types IncFIB, IncFIC(FII), IncI2, and IncQ1 were predicted in PK-3225. The strain belonged to the O101:H10 serotype and carried multiple virulence genes cvaC, etsC, gad, hlyF, hra, iha, iroN, iss, iucC, iutA, mchB, mchC, mchF, ompT, sitA, terC, and traT.
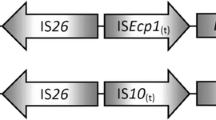
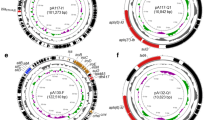
The mcr-1.1 gene was found to be encoding by the IncI2 conjugative plasmid, which has been globally associated with mcr-1 (Ovejero et al. 2017; Sun et al. 2017; Wang et al. 2017). The complete nucleotide sequence of pPK-3225mcr was 59.630 bp and contains 81 coding sequences (CDS) and a G+C average content of 42.26%. The nikb-mcr-1-pap2 gene array was found in pPK-3225. Interestingly, the same genetic context of mcr-1 was also found in colistin-resistant E. coli from dairy cows in China (Zheng et al. 2019). On the other hand, nikB (relaxase) and pap2 gene cassettes and been identified in mcr-1-positive E. coli isolates from outpatients, in Chile (Gutiérrez et al. 2019). More recently, it was discovered in China that the pap2 gene along with mcr-1 was necessary to reduce colistin susceptibility (Choi et al. 2020). Comparative plasmid analysis using BLASTN revealed that pPK-3225mcr is highly similar (≥ 99% pairwise identity) to other mcr-1-harboring IncI2 plasmids identified in E. coli and Salmonella enterica isolates from humans, livestock, and chicken meat in China, South Korea, and the Netherlands, respectively (Fig. 1).
Circular view and alignment comparison of closely related IncI2 plasmids harboring the mcr-1 gene. IncI2/mcr-1 genetic arrays have been identified in E. coli strains isolated from farm wastewater in Pakistan (pPK-3225mcr, this study), human hosts in China (MK574667; CP029184) and the Netherlands (LR882922); Salmonella Typhimurium from pig in South Korea (CP065423), and Salmonella enterica from chicken meat (CP033355) and E. coli from poultry (CP069705) in China. In outer circles, arrows indicate the positions and directions of genes
The colistin-resistant E. coli strain PK-3225 belongs to the pandemic MLST sequence type ST10, clonal complex CC10, serotype O101:H10, phylogroup A, and fimH30 allele. In this regard, the adhesin fimH allele H30 has been rarely identified in non-ST131 E. coli lineages. In fact, E. coli ST131-H30 subclade has emerged in recent decades as an epidemic clonal group responsible for a large amount of human extraintestinal infections worldwide, and thus the, fimH allele H30 could be conferring advantage to human host adaptation (Stoesser et al. 2016). Moreover, virulome analysis revealed the presence of genes related to adherence (iha), iron uptake (iroN, sitA, iutA, iucC, hra), colicins (cvaC, mchBCF), serum resistance (iss, ompT, traT), hemolysin (hlyF), glutamate decarboxylase (gad), and tellurium resistance (terC).
The phylogenomic analysis of PK-3225 was performed with 103 genomic sequences of E. coli strains belonging to the international ST10 (Fig. 2). In this regard, the environmental E. coli strain PK-3225 was closely related (SNP difference = 196) to the human E. coli strain ESC_FB9722AA, isolated from an urinary tract infection case, in Brazil (Campos et al. 2018) and SNP difference of 1270 from a clinical E. coli strain ESC_BA5361AA originated in USA. While tetB, sul2, dfrA17, blaTEM-IB, aadA5, aph(3”)-Ib, and aph(6)-Id resistance genes were shared by PK-3225 and ESC_FB9722AA E. coli strains, common resistance genes among 103 E. coli ST10 genomes analyzed conferred resistance to tetracycline, aminoglycosides, sulphonamide, and trimethoprim (Fig. 2). The presence of the clinically relevant mcr-1-positive ST10 E. coli in wastewater with multidrug resistance is concerning.
Colistin-resistant E. coli strains have been reported from clinical, broiler, and wild migratory bird samples, in Pakistan (Mohsin et al. 2016, 2017; Azam et al. 2017; Lv et al. 2018); however, it has not been reported from farm environments. Therefore, in this study, we report the first draft genome sequence of a multidrug-resistant IncI2/mcr-1.1-positive E. coli strain belonging to the pandemic ST10, identified in a farm wastewater source in Pakistan. Considering the high burden of colistin resistance in Pakistan, presence of priority high-risk E. coli clones in the environment requires strict surveillance.
References
Arnemann J (2018) NCBI. In: Gressner A, Arndt T (eds) Lexikon der Medizinischen Laboratoriumsdiagnostik. Springer Reference Medizin. Springer, Berlin, Heidelberg. https://doi.org/10.1007/978-3-662-49054-9_3540-1
Azam M, Ehsan I, Sajjad-ur-Rahman et al (2017) Detection of the colistin resistance gene mcr-1 in avian pathogenic Escherichia coli in Pakistan. J Glob Antimicrob Resist 11:152–153. https://doi.org/10.1016/j.jgar.2017.10.012
Azam M, Mohsin M, Johnson TJ, Smith EA, Johnson A, Umair M, Saleemi MK, Sajjad-ur-Rahman (2020) Genomic landscape of multi-drug resistant avian pathogenic Escherichia coli recovered from broilers. Vet Microbiol 247:108766. https://doi.org/10.1016/j.vetmic.2020.108766
Campos ACC, Andrade NL, Ferdous M, Chlebowicz MA, Santos CC, Correal JCD, Lo ten Foe JR, Rosa ACP, Damasco PV, Friedrich AW, Rossen JWA (2018) Comprehensive molecular characterization of Escherichia coli isolates from urine samples of hospitalized patients in Rio de Janeiro, Brazil. Front Microbiol 9:1–12. https://doi.org/10.3389/fmicb.2018.00243
Carattoli A, Zankari E, Garciá-Fernández A et al (2014) In silico detection and typing of plasmids using plasmidfinder and plasmid multilocus sequence typing. Antimicrob Agents Chemother 58:3895–3903. https://doi.org/10.1128/AAC.02412-14
Choi Y, Lee JY, Lim SK, Ko KS (2020) Intact pap2 downstream of mcr-1 appears to be required for colistin resistance: pap2 in colistin resistance. Diagn Microbiol Infect Dis 97:114997. https://doi.org/10.1016/j.diagmicrobio.2020.114997
CLSI (2020) Performance Standards for Antimicrobial Susceptibility Testing. CLSI Publishing. https://clsi.org/about/press-releases/clsi-s-publishes-30th-edition-of-m100. Accessed 23 Jan 2020
CLSI (2021) Performance Standards for Antimicrobial Susceptibility Testing. CLSI Publishing. http://clsi.org/standards/products/microbiology/documents/m100. Accessed 23 Mar 2021
EUCAST (2021) European Committee on Antimicrobial Susceptibility Testing Breakpoint tables for interpretation of MICs and zone diameters European Committee on Antimicrobial Susceptibility Testing Breakpoint tables for interpretation of MICs and zone diameters. http://www.eucast.org/fileadmin/src/media/PDFs/EUCAST_files/Breakpoint_tables/v_50_Breakpoint_Table_01.pdf 0–77. Accessed 01 Jan 2021
Fuga B, Cerdeira L, Moura Q, Fontana H, Fuentes-Castillo D, Carvalho AC, Lincopan N (2021) Genomic data reveals the emergence of an IncQ1 small plasmid carrying blaKPC-2 in Escherichia coli of the pandemic sequence type 648. J Glob Antimicrob Resist 25:8–13. https://doi.org/10.1016/j.jgar.2021.02.014
Gutiérrez C, Zenis J, Legarraga P, Cabrera-Pardo JR, García P, Bello-Toledo H, Opazo-Capurro A, González-Rocha G (2019) Genetic analysis of the first mcr-1 positive Escherichia coli isolate collected from an outpatient in Chile. Braz J Infect Dis 23:203–206. https://doi.org/10.1016/j.bjid.2019.05.008
Hussein NH, AL-Kadmy IMS, Taha BM, Hussein JD (2021) Mobilized colistin resistance (mcr) genes from 1 to 10: a comprehensive review. Mol Biol Rep 48:2897–2907. https://doi.org/10.1007/s11033-021-06307-y
Joensen KG, Tetzschner AMM, Iguchi A, Aarestrup FM, Scheutz F (2015) Rapid and easy in silico serotyping of Escherichia coli isolates by use of whole-genome sequencing data. J Clin Microbiol 53:2410–2426. https://doi.org/10.1128/JCM.00008-15
Kola M, Trimble MJ, Mlyna P (2016) Polymyxin: Alternative Mechanisms of Action. Cold Spring Harb Perspect Med 6:1–22. https://doi.org/10.1101/cshperspect.a025288
Larsen MV, Cosentino S, Rasmussen S, Friis C, Hasman H, Marvig RL, Jelsbak L, Sicheritz-Ponten T, Ussery DW, Aarestrup FM, Lund O (2012) Multilocus sequence typing of total-genome-sequenced bacteria. J Clin Microbiol 50:1355–1361. https://doi.org/10.1128/JCM.06094-11
Li R, Xie M, Zhang J, Yang Z, Liu L, Liu X, Zheng Z, Chan EWC, Chen S (2017) Genetic characterization of mcr-1-bearing plasmids to depict molecular mechanisms underlying dissemination of the colistin resistance determinant. J Antimicrob Chemother 72:393–401. https://doi.org/10.1093/jac/dkw411
Liu YY, Wang Y, Walsh TR, Yi LX, Zhang R, Spencer J, Doi Y, Tian G, Dong B, Huang X, Yu LF, Gu D, Ren H, Chen X, Lv L, He D, Zhou H, Liang Z, Liu JH, Shen J (2016) Emergence of plasmid-mediated colistin resistance mechanism MCR-1 in animals and human beings in China: a microbiological and molecular biological study. Lancet Infect Dis 16:161–168. https://doi.org/10.1016/S1473-3099(15)00424-7
Lv J, Mohsin M, Lei S, Srinivas S, Wiqar RT, Lin J, Feng Y (2018) Discovery of a mcr-1-bearing plasmid in commensal colistin-resistant escherichia coli from healthy broilers in faisalabad, pakistan. Virulence 9:994–999. https://doi.org/10.1080/21505594.2018.1462060
Mohsin M, Raza S, Roschanski N, Schaufler K, Guenther S (2016) First description of plasmid-mediated colistin-resistant extended-spectrum β-lactamase-producing Escherichia coli in a wild migratory bird from Asia. Int J Antimicrob Agents 48:463–464. https://doi.org/10.1016/j.ijantimicag.2016.07.001
Mohsin M, Raza S, Roschanski N et al (2017) Description of the First Escherichia coli Resistance Gene mcr-1 from the Indian. Antimicrob Agents Chemother 61:2016–2017. https://doi.org/10.1128/aac.01945-16
Ovejero CM, Delgado-Blas JF, Calero-Caceres W, Muniesa M, Gonzalez-Zorn B (2017) Spread of mcr-1-carrying Enterobacteriaceae in sewage water from Spain. J Antimicrob Chemother 72:1050–1053. https://doi.org/10.1093/jac/dkw533
Poirel L, Aurélie J, Nordmann P (2017) Polymyxins: antibacterial activity, susceptibility testing, and resistance mechanisms encoded by plasmids or chromosomes Laurent Poirela,b,c, Aurélie Jayola,b,c and Patrice Nordmann. Clin Microbiol Rev 30:557–596. https://doi.org/10.1128/CMR.00064-16
Paterson DL, Bonomo RA (2019) Multidrug-resistant gram- negative pathogens: the urgent need for “Old” polymyxins. Adv Exp Med Biol 9–13. https://doi.org/10.1007/978-3-030-16373-0_2
Rafique M, Potter RF, Ferreiro A, Wallace MA, Rahim A, Malik AA, Siddique N, Abbas MA, Souza AWD, CAD B, Ali N, Gautam D (2020) Genomic characterization of antibiotic resistant Escherichia coli isolated from domestic chickens in Pakistan. Front Microbiol 10:1–10. https://doi.org/10.3389/fmicb.2019.03052
Stoesser N, Sheppard AE, Pankhurst L, de Maio N, Moore CE, Sebra R, Turner P, Anson LW, Kasarskis A, Batty EM, Kos V, Wilson DJ, Phetsouvanh R, Wyllie D, Sokurenko E, Manges AR, Johnson TJ, Price LB, Peto TEA, Johnson JR, Didelot X, Walker AS, Crook DW, Modernizing Medical Microbiology Informatics Group (MMMIG) (2016) Evolutionary history of the global emergence of the Escherichia coli epidemic clone ST131. MBio 7:1–15. https://doi.org/10.1128/mBio.02162-15
Sun J, Fang LX, Wu Z, Deng H, Yang RS, Li XP, Li SM, Liao XP, Feng Y, Liu YH (2017) Genetic analysis of the IncX4 plasmids: implications for a unique pattern in the mcr-1 acquisition. Sci Rep 7:1–9. https://doi.org/10.1038/s41598-017-00095-x
Tanizawa Y, Fujisawa T, Arita M, Nakamura Y (2019) Generating publication-ready prokaryotic genome annotations with DFAST. Methods in Molecular Biology, pp 215–226
Wang Q, Sun J, Ding Y, Li XP, Liu YH, Feng Y (2017) Genomic insights into mcr-1-positive plasmids carried by colistin-resistant Escherichia coli isolates from inpatients. Antimicrob Agents Chemother 61:1–4. https://doi.org/10.1128/AAC.00361-17
Wick RR, Judd LM, Gorrie CL, Holt KE (2017) Unicycler: resolving bacterial genome assemblies from short and long sequencing reads. PLoS Comput Biol 13:1–22. https://doi.org/10.1371/journal.pcbi.1005595
Yao X, Doi Y, Zeng L, Lv L, Liu JH (2016) Carbapenem-resistant and colistin-resistant Escherichia coli co-producing NDM-9 and MCR-1. Lancet Infect Dis 16:288–289. https://doi.org/10.1016/S1473-3099(16)00057-8
Zankari E, Hasman H, Cosentino S, Vestergaard M, Rasmussen S, Lund O, Aarestrup FM, Larsen MV (2012) Identification of acquired antimicrobial resistance genes. J Antimicrob Chemother 67:2640–2644. https://doi.org/10.1093/jac/dks261
Zheng B, Feng C, Xu H, Yu X, Guo L, Jiang X, Song X (2019) Detection and characterization of ESBL-producing Escherichia coli expressing mcr-1 from dairy cows in China. J Antimicrob Chemother 74:321–325. https://doi.org/10.1093/jac/dky446
Acknowledgements
The author would like to thank dairy farm owners for the cooperation during sample collection.
Availability of data
All data generated or analyzed during this study are included in this published article.
Funding
The study is funded by Higher Education Commission (grant no. HEC-NRPU- 4681). N.L. is a research fellow of Conselho Nacional de Desenvolvimento Científico e Tecnológico [CNPq grant 312249/2017-9].
Author information
Authors and Affiliations
Contributions
AA genomic analysis and wrote the paper, HF and ES analyzed the data, RL reviewed and edited, MH sample collection and PCR, SR reviewed the study, and NL and MM conceptualization and wrote the paper.
Corresponding authors
Ethics declarations
Ethical approval and consent to participate
Not applicable.
Consent for publication
The author assigns publication rights to the journal, we assured that this is a unique publication, and we have full authority to make this grant.
Competing interests
The authors declare no competing interests.
Additional information
Responsible Editor: Diane Purchase
Nucleotide sequence accession number
The genome sequence was submitted to NCBI/DDBJ/ENA under BioProject ID PRJNA591297 and accession number WTCP00000000. The version described in this paper is WTCP01000000.
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Ali, ., Fontana, H., Sano, E. et al. Genomic features of a high-risk mcr-1.1-positive Escherichia coli ST10 isolated from cattle farm environment. Environ Sci Pollut Res 28, 54147–54152 (2021). https://doi.org/10.1007/s11356-021-15437-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11356-021-15437-6