Abstract
Antibiotic resistance genes (ARGs) in rural environments have been poorly characterized in the literature. In this study, the diversity, abundance, and distribution of ARGs in surface waters, soils, and sediments of a typical hilly rural area in the Upper Yangtze River watershed were investigated using the high-throughput quantitative polymerase chain reaction, and their relationships with chemical properties of the samples were analyzed. No significant differences in the diversity and abundance of ARGs were observed among the three medium types while the ARG distribution pattern in the sediments was obviously different from that of the surface waters. According to the co-occurrence pattern of ARGs subtypes obtained by network analysis, blaOXA10-02, blaPSE, lnuB-02, and qacEΔ1-01 can be used to estimate the relative abundance of total ARGs for the study area. It appeared that the prevalence of ARGs in the sediments was promoted by the horizontal gene transfer (HGT) and vertical gene transfer together, while their spread in the surface waters and soils were facilitated by the supply of biogenic elements and HGT, respectively. Mobile genetic elements (MGEs) were abundant and detected in all samples, and their abundance was significantly and positively correlated with that of ARGs, implying that the potential horizontal transfer of ARGs to other bacteria and pathogens in rural environments should not be overlooked.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
The rapid dissemination of antibiotic resistance genes (ARGs) and their potential transmission to clinical pathogens are identified as a severe threat to public health in the twenty-first century (Forsberg et al. 2012; WHO 2014). These emerging environmental pollutants have attracted world-wide attention and great efforts have been made to address their occurrence and spread in various environmental settings (Pruden et al. 2006; Zhu et al. 2013; Ou-yang et al. 2015; Xu et al. 2016b). The use of clinical and veterinary antibiotics generally results in dispersal and propagation of ARGs in natural environments (Sidrach-Cardona et al. 2014; Ou-yang et al. 2015; Chen et al. 2018). Additionally, environmental tolerant bacteria were reported to be phylogenetically close to clinical pathogens (Dantas et al. 2008), indicating the potential transfer of ARGs from environmental bacteria to human pathogens via horizontal gene transfer (HGT) (Forsberg et al. 2012). Therefore, the comprehensive examination of the abundance and diversity of ARGs in multiple environments is essential to control the propagation of environmental ARGs (Li et al. 2015; Chen et al. 2019a).
China is the largest antibiotic producer and consumer in the world. The national antibiotic consumption (normalized by defined daily doses) in China was nearly 6 times that in the UK and USA (Zhang et al. 2015). The antibiotic usage in food animals of China accounted for 20% of the global consumption in 2010 (Van Boeckel et al. 2015). The overuse and misuse of antibiotics have ultimately contributed to the obvious increase in antibiotic resistance in China (Qiao et al. 2017). Thereby, diverse and abundant ARGs are frequently detected in diverse environmental locals (e.g., soils, surface waters, and sediments) (Luo et al. 2010; Zhu et al. 2013; Ou-yang et al. 2015). However, most of these studies are conducted in the hotspots with intensive anthropogenic activities, like urban cities, animal feedlots, and estuaries (Chen et al. 2013; He et al. 2014; Xu et al. 2016b). Information about the occurrence of ARGs in less-impacted rural areas was limited and reported as the low human activity control, focusing on individual environmental medium or a limited number of resistance genes (Luo et al. 2010; Jiang et al. 2013; Ou-yang et al. 2015; Xiang et al. 2018; Zheng et al. 2018). Considering China’s large rural population, a comprehensive and integral investigation to profile the antibiotic resistomes in rural areas is needed. In particular, with the rapid socioeconomic development of rural areas in China in the past decades, increasing amounts of various wastewaters such as household sewage and wastewater from small-scale livestock farms have been produced. These rural wastewaters likely contain not only abundant nutrients (Kumwimba et al. 2016), but also various microbial contaminates like pathogens (Xue et al. 2018) and ARGs (Chen and Zhang 2013; Chen et al. 2015). In Hangzhou located in eastern China, four tetracycline resistance genes (tetM, tetO, tetQ, and tetW) and two sulfonamide genes (sulI and sulII) were detected in all influent samples from eight rural domestic sewage treatment systems and four municipal wastewater treatment plants (Chen and Zhang 2013). There were no significant differences in sul gene abundances between the two types of influent samples; in contrast, the abundances of tet genes were higher and showed less variation in municipal wastewaters compared to rural domestic sewage (Chen and Zhang 2013). Likewise, in Shanghai, the antibiotic and ARG contaminations of some raw drinking water sources located in or near its suburban rural areas were more severe than those in its urban sites (Jiang et al. 2011, 2013). In East Tiaoxi River, higher levels of bacterial pathogen were detected in rural areas than those in urban areas, and rural areas, especially arable land, trended to be high spots for ARG shift and pathogen propagation (Zheng et al. 2017). Moreover, waterborne ARGs could spread into river sediments through sorption and deposition in association with particles or enter agricultural soils through irrigation, resulting in the dispersal and propagation of ARGs in receiving sediments and soils (Luo et al. 2010; Wang et al. 2014). Therefore, knowledge of the resistomes in soils, surface waters, and sediments surrounding residential areas and livestock farms in China’s rural areas is crucial to developing appropriate mitigation strategies and countermeasures for controlling ARG contamination in rural environments.
The vast hilly region (160,000 km2) in central Sichuan Basin in the upper Yangtze River watershed is dominated by agriculture. The region, which is characterized by a thin layer of easily erodible and poorly structured purple soil (an entisol according to USDA soil taxonomy) on slopes, is the primary source of sediments in the upper reaches of the Yangtze River (Zhu et al. 2009). In addition to non-point agricultural sources of contaminants, poorly treated domestic sewage from dispersed town residents and family livestock farms contributed substantially to the pollution of rural surface waters and groundwater in this region (Kumwimba et al. 2016; Xue et al. 2018). Thereby, large amounts of abiotic and biotic contaminants, either in dissolved/free forms or in association with sediments, could be transported toward the downstream fluvial system via runoff and thus may deteriorate the water quality of the upper reaches of the Yangtze River and the Three Gorges Reservoir. As an emerging pollutant, however, the occurrence and distribution of ARGs in different rural environments in this hilly region have not been reported in the literature. It should be noted that there have also been limited studies on ARG occurrence in rural environmental locals in other countries (Guo et al. 2018a; McConnell et al. 2018). Most of the small number of relevant studies in rural areas reported the phenotypic data of antibiotic resistant bacteria and pathogens isolated from human, animal, and natural environments (Hagedorn et al. 1999; Braykov et al. 2016), and a few others evaluated the prevalence of ARGs in water bodies (Bergeron et al. 2017; McConnell et al. 2018) and feces and production chickens of small-scale farms (Guo et al. 2018a). To our knowledge, no investigations linking the occurrence and distribution of ARGs in multiple environmental media of rural areas have been reported around the globe.
A field survey was carried out in a hilly rural area of the upper Yangtze River watershed to examine the diversities and abundances of ARGs in various environmental media. The main objectives of this study were (1) to screen a broad-spectrum of ARGs in surface waters, soils, and sediments at various potential hot spots using high-throughput quantitative polymerase chain reaction (HT-qPCR); (2) to assess the similarity/difference of ARGs distribution patterns among environmental compartments using principal coordinate analysis (PCoA); and (3) to reveal the co-occurrence pattern between ARGs subtypes using network analysis.
Materials and methods
Study area and sample collection
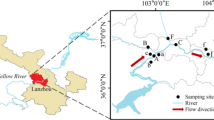
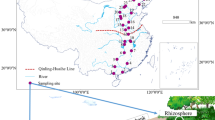
The study area (31°16′ N, 105°28′ E) is located at Yanting, Sichuan Province, Southwest China, where the Yanting Agro-Ecological Experimental Station of Purple Soil, Chinese Academy of Sciences, was established in 1980. The study area’s altitude ranges from 400 to 600 m above sea level. It features a moderate subtropical monsoon climate with an annual mean temperature of 17.3 °C and an average annual precipitation of 826 mm (1981 to 2009). The annual precipitation is unevenly distributed, with approximately 70% of annual precipitation occurring in the period from May to September. The study area has a high population density of 120 residents per km2 (Xue et al. 2018).
A total of 27 samples were collected from 14 sampling locations in the study area in September 2017. The sampling sites include Wanan weir (WAW), Daxing weir (DXW), Jieliu weir (JLW), and Surong weir (SRW) of Linshan river, domestic sewage weir of Linshan Town (DSW), outlet of Gongyu river which is a tributary to Linshan river (GYR), Zhangjia pig feedlot (PF) and bovine feedlot (BF), Longquan chicken feedlot (CF), Wujia fishpond (FP), an ecological ditch (ED), a domestic sewage irrigated vegetable field (DSIF), a human excrement irrigated vegetable field (HEIF), and a small orchard with chicken raising under the trees (OC) (Fig. S1). Overall, 7 surface water samples, 6 soil samples, and 14 sediment samples were collected in this study (Table S1). Along the flow direction in the ecological ditch, five sediment samples (ED-1 to ED-5) were taken from the five ditch segments planted with Zizania latifolia, Thalia dealbata, Hydrocotyle vulgaris, Canna indica, and Acorus calamus, respectively. At the time of sampling, no flowing waters presented in the ED, SRW, and DSW, and thus, no surface water samples were available at these sites for analysis. At each location, three subsamples were randomly collected and mixed thoroughly to obtain one composite sample (each 2 L of water, 200 g of soil or sediment). All composite samples were transported to the laboratory on ice immediately and pretreated within 24 h for further processing.
Five hundred milliliters of each surface water sample was filtered through a 0.22-μm mixed cellulose ester membrane filter (Millipore, USA) to entrap ARG-containing materials on the membrane. Total DNA was extracted from each freeze-dried soil/sediment sample (0.25 g) and each filtered membrane for water sample using the PowerSoil DNA Isolation Kit (MoBio Laborataries Inc., USA). Each sample was extracted twice and the eluted DNA solutions were combined. The concentration and quality of extracted DNA were examined by spectrophotometric analysis with a NanoDrop ND-2000 (NanoDrop Technologies Inc., USA), diluted to 35 ng/μL, and then stored at − 20 °C for further analysis.
Chemical analysis
Selected chemical parameters of the surface water samples, including total nitrogen (TN), nitrate (NO3−-N), ammonium (NH4+-N), and dissolved organic carbon (DOC), were measured using a continuous flow analyzer (Auto Analyzer 3, SEAL Analytical, Norderstedt, Germany). Before analysis, soil and sediment samples were freeze-dried using a vacuum freeze dryer (FD-1A-50, Biocool, China) and ground. Total nitrogen (TN) and total organic carbon (TOC) of soil and sediment samples were determined using a CN analyzer (Varia EL, Elementar GmbH, Germany) after removing carbonates with 1 mol L−1 HCl (Sinopharm, China) at a 1:2.5 soil to water ratio (Su et al. 2014). Inorganic nitrogen (including NO3−-N and NH4+-N) in soil and sediment samples was extracted with 2 mol L−1 KCl (Sinopharm, China) at a soil to water ratio of 1:10 and then determined using an AA3 continuous flow analyzer (SEAL Analytical, Norderstedt, Germany).
Quantitative analysis of ARGs by HT-qPCR array
HT-qPCR array approach was employed to evaluate the diversity and abundance of ARGs and MGEs using the SmartChip Real-time PCR system (Warfergen Inc. USA). A total of 296 primer sets were used to investigate the antibiotic resistome (283 primer sets), transposase genes (8 primer sets), class 1 integron-integrase genes (4 primer sets), and 16S rRNA gene (Xiang et al. 2018; Li et al. 2019). The analyzed 283 ARGs in this study encode resistance to almost all major classes of antibiotics, including fluoroquinolone/quinolone/florfenicol/chloramphenicol/amphenicol (FCA), multidrug, aminoglycoside, macrolide/lincosamide/streptogramin B (MLSB), beta-lactamase, sulfonamide, tetracycline, vancomycin, and other. A detailed description about the qPCR conditions and amplification procedure can be found in previous studies (Zheng et al. 2017, 2018).
The results of the HT-qPCR array were analyzed using SmartChip qPCR software (V 2.7.0.1). A threshold cycle (CT) of 31 was used as the detection limit (Ou-yang et al. 2015). Reactions with amplification efficiencies beyond the range (1.8–2.2) or with multiple melting peaks were discarded, and the remained wells were regarded as positive amplification. Only the data with three positive technical replicates were used in subsequent analysis. Gene copy number was calculated as described in a previous study (Zheng et al. 2018): gene copy number = \( {10}^{\left(31-{\mathrm{C}}_{\mathrm{T}}\right)\left(10/3\right)} \), where CT refers to quantitative PCR results. The relative abundance of ARGs was calculated by normalizing each gene’s copy number to the 16S rRNA’s copy number. The absolute abundance of ARGs was recorded as the product of the relative abundance and the 16S rRNA’s absolute copy number that was quantified separately using a Roche 480 system (Roche Molecular Systems Inc., USA) as described previously (Ou-yang et al. 2015). For individual samples, ARG diversity was represented by the number of detected ARG subtypes. The genes observed in at least one sample of all the 27 samples or all the samples of each environmental medium type were used to assess the ARG diversity (represented by numbers of detected ARG subtypes) across all the samples of different environmental media or the samples of each environmental medium, respectively. The diversity was also calculated in percentage for ARG classes of different resistance mechanisms or corresponding antibiotics across all the 27 samples.
Statistical analysis
Basic statistical analyses (sum, mean, etc) on raw data were conducted in Excel 2010 (Microsoft office 2010, Microsoft, USA). Principal coordinate analysis (PCoA) based on Bray−Curtis distances was used to evaluate the overall pattern of ARGs among the samples. Pearson correlation analysis was applied to explore the relationships between ARGs, mobile genetic elements (MGEs), and chemical properties. PCoA, Adonis test, Pearson correlation, heatmap, and Circos graph were performed in R 3.5.3 (R Core Team 2019) with the packages “vegan” (Oksanen et al. 2018), “psych” (Revelle 2018), “pheatmap” (Kolde 2018), and “circlize” (Gu et al. 2014). All statistical tests were considered significant at P < 0.05. Other diagrams were generated using “ggplot2” (Wickham 2016) and “ggpubr” (Kassambara 2018) packages in R 3.5.3 (R Core Team 2019). To evaluate the co-occurrence patterns of 77 ARG and MGE subtypes that occurred in at least six samples out of all samples, network analysis based on statistically robust correlations (Spearman’s correlation coefficient > 0.80 and P < 0.01) was conducted in R environment and visualized using the Gephi 0.9.2 (Bastian et al. 2009) as described by Li et al. (2015). Likewise, bipartite network visualization that depicted the shared and unique ARGs between different environmental media was also conducted on Gephi 0.9.2 (Bastian et al. 2009). In bipartite network analysis, a gene was considered to be present in an environmental medium if it was present in at least half of the samples of this medium type.
Results
Chemical properties of surface waters, soils, and sediments
Selected chemical properties of the 27 samples are shown in Tables S2 and S3. In surface water samples, the NO3−-N concentration differed greatly among the sampling sites, while the TN, NH4+-N, and DOC concentrations were similar at different locations, except the PF site where the highest values of these properties were detected (Table S2). In soil samples, the highest TN, NH4+-N, and TOC contents were observed at the OC site while the highest NO3−-N content was detected at the HEIF site (Table S3). All the sediment samples collected from different sites displayed comparable TN and TOC contents. The NH4+-N contents in the ED-1 to ED-5 sediment samples were lower than those of the sediment samples collected from the other sites. Contrastingly, the highest NO3−-N contents were detected in the ED-1 to ED-5 sediment samples (Table S3).
Diversity and abundance of antibiotic resistance genes
A total of 163 ARGs were detected across all the samples. The detected ARGs potentially conferred resistance to almost all major antibiotics widely used for human medicines and animal production and represented three major resistance mechanisms: antibiotic deactivation (44.79%), efflux pump (33.74%), and cellular protection (19.63%) (Fig. S2a). Overall, multidrug and aminoglycoside resistance genes accounted for 20.25% and 18.40%, respectively, followed by beta-lactamase (17.79%), tetracycline (16.56%), and MLSB (13.50%) resistance genes (Fig. S2b). Regarding individual environmental medium types, the surface waters, soils, and sediments harbored 117, 108, and 139 ARGs, respectively. Analysis of variance showed that the detected ARG numbers were comparable between the soils and sediment samples, and observably but insignificantly (P > 0.05) lower numbers of ARGs were detected in surface water samples than the soil and sediment samples (Fig. S3). The numbers of ARGs detected in individual samples ranged from 15 to 72, 16 to 57, and 12 to 71 for the surface water, soil, and sediment samples, respectively (Fig. 1).
The abundances of ARGs in individual samples from different environmental habitats are illustrated in Fig. S4. ARG absolute abundances in the surface water, soil, and sediment samples ranged from 5.96 × 107 to 6.69 × 109 copies L−1, 4.27 × 108 to 2.64 × 1010 copies g−1, 2.62 × 108 to 8.67 × 1010 copies g−1, respectively. Being similar to the above results in terms of detected numbers of ARGs, resistance genes to multidrug, aminoglycoside, beta-lactamase, and tetracycline were the most abundant ARG classes in rural samples. To minimize potential variations in background bacterial load among different samples, the absolute abundances of ARGs were normalized by 16S rRNA (Fig. S4). Apparently, the relative abundances of ARGs in different environmental media followed this order: sediments > soils > surface waters (Fig. 2a), although the differences did not reach a statistically significant level (ANOVA, P > 0.05). Moreover, 10 out of the 12 mobile genetic elements targeted on our arrays were detected, with the clinical and universal class 1 integron-integrase genes being the most abundant MGEs (Fig. S5). Noteworthy, the occurrence of MGEs across all tested samples, in terms of both absolute and relative abundance, displayed a similar trend to that of ARGs (Fig. S4).
The overall patterns of ARGs in the surface waters were significantly different from that of the sediments (Adonis, P < 0.05), while ARG patterns in the soils were close to those in the surface waters and sediments (Adonis, P > 0.05). Results of PCoA using Bray−Curtis distances based on the relative abundances of ARGs (Fig. 2b) showed that the soil samples were neither clustered together nor separated from the samples of the other two medium types. Different separation patterns were observed between the surface waters and sediments despite their dispersed distributions. Specifically, the surface waters were separated primarily along PC2 (explained 16.50% of the variation of ARGs) while the sediments were separated mostly along PC1 (explained 24.81% of the variation).
Shared major antibiotic resistome among surface waters, soils, and sediments
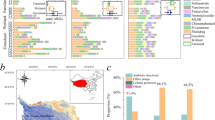
A total of 36 major ARGs (detected in at least half of the samples within any one of the three environmental media) were identified in all test samples. These genes were used to perform a bipartite network analysis for identification of shared and unique ARGs in different environmental media. The identified major antibiotic resistome can be categorized into six groups, as described clockwise in Fig. 3: (a) five ARGs and one integrase (intI3) that were found only in the surface waters; (b) four ARGs (qacEΔ1-01, sul2, aadA-01, and cphA-01) and three MGEs (intI-1LC, tnpA-04, and tnpA-05) that were concurrently detected in the surface waters, soils, and sediments; (c) 8 ARGs that were exclusively detected in the soils; (d) integrase intI-1(clinic) and 12 ARGs that were shared between the soils and sediments; (e) 3 ARGs that were detected only in the sediments; (f) 4 ARGs that were shared between the surface waters and sediments. Resistome groups b, d, and f, which concurrently harbored in more than one environmental medium, were of major concern. Notably, there were no genes shared only between soil and water samples. The total abundance of the four shared major ARGs in group B accounted for 40–73% of the total ARG abundance in three environmental media (Table S4).
Co-occurrence pattern among ARGs subtypes
A co-occurrence network among ARGs containing 28 nodes and 34 edges with a modularity index of 0.524 and a clustering coefficient of 0.390 was constructed. The entire topology could be parsed into seven modules based on modularity (Fig. 4). Modules I and II were the two largest ones. The most densely connected nodes in their module were defined as module hub genes. For instance, blaXOA10-02 and qacEΔ1-01 were the hub genes of the module I, and lnuB-02 and blaPSE were the hub genes of the model II. These hub genes might be served as markers for co-occurring ARGs in the same module. Additionally, MGEs also show significant connections with ARGs. Specifically, integrase intI-1LC and transposase tnpA-01 co-occurred with aadA2-02 and lnuB-01, respectively, while tnpA-07 co-occurred with more than one unique ARGs, including catB3 (FCA), dfrA1(sulfonamide), lnuB-01 (MLSB), and blaPSE (beta-lactamase) (Fig. 4).
Relationships among ARGs, MGEs, and environmental factors
On the whole, both the absolute and relative abundance of MGEs were significantly (P < 0.01) and positively correlated with those of total ARGs, FCA, aminoglycoside, MLSB, multidrug, and sulfonamide resistance genes (Table 1). Correlations of the absolute abundance of total ARGs with those of MGEs and 16S rRNA genes (Fig. 5) showed that the absolute abundance of total ARGs was correlated significantly (P < 0.01) with that of total MGEs (r = 0.847) and that of 16S rRNA genes (r = 0.745) in the sediments, while the absolute abundance of total ARGs in the soils was significantly correlated with MGEs only (r = 0.814, P < 0.05); no significant relationships between the absolute abundances of total ARGs and those of 16S rRNA and total MGEs were observed in the surface waters.
The relative abundances of total ARGs (P < 0.05) and some ARG types (P < 0.01) in the surface waters were positively correlated with TN, NH4+-N, and DOC concentrations (Table S5). The relative abundances of total ARGs and some ARG types were positively correlated with TOC content in the sediments (P < 0.01) but not in the soils. Notably, in all three environmental media, NO3−-N level exhibited no significant correlations with the relative abundances of total ARGs and individual ARG subtypes (P > 0.05).
Discussion
Occurrence of ARGs in various rural environmental media
In this study, antibiotic deactivation and efflux pumps, via transforming antibiotics into low-activity metabolites and extruding antibiotics out of cells, respectively (Chen et al. 2018), were the two major antibiotic resistance mechanisms. This finding is congruent with the observations in soils of 32 wetlands on the Qing-Tibet Plateau (Yang et al. 2019), soils of Chinese natural forests (Hu et al. 2018), manures, composts, and soils of three large-scale Chinese swine farms (Zhu et al. 2013), and previously investigated manured upland soils in the same area as the present study (Cheng et al. 2019). Both mechanisms can mitigate the intracellular selective pressure and therefore prevail in both pristine and polluted circumstances (Yang et al. 2019). Being similar to the results of previous screening of ARGs across Chinese water bodies and estuarine sediments (Zhu et al. 2017; Liu et al. 2018), the genes conferring resistance to multidrug in this study were also highly diverse and abundant, which could be caused by the extensive use of multiple clinical and veterinary antibiotics. The presence and spread of multidrug resistance genes may pose a potential human health risk; therefore, it is of great significance to control the development and dispersal of multidrug resistance, especially its potential to be acquired by pathogens.
The number of detected ARGs in the surface waters in this rural study area was comparable to that in Chinese lakes and reservoirs (15–57 ARGs per sample) (Liu et al. 2018), but lower than that in the East Tiaoxi River (84–105 ARGs per sample) (Zheng et al. 2017) and Zhangxi River (46–154 ARGs per sample) (Zheng et al. 2018). The absolute abundances of detected ARGs were at a similar level to those in inland water bodies (106 to 109 copied L−1) (Liu et al. 2018; Chen et al. 2019a), but much lower than those in urban rivers, like Jiulong River (108 to 1011 copied L−1) (Ou-yang et al. 2015), Qiantang River (109 to 1011 copied L−1) (Xu et al. 2016a), and Wenruitang River (108 to 1011 copied L−1) (Zhou et al. 2017). The lower ARG levels observed in the present study than those of urban/peri-urban rivers can be attributed to the less exogenous input in this rural area due to the lower anthropogenic activities. Likewise, the diversity (26–57 ARGs per sample) and absolute abundance (108 to 1010 copies g−1) of ARGs in the soils were lower than those previously observed in the peri-urban soils (70–128 ARGs per sample and 108 to 1012 copies g−1) of Ningbo in eastern China (Xiang et al. 2018) and the urban soils collected from Melbourne (210) in Australia and Belfast (164) in Northern Ireland (Yan et al. 2019; Zhao et al. 2019). The number of detected ARGs in the sediments (139) was almost half of that in Chinese estuarine sediments (248) with intensive anthropogenic disturbance; however, the absolute abundances of ARGs (105 to 108 copies g−1) in these estuarine sediments were lower than those in this study (Zhu et al. 2017). Interestingly, the river sediments impacted by an urban wastewater network in Spain (Quintela-Baluja et al. 2019) showed a similar absolute abundance (5.40 × 1010 copies g−1) but a much higher relative abundance (1.4 copies cell−1) of ARGs, as compared to our study (108 to 1010 copies g−1 and 0.04–0.41 copies cell−1). These indicate that, compared to the relative abundances of ARGs, the absolute abundances of ARGs in sediments may be less influenced by anthropogenic activities (Chen et al. 2019a; Yang et al. 2019).
It is noteworthy that the samples strongly affected by waste discharges from animal feedlots and residential areas generally showed higher levels of ARGs (Fig. S4). The highest absolute abundances of ARGs in the surface waters, soils, and sediments were observed in the ditch near the pig feedlot, the domestic sewage irrigated vegetable field, and the ditch near the bovine feedlot, respectively. The ubiquitous presence and high relative abundance of the clinical class 1 integron-integrase gene, intI-1(clinic), in the samples with high abundances of total ARGs reflect the strong impact of human activities on ARG distribution and these genes have the potential to be used as generic markers for ARG pollution (Figs. 3 and S5). Similarly, it was proposed in a recent review that the integrase intI-1(clinic) can serve as a universal proxy for anthropogenic pollution (Gillings et al. 2015). It can be inferred that the prevalence and dissemination of ARGs at these potential hotspots in the hilly rural area resulted not only from the direct input of ARGs from the nearby sources but also from the in situ ARG propagation in environmental media under the selective pressure of antibiotic loadings, via wastewater discharges or irrigation. The observed highest relative abundances of total ARGs in the sediment near the inlet of the ecological ditch receiving domestic sewage from the town and the soil of vegetable field irrigated with domestic sewage (Fig. S4f) agrees with the finding of a previous investigation in the same area that domestic sewage discharge from residential areas contributed most of the fecal bacteria (i.e., total coliform and Escherichia coli) load to the Linshan River system (Xue et al. 2018), as it was very likely that these human fecal bacteria were the major transporters of ARGs in the river network (Sidrach-Cardona et al. 2014; Karkman et al. 2019).
A 300-m long vegetated ecological ditch, which was constructed originally for the purpose of removing nutrients in domestic sewage (Kumwimba et al. 2016), also showed great removal efficiencies of fecal bacteria (Xue et al. 2018). Previous studies have shown that sorption and deposition onto surface soil or medium are the main mechanisms for ARG elimination in the constructed wetland, with ARG abundances in wetland substrates being elevated following its operation to remove ARGs from the wastewater (Chen et al. 2015; Huang et al. 2017). Therefore, the very high abundances of ARGs in the sediment sample collected near the ditch inlet, which was almost 1 to 2 orders of magnitude greater than those of the other sediment samples collected from downstream locations in the ditch, indicate that the ecological ditch has the potential to serve as an effective ARG trap. Future studies could devote more efforts to explore the dynamics of ARG removal from wastewaters of different chemical compositions by ecological ditches under varying hydro-climate conditions.
Distribution and co-occurrence patterns of ARGs
Despite the insignificant differences in the number and abundances of detected ARGs, the significant differences in ARG distribution pattern observed between the surface waters and sediments (Fig. 2b) illustrated that the pattern of ARG occurrence was strongly influenced by the source of samples, which is in agreement with previous studies (Li et al. 2015; Chen et al. 2019a). The distinctive separation phenomenon between different sample sources was most pronounced between the paired samples of surface water and sediment collected from the same site except the fishpond. The occurrence of similar ARG compositions of fishpond samples may be attributed to the full migration/exchange of ARGs and their host bacteria between the standing water and sediment given the almost quiescent conditions in the fishpond, as compared to the other sites that experienced more dynamic hydrological processes and resultant larger temporal variations in loadings and exports of various materials (e.g., water, soil/sediment, microbes, biogenic elements).
By utilizing conventional quantitative PCR methods, previous studies have claimed that the levels of ARGs in the sediments are notably higher than their paired surface waters in some aquatic environments (Xu et al. 2016b; Guan et al. 2018; Lu et al. 2019), which could be attributed to the higher population of bacteria in the sediments (Chen et al. 2013). The total concentration of sul genes (sul1 and sul2) was approximately 120 to 2000 times higher in the sediments than that in the water of the Haihe River (Luo et al. 2010). Similarly, the total absolute abundance of 21 ARGs in the sediments of the Ba River was 1 to 2 orders of magnitude higher than that of the water (Guan et al. 2018), whereas, in this study, the diversity and abundance of ARGs in the sediments were only slightly higher than those in the surface waters. This discrepancy between previously reported discoveries and our results could be attributed to two reasons. Methodology difference may be a major reason for this inconsistence. Only a very limited number of ARGs (5–21) were examined in the previous studies due to the low throughput of the common qPCR used, while a total of 283 primer sets targeting almost all major classes of ARGs were used in the present study. It was possible that some ARGs were present at lower abundances in the sediments compared to the surface waters (Lu et al. 2019). Therefore, a comprehensive assessment, instead of partial analysis for a limited number of well-studied genes, should be used to evaluate the distribution behaviors of ARGs between surface waters and sediments. ARGs were found to be more diverse and abundant in surface waters than those in sediments in recent studies by metagenomic analysis (Jiang et al. 2018) and HT-qPCR array (Chen et al. 2019a). On the other hand, the fate and migration of ARGs in surface water-sediment systems could be affected by environmental conditions, such as water depth, hydraulic retention time, and sample properties (Chen et al. 2019b; Lu et al. 2019). The observed decoupling relationship between ARG abundance of sediments and that of their corresponding overlying waters at most sampling sites of this study implies that the exchange and transport of ARGs between the two environmental media may be relatively weak. Similarly, obvious ARG separations between the surface water and sediment were reported in the Fengshuba Reservoir (a mainstream reservoir) of Guangdong Province, south China (Chen et al. 2019a).
The interactions among ARG subtypes were weak as indicated by the low clustering coefficient (0.390), which may be due to the strict robustness level (Spearman’s correlation coefficient > 0.80, P < 0.01) of network analysis used in this study. However, the modularity index of 0.524 implies that the constructed network had an obvious modular structure (Newman 2006), suggesting that the co-occurring ARGs in each module were potentially harbored in some specific bacterial species or MGEs (Li et al. 2015; Zheng et al. 2018). Indeed, the co-occurrence of transposon tnpA-07 and some ARGs targeting different antibiotic classes were observed, implying that these genes could be synchronously transferred by the transposase (Zhu et al. 2018), and thus likely rendered a complex proliferation of multiple ARGs when exposed to a single antibiotic class (Li et al. 2015; Chen et al. 2018; Qian et al. 2018). In addition, previous studies have suggested that the hub genes could be used as indicators to estimate the abundance of the co-occurring genes (Zheng et al. 2018; Chen et al. 2019a) or total ARGs (Qian et al. 2018). In this study, with the relatively small dataset of all 27 samples, four equations relating the relative abundance of total ARGs with that of the four identified indicator ARGs (i.e., blaXOA10-02, qacEΔ1-01, lnuB-02, and blaPSE) individually were obtained satisfactorily with the R2 value ranging from 0.701 to 0.952 (P < 0.001) (Fig. S6), despite that the ARG co-occurrence pattern may be different in different environmental habitats (Chen et al. 2019a). The accuracy and reliability of such predictive equations or models may be improved if a larger dataset for each environmental medium will become available for use in future.
Shared major ARGs among different environmental media
In this study, four ARGs, qacEΔ1-01, sul2, aadA-01, and cphA-01, were detected in all three environmental media. qacEΔ1-01 was the most abundant resistance gene (Fig. S5), which represents a mechanism for an efflux pump and is an ancestral part of clinical class 1 integrons (intI1) commonly found in pathogens or commensal bacteria of humans and domestic animals (Gillings 2014). qacEΔ1 has been reported to universally present and highly abundant in soils (Li et al. 2019), sediments (Zhu et al. 2017; Yang et al. 2019), wastewaters (An et al. 2018), and lakes/reservoirs (Liu et al. 2018). Kazama et al. (1998) observed qacEΔ1 genes in 41 of 63 clinical isolates of Pseudomonas aeruginosa, 3 of 5 clinical isolates of Vibrio parahaemolyticus, and 6 of 7 clinical isolates of Vibrio cholerae non-O1. sul2 genes were detected in 80.00%, 76.31%, and 50.03% of the sulphonamide-resistant Escherichia coli isolates from humans, pork, and pigs, respectively (Hammerum et al. 2006). A total of 38.57% of all 70 Escherichia coli isolates from bovine mastitis carried aadA resistance genes (Fazel et al. 2019). cphA genes were detected in 77 of 100 Aeromonas isolates obtained from clinical specimens (Sinclair et al. 2016). Apart from resistance genes, three MGEs including one integrase (intI-1LC) and two transposases (tnpA-04 and tnpA-05) were observed in all three environmental media with relative high abundances. intI-1 genes were present in about 46.08% of the isolates from the family Enterobacteriaceae (Goldstein et al. 2001). Carnelli et al. (2017) reported that 22 Aeromonas spp. and 14 fecal coliforms were tnpA hosts in polluted water. Recent studies using the HT-qPCR array detected a high concentration of intI-1 in inland waterbodies (Liu et al. 2018) and estuarine sediments (Zhu et al. 2017), while tnpA-04 and tnpA-05 were found to be the most enriched transposase genes in urban park soils with reclaimed water irrigation (Wang et al. 2014). The HGT mediated by MGEs is a crucial pathway for the exchange of genes among different microorganisms, even between pathogens and non-pathogens (Smillie et al. 2011; Forsberg et al. 2012). Numerous studies have shown that multiple ARG cassettes can be captured by class 1 integrons and transported by their transmissible carriers (e.g., plasmids and transposons) (Partridge et al. 2009; Heuer et al. 2011). Significant and positive correlations between the abundances of MGEs and total ARGs observed in this study (Table 1) were also reported by previous studies in various environments elsewhere (Zhu et al. 2013, 2017; Chen et al. 2019a). Taken together, these results suggest that the risk of horizontal transfer, dispersal of resistance genes, and transmission of ARGs to human pathogens in/among various environmental media at potential hotspots in hilly rural area affected by human activities should be evaluated in future.
Factors influencing the profile of ARGs in rural environmental media
Microorganisms can develop antibiotic resistance through horizontal gene transfer (HGT) and vertical gene transfer (VGT) in their evolutionary process (Dantas and Sommer 2014). Recently, relationships between the absolute abundance of ARGs and those of MGEs and 16S rRNA genes have been used to evaluate the role of HGT and VGT in the spread of ARGs, respectively (Guo et al. 2018b; Chen et al. 2019a). It can be inferred from the results of correlation analyses obtained in this study (Fig. 5) that the HGT and VGT together could facilitate the propagation and dissemination of ARGs in the sediments, while the HGT alone might be the dominant pathway for the dispersal of ARGs in the soils; neither of these two fundamental biological pathways played significant roles on ARG transfer in the surface waters. This medium specific phenomenon was also reported in a mainstream reservoir by a previous study (Chen et al. 2019a). The results of correlation analysis between the relative abundances of ARGs and chemical properties of environmental media obtained in this study (Table S5) suggested that biogenic elements (e.g., C, N, P) could be important factors regulating the occurrence and spread of ARGs in the surface waters, and similar findings were also reported in various surface water bodies elsewhere (Zheng et al. 2017; Zhou et al. 2017; Guo et al. 2018b).
It should be noted that the sampling campaign of this study was performed in a single month, and the obtained results may be applicable to the time-frame of sampling only. Growing evidence suggests that obvious seasonal variations of ARGs and MGEs could occur in various environmental locals, such as soils (Xiang et al. 2018), sediments (Luo et al. 2010; Su et al. 2014), rivers (Di Cesare et al. 2017; Zheng et al. 2018), and reservoir waters (Fang et al. 2019). Moreover, a number of previous studies have revealed that stormwater runoff induced by rainfall could deliver abundant ARGs and MGEs in discharges from residential and agricultural areas or in association with suspended soil/sediment particles (Di Cesare et al. 2017), and probably resulted in elevated levels of ARGs in the receiving rivers and reservoirs (Chen et al. 2019b; Fang et al. 2019). Therefore, given the high but unevenly distributed rainfall in this hilly study area, future studies could be dedicated to evaluate the temporal fluctuation and spatial redistribution of ARGs under varying hydro-climate conditions by conducting seasonal or rain event based sampling campaigns.
Conclusions
Occurrence and distribution of ARGs in various environmental media of a hilly rural area in the upper Yangtze River watershed were revealed using HT-qPCR array in the present study. Although the detected ARG levels were much lower than those previously reported at intensively impacted hotpots elsewhere, it was found that rural anthropogenic activities such as discharges of human and animal wastes may still promote the dissemination of ARGs in the receiving surface waters, soils, and sediments, as reflected by the universal presence and high abundance of intI-1(clinic) and qacEΔ1-01 genes in all three environmental media. Distinctive separation of ARG patterns between the paired samples of sediments and overlying surface waters could occur at sites subject to highly dynamic material loadings and exports via hydrological processes under varying hydro-climate conditions. Four resistance genes including blaOXA10-02, blaPSE, lnuB-02, and qacEΔ1-01 can be used to estimate the relative abundance of total ARGs in this study area. In order to support the development of effective measures to control the potential delivering of ARGs to surface waters via runoff in hilly rural areas, future investigations involving seasonal and rain event-based field sampling campaigns at multiple locations, are needed to reveal the temporal and spatial distributions of ARGs in various environmental media and identify the major influential factors.
References
An XL, Su JQ, Li B, Ouyang WY, Zhao Y, Chen QL, Cui L, Chen H, Gillings MR, Zhang T (2018) Tracking antibiotic resistome during wastewater treatment using high throughput quantitative PCR. Environ Int 117:146–153
Bastian M, Heymann S, Jacomy M (2009) Gephi: an open source software for exploring and manipulating networks. International AAAI Conference on Weblogs and Social Media
Bergeron S, Raj B, Nathaniel R, Corbin A, LaFleur G (2017) Presence of antibiotic resistance genes in raw source water of a drinking water treatment plant in a rural community of USA. Int Biodeterior Biodegradation 124:3–9
Braykov NP, Eisenberg JN, Grossman M, Zhang LX, Vasco K, Cevallos W, Muñoz D, Acevedo A, Moser KA, Marrs CF (2016) Antibiotic resistance in animal and environmental samples associated with small-scale poultry farming in northwestern Ecuador. mSphere 1:e00021–e00015
Carnelli A, Mauri F, Demarta A (2017) Characterization of genetic determinants involved in antibiotic resistance in Aeromonas spp. and fecal coliforms isolated from different aquatic environments. Res Microbiol 168:461–471
Chen H, Zhang MM (2013) Occurrence and removal of antibiotic resistance genes in municipal wastewater and rural domestic sewage treatment systems in eastern China. Environ Int 55:9–14
Chen BW, Liang XM, Huang XP, Zhang T, Li XD (2013) Differentiating anthropogenic impacts on ARGs in the Pearl River Estuary by using suitable gene indicators. Water Res 47:2811–2820
Chen J, Liu YS, Su HC, Ying GG, Liu F, Liu SS, He LY, Chen ZF, Yang YQ, Chen FR (2015) Removal of antibiotics and antibiotic resistance genes in rural wastewater by an integrated constructed wetland. Environ Sci Pollut Res 22:1794–1803
Chen BW, Lin L, Fang L, Yang Y, Chen E, Yuan K, Zou S, Wang X, Luan T (2018) Complex pollution of antibiotic resistance genes due to beta-lactam and aminoglycoside use in aquaculture farming. Water Res 134:200–208
Chen Y, Li P, Huang Y, Yu K, Chen H, Cui K, Huang Q, Zhang J, Yew-Hoong Gin K, He Y (2019a) Environmental media exert a bottleneck in driving the dynamics of antibiotic resistance genes in modern aquatic environment. Water Res 162:127–138
Chen Y, Su JQ, Zhang J, Li P, Chen H, Zhang B, Gin KY, He Y (2019b) High-throughput profiling of antibiotic resistance gene dynamic in a drinking water river-reservoir system. Water Res 149:179–189
Cheng JH, Tang XY, Cui JF (2019) Effect of long-term manure slurry application on the occurrence of antibiotic resistance genes in arable purple soil (entisol). Sci Total Environ 647:853–861
Dantas G, Sommer MOA (2014) How to fight back against antibiotic resistance. Am Sci 102:42–51
Dantas G, Sommer MOA, Oluwasegun RD, Church GM (2008) Bacteria subsisting on antibiotics. Science 320:100–103
Di Cesare A, Eckert EM, Rogora M, Corno G (2017) Rainfall increases the abundance of antibiotic resistance genes within a riverine microbial community. Environ Pollut 226:473–478
Fang PJ, Peng F, Gao XF, Xiao P, Yang J (2019) Decoupling the dynamics of bacterial taxonomy and antibiotic resistance function in a subtropical urban reservoir as revealed by high-frequency sampling. Front Microbiol 10:1448
Fazel F, Jamshidi A, Khoramian B (2019) Phenotypic and genotypic study on antimicrobial resistance patterns of E. coli isolates from bovine mastitis. Microb Pathog 132:355–361
Forsberg KJ, Reyes A, Wang B, Selleck EM, Sommer MO, Dantas G (2012) The shared antibiotic resistome of soil bacteria and human pathogens. Science 337:1107–1111
Gillings MR (2014) Integrons: past, present, and future. Microbiol Mol Biol Rev 78:257–277
Gillings MR, Gaze WH, Pruden A, Smalla K, Tiedje JM, Zhu YG (2015) Using the class 1 integron-integrase gene as a proxy for anthropogenic pollution. ISME J 9:1269–1279
Goldstein C, Lee MD, Sanchez S, Hudson C, Phillips B, Register B, Grady M, Liebert C, Summers AO, White DG (2001) Incidence of class 1 and 2 integrases in clinical and commensal bacteria from livestock, companion animals, and exotics. Antimicrob Agents Chemother 45:723–726
Gu ZG, Gu L, Eils R, Schlesner M, Brors B (2014) circlize implements and enhances circular visualization in R. Bioinformatics. 30:2811–2812
Guan Y, Jia J, Wu L, Xue X, Zhang G, Wang Z (2018) Analysis of bacterial community characteristics, abundance of antibiotics and antibiotic resistance genes along a pollution gradient of Ba River in Xi'an, China. Front Microbiol 9:3191
Guo XP, Stedtfeld RD, Hedman H, Eisenberg JNS, Truebac G, Yin DQ, Tiedje JM, Zhang LX (2018a) Antibiotic resistome associated with small-scale poultry production in rural Ecuador. Environ Sci Technol 52:8165–8172
Guo XP, Yang Y, Lu DP, Niu ZS, Feng JN, Chen YR, Tou FY, Garner E, Xu J, Liu M, Hochella MF Jr (2018b) Biofilms as a sink for antibiotic resistance genes (ARGs) in the Yangtze Estuary. Water Res 129:277–286
Hagedorn C, Robinson SL, Filtz JR, Grubbs SM, Angier TA, Reneau RB (1999) Determining sources of fecal pollution in a rural Virginia watershed with antibiotic resistance patterns in fecal streptococci. Appl Environ Microbiol 65:5522–5531
Hammerum AM, Sandvang D, Andersen SR, Seyfarth AM, Porsbo LJ, Frimodt-Møller N, Heuer OE (2006) Detection of sul1, sul2 and sul3 in sulphonamide resistant Escherichia coli isolates obtained from healthy humans, pork and pigs in Denmark. Int J Food Microbiol 106:235–237
He LY, Liu YS, Su HC, Zhao JL, Liu SS, Chen J, Liu WR, Ying GG (2014) Dissemination of antibiotic resistance genes in representative broiler feedlots environments: identification of indicator ARGs and correlations with environmental variables. Environ Sci Technol 48:13120–13129
Heuer H, Schmitt H, Smalla K (2011) Antibiotic resistance gene spread due to manure application on agricultural fields. Curr Opin Microbiol 14:236–243
Hu HW, Wang JT, Singh BK, Liu YR, Chen YL, Zhang YJ, He JZ (2018) Diversity of herbaceous plants and bacterial communities regulates soil resistome across forest biomes. Environ Microbiol 20:3186–3200
Huang X, Zheng JL, Liu CX, Liu L, Liu Y, Fan H (2017) Removal of antibiotics and resistance genes from swine wastewater using vertical flow constructed wetlands: effect of hydraulic flow direction and substrate type. Chem Eng J 308:692–699
Jiang L, Hu X, Yin D, Zhang H, Yu Z (2011) Occurrence, distribution and seasonal variation of antibiotics in the Huangpu River, Shanghai, China. Chemosphere 82:822–828
Jiang L, Hu X, Xu T, Zhang H, Sheng D, Yin D (2013) Prevalence of antibiotic resistance genes and their relationship with antibiotics in the Huangpu River and the drinking water sources, Shanghai, China. Sci Total Environ 458-460:267–272
Jiang H, Zhou R, Zhang M, Cheng Z, Li J, Zhang G, Chen B, Zou S, Yang Y (2018) Exploring the differences of antibiotic resistance genes profiles between river surface water and sediments using metagenomic approach. Ecotoxicol Environ Saf 161:64–69
Karkman A, Pärnänen K, Larsson DGJ (2019) Fecal pollution can explain antibiotic resistance gene abundances in anthropogenically impacted environments. Nat Commun 10:1–8
Kassambara A (2018) ggpubr: ‘ggplot2’ based publication ready plots. R Package Version 0.1.9.https://cran.r-project.org/src/contrib/Archive/ggpubr/ggpubr_0.1.9.tar.gz. Accessed 10 Apr 2020
Kazama H, Hamashima H, Sasatsu M, Arai T (1998) Distribution of the antiseptic-resistance genes qacE and qacEΔ1 in Gram-negative bacteria. FEMS Microbiol Lett 159:173–178
Kolde R (2018) Pheatmap: pretty Heatmaps. R Package Version 1.0.10. https://cran.r-project.org/src/contrib/Archive/pheatmap/pheatmap_1.0.10.tar.gz. Accessed 10 Apr 2020
Kumwimba MN, Dzakpasu M, Zhu B, Wang T, Muyembe DK (2016) Nutrient removal in a trapezoidal vegetated drainage ditch used to treat primary domestic sewage in a small catchment of the upper Yangtze River. Water Environ J 31:72–79
Li B, Yang Y, Ma LP, Ju F, Guo F, Tiedje JM, Zhang T (2015) Metagenomic and network analysis reveal wide distribution and co-occurrence of environmental antibiotic resistance genes. ISME J 9:2490–2502
Li S, Yao Q, Liu JJ, Wei D, Zhou BK, Zhu P, Cui XA, Jin J, Liu XB, Wang GH (2019) Profiles of antibiotic resistome with animal manure application in black soils of Northeast China. J Hazard Mater 121216
Liu LM, Su JQ, Guo YY, Wilkinson DM, Liu ZW, Zhu YG, Yang J (2018) Large-scale biogeographical patterns of bacterial antibiotic resistome in the waterbodies of China. Environ Int 117:292–299
Lu J, Zhang Y, Wu J, Wang J, Zhang C, Lin Y (2019) Occurrence and spatial distribution of antibiotic resistance genes in the Bohai Sea and Yellow Sea areas, China. Environ Pollut 252:450–460
Luo Y, Mao DQ, Rysz M, Zhou QX, Zhang HJ, Xu L, Alvarez PJJ (2010) Trends in antibiotic resistance genes occurrence in the Haihe River, China. Environ Sci Technol 44:7220–7225
McConnell MM, Hansen LT, Neudorf KD, Hayward JL, Jamieson RC, Yost CK, Anthony T (2018) Sources of antibiotic resistance genes in a rural river system. J Environ Qual 47:997–1005
Newman ME (2006) Modularity and community structure in networks. Proc Natl Acad Sci U S A 103:8577–8582
Oksanen J, Blanchet FG, Friendly M, Kindt R, Legendre P, McGlinn D, Minchin RP, O'Hara BR, Simpson GL, Solymos P, Stevens HHM, Szoecs E, Wagner H (2018) Vegan: community ecology package. R package version 2.5–3. https://cran.r-project.org/src/contrib/Archive/vegan/vegan_2.5-3.tar.gz. Accessed 10 Apr 2020
Ou-yang WY, Huang FY, Zhao Y, Li H, Su JQ (2015) Increased levels of antibiotic resistance in urban stream of Jiulongjiang River, China. Appl Microbiol Biotechnol 99:5697–5707
Partridge SR, Tsafnat G, Coiera E, Iredell JR (2009) Gene cassettes and cassette arrays in mobile resistance integrons. FEMS Microbiol Rev 33:757–784
Pruden A, Pei R, Storteboom H, Carlson KH (2006) Antibiotic resistance genes as emerging contaminants: studies in northern Colorado†. Environ Sci Technol 40:7445–7450
Qian X, Gu J, Sun W, Wang XJ, Su JQ, Stedfeld R (2018) Diversity, abundance, and persistence of antibiotic resistance genes in various types of animal manure following industrial composting. J Hazard Mater 344:716–722
Qiao M, Ying GG, Singer AC, Zhu YG (2017) Review of antibiotic resistance in China and its environment. Environ Int 110:160–172
Quintela-Baluja M, Abouelnaga M, Romalde J, Su JQ, Yu Y, Gomez-Lopez M, Smets B, Zhu YG, Graham DW (2019) Spatial ecology of a wastewater network defines the antibiotic resistance genes in downstream receiving waters. Water Res 162:347–357
R Core Team (2019) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna. https://cran.r-project.org/src/base/R-3/R-3.5.3.tar.gz. Accessed 10 Apr 2020
Revelle M (2018) Psych: procedures for psychological, psychometric, and personality research. R Package Version 1.8.10. https://cran.r-project.org/src/contrib/Archive/psych/psych_1.8.10.tar.gz. Accessed 10 Apr 2020
Sidrach-Cardona R, Hijosa-Valsero M, Marti E, Balcázar JL, Becares E (2014) Prevalence of antibiotic-resistant fecal bacteria in a river impacted by both an antibiotic production plant and urban treated discharges. Sci Total Environ 488-489:220–227
Sinclair HA, Heney C, Sidjabat HE, George NM, Bergh H, Anuj SN, Nimmo GR, Paterson DL (2016) Genotypic and phenotypic identification of Aeromonas species and CphA-mediated carbapenem resistance in Queensland, Australia. Diagn Microbiol Infect Dis 85:98–101
Smillie CS, Smith MB, Friedman J, Cordero OX, David LA, Alm EJ (2011) Ecology drives a global network of gene exchange connecting the human microbiome. Nature 480:241–244
Su HC, Pan CG, Ying GG, Zhao JL, Zhou LJ, Liu YS, Tao R, Zhang RQ, He LY (2014) Contamination profiles of antibiotic resistance genes in the sediments at a catchment scale. Sci Total Environ 490:708–714
Van Boeckel TP, Brower C, Gilbert M, Grenfell BT, Levin SA, Robinson TP, Teillant A, Laxminarayan R (2015) Global trends in antimicrobial use in food animals. Proc Natl Acad Sci U S A 112:5649–5694
Wang FH, Qiao M, Su JQ, Chen Z, Zhou X, Zhu YG (2014) High throughput profiling of antibiotic resistance genes in urban park soils with reclaimed water irrigation. Environ Sci Technol 48:9079–9085
Wickham H (2016) ggplot2: elegant graphics for data analysis. Springer-Verlag, New York
World Health Organization (2014) Antimicrobial resistance: global report on surveillance. World Health Organization, Geneva
Xiang Q, Chen QL, Zhu D, An XL, Yang XR, Su JQ, Qiao M, Zhu YG (2018) Spatial and temporal distribution of antibiotic resistomes in a peri-urban area is associated significantly with anthropogenic activities ☆. Environ Pollut 235:525–533
Xu LK, Ou-yang WY, Qian YY, Su C, Su J, Chen H (2016a) High-throughput profiling of antibiotic resistance genes in drinking water treatment plants and distribution systems ☆. Environ Pollut 213:119–126
Xu Y, Guo C, Luo Y, Lv J, Zhang Y, Lin H, Wang L, Xu J (2016b) Occurrence and distribution of antibiotics, antibiotic resistance genes in the urban rivers in Beijing, China. Environ Pollut 213:833–840
Xue F, Tang J, Dong Z, Shen D, Liu H, Zhang X, Holden NM (2018) Tempo-spatial controls of total coliform and E. coli contamination in a subtropical hilly agricultural catchment. Agric Water Manag 200:10–18
Yan ZZ, Chen QL, Zhang YJ, He JZ, Hu HW (2019) Antibiotic resistance in urban green spaces mirrors the pattern of industrial distribution. Environ Int 132:105106
Yang Y, Liu G, Ye C, Liu W (2019) Bacterial community and climate change implication affected the diversity and abundance of antibiotic resistance genes in wetlands on the Qinghai-Tibetan Plateau. J Hazard Mater 361:283–293
Zhang QQ, Ying GG, Pan CG, Liu YS, Zhao JL (2015) Comprehensive evaluation of antibiotics emission and fate in the river basins of China: source analysis, multimedia modeling, and linkage to bacterial resistance. Environ Sci Technol 49:6772–6782
Zhao Y, Cocerva T, Cox S, Tardif S, Su JQ, Zhu YG, Brandt KK (2019) Evidence for co-selection of antibiotic resistance genes and mobile genetic elements in metal polluted urban soils. Sci Total Environ 656:512–520
Zheng J, Gao R, Wei Y, Chen T, Fan J, Zhou Z, Makimilua TB, Jiao Y, Chen H (2017) High-throughput profiling and analysis of antibiotic resistance genes in East Tiaoxi River, China. Environ Pollut 230:648–654
Zheng J, Zhou Z, Wei Y, Chen T, Feng W, Chen H (2018) High-throughput profiling of seasonal variations of antibiotic resistance gene transport in a peri-urban river. Environ Int 114:87–94
Zhou ZC, Zheng J, Wei YY, Chen T, Dahlgren RA, Shang X, Chen H (2017) Antibiotic resistance genes in an urban river as impacted by bacterial community and physicochemical parameters. Environ Sci Pollut Res 24:23753–23762
Zhu B, Wang T, Kuang FH, Luo ZX, Tang JL, Xu TP (2009) Measurements of nitrate leaching from a hillslope cropland in the Central Sichuan Basin, China. Soil Sci Soc Am J 73:1419–1426
Zhu YG, Johnson TA, Su JQ, Qiao M, Guo GX, Stedtfeld RD, Hashsham SA, Tiedje JM (2013) Diverse and abundant antibiotic resistance genes in Chinese swine farms. Proc Natl Acad Sci U S A 110:3435–3440
Zhu YG, Zhao Y, Li B, Huang CL, Zhang SY, Yu S, Chen YS, Zhang T, Gillings MR, Su JQ (2017) Continental-scale pollution of estuaries with antibiotic resistance genes. Nat Microbiol 2:16270
Zhu D, Chen QL, Li H, Yang XR, Christie P, Ke X, Zhu YG (2018) Land use influences antibiotic resistance in the microbiome of soil collembolans Orchesellides sinensis. Environ Sci Technol 52:14088–14098
Funding
This research was funded by the 135 Strategic Program of the Institute of Mountain Hazards and Environment, CAS (Grant No. SDS-135-1702) and the National Natural Science Foundation of China (Grant Nos. 41790431 and 41771521).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Responsible editor: Robert Duran
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(DOCX 2347 kb).
Rights and permissions
About this article
Cite this article
Cheng, J., Tang, X. & Liu, C. Occurrence and distribution of antibiotic resistance genes in various rural environmental media. Environ Sci Pollut Res 27, 29191–29203 (2020). https://doi.org/10.1007/s11356-020-09287-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11356-020-09287-x