Abstract
Microbial community analysis of crude oil containing sludge collected from Duliajan oil field, Assam, India, showed the predominance of hydrocarbon-degrading bacteria such as Pseudomonas (20.1%), Pseudoxanthomonas (15.8%), Brevundimonas (1.6%), and Bacillus (0.8%) alongwith anaerobic, fermentative, nitrogen-fixing, nitrate-, sulfate-, and metal-reducing, syntrophic bacteria, and methanogenic archaea. Phylogenetic Investigation of Communities by Reconstruction of Unobserved States (PICRUSt) analysis indicated gene collection for potential hydrocarbon degradation, lipid, nitrogen, sulfur, and methane metabolism. The culturable microbial community was predominated by Pseudomonas and Bacillus with the metabolic potential for utilizing diverse hydrocarbons, crude oil, and actual petroleum sludge as sole carbon source during growth and tolerating various environmental stresses prevailing in such contaminated sites. More than 90% of the isolated strains could produce biosurfactant and exhibit catechol 2,3-dioxygenase activity. Nearly 30% of the isolates showed alkane hydroxylase activity with the maximum specific activity of 0.54 μmol min−1 mg−1. The study provided better insights into the microbial diversity and functional potential within the crude oil containing sludge which could be exploited for in situ bioremediation of contaminated sites.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Over the last few decades, the expansion of petroleum and allied industries has resulted in the generation of huge amount of hazardous petroleum containing waste, whose management has become a major concern worldwide (Sarkar et al. 2017). With an anticipated rise in global demand for oil up to 123 million barrels per day by the year 2025, production of oil and its processing are continuously expanding to meet the accelerated growth in global energy demand leading to serious pollution of the environment (Ismail et al. 2017). Aliphatic compounds are the major constituents derived from petroleum hydrocarbons. Apart from that, the mixture also contains aromatics, asphaltenes, nitrogen-, sulfur-, oxygen (NSO)-containing compounds and metals (Reddy et al. 2011; Sarkar et al. 2016). Improper handling of petroleum hydrocarbons severely polluted the environment because of its complex composition, persistence nature, and toxicity (Reddy et al. 2011; Fuentes et al. 2014). Bioremediation of petroleum hydrocarbons has been in use for the last five decades; however, it captured tremendous interest after successful clean-up of the Exxon Valdez oil spill (Chandra et al. 2013). Bioremediation relies on the diverse metabolic capabilities of microorganisms to remove wide range of pollutants leading to environmental decontamination (Fuentes et al. 2014). However, there are many practical constraints in implementing efficient bioremediation technology due to the complex nature of contaminated sites and to the lack of knowledge about the adaptation and survival strategies of microorganisms and about the various interacting environmental factors regulating bioremediation processes in such adverse environments (Kostka et al. 2014). Microbial population in the soils of contaminated sites are determined by the physical and chemical conditions available, in particular, at contaminated sites. In sites, adopted microoganisms require carbon and other elements from the contaminants for survival. Therefore, management of such contaminated environments requires proper understanding of the native microbial activities (Kleinsteuber et al. 2012). Culture-independent molecular investigations of different oil-associated environments have been extensively used to reveal the complexity in coexistence of hydrocarbon-degrading, fermentative, syntrophic, and methane metabolizing microorganisms (Hazen et al. 2013). In recent times, PICRUSt (Phylogenetic Investigation of Communities by Reconstruction of Unobserved States) tool has been employed by various investigators to comprehensively describe phylogenetic and functional compositions of these contaminated habitats (Langille et al. 2013; Wang et al. 2016; Mukherjee et al. 2017; Omrani et al. 2018; Roy et al. 2018a). This have allowed for the metabolomics study of microbial communities that are complex in nature with reasonable precision and high rate of taxonomic resolution. The application of extensive culture-dependent approaches has helped in assessing the physiology and metabolic potential of isolated microorganisms in order to exploit them for successful bioaugmentation-based bioremediation strategy (Kostka et al. 2011; Poi et al. 2018). The complex process of bioremediation involves co-metabolism among microbes along with cross-induction, inhibition, and non-interaction, possibly because of the nature of petroleum hydrocarbons which is utilized differently by different microbes to support their growth and survival (Van Hamme et al. 2003; Yang et al. 2016). Various research performed on the microbial community of petroleum hydrocarbon-contaminated environments (such as contaminated soil, sludge, or sediments) revealed the prevalence of Proteobacteria, Firmicutes, Actinobacteria, and Euryarchaeota (Kostka et al. 2011; Yergeau et al. 2012; Head et al. 2014; Gao et al. 2015; Yang et al. 2016; Tian et al. 2019). Archaeal population have been shown to contribute significantly to petroleum contaminant restoration through methanogenesis (Gray et al. 2010; Gieg et al. 2014; Head et al. 2014; Tan et al. 2015; Fowler et al. 2016; Toth and Gieg 2018). Hydrocarbon mineralization through methanogenesis could be important in degradation and considered to be the most dominant terminal process in anaerobic environments (Li et al. 2017). Members of Deltaproteobacteria have been reported to carry out syntrophic hydrocarbon or crude oil alkane degradation in strict anaerobic or methanogenic conditions (Gray et al. 2011; Yang et al. 2016; Sampaio et al. 2017; Ji et al. 2019). However, syntrophic relationship of methanogens with Anaerolineaceae members during hydrocarbon degradation has also been well documented (Liang et al. 2015).
Culture-dependent and advanced meta-omics technologies have documented hydrocarbon metabolizing bacteria related to Pseudomonas, Acinetobacter, Burkholderia, Sphingomonas, Bacillus, Rhodococcus, and Alcanivorax from various hydrocarbon-associated environments (Kostka et al. 2011; Head et al. 2014; Gao et al. 2015). Hydrocarbon-degrading genes can act as useful biomarkers for functional characterization of microbial community, thus presenting a broader estimation of contaminant bioremediation (Yang et al. 2014). The present study was aimed to explore the diversity and hydrocarbon bioremediation potential of microbial (bacterial and archaeal) community in oil containing sludge of Duliajan oil field, Assam, India. The Ion Torrent platform-based next-generation sequencing and analysis of sludge metagenome shed light on the microbial diversity inhabiting the petroleum containing sludge thereby predicting their functional role through PICRUSt. Culturable bacterial community was also identified and characterized based on their physiological and metabolic characteristics. Valuable information regarding the microbial community composition and function could be obtained from this particular study that might assist in designing technologies for in situ restoration of contaminated sites.
Materials and methods
Sample collection, physicochemical analysis, and enumeration of microorganisms
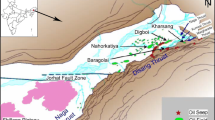
Crude oil-contaminated waste sludge was collected in pre-sterilized DEPC-treated screw capped bottles (1 L capacity) from oil exploration sites of Duliajan oil field, Assam, India (27.3667° N, 95.3167° E). Samples were immediately stored in ice. Physicochemical parameters like temperature, pH, oxidation-reduction potential (ORP), conductivity, salinity, and moisture content were measured as per the method of Roy et al. (2018b). Collected samples were immediately transferred to 4 °C after reaching the laboratory and aliquots were sent for further analysis within 1 week. Chemical analysis of sludge sample for total petroleum hydrocarbon (TPH), aliphatic and aromatic components, metals, and ions was done using standard analytical techniques by S. K. Mitra Pvt. Ltd., Kolkata, India. Total aliphatic components and aromatic components in the sample were estimated by GC-MS analysis (Shimadzu QP2010SE GC-MS system) by following the method of Maiti et al. (2014).
Bacterial enumeration of aerobic bacteria was done on R2A and mineral salt medium (MSM). Both the medium was prepared following the method of Das and Kazy (2014). MSM was supplemented with 0.1% of trace element and 0.2% vitamin solution. Enumeration of anaerobic bacteria was performed following the method of Roy et al. (2018b).
Metagenome extraction, 16S rRNA gene sequencing, and phylogenetic analysis
Total community DNA extraction from 250 mg sample was done in triplicates by using MoBio PowerSoilTM DNA extraction kit (MoBio, USA) by following the manufacturer’s protocol. To increase the DNA yield, extracted metagenome from each set was pooled in a single tube prior to PCR and was visualized by agarose gel electrophoresis followed by quantification using Nanodrop spectrophotometer (Nano 2000 Thermo Fischer Scientific, USA) in the laboratory (Sarkar et al. 2016; Roy et al. 2018b). Gene for the 16S rRNA was amplified using universal V4 region specific conserved primer set of 515F and 806R to construct amplicon library (Caporaso et al. 2011). Ion Torrent-based NGS services were provided by SciGenom Labs Pvt. Ltd., Cochin, Kerala, India. Single end sequencing of the V4 region was done in a 530 Ion Express chip in Ion S5 (Life Technologies, USA) following the user manual protocol. Obtained raw reads were analyzed through Quantitative Insights Into Microbial Ecology (QIIME) pipeline, version 1.7.0 (Caporaso et al. 2010). After quality filtering, identification of operational taxonomic units (OTUs) at an identity level of 97% and its taxonomy assignment was performed in QIIME as per described by Roy et al. (2018b), by considering SILVAngs (version 1.7.0) as a reference database. Estimation of the alpha diversity indices (Shannon index, Simpson index, Chao1 index, and Good’s coverage) was also performed as mentioned by Roy et al. (2018b).
Isolation of culturable bacteria and their identification based on partial sequence of the bacterial 16S rRNA gene
Forty-three aerobic and nine anaerobic strains of bacteria were isolated from the petroleum sludge sample on R2A and anaerobic agar medium using serial dilution technique. Anaerobic experiments were executed as per the protocol of Das and Kazy (2014). Incubation of the inoculated plates was done at 30 °C for 7 days under both aerobic and anaerobic conditions. Aerobically isolated colonies were purified with repetitive subculturing on agar plates. For extended period of preservation, 15% glycerol was used to store the strains at − 80 °C. Anaerobic cultures were maintained by subculturing after certain time intervals.
Genomic DNA extraction of bacterial isolates was done using genomic DNA purification kit (Promega, USA) by following manufacturer’s instructions. The primers and method for amplifying 16S rRNA gene by polymerase chain reaction (PCR) was carried out following the protocol of Paul et al. (2015). Amplified 16S rRNA gene of the isolates was sequenced using 518R (“5-ATTACCGCGGCTGCTGG-3”) primer by Sanger sequencing method by Eurofins Genomics India Pvt. Ltd., Bengaluru, India (Røberg et al. 2011). The sequences were analyzed using NCBI BLASTn and RDP (Ribosomal Database Project) classifier program. MEGA 5 was used to draw the phylogenetic tree by employing the neighbor-joining method with 1000 bootstrap value (Tamura et al. 2011).
Metabolic characterization of sludge sample through PICRUSt analysis
The genomic inventory of sludge metagenome was predicted via PICRUSt analysis (Langille et al. 2013). The analysis was performed through QIIME by using the method as described by Roy et al. (2018b). Functional predictions of metagenomes were done with Nearest Sequenced Taxon Index (NSTI) values followed by reconstruction of the metabolic pathways using KEGG (Kyoto Encyclopedia of Genes and Genomes) database.
Metabolic characterization of bacterial isolates
Metabolic potential of the aerobic strains was determined by their growth at various temperatures (4–50 °C), at different NaCl concentrations (0–10%), and at different pH range (1.0–11.0) in mineral salt medium (MSM) (Das and Kazy 2014). Sensitivity toward heavy metals was examined by growing the isolates on MSM agar plates at 30 °C for 7 days. Yeast extract (5.0 g L−1) was added to MSM as carbon source and different heavy metal salts (CdNO3, Pb(NO3)2, and NiCl2) were added at concentrations ranging from 0.1 to 5 mM. Growth in anaerobic condition was tested in anaerobic agar medium (HiMedia, India). Utilization of multiple electron acceptors during growth other than oxygen was investigated by providing nitrate-, sulfate-, and iron-reducing conditions in MSM agar plates amended with various electron acceptors (NaNO3, Na2SO4, and FeCl3; 20 mM each) and electron donors (glucose, 0.1% and yeast extract, 0.1%) (Das and Kazy 2014). Surfactant production ability of the isolates was examined by adding equal volume diesel with culture supernatant (2 mL) in a culture tube and vortexed vigorously to form an emulsion. The tube was allowed to stand for 24 h. The emulsion stability was determined after 24 h. The emulsification index (E24) was estimated as the percentage height of the emulsion layer to the total liquid column height (Cerqueira et al. 2011; Pacwa-Plociniczak et al. 2014).
The potential of the isolates to utilize various hydrocarbons as sole carbon source was assessed in MSM. The cultures were incubated at 30 °C for 3 days in presence of BTEX individually at concentration of 50 mg L−1 and BTEX mixture at 200 mg L−1. Total aliphatic component of the sludge was quite high (Table 1) and GC-MS analysis indicated the abundance of medium chain length alkanes in the sample (Table 2). Therefore, biodegradation potential of medium chain length alkanes (C15 (pentadecane) and C16 (hexadecane)) by the bacterial isolates was assessed. Utilization of alkanes as sole source of carbon by the bacterium was carried out during growth in 100 mL flasks containing 20 mL MSM supplemented with pentadecane (C15) or hexadecane (C16) at a concentration of 250 mg L−1. Inoculation was done from an overnight grown cell in the range of 107–108 CFU mL−1 and incubated at 30 °C with 150 rpm shaking. Cell numbers were determined after every 7 days and the residual n-alkane was analyzed by gas chromatography. The residual n-alkane from the medium was extracted by adding equal volume of n-hexane and deep freezing the lower water layer to collect the upper organic phase. Remaining water was absorbed by adding Na2SO4. The residual concentrations of the alkanes were analyzed using a GC (Agilent 7820A) equipped with a split/splitless injector, FID detector, and HP-5 column (30 m × 0.32 mm and i.d. 0.25 μm thickness). Carrier gas was nitrogen with flow rate of 25 mL min−1. The oven program was set initially at 80 °C for 2 min, followed by increasing to 210 °C at 10 °C rise per minute. Crude oil and oil containing actual sludge were also used as sole carbon source (1% w/v) and growth was measured by corresponding CFU count. Unweighted pair group method with arithmetic mean (UPGMA) analysis was performed to identify the correlation among the strains based on their metabolic and physiological properties. Percentage identity matrix was used to draw the resemblance dendrogram. Multivariate Statistical Package (MVSP, version 3.1) was used for UPGMA analysis and to generate the dendrogram.
Enzyme assay
Assay of alkane hydroxylase
Alkane hydroxylase enzyme was estimated spectrophotometrically following the method as described by Mishra and Singh (2012). Bacterial isolates were grown in 50 mL MSM with 250 ppm hexadecane. The culture was centrifuged at 10,000 rpm (5 min at 4 °C). Resuspension of the cell pellet was done in 2 mL Tris-HCl buffer (20 mM, pH 7.4) containing 0.15% CHAPS buffer (pH 7.4) and disrupted by sonication at 20 kHz for 1 min five times with an interval of 30 s. The disrupted cell suspension was centrifuged at 10,000 rpm (30 min at 4 °C). Alkane hydroxylase activity of the cell free supernatant was assayed. The reaction mixture contained 20 mM Tris-HCl, 0.15% CHAPS buffer (pH 7.4), 0.1 mM NADH and 0.1 mM alkane (mixture of 1% alkane and 80% DMSO), and 100 μl crude extract in 1 mL volume. Enzyme activity was measured by a decrease in NADH absorbance with time at 2, 5, and 10 min as compared to 0 min by spectrophotometric analysis at 340 nm (Maeng et al. 1996; Mishra and Singh 2012). One unit of enzyme activity is the amount of enzyme required to generate 1 μmol of product per minute. Bradford method was used to determine the protein concentration present in the cellular crude extract (Bradford 1976). Bovine serum albumin was used as standard.
Catechol 2,3-dioxygenase and catechol 1,2-dioxygenase assay
Qualitative assay for catechol 2,3-dioxygenase and catechol 1,2-dioxygenase was performed on naphthalene grown cells (100 ppm) for 24 to 48 h by following the protocol of Margesin et al. (2003). Naphthalene grown cells were precipitated by centrifugation and dissolved in 100 μl of phosphate buffered saline (PBS). Liquid culture of 50 μl was added to a solution of 150 μl, which contained 90 mM of catechol dissolved in 50 mM Tris-acetate buffer (pH 7.5). Development of green-brownish color at room temperature (25 °C) within a period of 2 h in the dark indicated the presence of catechol 2,3-dioxygenase. Presence of catechol 1,2-dioxygenase was estimated with the remaining 50 μl of culture supernatant which was added with 150 μl of solution containing phenol red (0.004%), EDTA (1 mM), and catechol (10 mM). Ammonium hydroxide was added to maintain the pH at 7.5. Presence of catechol 1,2-dioxygenase resulted in change of color from red to yellow-orange when kept in dark for 10 min at 25 °C.
Nucleotide sequence and strain submission
The raw reads of 16S rRNA gene of metagenome was deposited in NCBI under BioProject ID: PRJNA498582 and sequencing data was submitted bearing ID: SRR8235268. The sequences of 16S rRNA genes of bacterial isolates obtained from this study have been deposited in the NCBI bearing accession numbers KM054633–KM054684.
Results and discussion
Physicochemical analysis and enumeration of microorganisms
Physicochemical parameters of the sludge were analyzed for assessing the environmental conditions available to the resident microbial community which has been illustrated in Table 1. The pH and temperature of the oily sludge were favorable for the survival and activity of most of the microorganisms. The moisture content and salinity were very low at 1.6% and 14 ppm, respectively. High conductivity (249.6 μS cm−1) was due to the abundance of cations and anions in the sludge sample, which is also supported by the presence of metal ions including heavy metals like Pb, Cd, Ni, Co, Zn, and As (Das and Kazy 2014). Although the ORP value was on the higher side, presence of nitrate and sulfate suggested the available electron accepting regimes, especially when oxygen concentration becomes limiting (Das and Kazy 2014; Roy et al. 2018b). The GC-MS results ascertained the abundance of cyclohexane, dodecane, pentadecane, hexadecane, naphthalene, and their derivatives in the petroleum sludge sample (Table 2). The abundance of C12–C20 compounds indicated a poor level of ongoing biodegradation within the sludge (Roy et al. 2018b). Lack of suitable condition, presence of inhibitors like heavy metals, as observed within DJ3 sample could lead to non-appreciable amount of biodegradation. Aerobically cultivable bacterial counts were nearly similar on R2A and MS Media whereas anaerobic CFU count on anaerobic agar was of one order less (Table 1). Heterotrophic bacterial counts were in the range as observed in numerous petroleum sludge samples where various physicochemical parameters play a driving force in shaping the overall bacterial community (Das and Kazy 2014; Sarkar et al. 2016; Roy et al. 2018b).
Microbial community composition in the sludge
Culture-independent analysis
Microbial community composition within the petroleum sludge was successfully explored by amplicon sequencing of 16S rRNA gene derived from the metagenome. Total sequence reads (sequence similarity > 97%) was obtained while sequencing the V4 region of the 16S rRNA gene of the sludge metagenome. Shannon diversity index of 6.36 was within the range of various other values previously reported from petroleum-associated samples (Supplementary data, Table 1). Higher Simpson’s index of 0.93 could explain the efficacy of the sequencing results (Kostka et al. 2011; Silva et al. 2013; Gao et al. 2015).
The majority of the most abundant bacterial OTUs were affiliated to Proteobacteria (almost 54%) at the phylum level (Fig. 1) which further indicated the abundance of Proteobacterial classes like γ-Proteobacteria (36.9%) followed by δ-Proteobacteria (9.2%), α-Proteobacteria (5.9%), and β-Proteobacteria (2.1%) (Supplementary data, Fig. S2). Among the other phyla, Candidate division KB1 (5.5%), Bacteroidetes (4.7%), Actinobacteria (2.4%), Firmicutes (2.3%), Synergistetes (1.2%), Spirochaetae (1.2%), and Chloroflexi (1.1%) were also detected as major groups. High abundance of such phyla have been observed in various hydrocarbon-impacted samples (Silva et al. 2013; Tan et al. 2015; Yuan et al. 2017; Galazka et al. 2018; Liu et al. 2019; Wang et al. 2019). Archaeal phylum Euryarchaeota represented 21% of the total reads mainly dominated by hydrogenotrophic and acetoclastic methanogens, which could play important roles in anaerobic alkane metabolism (Mbadinga et al. 2012). Planctomycetes, Verrucomicrobia, Elusimicrobia, Acidobacteria, candidate divisions OD1, TA06, TM7, and OP3 were also detected in low numbers and were thus considered as minor groups in this sample (Supplementary data, Fig. S1). Bacteroidia (4.1%), Clostridia (1.5%), Synergistia (1.3%), Spirochaetes (1.2%), Bacilli (0.9%), and Anaerolineae (0.5%) were the other major bacterial classes. Methanobacteria (13.1%) and Methanomicrobia (8%) were the major archaeal classes present in the sludge sample (Supplementary data, Fig. S2). Elusimicrobia, Thermoplasmata, Acidobacteria, Cyanobacteri, and unclassified Verrucomicrobia were other important classes identified in relatively lower abundance (< 0.1%) (Supplementary data, Fig. S3a and Fig. S3b).
Taxonomic distribution at the family level indicated the predominance of Pseudomonadaceae (20.2%) and Xanthomonadaceae (16.1%), both belonging to the class γ-Proteobacteria (Fig. 2). Apart from that, Porphyromonadaceae (3.9%), Erythrobacteraceae (2.1%), Caulobacteraceae (1.6%), Rhodobacteraceae (1.4%), Synergistaceae (1.3%), Bacillaceae (0.8%), Rhodocyclaceae (0.7%), Anaerolineaceae (0.5%), Syntrophaceae (0.5%), Methanobacteriaceae (13.1%), and Methanosaetaceae (5.6%) were other major bacterial and archaeal families observed in the test sample. Some of the minor (0.1% > abundance ≥ 0.01%) and rare (< 0.01%) families identified in this sample were Ruminococcaceae, Sphingobacteriaceae, Syntrophobacteraceae, Sphingomonadaceae, Syntrophorhabdaceae, Methanomicrobiaceae, and Enterobacteriaceae.
More than 300 genera have been identified to be present in DJ3 petroleum sludge sample. Pseudomonas (20.1%) was the most abundant genus followed by Pseudoxanthomonas (15.8%), Brevundimonas (1.6%), Paludibacter (1.5%), Bacillus (0.8%), Thermovirga (0.4%), Anaerobaculum (0.3%), Janthinobacterium (0.3%), Coprothermobacter (0.3%), Proteiniphilum (0.2%), and Thermotoga (0.1%) (Fig. 3). However, Smithella, Thauera, Sedimentibacter, Azoarcus, Rhizobium, Syntrophus, Burkholderia, Stenotrophomonas, and Paenibacillus were also detected as minor (0.1% > abundance ≥ 0.01%) and rare (< 0.01%) genera due to their lesser abundance. Archaeal members like Methanobacterium (9.1%), Methanosaeta (5.6%), and Methanothermobacter (3.6%) were abundant in the sludge sample followed by lower abundances of Methanolinea, Methanoculleus, Methanosarcina, Methanospirillum, Methanomethylovorans, and Methanolobus.
Isolation of culturable bacteria and their identification based on partial sequence of the bacterial 16S rRNA gene
Fifty-two bacterial isolates were obtained from the crude oil-contaminated sludge of Duliajan oil field. Phylogenetic affiliation of 52 strains was determined through 16S rRNA partial gene sequence analysis (Table 3; Fig. 4). Among the aerobically isolated 43 strains and anaerobically isolated 9 strains, two phyla were most dominant, Proteobacteria and Firmicutes. Among Proteobacteria, Gammaproteobacteria was dominant. Alphaproteobacteria and Betaproteobacteria were present only in low numbers. The dominant genera in Gammaproteobacteria were Pseudomonas (15/43) followed by Stenotrophomonas (3/43) and a single strain of Franconibacter. Rhizobium was the only genus that represented Alphaproteobacteria in this sample (2/43) and Burkholderia represented Betaproteobacteria (1/43). The phylum Firmicutes were represented by Bacillus (20/43) and Paenibacillus (1/43). Clostridium (5/9) and Serratia (2/9) were the two main genera identified through anaerobic culturing of the sludge. Pseudomonas was the most abundant genus according to the culture-independent analysis and almost 36% of the cultivable isolates represented Pseudomonas. Presence of Bacillus was also identified in high numbers in both the techniques. However, Clostridium and Serratia have not been identified through culture-independent study. Clostridium has been found as highly abundant group in methanogenic toluene-degrading sample and benzene-degrading microcosms and has been known to play vital role in methanogenic hydrocarbon degradation (Fowler et al. 2012; Gieg et al. 2014).
Distribution of culturable diversity (a); Phylogenetic tree based on bacterial 16S rRNA gene sequences was constructed using the neighbor-joining method incorporating Jukes-Cantor distance corrections (b). Sequence of Methanosarcina barkeri CM1 was used as the out group. Numbers at nodes represent bootstrap values obtained with 1000 replications. Bar 0.05 indicates 5% nucleotide substitution
Proteobacteria
The proteobacterial group was mainly dominated by Pseudomonas and Pseudoxanthomonas, both belonging to Gammaproteobacteria. Pseudomonas spp. have been reported earlier from soil contaminated with crude oil or hydrocarbons, water flooding reservoirs, and oil sands. The role of Pseudomonas in effectively remediating polycyclic hydrocarbons had long been investigated in contaminated soils, reservoirs, etc., especially in aerobic condition as well as anaerobically, where they can utilize nitrate as a potent source of terminal electron acceptor (Gao et al. 2015; Isaac et al. 2015). Many strains of Pseudomonas showed the ability to produce biosurfactant that could facilitate crude oil degradation (Chebbi et al. 2017). Zhang et al. (2011) reported P. aeruginosa strain DQ8 capable of degrading n-tetrodecane, n-docosane, n-triacontane, and n-tetrocontane. Various studies have successfully detected alkane hydroxylase gene from different strains of Pseudomonas (van Beilen et al. 2001). Pseudoxanthomonas was the second most dominant genera after Pseudomonas as found in this sample. Pseudoxanthomonas strains have been previously reported from oil-, gasoline-, and hydrocarbon-contaminated soils and sediments. Sarkar et al. (2016) reported the increase in abundance of Pseudoxanthomonas in nitrate-reducing environment during the bioremediation of petroleum refinery sludge. Thermomonas were found in good numbers in DJ3 sample, which have been reported to be slightly thermophilic and aerobic but can carry out metabolism in nitrate-reducing condition. Stenotrophomonas, which constituted nearly 5% of the culturable community isolated from DJ3 sample, have also been widely reported from oil-contaminated environment, which could perform hydrocarbon degradation, particularly of low and high molecular weight PAHs (Cerqueira et al. 2011). Franconibacter pulveris strain DJ34 was the first of its kind reported from petroleum-contaminated environment through culture-dependent study (Pal et al. 2017).
Gammaproteobacteria was followed by Deltaproteobacteria (9.2%) in DJ3 sample as the second most abundant class within the bacterial domain. Unclassified members of the order Desulfuromonadales were most abundant followed by uncultured Bdellovibrionales. Desulfuromonas, a dominant member of Deltaproteobacteria in DJ3 sample, was mostly reported as iron- or sulfur-reducing bacterium (An and Picardal 2015). A 13C-toluene-based Stable Isotope Probing (SIP) experiment showed the importance of Desulfuromonas for toluene degradation in oil-polluted tidal flats (Sampaio et al. 2017). Members of Syntrophaceae were also abundant in the test sludge sample. In the presence of methanogenic condition, this group has a direct role in activating crude oil alkane oxidation to form acetate and hydrogen (Gray et al. 2011). Syntrophaceae mainly composed of the genera Smithella and Syntrophus, which have also been previously identified from methanogenic environments containing hydrocarbons, implying their ability for syntrophic hydrocarbon metabolism. The roles of Smithella and Syntrophus in methanogenic crude oil degradation have been indicated in various studies (Gray et al. 2011; Berdugo-Clavijo and Gieg 2014; Yang et al. 2016). Short chain fatty acids like acetate, propionate, butyrate, and benzoate have been found as ubiquitous intermediates from organic compound degradation in anoxic environments. Syntrophus were also well known for the conversion of such intermediates into H2, CO2, and formate, which could further be consumed by hydrogenotrophic methanogens for methane production (Allen et al. 2007). Desulfovibrio, under the class of sulfate-reducing bacteria (SRB), could reduce sulfate by using hydrogen, organic acids, or alcohols as electron donors (Heidelberg et al. 2004). They have been previously isolated from anoxic layers of hydrocarbon-polluted microbial mats and have been enriched from production water of oil reservoirs, where they could exist in syntrophic association with members of hydrogenotrophic methanogens or Firmicutes (Abed et al. 2011; Berdugo-Clavijo and Gieg 2014).
Brevundimonas, a member of Caulobacteraceae, was the most dominant among Alphaproteobacteria in the DJ3 sample. Members of this genus have been isolated from crude oil-contaminated sludge and have been known as crude oil and hydrocarbon degraders (Mansur et al. 2014). More than 1% OTUs in the test sludge sample showed strong lineage with unclassified Rhodobacteraceae. Members of this family were found to play key roles in biodegradation of crude oil in the Deepwater Horizon oil spill-impacted beach sands of Gulf of Mexico (Kostka et al. 2011). Nitrogen fixers were abundant in the DJ3 sample, which included Rhizobium, Xanthobacter, and Azospirillum. Many strains of Rhizobium, isolated from petroleum-contaminated sludge samples, have been well known as phenanthrene degraders (Wen et al. 2011).
Betaproteobacteria in the DJ3 sample was mainly dominated by the members of Rhodocyclaceae family. Thauera and Azoarcus, two important members of this group found in this sample, have been known to degrade aromatic compounds both aerobically and anaerobically, especially in nitrate-reducing environment (Krieger et al. 1999; Shinoda et al. 2004; Das and Kazy 2014). Janthinobacterium, a member of Oxalobacteraceae, has been reported from heavily petroleum-contaminated environments, which was capable of metabolizing PAHs (Das and Kazy 2014). Burkholderia, identified in both culture-dependent and independent study of DJ3 sample, has been isolated from many petroleum-impacted habitats which showed the ability to utilize diverse hydrocarbon compounds such as MAHs, PAHs, aliphatics, and even nitro- and chloroaromatics (Sarkar et al. 2017; Yuan et al. 2017).
Bacteroidetes
This group has also been previously reported from crude petroleum sludge, hydrocarbon-contaminated soils, contaminated wetlands, marine sediments, and petroleum reservoir and was considered to be composed of organisms associated with the degradation of many organic compounds, benzene, toluene, and other hydrocarbons in sulfate-reducing anaerobic environments (Kostka et al. 2011; Das and Kazy 2014; Wang et al. 2019). Paludibacter and Proteiniphilum are strict anaerobic bacteria, which could be involved in soil carbon cycling by fermenting plant polymers and dead microbial biomass with the subsequent production of short chain fatty acids and alcohols along with CO2, H2O, and organic acids like acetates and propionates (Müller et al. 2009; Yang et al. 2016). Sphingobacterium were known to produce biosufactants during growth in organic materials. Their biodegradation ability, especially for aromatic hydrocarbons, have also been determined (Noparat et al. 2014).
Firmicutes
The phylum Firmicutes included both aerobic and anaerobic microorganisms in different petroleum-contaminated habitats, which were known to utilize different hydrocarbons during growth. Firmicutes have been successfully isolated or identified from arctic or permafrost regions affected by hydrocarbon contamination, contaminated soil from oil reservoirs, deep subsurface petroleum reservoirs, petroleum sludge, river, coastal, and salt march sediments impacted by oil spills (Das and Kazy 2014; Yang et al. 2016; Sierra-Garcia et al. 2017; Yuan et al. 2017; Liu et al. 2019). Microbial community data from a wide range of studies have demonstrated Firmicutes to be present in the highest relative abundance, especially from oil- and hydrocarbon-contaminated anoxic environments (Head et al. 2014). Clostridia, the well-known Firmicutes member, were found to play vital role in fermenting alkanes, especially in high temperatures (Mbadinga et al. 2012; Head et al. 2014). The genus Bacillus, with an abundance of 0.8%, was the major genus found within DJ3 sample followed by Coprothermobacter. Metabolically diverse members of Bacillus were reported to be involved in degradation of crude oil and its components including various PAHs viz. phenanthrene, fluorene, pyrene, and acenaphthalene, as well as short to long chain aliphatics along with concomitant production of biosurfactant (Tao et al. 2017; Das and Kazy 2014; Yuan et al. 2017). Coprothermobacter, affiliated to the class Clostridia and order Thermoanaerobacterales, has been reported as anaerobic, thermophilic, fermentative organism, capable of degrading proteins or peptides, and mostly found in syntrophic association with hydrogenotrophic methanogenic archaea (Gagliano et al. 2015). Fusibacter has been found to be associated with oil producing wells with the ability to degrade PAHs under anaerobic condition (Kappell et al. 2014). Paenibacillus spp. have been well known for inhabiting petroleum-contaminated soil, oil sludge, and marsh lands (Das and Kazy 2014). Several recent studies also confirmed the potential of Paenibacillus in utilizing various other PAHs, crude oil, and alkanes along with their ability to produce bioemulsifiers (Das and Kazy 2014; Reddy et al. 2017).
Spirochaetes, Chloroflexi, Verrucomicrobia, and Planctomycetes
Members of Spirochaetes, Chloroflexi, Verrucomicrobia, and Planctomycetes were also detected in high or low abundance in hydrocarbon-impacted habitats. Spirochaetes, Chloroflexi, and Bacteroidetes members have been frequently found in methanogenic hydrocarbon-associated environments, where they could perform an essential role in the transformation of hydrocarbon degradation intermediates, mainly the fatty acids to methanogenic substrates, thus contributing to the recycling of waste products and maintaining a low redox potential within the enrichment.
Members of Spirochaetaceae have previously been reported from hexadecane-degrading methanogenic consortium obtained from Shengli oilfields, crude oil-degrading methanogenic enrichments, flooding water samples of petroleum reservoirs, hydrocarbon-contaminated oily sludge, and soil (Das and Kazy 2014; Gao et al. 2015; Ma et al. 2017). Thermotogaceae, Synergistaceae, and Chlorofexi along with Spirochaetaceae have been considered to play key roles either as secondary degraders during hexadecane degradation or as scavengers of anabolic products (lipids and proteins), probably derivative of detrital microbial biomass (Ma et al. 2017). The most abundant Thermotogae member found in DJ3 sample was Fervidobacterium, a fermentative thermophilic bacterium often isolated from petroleum reservoirs, which showed huge biotechnological applications due to production of thermostable enzymes applied for conversion of biomass to biofuel (Saxena et al. 2017). The most abundant Nitrospirae member in the test DJ3 sample was Thermodesulfovibrio, which were known as sulfate-reducing microorganisms that could cooperate with other bacteria or methanogens syntrophically in alkane-degrading processes (Liang et al. 2016). Nitrospira reported from hydrocarbon-contaminated aquifer, soil, and thermophilic methanogenic alkane-degrading consortium might play a key role in the two-step nitrification process in soil and in activated sludge (Daims et al. 2015; Liang et al. 2016).
Candidate phyla
The second most abundant candidate division in the test sample was JS1, the highest being KB1. KB1 has no such reports to be identified from hydrocarbon-impacted environments but has been known to possess osmoregulatory mechanisms thereby adapting to high saline concentrations and diverse metabolic routes to counteract to various nutritional conditions (Nigro et al. 2016). JS1 has proposed to be a member of candidate phylum Atribacteria along with OP9. JS1 and OP9 have previously been observed from petroleum reservoirs and other related environments such as sediments associated with methane hydrates and hydrocarbon seeps; hypersaline microbial mats, and geothermal environments (Nobu et al. 2016). OP8 have been affiliated to the member of Aminicenantes, which appeared to be the most abundant in hydrocarbon-impacted environments, especially in anoxic habitats with slightly high temperatures (Farag et al. 2014).
Euryarchaeota
Euryarchaeota have been observed as a dominant archaeal group in heavily oil-contaminated environments, oil reservoirs, oil-contaminated soil, oil water mixture of petroleum reservoirs, and petroleum-contaminated sludge (Liu et al. 2009; Das and Kazy 2014; Gao et al. 2015; Sierra-Garcia et al. 2017). Methanogenesis has been considered as the most dominant terminal process in such anaerobic environments, for significant degradation and considerable mineralization of hydrocarbons (Li et al. 2017).
Members of Euryarchaeota in DJ3 sample were mostly characterized by sequences closely related to methanogens of the order Methanobacteriales (Methanobacterium, Methanothermobacter), Methanosarcinales (Methanosaeta, Methanomethylovorans, Methanosarcina, Methanolobus), and Methanomicrobiales (Methanolinea, Methanoculleus, Methanospirillum). The most abundant archaeon in the test sample was related to Methanobacterium, a hydrogenotrophic methanogen utilizing hydrogen and formate as substrate for methanogenesis (Gao et al. 2014). Methanothermobacter, another hydrogenotrophic methanogen, abundant in DJ3 sample, has also been frequently recovered from high temperature oil reservoirs that could metabolize hydrocarbon and carbon dioxide into methane (Ren et al. 2011; Zhao et al. 2012; Chen et al. 2019). H2, CO2, and formate are the final products of short chain fatty acid fermentation carried out by fermentative bacteria (e.g., Clostridium, Syntrophus, Acidiphilium, and Anaerobaculum). Hydrogenotrophic methanogens living in close association with such fermentative bacteria could convert CO2 and H2 into CH4 (Liu et al. 2016).
Methanosaeta and Methanosarcina, important members of Methanosarcinales, have been known as acetoclastic methanogens, which take part in the process of converting acetate into methane and CO2 (Li et al. 2017). Methanosaeta members have been detected from various hydrocarbon-impacted environments in high abundance and observed to be capable of performing anaerobic mineralization of long chain alkanes in consortium with Anaerolineae or other Anaerolineaceae members (Liang et al. 2015; Toth and Gieg 2018). Presence of both Methanosaeta and Anaerolineae in DJ3 sample might suggest the possibility of natural attenuation taking place in this particular site. Moreover, Zhao et al. (2012) suggested the abundance of methanogens might be closely related to bacterial community dominated by Clostridia, in which several species could be involved in anaerobic crude oil degradation, thus producing various low molecular weight organic acids (such as formic acid, acetic acid, and methyl amine) for the utilization by hydrogenotrophic, acetoclastic, or methylotrophic methanogens to produce CH4. Biodegradation of hydrocarbons through methanogenesis is important in many environments, especially contaminated groundwater, sediments, and oil reservoirs, where electron acceptors such as oxygen, nitrate, sulfate, and iron (III) are lacking (Head et al. 2014; Tan et al. 2015). Methanogenic hydrocarbon metabolism involves a metabolic cooperation of syntrophic fermentative bacteria that catalyzes the initial attack on the hydrocarbon substrates thereby forming methanogenic substrates (such as hydrogen, acetate, propionate, and formate), which might be used by the methanogens to produce CH4 (Gray et al. 2010). The initial anaerobic oxidation of substrates is thermodynamically unfavorable under standard condition. The methanogens thus play a key role in making this process favorable by keeping those intermediates in low concentrations (Gieg et al. 2014).
Analysis of metabolic potential of sludge microbial community using PICRUSt tool
PICRUSt analysis predicted the putative metabolic inventory of native microbial community in the DJ3 petroleum sludge sample (Fig. 5). Broad classes of metabolic pathway genes within the sample were classified into metabolism (49.7%); genetic information processing (17.1%); environmental information processing (13.1%); cellular processes (4.8%), and unclassified group (15.2%). A comprehensive analysis of metabolic category showed highest allocation of genes related to amino acid metabolism (10.6%) followed by carbohydrate metabolism (9.6%) and energy metabolism (6.3%). Genes related to xenobiotic biodegradation and metabolism also accounted a significant proportion (3.4%). Allocation of various genes related to metabolism, survival of microorganisms, and genes involved directly or indirectly in hydrocarbon degradation have been listed in Supplementary data, Table 2.
The most abundant within the xenobiotic biodegradation and metabolism was the genes involved in benzoate metabolism (0.51%) followed by aminobenzoate metabolism (0.34%). Significant proportion of genes was also related to degradation of monoaromatics, polyaromatics, and chlorinated hydrocarbons. Cytochrome P450 family of genes involved in the aerobic degradation of aliphatic hydrocarbons and cyclo-alkanes have been detected in this sample. These genes have previously been reported from members of Acinetobacter, Alcanivorax, Caulobacter, and Mycobacterium from various oil-polluted environments (Omrani et al. 2018; Neethu et al. 2019). These groups of genes are quite evident in degradation of low to high molecular weight PAHs (Crampon et al. 2018). Other noteworthy observations included the abundance of naphthalene (0.25%) and polycyclic aromatic hydrocarbon (0.11%) degrading genes. This was evident since this particular sample consisted of high amounts of alkanes and aromatics like naphthalene. PICRUSt also predicted a large number of genes able to carry out BTEX degradation (0.4%), chloroalkane (0.24%), chlorocyclohexane, and chlorobenzene (0.1%) degradation. The abundance of genes related to xenobiotic degradation especially from samples contaminated with petroleum that harbor a large number of hydrocarbonoclastic organisms was also evident from various previous studies (Wang et al. 2016; Mukherjee et al. 2017; Roy et al. 2018a). Such observation was in agreement with the study of Roy et al. (2018a), where a similar percentage of xenobiotic-degrading gene abundance was demonstrated in the petroleum sludge samples showing maximum TPH degradation. Reconstruction of the metabolic pathways using the KEGG database showed the presence of naphthalene 1,2-dioxygenase and 1,2-dihydroxynaphthalene dioxygenase responsible to carry out naphthalene degradation especially by Pseudomonas spp. which was also abundant in this particular sample. Gentisate is a key intermediary metabolite in the degradation of polycyclic or monocyclic aromatic hydrocarbons and cresols. Members of Pseudomonas and Sphingomonas have been detected with gentisate 1,2-dioxygenase gene (Yeo et al. 2007). Yergeau et al. (2012) showed the rise in gentisate 1,2-dioxygenase gene especially in members of Betaproteobacteria and Acidobacteria during the degradation of diesel-contaminated soil, whereas homogentisate 1,2-dioxygenase increased in all Proteobacterial members. Catechol and protocatechuates could be detected as intermediate products of aerobic aromatic hydrocarbon degradation. Genes encoding for catechol 2,3-dioxygenase and catechol 1,2-dioxygenase have also been detected in large numbers in the test sample and their roles in biodegradation have been illustrated in various studies. The study also predicted the presence of benzene 1,2-dioxygenase, ethylbenzene dioxygenase, 2,3-dihydroxyethylbenzene, and 1,2-dioxygenase for the degradation of monoaromatic hydrocarbons. Genes encoding for benzylsuccinate synthase (bssA) have also been detected by previous investigators, which highlighted the probability of monoaromatic hydrocarbon degradation in anaerobic condition (Callaghan et al. 2010; Tan et al. 2015). Presence of alkane-1-monooxygenes further suggested the abundance of alkane-degrading microbial population in this alkane rich test sample. Alkanesulfonate monooxygenase ensured the degradation of sulfur containing alkane compounds (Pal et al. 2017). Even the detected cytochrome P450 hydroxylase could also contribute toward alkane degradation. More than 1% of the total gene counts have been assigned for methane metabolism (~1.5%) in the test DJ3 sample. This could properly match with the fact that the DJ3 sample has a high percentage of Euryarchaeota, thus performing the process of methanogenesis as well as methylotrophy. Methanobacterium, Methanosaeta, and Methanothermobacter were known to carry out the process of methanogenesis, whereas members of Verrucomicrobia, Paracoccus, Methylobacterium, Methylophilus, Methylophaga, Methanomethylovorans, and Methanolobus, as observed in this sample, could carry out the process of methanotrophy and methylotrophy (Chistoserdova et al. 2009; Li et al. 2017). These two processes have been found to be important in hydrocarbon mineralization. PICRUSt analysis has been used to illustrate high abundance of genes involved in methanogenesis and methylotrophy in oil-contaminated environments (Mukherjee et al. 2017; Roy et al. 2018a). A high abundance of methane monooxygenase gene as observed from the KEGG analysis also supported the fact that methane thus produced by the methanogens could be used up by the aerobic methanotrophic bacteria as this particular gene has been reported to be present in all aerobic methanotrophic bacteria with minor exception and this could help in explaining the methane flux dynamics in such contaminated environments, where the deep anoxic layer is connected with the upper oxic layer through the association of the methanogens and the aerobic methanotrophs (Zhou et al. 2015).
Metabolic characterization of bacterial isolates
Characterization of the aerobic isolates was further performed based on their metabolic ability of utilizing various hydrocarbons as sole carbon source during growth, ability of producing biosurfactants, and tolerate various environmental stress conditions (Supplementary data, Fig. S4). Almost all the strains could grow optimally at ambient temperature range of 25–35 °C, but their growth declined at temperature below 15 °C and above 40 °C. Similar observation was recorded for pH variation. pH range of 5–7 was the most suitable for bacterial growth. However, their growth declined below pH 5 and above pH 9. Good growth was observed for most of the strains above 5% NaCl condition suggesting the ability of the strains to resist high salinity. Production of biosurfactant by microorganisms enhances their ability to reduce oil viscocity by facilitating emulsification and thereby increasing biodegradation of hydrocarbon compounds (Pacwa-Plociniczak et al. 2014). Based on the E24 value (emulsification index after 24 h), it was observed that almost 90% of the strains could produce biosurfactants. Bacterial resistance to Pb, Ni, and Cd was also tested at 1 mM concentration of each heavy metal. Although almost all the strains could grow in presence of Pb and Ni, Cd proved to be toxic to the cells as only 30% isolates were able to tolerate it.
Growth in presence of BTEX (200 ppm) showed that almost 80% of the strains could utilize these compounds. Majority of the strains (95%) could also utilize and degrade pentadecane, while near about 80% of the strains could do the same with hexadecane. However, the proportion of alkane degradation varied for different strains (Fig. 6). Strains DJ25, DJ30, DJ31, DJ32, and DJ34 effectively degraded both the alkanes. The isolates preferred pentadecane over hexadecane as sole carbon source probably due to the shorter chain length of the former. Similar results have also been previously demonstrated wherein greater cell yield was obtained when pentadecane was used as carbon source rather than hexadecane (Wang et al. 2006). Almost all the strains could utilize petroleum sludge (1%, w/v) as carbon source and crude oil (1%, v/v) was utilized by 80% of the population. Growth of the microorganisms could be supported due to the complex nature of lighter to heavier fractions of petroleum hydrocarbons. It has been observed that the lighter fractions of petroleum hydrocarbons have been utilized by microorganisms in faster rates as compared to the medium to heavier fractions. The enzyme assay suggested the expression of alkane hydroxylase enzyme by many strains while growing in presence of alkane (pentadecane and hexadecane). The strains that showed the highest growth in and degradation of alkanes also showed higher activity level for this particular enzyme. The highest activity (~0.54 μmol/min/mg) was shown by whole cell lysate of DJ34 strain followed by DJ8, DJ11, DJ15, DJ16, DJ19, DJ20, DJ30, DJ32, and DJ-E10 strains (Supplementary data, Table 3). Similar kind of alkane hydroxylases enzyme activity was observed in Pseudomonas putida GPo1 during the hydroxylation of n-octane (Alonso and Roujeinikova 2012). The result of the enzyme assay was also comparable to the study of Maeng et al. (1996) which showed that the specific activity of whole cell lysate was approximately 0.09–0.1 μmol/min/mg. Most of the strains were positive for catechol 2,3-dioxygenase production except DJ11, DJ17, and DJ-E2 (Supplementary data, Table 3). However, catechol 1,2-dioxygenase was poorly activated and only few strains listed in the table have shown a moderately positive result. This might be due to higher concentration of aromatic compound that has been used for bacterial growth as it could inhibit the expression of catechol 1,2-dioxygenase (Hupert-Kocurek et al. 2012).
UPGMA (Unweighted Pair Group Method with Arithmetic mean)-based statistical analysis was carried out to investigate relationship among the bacterial isolates with respect to their different metabolic activities like hydrocarbon degradation, tolerance to various environmental stress conditions and heavy metals, growth under oxic/anoxic conditions by utilizing multiple electron acceptors, and biosurfactant production (Fig. 7). The non-taxonomic clustering based on the above-mentioned properties represented small clades of Bacillus spp. and Pseudomonas spp., which could be due to their versatile metabolic abilities in hydrocarbon-associated environments. However, an interesting observation was the cladding of enriched strains suggesting the strains to be metabolically similar considering the above-mentioned parameters. The strain DJ34 exhibited the best result for diverse petroleum hydrocarbon utilization and growth in presence of different electron acceptors. The strain also showed heavy metal resistance, biosurfactant production, and could tolerate various physicochemical conditions prevailing in oil-contaminated habitats.
Conclusion
Culture-independent analysis of the petroleum sludge microbial community obtained through NGS-based sequencing technique revealed the presence of diverse group of microorganisms capable of hydrocarbon degradation in both aerobic and anaerobic conditions. Nitrate-reducing, sulfate-reducing, metal-reducing, fermentative, syntrophic, methanogenic, N2-fixing, CO2-fixing, and biosurfactant-producing microorganisms have also been detected in abundance which could assist the process of hydrocarbon degradation. PICRUSt analysis predicted putative genetic collection within the sludge microbiome and its potential for various hydrocarbon and xenobiotic compound degradation, lipid, nitrogen, sulfur, and methane metabolism. Cultivable bacterial community has also supported the culture-independent method as Pseudomonas strains proved to be dominant in both the methods of determining the microbial community. Most of the cultivable isolates indicated their ability to degrade petroleum hydrocarbons and thereby sustain the harsh environmental conditions prevailing in those contaminated habitats. The overall community composition could indicate an associated interaction and possible metabolic interplay between the microorganisms which could potentially be employed for in situ oily sludge bioremediation.
References
Abed RMM, Musat N, Musat F, Mußmann M (2011) Structure of microbial communities and hydrocarbon-dependent sulfate reduction in the anoxic layer of a polluted microbial mat. Mar Pollut Bull 62:539–546. https://doi.org/10.1016/j.marpolbul.2010.11.030
Allen JP, Atekwana EA, Atekwana EA, Duris JW, Rossbach S (2007) The microbial community structure in petroleum-contaminated sediments corresponds to geophysical signatures. Appl Environ Microbiol 73:2860–2870
Alonso H, Roujeinikova A (2012) Characterization and two-dimensional crystallization of membrane component AlkB of the medium-chain alkane hydroxylase system from Pseudomonas putida GPo1. Appl Environ Microbiol 78(22):7946–7953
An TT, Picardal FW (2015) Desulfuromonas carbonis sp. nov., an Fe(III)-, S0- and Mn(IV)-reducing bacterium isolated from an active coalbed methane gas well. Int J Syst Evol Microbiol 65:686–1693
Berdugo-Clavijo C, Gieg LM (2014) Conversion of crude oil to methane by a microbial consortium enriched from oil reservoir production waters. Front Microbiol 5:197. https://doi.org/10.3389/fmicb.2014.00197
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72:248–254. https://doi.org/10.1016/0003-2697(76)90527-3
Callaghan AV, Davidova IA, Savage-Ashlock K, Parisi VA, Gieg LM, Suflita JM, Kukor JJ, Wawrik B (2010) Diversity of benzyl- and alkylsuccinate synthase genes in hydrocarbon-impacted environments and enrichment cultures. Environ Sci Technol 44:7287–7294. https://doi.org/10.1021/es1002023
Caporaso JG, Kuczynski J, Stombaugh J, Bittinger K, Bushman FD, Costello EK, Fierer N, Peña AG, Goodrich JK, Gordon JI, Huttley GA, Kelley ST, Knights D, Koenig JE, Ley RE, Lozupone CA, McDonald D, Muegge BD, Pirrung M, Reeder J, Sevinsky JR, Turnbaugh PJ, Walters WA, Widmann J, Yatsunenko T, Zaneveld J, Knight R (2010) QIIME allows analysis of high throughput community sequencing data. Nat Methods 7:335–336. https://doi.org/10.1038/nmeth.f.303
Caporaso JG, Lauber CL, Walters WA, Berg-Lyons D, Lozupone CA, Turnbaugh PJ, Noah Fierer N, Knight R (2011) Global patterns of 16S rRNA diversity at a depth of millions of sequences per sample. Proc Natl Acad Sci U S A 108:4516–4522. https://doi.org/10.1073/pnas.1000080107
Cerqueira VS, Hollenbach EB, Maboni F, Vainstein MH, Camargo FAO, Peralba MCR, Bento FM (2011) Biodegradation potential of oily sludge by pure and mixed bacterial cultures. Bioresour Technol 102:11003–11010. https://doi.org/10.1016/j.biortech.2011.09.074
Chandra S, Sharma R, Singh K, Sharma A (2013) Application of bioremediation technology in the environment contaminated with petroleum hydrocarbon. Ann Microbiol 63:417–431. https://doi.org/10.1007/s13213-012-0543-3
Chebbi A, Hentati D, Zaghden H, Baccar N, Rezgui F, Chalbi M, Sayadi S, Chamkha M (2017) Polycyclic aromatic hydrocarbon degradation and biosurfactant production by a newly isolated Pseudomonas sp. strain from used motor oil-contaminated soil. Int Biodeterior Biodegrad 122:128–140. https://doi.org/10.1016/j.ibiod.2017.05.006
Chen J, Liu Y-F, Zhou L, Mbadinga SM, Yang T, Zhou J, Liu J-F, Yang S-Z, Gu J-D, Mu B-Z (2019) Methanogenic degradation of branched alkanes in enrichment cultures of production water from a high-temperature petroleum reservoir. Appl Microbiol Biotechnol 103(5):2391–2401
Chistoserdova L, Kalyuzhnaya MG, Lidstrom ME (2009) The expanding world of methylotrophic metabolism. Annu Rev Microbiol 63:477–499. https://doi.org/10.1146/annurev.micro.091208.073600
Crampon M, Bodilis J, Portet-Koltalo F (2018) Linking initial soil bacterial diversity and polycyclic aromatic hydrocarbons (PAHs) degradation potential. J Hazard Mater 359:500–509. https://doi.org/10.1016/j.jhazmat.2018.07.088
Daims H, Lebedeva EV, Pjevac P, Han P, Herbold C, Albertsen M, Jehmlich N, Palatinszky M, Vierheilig J, Bulaev A, Kirkegaard RH, von Bergen M, Rattei T, Bendinger B, Nielsen PH, Wagner M (2015) Complete nitrification by Nitrospira bacteria. Nature 528:504–509. https://doi.org/10.1038/nature16461
Das R, Kazy SK (2014) Microbial diversity, community composition and metabolic potential in hydrocarbon contaminated oily sludge: prospects for in situ bioremediation. Environ Sci Pollut Res 21:7369–7389. https://doi.org/10.1007/s11356-014-2640-2
Farag IF, Davis JP, Youssef NH, Elshahed MS (2014) Global patterns of abundance, diversity and community structure of the Aminicenantes (Candidate Phylum OP8). PLoS One 9:e92139. https://doi.org/10.1371/journal.pone.0092139
Fowler SJ, Dong X, Sensen CW, Suflita JM, Gieg LM (2012) Methanogenic toluene metabolism: community structure and intermediates. Environ Microbiol 14:754–764. https://doi.org/10.1111/j.1462-2920.2011.02631.x
Fowler SJ, Toth CRA, Gieg LM (2016) Community structure in methanogenic enrichments provides insight into syntrophic interactions in hydrocarbon-impacted environments. Front Microbiol 7:562. https://doi.org/10.3389/fmicb.2016.00562
Fuentes S, Méndez V, Aguila P, Seeger M (2014) Bioremediation of petroleum hydrocarbons: catabolic genes, microbial communities and applications. Appl Microbiol Biotechnol 98:4781–4794. https://doi.org/10.1007/s00253-014-5684-9
Gagliano MC, Braguglia CM, Gianico A, Mininni G, Nakamura K, Rossetti S (2015) Thermophilic anaerobic digestion of thermal pretreated sludge: role of microbial community structure and correlation with process performances. Water Res 68:498–509. https://doi.org/10.1016/j.watres.2014.10.031
Galazka A, Grzadziel J, Galazka R, Ukalska-Jaruga A, Strzelecka J, Smreczak B (2018) Genetic and functional diversity of bacterial microbiome in soils with long term impacts of petroleum hydrocarbons. Front Microbiol 9:1923. https://doi.org/10.3389/fmicb.2018.01923
Gao P-K, Li G-Q, Zhao L-X, Dai X-C, Tian H-M, Dai L-B, Wang HB, Huang HD, Chen YH, Ma T (2014) Dynamic processes of indigenous microorganisms from a low-temperature petroleum reservoir during nutrient stimulation. J Biosci Bioeng 117:215–221. https://doi.org/10.1016/j.jbiosc.2013.07.009
Gao P, Tian H, Li G, Sun H, Ma T (2015) Microbial diversity and abundance in the Xinjiang Luliang long-term water-flooding petroleum reservoir. MicrobiolOpen 42:332–342. https://doi.org/10.1002/mbo3.241
Gieg LM, Fowler SJ, Berdugo-Clavijo C (2014) Syntrophic biodegradation of hydrocarbon contaminants. Curr Opin Biotechnol 27:21–29. https://doi.org/10.1016/j.copbio.2013.09.002
Gray ND, Sherry A, Hubert C, Dolfing J, Head IM (2010) Methanogenic degradation of petroleum hydrocarbons in subsurface environments: remediation, heavy oil formation, and energy recovery. Adv Appl Microbiol 72:137–161. https://doi.org/10.1016/S0065-2164(10)72005-0
Gray ND, Sherry A, Grant RJ, Rowan AK, Hubert CRJ, Callbeck CM, Aitken CM, Jones DM, Adams JJ, Larter SR, Head IM (2011) The quantitative significance of Syntrophaceae and syntrophic partnerships in methanogenic degradation of crude oil alkanes. Environ Microbiol 13:2957–2975. https://doi.org/10.1111/j.1462-2920.2011.02570.x
Hazen TC, Rocha AM, Techtmann SM (2013) Advances in monitoring environmental microbes. Curr Opin Biotechnol 24:526–533. https://doi.org/10.1016/j.copbio.2012.10.020
Head IM, Gray ND, Larter SR (2014) Life in the slow lane; biogeochemistry of biodegraded petroleum containing reservoirs and implications for energy recovery and carbon management. Front Microbiol 5:566. https://doi.org/10.3389/fmicb.2014.00566
Heidelberg JF, Seshadri R, Haveman SA, Hemme CL, Paulsen IT, Kolonay JF, Eisen JA, Ward N, Methe B, Brinkac LM, Daugherty SC, Deboy RT, Dodson RJ, Durkin AS, Madupu R, Nelson WC, Sullivan SA, Fouts D, Haft DH, Selengut J, Peterson JD, Davidsen TM, Zafar N, Zhou L, Radune D, Dimitrov G, Hance M, Tran K, Khouri H, Gill J, Utterback TR, Feldblyum TV, Wall JD, Voordouw G, Fraser CM (2004) The genome sequence of the anaerobic, sulfate-reducing bacterium Desulfovibrio vulgaris Hildenborough. Nat Biotechnol 22:554–559. https://doi.org/10.1038/nbt959
Hupert-Kocurek K, Guzik U, Wojcieszyńska D (2012) Characterization of catechol 2,3-dioxygenase from Planococcus sp. strain S5 induced by high phenol concentration. Acta Biochim Pol 59:345–351
Isaac P, Martinez FL, Bourguignon N, Sanchez LA, Ferrero MA (2015) Improved PAHs removal performance by a defined bacterial consortium of indigenous Pseudomonas and actinobacteria from Patagonia, Argentina. Int Biodeterior Biodegrad 101:23–31. https://doi.org/10.1016/j.ibiod.2015.03.014
Ismail WA, Van Hamme DJ, Kilbane JJ, Gu DJ (2017) Editorial: petroleum microbial biotechnology: challenges and prospects. Front Microbiol 8:833. https://doi.org/10.3389/fmicb.2017.00833
Ji J-H, Liu Y-F, Zhou L, Mbadinga SM, Pan P, Chen J, Liu J-F, Yang S-Z, Sand W, Gu J-D, Mu BZ (2019) Methanogenic degradation of long n-alkanes requires fumarate-dependent activation. Appl Environ Microbiol 85(16):e00985–e00919. https://doi.org/10.1128/AEM.00985-19
Kappell AD, Wei Y, Newton RJ, Nostrand JDV, Zhou J, McLellan SL, Hristova KR (2014) The polycyclic aromatic hydrocarbon degradation potential of Gulf of Mexico native coastal microbial communities after the Deepwater Horizon oil spill. Front Microbiol 5:1–13. https://doi.org/10.3389/fmicb.2014.00205
Kleinsteuber S, Schleinitz KM, Vogt C (2012) Key players and team play: anaerobic microbial communities in hydrocarbon-contaminated aquifers. Appl Microbiol Biotechnol 94:851–873. https://doi.org/10.1007/s00253-012-4025-0
Kostka JE, Prakash O, Overholt WA, Green SJ, Freyer G, Canion A, Delgardio J, Norton N, Hazen TC, Huettel M (2011) Hydrocarbon degrading bacteria and the bacterial community response in Gulf of Mexico beach sands impacted by the deepwater horizon oil spill. Appl Environ Microbiol 77:7962–7974. https://doi.org/10.1128/AEM.05402-11
Kostka JE, Teske AP, Joye SB, Head IM (2014) The metabolic pathways and environmental controls of hydrocarbon biodegradation in marine ecosystems. Front Microbiol 5:471. https://doi.org/10.3389/fmicb.2014.00471
Krieger CJ, Beller HR, Reinhard M, Spormann AM (1999) Initial reactions in anaerobic oxidation of m-Xylene by the denitrifying bacterium Azoarcus sp. strain T. J Bacteriol 181:6403–6410
Langille MG, Zaneveld J, Caporaso JG, McDonald D, Knights D, Reyes JA, Beiko RG (2013) Predictive functional profiling of microbial communities using 16S rRNA marker gene sequences. Nat Biotechnol 319:814–821. https://doi.org/10.1038/nbt.2676
Li X-X, Mbadinga SM, Liu J-F, Zhou L, Yang S-Z, Gu J-D, Mu B-Z (2017) Microbiota and their affiliation with physiochemical characteristics of different subsurface petroleum reservoirs. Int Biodeterior Biodegrad 120:170–185. https://doi.org/10.1016/j.ibiod.2017.02.005
Liang B, Wang LY, Mbadinga SM, Liu JF, Yang SZ, Gu JD, Mu BZ (2015) Anaerolineaceae and Methanosaeta turned to be the dominant microorganisms in alkanes-dependent methanogenic culture after long-term of incubation. AMB Express 5:1–13. https://doi.org/10.1186/s13568-015-0117-4
Liang B, Wang L-Y, Zhou Z, Mbadinga SM, Zhou L, Liu J-F et al (2016) High frequency of Thermodesulfovibrio spp. and Anaerolineaceae in Association with Methanoculleus spp. in a long-term incubation of n-alkanes-degrading methanogenic enrichment culture. Front Microbiol 7:1431. https://doi.org/10.3389/fmicb.2016.01431
Liu R, Zhang Y, Ding R, Li D, Gao Y, Yang M (2009) Comparison of archaeal and bacterial community structures in heavily oil-contaminated and pristine soils. J Biosci Bioeng 108:400–407. https://doi.org/10.1016/j.jbiosc.2009.05.010
Liu J-F, Mbadinga SM, Sun X-B, Yang G-C, Yang S-Z, Gu J-D, Mu BZ (2016) Microbial communities responsible for fixation of CO2 revealed by using mcrA, cbbM, cbbL, fthfs, fefe-hydrogenase genes as molecular biomarkers in petroleum reservoirs of different temperatures. Int Biodeterior Biodegrad 114:164–175. https://doi.org/10.1016/j.ibiod.2016.06.019
Liu J, Wu J, Lin J, Zhao J, Xu T, Yang Q, Zhao J, Zhao Z, Song X (2019) Changes in the microbial community diversity of oil exploitation. Genes 10:556. https://doi.org/10.3390/genes10080556
Ma T-T, Liu L-Y, Rui J-P, Yuan Q, Feng D-S, Zhou Z, Dai LR, Zeng WQ, Zhang H, Cheng L (2017) Coexistence and competition of sulfate-reducing and methanogenic populations in an anaerobic hexadecane-degrading culture. Biotechnol Biofuels 10:207. https://doi.org/10.1186/s13068-017-0895-9
Maeng JH, Sakai Y, Tani Y, Kato N (1996) Isolation and Characterization of a novel oxygenase that catalyzes the first step of n-alkane oxidation in Acinetobacter sp. Strain M-1. J Bacteriol 178:3695–3700. https://doi.org/10.1128/jb.178.13.3695-3700.1996
Maiti S, Moon UR, Bera P, Samanta T, Mitra A (2014) The in vitro antioxidant capacities of Polianthes tuberosa L. flower extracts. Acta Physiol Plant 36:2597–2605. https://doi.org/10.1007/s11738-014-1630-9
Mansur AA, Adetutu EM, Kadali KK, Morrison PD, Nurulita Y, Ball AS (2014) Assessing the hydrocarbon degrading potential of indigenous bacteria isolated from crude oil tank bottom sludge and hydrocarbon-contaminated soil of Azzawiya oil refinery, Libya. Environ Sci Pollut Res 21:10725–10735. https://doi.org/10.1007/s11356-014-3018-1
Margesin R, Gander S, Zacke G, Gounot AM, Schinner F (2003) Hydrocarbon degradation and enzyme activities of cold-adapted bacteria and yeasts. Extremophiles 7:451–458. https://doi.org/10.1007/s00792-003-0347-2
Mbadinga SM, Li KP, Zhou L, Wang LY, Yang SZ, Liu JF, Gu JD, Mu BZ (2012) Analysis of alkane-dependent methanogenic community derived from production water of a high-temperature petroleum reservoir. Appl Microbiol Biotechnol 962:531–542. https://doi.org/10.1007/s00253-011-3828-8
Mishra S, Singh SN (2012) Microbial degradation of n-hexadecane in mineral salt medium as mediated by degradative enzymes. Bioresour Technol 111:148–154. https://doi.org/10.1016/j.biortech.2012.02.049
Mukherjee A, Chettri B, Langpoklakpam JS, Basak P, Prasad A, Mukherjee AK (2017) Bioinformatic approaches including predictive metagenomics profiling reveal characteristics of bacterial response to petroleum hydrocarbon contamination in diverse environments. Sci Rep 7:1108. https://doi.org/10.1038/s41598-017-01126-3
Müller S, Vogt C, Laube M, Harms H, Kleinsteuber S (2009) Community dynamics within a bacterial consortium during growth on toluene under sulfate-reducing conditions. FEMS Microbiol Ecol 70:586–596. https://doi.org/10.1111/j.1574-6941.2009.00768.x
Neethu CS, Saravanakumar C, Purvaja R, Robin RS, Ramesh R (2019) Oil-spill triggered shift in indigenous microbial structure and functional dynamics in different marine environmental matrices. Sci Rep 9:1354. https://doi.org/10.1038/s41598-018-37903-x
Nigro LM, Hyde AS, MacGregor BJ, Teske A (2016) Phylogeography, salinity adaptations and metabolic potential of the candidate division KB1 bacteria based on a partial single cell genome. Front Microbiol 7:1266. https://doi.org/10.3389/fmicb.2016.01266
Nobu MK, Dodsworth JA, Murugapiran SK, Rinke C, Gies EA, Webster G, Schwientek P, Kille P, Parkes RJ, Sass H, Jørgensen BB, Weightman AJ, Liu WT, Hallam SJ, Tsiamis G, Woyke T, Hedlund BP (2016) Phylogeny and physiology of candidate phylum ‘Atribacteria’ (OP9/JS1) inferred from cultivation independent genomics. ISME J 10:273–286. https://doi.org/10.1038/ismej.2015.97
Noparat P, Maneerat S, Saimmai A (2014) Application of biosurfactant from Sphingobacterium spiritivorum AS43 in the biodegradation of used lubricating oil. Appl Biochem Biotechnol 172:3949–3963. https://doi.org/10.1007/s12010-014-0829-y
Omrani R, Spini G, Puglisi E, Saidane D (2018) Modulation of microbial consortia enriched from different polluted environments during petroleum biodegradation. Biodegradation 29:187–209. https://doi.org/10.1007/s10532-018-9823-3
Pacwa-Plociniczak MP, Plaza GN, Poliwoda A, Seget ZP (2014) Characterization of hydrocarbon-degrading and biosurfactant-producing Pseudomonas sp. P-1 strain as a potential tool for bioremediation of petroleum-contaminated soil. Environ Sci Pollut Res 21:9385–9395. https://doi.org/10.1007/s11356-014-2872-1
Pal S, Kundu A, Banerjee TD, Mohapatra B, Roy A, Manna R, Sar P, Kazy SK (2017) Genome analysis of crude oil degrading Franconibacter pulveris strain DJ34 revealed its genetic basis for hydrocarbon degradation and survival in oil contaminated environment. Genomics 109:374–382
Paul D, Kazy SK, Gupta AK, Pal T, Sar P (2015) Diversity, metabolic properties and arsenic mobilization potential of indigenous bacteria in arsenic contaminated groundwater of West Bengal. India. PLoS One 10(3):e0118735. https://doi.org/10.1371/journal.pone.0118735
Poi G, Shahsavari E, Aburto-Medina A, Mok PC, Ball AS (2018) Large scale treatment of total petroleum-hydrocarbon contaminated groundwater using bioaugmentation. J Environ Manag 214:157–163. https://doi.org/10.1016/j.jenvman.2018.02.079
Reddy MV, Devi MP, Chandrasekhar K, Goud RK, Mohan SV (2011) Aerobic remediation of petroleum sludge through soil supplementation: microbial community analysis. J Hazard Mater 197:80–87. https://doi.org/10.1016/j.jhazmat.2011.09.061
Reddy PV, Karegoudar TB, Monisha TR, Nayak AS (2017) Catabolism of fluorene through 2,3-dihydroxy indanone in Paenibacillus sp. PRNK-6. Int Biodeterior Biodegrad 123:156–163. https://doi.org/10.1016/j.ibiod.2017.05.019
Ren H-Y, Zhang X-J, Song Z-Y, Rupert W, Gao G-J, Guo S-X, Zhao L-P (2011) Comparison of microbial community compositions of injection and production well samples in a long-term water-flooded petroleum reservoir. PLoS One 6:e23258. https://doi.org/10.1371/journal.pone.0023258
Røberg S, Østerhus JI, Landfald B (2011) Dynamics of bacterial community exposed to hydrocarbons and oleophilic fertilizer in high-Artic intertidal beach. Polar Biol 34:1455–1465. https://doi.org/10.1007/s00300-011-1003-4
Roy A, Dutta A, Pal S, Gupta A, Sarkar J, Chatterjee A, Saha A, Sarkar P, Sar P, Kazy SK (2018a) Biostimulation and bioaugmentation of native microbial community accelerated bioremediation of oil refinery sludge. Bioresour Technol 253:22–32. https://doi.org/10.1016/j.biortech.2018.01.004
Roy A, Sar P, Sarkar J, Dutta A, Sarkar P, Gupta A, Mohapatra B, Pal S, Kazy SK (2018b) Petroleum hydrocarbon rich oil refinery sludge of North-East India harbours anaerobic, fermentative, sulfate-reducing, syntrophic and methanogenic microbial populations. BMC Microbiol 18:151. https://doi.org/10.1186/s12866-018-1275-8
Sampaio DS, Almeida JRB, de Jesus HE, Rosado AS, Seldin L, Jurelevicius D (2017) Distribution of anaerobic hydrocarbon-degrading bacteria in soils from King George Island, Maritime Antarctica. Microb Ecol 744:810–820
Sarkar J, Kazy SK, Gupta A, Dutta A, Mohapatra B, Roy A, Bera P, Mitra A, Sar P (2016) Biostimulation of indigenous microbial community for bioremediation of petroleum refinery sludge. Front Microbiol 7:1407. https://doi.org/10.3389/fmicb.2016.01407
Sarkar P, Roy A, Pal S, Mohapatra B, Kazy SK, Maiti MK, Sar P (2017) Enrichment and characterization of hydrocarbon-degrading bacteria from petroleum refinery waste as potent bioaugmentation agent for in situ bioremediation. Bioresour Technol 242:15–27. https://doi.org/10.1016/j.biortech.2017.05.010
Saxena R, Dhakan DB, Mittal P, Waiker P, Chowdhury A, Ghatak A, Sharma VK (2017) Metagenomic analysis of hot springs in Central India reveals hydrocarbon degrading thermophiles and pathways essential for survival in extreme environments. Front Microbiol 7:2123. https://doi.org/10.3389/fmicb.2016.02123
Shinoda Y, Sakai Y, Uenishi H, Uchihashi Y, Hiraishi A, Yukawa H (2004) Aerobic and anaerobic toluene degradation by a newly isolated denitrifying bacterium, Thauera sp. strain DNT-1. Appl Environ Microbiol 70:1385–1392. https://doi.org/10.1128/aem.70.3.1385-1392.2004
Sierra-Garcia IN, Dellagnezze BM, Santos VP, Chaves BMR, Capilla R, Neto EVS (2017) Microbial diversity in degraded and non-degraded petroleum samples and comparison across oil reservoirs at local and global scales. Extremophiles 21:211–229. https://doi.org/10.1007/s00792-016-0897-8
Silva TR, Verde LCL, Santos Neto EV, Oliveira VM (2013) Diversity analyses of microbial communities in petroleum samples from Brazilian oil fields. Int Biodeterior Biodegrad 81:57–70. https://doi.org/10.1016/j.ibiod.2012.05.005
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731–2739. https://doi.org/10.1093/molbev/msr121
Tan B, Fowler SJ, Laban NA, Dong X, Sensen CW, Foght J, Gieg LM (2015) Comparative analysis of metagenomes from three methanogenic hydrocarbon-degrading enrichment cultures with 41 environmental samples. ISME J 9:2028–2045. https://doi.org/10.1038/ismej.2015.22
Tao K, Liu X, Chen X, Hu X, Cao L, Yuan X (2017) Biodegradation of crude oil by a defined co-culture of indigenous bacterial consortium and exogenous Bacillus subtilis. Bioresour Technol 224:327–332. https://doi.org/10.1016/j.biortech.2016.10.073
Tian Y, Wan Y-Y, Mu H-M, Dong H-L, Briggs BR, Zhang Z-H (2019) Microbial diversity in high-temperature heavy oil reservoirs. Geomicrobiol J 37:59–66. https://doi.org/10.1080/01490451.2019.1662523
Toth CRA, Gieg LM (2018) Time course-dependent methanogenic crude oil biodegradation: dynamics of fumarate addition metabolites, biodegradative genes, and microbial community composition. Front Microbiol 8:2610. https://doi.org/10.3389/fmicb.2017.02610
van Beilen JB, Panke S, Lucchini S, Franchini AG, Röthlisberger M, Witholt B (2001) Analysis of Pseudomonas putida alkane-degradation gene clusters and flanking insertion sequences: evolution and regulation of the alk genes. Microbiology 147:1621–1630
Van Hamme JD, Singh A, Ward OP (2003) Recent advances in petroleum microbiology. Microbiol Mol Biol Rev 67:503–549. https://doi.org/10.1128/MMBR.67.4.503-549.2003
Wang L, Tang Y, Wang S, Liu RL, Liu MZ, Zhang Y, Liang FL, Feng L (2006) Isolation and characterization of a novel thermophilic Bacillus strain degrading long-chain n-alkanes. Extremophiles 10:347–356. https://doi.org/10.1007/s00792-006-0505-4
Wang H, Wang B, Dong W, Hu X (2016) Co-acclimation of bacterial communities under stresses of hydrocarbons with different structures. Sci Rep 6:34588. https://doi.org/10.1038/srep34588
Wang X, Li X, Yu L, Huang L, Xiu J, Lin W, Zhang Y (2019) Characterizing the microbiome in petroleum reservoir flooded by different water sources. Sci Total Environ 653:872–885. https://doi.org/10.1016/j.scitotenv.2018.10.410
Wen Y, Zhang J, Yan Q, Li S, Hong Q (2011) Rhizobium phenanthrenilyticum sp. nov., a novel phenanthrene-degrading bacterium isolated from a petroleum residue treatment system. J Gen Appl Microbiol 57:319–329. https://doi.org/10.2323/jgam.57.319
Yang S, Wen X, Zhao L, Shi Y, Jin H (2014) Crude oil treatment leads to shift of bacterial communities in soils from the deep active layer and upper permafrost along the China-Russia crude oil pipeline route. PLoS One 9:e96552. https://doi.org/10.1371/journal.pone.0096552
Yang S, Wen X, Shi Y, Liebner S, Jin H, Perfumo A (2016) Hydrocarbon degraders establish at the costs of microbial richness, abundance and keystone taxa after crude oil contamination in permafrost environments. Sci Rep 6:37473. https://doi.org/10.1038/srep37473
Yeo CC, Tan CL, Gao XL, Zhou B, Pho CL (2007) Characterization of hbzE-encoded gentisate 1,2-dioxygenase from Pseudomonas alcaligenes NCIMB 9867. Res Microbiol 158:608–616. https://doi.org/10.1016/j.resmic.2007.06.003
Yergeau E, Sanschagrin S, Beaumier D, Greer CW (2012) Metagenomic analysis of the bioremediation of diesel contaminated Canadian high arctic soils. PLoS One 7:e30058. https://doi.org/10.1371/journal.pone.0030058
Yuan K, Chen B, Qing Q, Zou S, Wang X (2017) Polycyclic aromatic hydrocarbons (PAHs) enrich their degrading genera and genes in human-impacted aquatic environments. Environ Pollut 230:936–944. https://doi.org/10.1016/j.envpol.2017.07.059
Zhang Z, Hou Z, Yang C, Ma C, Tao F, Xu P (2011) Degradation of n-alkanes and polycyclic aromatic hydrocarbons in petroleum by a newly isolated Pseudomonas aeruginosa DQ8. Bioresour Technol 102:4111–4116
Zhao L, Ma T, Gao M, Gao P, Cao M, Zhu X, Li G (2012) Characterization of microbial diversity and community in water flooding oil reservoirs in China. World J Microbiol Biotechnol 28:3039–3052. https://doi.org/10.1007/s11274-012-1114-2
Zhou Z, Chen J, Cao H, Han P, Gu J-D (2015) Analysis of methane-producing and metabolizing archaeal and bacterial communities in sediments of the northern South China Sea and coastal Mai Po Nature Reserve revealed by PCR amplification of mcrA and pmoA genes. Front Microbiol 5:1–15. https://doi.org/10.3389/fmicb.2014.00789
Acknowledgements
We gratefully acknowledge the support of Oil India Limited, Duliajan, Assam, in sample collection.
Availability of data and materials
Not applicable.
Funding
This work was supported by NER TWINNING Project (grant no. T/226/NE/TBP/2011) from the Department of Biotechnology, Government of India.
Author information
Authors and Affiliations
Contributions
Sufia Kazy and Pinaki Sar conceptualized and designed the work methodology, arranged funds and logistics, and supervised. Ajoy Roy and Pinaki Sar were responsible for sampling the crude oil containing sludge from Duliajan oil field and further physicochemical analysis. Siddhartha Pal, Avishek Dutta, and Jayeeta Sarkar performed the major experiments. Siddhartha Pal and Sufia Kazy were responsible for manuscript preparation. All authors read and approved the manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare no competing interests.
Additional information
Responsible Editor: Robert Duran
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Fig. S1
Relative abundance of minor bacterial and archaeal phyla with cumulative abundance <0.1 % in DJ3 sample; Uncls., Unclassified; div., division (PDF 52 kb)
Fig. S2
Relative abundance of major bacterial and archaeal classes with cumulative abundance ≥ 0.1 % in DJ3 sample; Uncls., Unclassified; div., division; Uncult., Uncultured; Misc., Miscellaneous (PDF 63 kb)
Fig. S3a
Relative abundance of minor bacterial and archaeal classes with cumulative abundance <0.1 % but ≥0.01 % in DJ3 sample; Uncls., Unclassified; div., division; Uncult., Uncultured; Misc., Miscellaneous. b Relative abundance of rare bacterial and archaeal classes with cumulative abundance <0.01 % in DJ3 sample; Uncls., Unclassified; div., division; Uncult., Uncultured; Misc., Miscellaneous (PDF 122 kb)
Fig. S4
Summary of aerobic isolates showing positive response towards various physiological and metabolic properties. Growth of percentage of isolates at different pH (a); salinity (b); temperature (c); electron acceptors (d); heavy metals (e); hydrocarbons (f) (PDF 77 kb)
Table S1
(PDF 78 kb)
Table S2
(PDF 79 kb)
Table S3
(PDF 95 kb)
Rights and permissions
About this article
Cite this article
Pal, S., Dutta, A., Sarkar, J. et al. Exploring the diversity and hydrocarbon bioremediation potential of microbial community in the waste sludge of Duliajan oil field, Assam, India. Environ Sci Pollut Res 28, 50074–50093 (2021). https://doi.org/10.1007/s11356-021-13744-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11356-021-13744-6