Abstract
Wastewater contains subinhibitory concentrations of different micropollutants such as antibiotics that create selective pressure on bacteria. This phenomenon is also caused by insufficient wastewater treatment technology leading to the development and spread of antibiotic-resistant bacteria and resistance genes into the environment. Therefore, this work focused on monitoring of antibiotic-resistant coliform bacteria and enterococci in influent and effluent wastewaters taken from the second biggest wastewater treatment plant (Petržalka) in the capital of Slovakia during 1 year. Antibiotic-resistant strains were isolated, identified, and characterized in terms of susceptibility and biofilm production. All of 27 antibiotic-resistant isolates were identified mainly as Morganella morganii, Citrobacter spp., and E. coli. Multidrug-resistance was detected in 58% of isolated strains. All tested isolates could form biofilm; two strains were very strong producers, and 74% formed biofilm by strong intensity. The flow rate of the influent wastewater had a more significant impact on the number of studied bacteria than the temperature.

Graphical abstract
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Water treatment infrastructure in the twenty-first century is insufficient, because of the growing quantities of micropollutants. Wastewaters receive different types of contaminants including surfactants, endocrine disruptors, pharmaceuticals, or personal care products. In recent years, attention has been focused on pharmaceuticals especially antibiotics as important emerging contaminants that are of concern due to their increasing worldwide use (Sharma et al. 2015).
Once antibiotics enter the ecosystems, they can affect the evolution of the microbial community structure, which accordingly influences the ecological function of water ecosystem (Bouki et al. 2013). In general, 30–90% of administered doses of water-soluble antibiotic substances are excreted in human urine and stool. On the other hand, large amounts of antibiotics get into wastewater due to disposal of unused antibiotics (Birošova et al. 2014; Bouki et al. 2013). Active molecules of such antibiotic residues and other pharmaceuticals with potential selective pressure are heavily discharged into the municipal sewage system and then flow to the wastewater treatment plant (WWTP) (Birošova et al. 2014; Novo et al. 2013). The main concern for the release of antibiotics into the environment is the development and spread of antibiotic resistance genes (ARG) and antibiotic-resistant bacteria (ARB), which potentially reduce the therapeutic potential against human and animal pathogens (Rizzo et al. 2013). One of the most common mechanisms of resistance acquisition is the formation of spontaneous mutations in the bacterial DNA with a significantly higher frequency compared with lethal concentrations of antibiotics (Aydin et al. 2015). In addition, many pathogenic bacteria released with wastewater harbor ARG that are eventually inserted into genetic mobile platforms that could be transferred into indigenous environmental bacteria via a horizontal gene transfer (Baquero et al. 2008; Cheng et al. 2016). Many ARB as well as ARG gets into wastewater with human stool. Feuerpfeil and Sterzel (1992) found that 80.5% of stool samples derived from healthy probands contained ARB, and after isolation, 98% of them were identified as E. coli. These bacteria can spread into the environment together with treated wastewater.
The World Health Organization (WHO) has classified the emergence and spread of antibiotic resistance and ARB as one of the three biggest threats to public health where increased antibiotic consumption is driving resistance (WHO 2016). Recent studies focus on pathogenic bacteria as well as fecal indicator microorganisms including representative strains of the family Enterobacteriaceae, Acinetobacter spp., Aeromonas spp., and E. coli, which are present in wastewater and have a tendency to accept genes encoding resistance to antibiotics (Bouki et al. 2013; Zhang et al. 2015). In aquatic environments, among the most common genes encoding resistance to different antibiotics belong 38 genes encoding resistance to tetracyclines (tet), 3 genes encoding resistance to oxytetracyclines (otr), sul1, sul2, sul3 genes encoding resistance to sulphonamides, erm, mef genes encoding resistance to erythromycin, mecA gene mediating resistance to β-lactams in case of methicillin-resistant Staphylococcus aureus (MRSA), blaTEM encoding resistance to extended-spectrum cephalosporins, and qnrS-related to fluoroquinolone resistance (Aydin et al. 2015; Rodriguez-Mozaz et al. 2015; Ou et al. 2015; Wan and Chou 2014). In addition, hospitals and health care facilities have a major role in resistance development because they harbor antibiotics, multidrug-resistant (MDR) bacteria, and resistance genes (Rodriguez-Mozaz et al. 2015).
Petržalka is the largest suburb of Bratislava with a population of 115,000 located on the right bank of the Danube River. Because Danube River is also the recipient of treated effluent wastewater released from WWTP Petržalka and can to contribute to the spread of antibiotic resistance phenomenon in the environment, the objective of our study was to understand the variations of antibiotic-resistant coliform bacteria and enterococci that occur in WWTP Petržalka along the year. The determination of the efficiency of the treatment in the removal of these groups of bacteria was also investigated. To contribute to the knowledge about the spread of ARB via wastewaters, resistant strains of coliform bacteria were isolated and identified. We also performed antibiotic susceptibility testing and biofilm formation assays. The characterization of antibiotic-resistant coliform isolates able to cause infections with increased mortality and morbidity may be a key step in the successful diagnosis.
Material and methods
Characterization of wastewater treatment plant and sampling
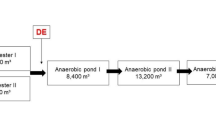
WWTP in Petržalka is dimensioned for 490,000 population equivalent (p.e.), but currently serves for 125,000 p.e. WWTP receives wastewater from the largest housing estate in Bratislava including one of the biggest hospitals as well as two small private hospitals (Bratislava Water Company 2013; Mackuľak et al. 2014). The mechanical step of WWTP consists of coarse and fine rakes, grit chamber and grease catcher. In the biological treatment, nitrification takes place, while sludge is anaerobically stabilized, and the generated biogas is energetically recovered. The water flow during the analyzed period ranged from 18,801 to 66,321 m3/day. The more detailed characterization of WWTP in Petržalka as well as sampling method has been published previously in Mackuľak et al. (2015). Sampling points were situated at the influent and effluent stream, while water samples were collected via an automatic sampler device every 15 min for 24 h (from 7:00 AM to 7:00 AM the next day). The samples of influent and effluent wastewater for microbiological analysis were immediately transferred into the laboratory in sterile falcon tubes. The wastewater samples were collected from January to December 2015 in 2-month intervals.
Detection of antibiotic-resistant coliform bacteria, E. coli, and enterococci
The detection and cultivation process of coliform bacteria and enterococci resistant to antibiotics was previously described in Mackuľak et al. (2015). Briefly, influent wastewater samples were diluted in physiological saline, and 0.1 mL of appropriate dilutions was spread on selective diagnostic agar plates. To determine the supposed low number of studied bacteria in effluent water, 10-mL aliquots of each sample were filtered through GH Polypro membrane (0.2 μm, Pall Corporation, USA) which was placed on antibiotic and antibiotic free agar plates. Resistance of coliform bacteria was tested on Chromocult Coliform agar (Merck, Germany) supplemented with ciprofloxacin, gentamicin, tetracycline, ampicillin, and chloramphenicol. Prevalence of ampicillin-, ciprofloxacin-, gentamicin- , and vancomycin-resistant Enterococcus spp. was detected on Slanetz-Bartley agar (Himedia, India). Concentrations of antibiotics were selected according to the resistance breakpoints established by European Committee on Antimicrobial Susceptibility Testing (EUCAST) for the European Union (EU) (EUCAST 2015). Because EUCAST did not establish resistance breakpoints for tetracycline, it was applied into diagnostic media for resistance testing in concentration established by Clinical Laboratory Standards Institute (CLSI) for United States (US) (CLSI 2014). Each experiment was run in triplicate. After cultivation, the colony forming units (CFU) of the observed bacteria were calculated, and the results were expressed as CFU per milliliter (CFU/mL) of wastewater sample. Data were statistically evaluated by ANOVA as well as Student’s t test. While colonies of E. coli on Chromocult Coliform agar (Merck, Germany) had blue or blue-violet color, other coliforms grew in form of pink colonies.
Isolation and identification of antibiotic-resistant coliform bacteria
Twenty-seven colonies of coliform bacteria were isolated from Chromocult Coliform agar (Merck, Germany) supplemented with antibiotics and transferred on Mueller–Hinton (Biolife, Italy) agar plates. After incubation at 37 °C for 24 h, the separated colonies of isolates were identified via a matrix-assisted laser desorption/ionization–time of flight (MALDI–TOF) mass spectrometer. One colony of each bacterium was spotted onto a steel target plate (Bruker Daltonics Inc., Billerica, MA) and covered with 1 μL of MALDI-TOF α-cyano-4-hydroxycinnamic acid matrix prepared as a saturated solution in 50% acetonitrile and 2.5% trifluoroacetic acid. A steel target plate with a solution matrix was dried at room temperature and analyzed via an AutoFlex I TOF-TOF apparatus (Bruker Daltonics Inc., Billerica, MA) in linear positive-ion mode across the m/z range of 2000 to 20,000 with gating of ions below m/z 400 and a delayed extraction time of 450 ns. Each spot was measured with 2000 laser shots at 25 Hz in groups of 50 shots per sampling area of the spot. A Bacterial Test Standard (Bruker Daltonics) was used for plate calibration. The spectra were analyzed using MALDI BioTyper software (v 2.0) (BioTyper Library v 3.0; Bruker Daltonics)—a proprietary algorithm for spectral pattern matching resulting in a logarithmic score from 0 to 3. Scores above 1.9 helped identify species, and scores ranging from 1.7 to 1.9 indicated genus identification; a score of < 1.7 indicated no identification.
Antibiotic susceptibility testing
Resistance of coliform bacteria isolates originating from Chromocult Coliform agar (Merck, Germany) plates to other antibiotic classes (β-lactams, fluoroquinolones, aminoglycosides, amphenicols, tetracyclines) was also observed. Mueller-Hinton agar (Biolife, Italy) plates containing ampicillin, gentamicin, ciprofloxacin, chloramphenicol, and tetracycline in concentrations defining resistance breakpoints given by EUCAST and CLSI were prepared for a plate dilution drop method (104 CFU)/drop) using the agar dilution method according to Olejníková et al. (2012) with slight modifications. The third concentration of antibiotics exceeded US standards and helped determine highly resistant coliform bacteria (Table 1). The visual evaluation of bacterial growth was performed after incubation at 37 °C for 24 h. Each experiment was performed in triplicate.
Detection of biofilm production by resistant isolates
The ability of resistant coliform bacteria isolates to form a biofilm was examined with a microtiter plate assay according to Beenken et al. (2003). Overnight cultures of isolates were diluted 1:200 in tryptic soy broth and transferred to the wells of a microtiter plate. After static incubation for 24 h and removing the overnight cultures, the wells were washed twice with phosphate buffered saline (PBS). To fix the adherent biofilm, 96% ethanol was used. These were subsequently stained with 0.41% crystal violet in 12% ethanol. Wells with crystal violet were rinsed twice with PBS and 96% ethanol at room temperature. Absorbance was measured at 570 nm using a plate reader device (BioTek, US). Experiments ran in six parallels and were repeated three times. The intensity of biofilm formation by coliform bacteria isolates was compared with the intensity of production by a positive control strain P. aeruginosa (CCM 3955) originating from the Check Collection of Bacterial Strains in Brno. Based on the absorbance values measured on spectrophotometer, a 27 isolates of coliform bacteria were ranked among weak, moderate, strong, and very strong biofilm producers according to Taniguchi et al. (2009).
Results and discussion
Total and antibiotic-resistant coliform bacteria prevalence in wastewaters during 1 year
Many studies document the occurrence of antibiotics along with bacteria resistant to them in all types of waters including mineral or drinking water, rivers, lakes, groundwater, and wastewater (Bouki et al. 2013; Azzam et al. 2017; Kumar and Pal 2018; Qiao et al. 2018). We monitored fecal indicator microorganism occurrence in wastewaters because many of these representative strains threaten public health. Furthermore, E. coli and other coliform bacteria receive resistance genes located on the plasmids relatively easily. This results in the spread of resistance to susceptible strains in WWTPs (Berrazeg et al. 2014).
Enterobacteriaceae are one of the most frequent producers of nosocomial infections in health care facilities as well as community infections (Khan et al. 2015). The emergence of antimicrobial resistance among members of this family leads to limited treatment options in cases of infections and has been recognized worldwide over the last decade (Badura et al. 2014). Coliform bacteria are a subgroup of the Enterobacteriaceae family that is characterized as indicator microorganisms for fecal contamination of drinking water and WWTP effluents (Naidoo and Olaniran 2014). The most significant representative strain of this group is E. coli, which is primarily located in the gastrointestinal tract of people including the healthy population. However, under certain conditions, it can cause disease due to virulence factors (Tenaillon et al. 2010).
Table 2 shows the total counts of bacteria studied at the inflow and outflow stream of WWTP Petržalka. The number of coliform bacteria in influent wastewater ranged from 4.6 to 5.9 log orders of CFU/mL. In the case of effluents, their prevalence was significantly lower (1.3 to 4.5 log CFU/mL). The highest value of coliform bacteria was determined in December. This can be caused by a massive discharge of sludge water at the time of sample collection. Sludge water is generated during the stabilization process of sewage sludge, and it contains many resistant coliforms. According to Mackuľak et al. (2015), part of these bacteria can be captured into sludge water, and this could be the reason for the increased bacterial number. In influent wastewater, the number of E. coli was 3.4 to 4.5 log CFU/mL, but only 0.9 to 1.8 log CFU/mL in effluent water.
Different classes of antibiotics were chosen for resistant coliform bacteria and E. coli detection. Resistance to ampicillin (ß-lactam) was tested, because of its frequent prescription rate in Slovakia (ECDC 2011). Although, a number of strains belonging to the group of coliform bacteria show intrinsic resistance to this antibiotic, in the case of E. coli the mechanism of ampicillin-resistance is acquired. Gentamicin (aminoglycoside) and ciprofloxacin (fluoroquinolone) can be found in wastewaters also in high concentrations as a consequence of their frequent use in both community and hospital sector. Even though the consumption of tetracycline and chloramphenicol is much lower than the consumption of abovementioned antibiotics, bacterial strains showing resistance against them are constantly emerging (Lépesová et al. 2018). These antibiotics except tetracycline were added to diagnostic agar plates (Chromocult Coliform agar) in concentrations established by EUCAST. Because resistance breakpoints for tetracycline are not defined by EUCAST, resistance of studied bacteria to this antibiotic was tested according to CLSI. According to EUCAST, MRSA, resistant Pseudomonas aeruginosa, Enterobacteriaceae producing extended-spectrum β-lactamases (ESBL), vancomycin-resistant enterococci, and Enterobacteriaceae resistant to carbapenems are the most important MDR bacterial strains (Garrett 2015; Picot-Guéraud et al. 2015). Antibiotic resistance of E. coli as well as horizontal gene transfer has been widely studied (Badura et al. 2014; Maal-Bared et al. 2013). Different types of ARGs are carried to E. coli mainly by plasmids such as the genes responsible for the production of TEM β-lactamases. Consequently, the TEM β-lactamases modify ESBL enzymes, and they developed extended resistance to cephalosporin antibiotics as well (Berrazeg et al. 2014; EUCAST 2013). According to Thenmozhi et al. (2014) up to 90% of ampicillin-resistance cases are caused by the production of TEM-1 enzyme.
Figure 1 plots the resistant coliform bacteria occurrence in influent wastewater samples during 1 year. Significant number of these bacteria was observed in all samples taken in different months. This is not surprising according to the composition and the diverse nature of wastewaters inflowing to the WWTP. In addition to the broad range of antibiotics in residual concentrations wastewaters also contain a significant proportion of bacteria showing resistance to antibiotics arriving from the human gastrointestinal tract, which when transported along the public sewer transmit their resistance determinants to susceptible species (Auguet et al. 2017). In general, the lowest number of resistant strains was registered in case of tetracycline and chloramphenicol. On the contrary, the highest number of coliforms was resistant to ampicillin, which is in accordance with its frequent use in Slovakia (ECDC 2017). Additionally, some coliform bacteria can be characterized by the intrinsic resistance to β-lactam antibiotics due to the presence of chromosomal ampC gene (Magiorakos et al. 2012). The lowest number of resistant coliforms in influent water was detected in July; however, these show resistance to all tested antibiotics according to EUCAST resistance breakpoints (Fig. 1). In addition, the percentage of resistant strains was over 50% suggesting the presence of MDR coliform bacteria (bacteria resistant to three or more antibiotic classes). Although the total number of coliform bacteria was the highest in influent wastewater in December (5.9 log CFU/mL), the nominal proportion of resistant strains was the lowest. On the other hand, most coliform bacteria in this sample were resistant to all antibiotics. Lamba and Ahammad (2017) analyzed the abundance of coliform bacteria as well as ESBL- and carbapenem-resistant bacteria in samples collected from various unit operations (raw influent wastewater, effluent from primary settling tank, effluent from aeration basin, effluent from final settling tank, and effluent from tertiary treatment) of WWTP in New Delhi during summer and winter season. They also found that in all of the collected samples the number of studied bacteria was higher in winter. It can be caused by a higher antibiotic usage in this season compared with summer which leads to increased amounts of antibiotic residues and selective pressure to present bacterial population. According to Ahammad et al. (2014) high amounts of ARB can be also resulted from lower flow rates and lower dilution of sewage in winter.
The prevalence of E. coli in influent wastewater samples was the highest in October (Fig. 2), while the percentage of these strains with resistance to all tested antibiotics was significantly higher compared with other samples. Wastewater samples taken in March, May, and December also had considerable presence of MDR E. coli except for the July sample where no tetracycline-resistant E. coli was detected. Although influent wastewater collected in January contained E. coli resistant to all applied antibiotics, their level was the lowest across all tested periods. The main reason for this phenomenon is likely the unfavorable climatic conditions for E. coli survival. Azuma et al. (2019) also studied the occurrence of ARB including E. coli (ESBL and carbapenem-resistant) among others in influent wastewater from WWTP in Japan. They registered the occurrence of ESBL- and carbapenem-resistant E. coli in influent wastewater sample; however, their concentrations were lower as in case of other studies (Hrenovic et al. 2017) indicating that the current rates of ARB spread varies among countries.
WWTPs should remove contaminants and eliminate microbial population (Di Fraia et al. 2018). Nevertheless, effluents from urban WWTPs are among the main anthropogenic sources for antibiotics, ARGs, and ARB (Rizzo et al. 2013). The Bratislava Water Company declares the efficiency of wastewater treatment in WWTP Petržalka to be 98% for monitored indicators (biochemical oxygen demand, chemical oxygen demand, insoluble substances), while bacteria are not monitored. The purified water is subsequently discharged into the Danube River at 0.5 m3/s that there is potential for contamination and spread of bacterial resistance to the environment (Bratislava Water Company 2013). In many studies was noted that effluents leaving the WWTP still contain a high amount of ARB (Huang et al. 2016; Neudorf et al. 2017; Qiao et al. 2018; Turolla et al. 2018; Wang et al. 2018). Thus, we monitored the prevalence of resistant coliform bacteria and E. coli in effluent wastewater. Coliform bacteria resistant to all tested antibiotics determined by EUCAST were observed mainly in effluents from March, May, October, and December (Fig. 3). Sample of effluent wastewater taken in January did not contain tetracycline-resistant coliforms, but in July only ampicillin-resistant coliform bacteria were detected. However, there is a strong likelihood that the majority of resistant strains presented in influent water were caught in sewage sludge. This phenomenon has been observed also by Lamba and Ahammad (2017) when coliforms and ESBL-producing bacteria showed a higher preference to stay attached to the solid phase compared with liquid phase. Their greater abundance was confirmed in sludge obtained from primary as well as final settling tank. They also indicate that less water consumption in winter leads to the 1.5–3-fold higher presence of solid content in sewage compared to summer season. It should be also noted that the rate of raw influent pollution plays an important role in wastewater treatment. In fact, high pollutant loads or increased flow rates increase the energy demand for the biological phase of treatment and result in spread of ARB into effluents (Di Fraia et al. 2018).
The highest prevalence of antibiotic-resistant E. coli based on EUCAST criteria (Fig. 4) was recorded in effluents from March, May, and December. In January, wastewater treatment survived only E. coli resistant to ampicillin and ciprofloxacin, while October sample showed only tetracycline-resistant E. coli. High levels of E. coli resistant to tetracycline and ampicillin were detected in other studies, too (Maal-Bared et al. 2013). Additionally, Azuma et al. (2019) found also a low number of ESBL- and carbapenem-resistant E. coli in effluent obtained from WWTP in Japan. Coliform bacteria resistant to ampicillin detected in effluent wastewater from July were not represented by E. coli.
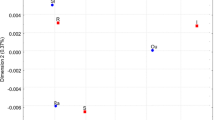
The concentration of pollutants in influents and effluents of WWTPs are routinely monitored in many countries (Fernández et al. 2014). However, seasonal variations in pharmaceuticals and resistant bacteria are rarely studied in wastewater. Only a small number of studies investigated the prevalence of pathogenic parasites and fecal indicator bacteria in WWTPs, their concentrations and removal rates during wastewater treatment, their seasonal variation and the impact of effluent discharges on the microbiological quality of drinking water reservoirs (Tolouei et al. 2019). The amount of coliform bacteria persisting in wastewater depends on different factors including the size, type, and capacity of treatment facilities (Tolouei et al. 2019). In this study, we also studied the effect of temperature of influent wastewater as well as the possibility of its dilution related to the wastewater flow. Figure 5 show that the temperature change in the wastewater did not influence the number of total coliform bacteria and E. coli. On the other hand, the number of total coliforms as well as E. coli decreased with decreasing flow of wastewater (Fig. 6). It is in accordance with the claim of Kay et al. (2008), by which secondary and tertiary treatments can be less effective for fecal indicator bacteria removal under high flow conditions. The flow rate of wastewaters inflowing to the WWTPs is affected mainly by weather conditions (rain, snow melting) (Tolouei et al. 2019). This implies that wastewater flow has a biggest impact on the prevalence of coliform bacteria than the temperature of the wastewater. This might be because of the rapid adaptability of coliform bacteria and their ability to survive in a wide range of temperatures. In addition, bacteria caught in the WWTP biofilms can be easily released into the effluent wastewater due to greater wastewater flow. Fernández et al. (2014) reported that the concentrations of pharmaceuticals in municipal wastewater and their treated effluents might vary along the year. The consumption of pharmaceuticals may be different in different seasons, and this has a massive influence on the prevalence of total and resistant bacteria in urban wastewater. During cold seasons (autumn-winter) the antibiotic content of wastewater is the highest. This may be a consequence of a higher incidence of flu as well as overuse of antibiotics during this period. On the other hand, the representation of other types of micropollutants in wastewaters varies seasonally as function of depression or allergy. Birošova et al. (2014) seasonally analyzed antibiotics of different classes as well as ARB in the influent and effluent water of two WWTPs in Bratislava. Their results also confirmed that seasonal changes in antibiotic consumption resulted in an increased number of coliforms and streptococci in sewage sludge.
Total and antibiotic resistant Enterococcus ssp. prevalence in wastewaters during 1 year
Even though Enterococcus spp. are commensal in the guts of animals including humans, they have virulence properties that could enable them to establish infections in certain categories of people whose immune systems have been compromised such as in the elderly, children, pregnant women, and long-term hospitalized patients (Iweriebor et al. 2015). Representatives of this genus are characterized by intrinsic resistance to penicillins, monobactams, and low levels of aminoglycosides, but tend to take resistance genes and build resistance to antimicrobial agents (Sukmawinata et al. 2018). We also measured total and resistant enterococci. Slanetz–Bartley media supplemented by ampicillin, gentamicin, ciprofloxacin, and vancomycin according to EUCAST resistance breakpoints was used as a selective diagnostic media for fecal streptococci. The use of vancomycin and teicoplanin for the control of enterococcal infections as well as MRSA was introduced into medical practice due to the increased resistance to aminoglycosides and β-lactams (Laishram et al. 2014; Ranotkar et al. 2014; Dubey and Padhy 2015). There is a currently rapid dissemination of vancomycin resistant enterococci worldwide—a global public health concern (Yang et al. 2015).
The mean value of the total number of Enterococcus ssp. in influent wastewaters was about 3.2 log CFU/mL except July when their presence was under the detection limit. Effluent sample collected in July did not contain fecal streptococci, but in January, March, May, October, and December their numbers ranged from 0.3 to 1.5 log CFU/mL (Table 2). Figure 7 shows the proportion of resistant enterococci in different months. The lowest number of resistant enterococci was recorded in January (as in case of coliform bacteria), but their prevalence in influent wastewaters taken in May, October, and December was comparable. The highest number of resistant enterococci was detected in March 2015. In all studied influent wastewater samples, we noted resistance to ampicillin, gentamicin, ciprofloxacin, and vancomycin, while in previous study only ampicillin- and ciprofloxacin-resistant enterococci in influent wastewater were detected (Lépesová et al. 2018). Figure 8 indicates that the growth of resistant enterococci was significantly reduced after wastewater treatment. No resistance was seen in the effluents from May and December, but those from January had ampicillin- and gentamicin-resistant strains. Taučer-Kapteijn et al. (2016) also found ampicillin-resistant enterococci in WWTP effluents. Enterococci resistant to all tested antibiotics including vancomycin in wastewater effluents taken in March and October were constantly present. Oravcová et al. (2017) has also managed to detect vancomycin-resistant enterococci in 32 from 37 collected effluent water samples in WWTP Brno. The most worrying feature of resistance to vancomycin are the high frequency of simultaneous resistance to several antibiotics as well as a putative risk of spreading resistance to more virulent bacteria, such as S. aureus including MRSA (Ramos et al. 2015; Hamiwe et al. 2019). However, vancomycin-resistant enterococci are most widespread in hospital environments; their dissemination takes place also outside of nosocomial environment (Braga et al. 2013).
The influence of temperature and flow rate of wastewater inflowing to the WWTP Petržalka on enterococci prevalence was also investigated. As in case of coliform bacteria, the number of enterococci depended not on the temperature but on the flow of wastewater (Figs. 5 and 6). The number of total enterococci showed increasing tendency with an increase in wastewater flow rate.
Isolation and identification of resistant coliform bacteria from wastewaters
Recently, proteomics based on the use of MALDI-TOF mass spectrometry has been shown to be a promising and reliable tool for identifying not only clinical (Clark et al. 2013; Croxatto et al. 2012) but also environmental isolates (Ruelle et al. 2004; Welker and Moore 2011; Santos et al. 2015). During annual monitoring of prevalence of coliform bacteria showing resistance to antibiotics in collected samples of influent wastewaters, 27 different bacterial strains were isolated and subsequently identified by MALDI–TOF (Table 3). The most frequently identified genus among coliform bacteria was Citrobacter spp., which was represented by strains Citrobacter freundii (11 strains), C. braakii (2 strains), and C. youngae (1 strain). These species belong to potential pathogens causing urinary tract infections in hospitals and are also widely distributed in water, soil, food, or the human gastrointestinal tract (Metri et al. 2013). Another frequent bacterial strain identified in influent wastewaters for a total of 12 isolates was Morganella morganii—an opportunistic pathogen widely represented in the environment. As Citrobacter spp., M. morganii also causes nosocomial, urinary tract, skin or soft tissue infections, meningitis, and bacteremia—often with fatal consequences (Tahrani et al. 2015). Only one strain among all isolates was identified as E. coli, which commonly causes bloodstream and urinary tract infections (ECDC 2017).
Antibiotic susceptibility determination of resistant isolates
All isolates are characterized as potential pathogenic microorganisms. These usually cause nosocomial infections in hospitals especially in immune compromised patients. In recent years, the medical community has witnessed an increase in infections caused by Gram-negative bacteria that are MDR to many types of antibiotics. According to European Centre for Disease Control MDR is defined as non-susceptibility to at least one agent in three or more antimicrobial categories (ECDC 2017). An increased incidence of these bacteria in health care facilities reduces the effectiveness of antibiotic therapy, which results in increased mortality and morbidity as well as high treatment costs (Bhatt et al. 2015). Magiorakos et al. (2012) created a list of specific microorganisms that are intrinsically resistant to certain antibiotic agents. According to their study, M. morganii is naturally resistant to penicillins (ampicillin), to combination of penicillins with β-lactamase inhibitors (amoxicillin–clavulanic acid or ampicillin–sulbactam), first and second generation of cephalosporins (cefazolin, cefuroxim), glycylcyclines (tigecycline), and polymyxins (colistin) as well as tetracyclines (tetracycline, doxycycline, minocycline). For the definition of “MDR bacteria”, only an acquired resistance was used. Therefore, the antibiotics mentioned above for M. morganii were not considered when assessing their MDR phenotype (Table 3). This means that 6 of 12 coliform isolates were declared as MDR. In case of Citrobacter ssp., the intrinsic resistance was determined to penicillin, to their combination with β-lactamase inhibitors, first and second generation of cephalosporins as well as cephamycins (cefoxitin, cefotetan) (Magiorakos et al. 2012). Thus, 12 isolates belonging to the genus of Citrobacter showed resistance to more than three antibiotic classes (Table 3). Unlike other coliform bacteria, E. coli do not possess intrinsic resistance to β-lactam character antibiotics (Magiorakos et al. 2012). Currently, the rate of resistance of E. coli is growing not only to penicillins and cephalosporins of first, second, third and fourth generation, but also to carbapenems. Such E. coli isolates can produce ESBL or carbapenemases. E. coli isolated on the influent stream in December was susceptible to ciprofloxacin and chloramphenicol, but it showed MDR character (Table 3). Maal-Bared et al. (2013) also found low levels of resistance to ciprofloxacin, which correspond to our results.
Biofilm detection assessment
In aquatic systems, the major fraction of bacterial communities are found in structurally organized surface associated aggregates called biofilms where bacterial cells are embedded in an extracellular matrix. Owing the proximity between bacteria, biofilms are ideal reaction chambers for horizontal gene transfer notably of ARGs. This results in accelerated evolution and spread of antibiotic resistance (Ory et al. 2016). Thus, changes in biofilm production by resistant isolates were detected. The absorbance values are directly proportional to the density of cells fixed in biofilm in case of both M. morganii and Citrobacter spp. According to Taniguchi et al. (2009), absorbance lower than 0.1 shows that bacterial strain is weak biofilm producer. If the absorbance value is 0.2–0.3, then the strain is moderate producer, and range 0.3–0.9 indicates a strong producer; if this value is over 1.0 then the bacteria is a very strong biofilm producer such as P. aeruginosa.
All isolated resistant coliforms could form biofilm. MDR strains of M. morganii and Citrobacter spp. were very strong producers of biofilm (Table 3). Eight more strains of M. morganii and 12 strains of Citrobacter spp. were characterized as strong biofilm producers resistant to all antibiotics. Only one isolate of Citrobacter ssp. was a moderate producer. Nevertheless, the single isolate of E. coli was susceptible to ciprofloxacin and chloramphenicol, and it also showed an ability to form biofilm with strong intensity. Smanthong et al. (2015) suggested that the ability of E. coli isolates to persist and grow as biofilm is an important factor involved in both the severity of urinary tract infections and antimicrobial resistance. A previous study on uropathogenic E. coli isolates showed a significant correlation between increased biofilm formation and resistance to multiple antimicrobial drugs such as ampicillin, amikacin, norfloxacin, and trimethoprim/sulfamethoxazole (Smanthong et al. 2015). Schwartz et al. (2003) investigated the presence of ARB in biofilms collected from municipal wastewater, hospital wastewater, and drinking as well as river water. They found that Enterobacteriaceae hydrolyzing β-lactams and vancomycin resistant enterococci were more frequent in wastewater biofilms than in river or drinking water. Maheshwari et al. (2016) also found that the frequency of blaCTX-M gene transfer was significantly higher in biofilms than in the planktonic mode of growth. The occurrence of ESBL-producing bacteria with biofilm-forming ability in hospital wastewater may have serious implications both on public health and infection control practices—the transmission of such strains to humans and animal populations can result in chronic and persistent infections.
Conclusions
This study focused on monitoring of prevalence of resistant coliform bacteria and enterococci in influent and effluent wastewaters from WWTP Petržalka during 2015. The number of antibiotic-resistant coliform bacteria and E. coli in wastewaters inflowing to the WWTP was high, and they were not completely eliminated after wastewater treatment. A high prevalence of MDR E. coli as well as coliform bacteria was detected in effluent samples taken mainly in March, May, and December. There was a massive prevalence of fecal streptococci resistant to ampicillin, gentamicin, ciprofloxacin, and vancomycin in all influent wastewaters samples. The most alarming observation is that effluents in March and October still contained epidemiologically dangerous vancomycin-resistant strains of Enterococcus spp.
The qualitative characterization of bacterial content of wastewaters inflowing to the WWTP was also performed. Isolates of resistant coliform bacteria were identified and belonged mainly to the Morganella and Citrobacter genera. The majority of isolated strains showed MDR character and could form biofilm with strong intensity.
However, results describing the quantity as well as the quality of ARB were observed in local municipal WWTP in Slovakia, more spatial design of study is needed, and further studies in municipal WWTPs of northern, western, and eastern Slovakia should be carried on.
References
Ahammad et al (2014) Increased waterborne blaNDM-1 resistance gene abundances associated with seasonal human pilgrimages to the upper ganges river. Environ Sci Technol 48:3014–3020
Auguet O, Pijuan M, Borrego CM, Rodriguez-Mozaz S, Triadó-Margarit X, Giustina SVD, Gutierrez O (2017) Sewers as potential reservoires of antibiotic resistance. Int Sci Total Environ 605-606:1047–1054
Aydin S, Ince B, Ince O (2015) Development of antibiotic resistance genes in microbial communities during long-term operation of anaerobic reactors in the treatment of pharmaceutical wastewater. Water Res 83:337–344
Azuma et al (2019) Environmental fate of pharmaceutical compounds and antimicrobial-resistant bacteria in hospital effluents, and contributions to pollutant loads in the surface waters in Japan. Sci Total Environ 657:476–484
Azzam MI, Ezzat SM, Othman BA, el-Dougdoug KA (2017) Antibiotics resistance phenomenon and virulence ability in bacteria from water environment. Water Sci 31:109–121
Badura A, Luxner J, Feierl G, Reinthaler FF, Zarfel G, Galler H, Pregartner G, Riedl R, Grisold AJ (2014) Prevalence, antibiotic resistance patterns and molecular characterization of Escherichia coli from Austrian sandpits. Environ Pollut 194:24–30
Baquero F, Martínez JL, Cantón R (2008) Antibiotics and antibiotic resistance in water environments. Curr Opin Biotechnol 19:260–265
Beenken KE, Blevins JS, Smeltzer MS (2003) Mutation in sarA in Staphylococcus aureus limits biofilm formation. Infect Immun 71(7):4206–4211
Berrazeg M et al (2014) New Delhi metallo-beta-lactamase around the world: an eReview using Google maps. Euro Surveill 19(20):1–14
Bhatt P, Tandel K, Shete V, Rathi KR (2015) Burden of extensively drug-resistant and pandrug-resistant Gram-negative bacteria at a tertiary-care centre. New Microbes New Infect 8:166–170
Birošova L et al (2014) Pilot study of seasonal occurrence and distribution of antibiotics and drug resistant bacteria in wastewater treatment plants in Slovakia. Sci Total Environ 490:440–444
Bouki CH, Venieri D, Diamadopoulos E (2013) Detection and fate of antibiotic resistant bacteria in wastewater treatment plants: a review. Ecotoxicol Environ Saf 91:1–9
Braga TM, Pomba C, Lopes MFS (2013) High-level vancomycin resistant Enterococcus faecium related to humans and pigs found in dust from pig breeding facilities. Vet Microbiol 161:344–349
Bratislava Water Company (2013) Press news. Modernization of the two largest wastewater treatment plants in Slovakia. (In Slovak) [http://www.bvsas.sk/sk/press/tlacove-spravy/modernizacia-dvoch-najvacsich-cistiarni-slovensku.html. Online 20.3.2017]
Cheng W, Li J, Wu Y, Xu L, Su C, Qian Y, Zhu YG, Chen H (2016) Behavior of antibiotics and antibiotic resistance genes in eco-agricultural systems: a case study. J Hazard Mater 304:18–25
Clark CG, Kruczkiewicz P, Guan C, McCorrister SJ, Chong P, Wylie J, van Caeseele P, Tabor HA, Snarr P, Gilmour MW, Taboada EN, Westmacott GR (2013) Evaluation of MALDI-TOF mass spectroscopy methods for determination of Escherichia coli pathotypes. J Microbiol Methods 94:180–191
Clinical Laboratory Standards Institute, CLSI (2014) M 100 – S 24 performance standards for antimicrobial susceptibility testing; twenty-fourth informational supplement
Croxatto A, Prod'hom G, Greub G (2012) Applications of MALDI-TOF mass spectrometry in clinical diagnostic microbiology. FEMS Microbiol Rev 36:380–407
Dubey D, Padhy RN (2015) Infection dynamics of vancomycin and inducible clindamycin resistant Enterococcus faecalis in an Indian teaching hospital. Asian Pac J Trop Dis 5:127–132. https://doi.org/10.1016/S2222-1808(15)60873-8
European Centre for Disease Prevention and Control, ECDC (2011) Surveillance of antimicrobial consumption in Europe 2011; Stockholm: 2014; 1–82
European Centre for Disease Prevention and Control, ECDC (2017) Antimicrobial resistance surveillance in Europe 2016. Annual Report of the European Antimicrobial Resistance Surveillance Network (EARS-Net). Stockholm, Sweden: ECDC; 2017, ISBN 978–92–9498-099-1, 1–11
European Committee on Antimicrobial Susceptibility Testing, EUCAST (2013) EUCAST guidelines for detection of resistance mechanisms and specific resistances of clinical and/or epidemiological importance
European Committee on Antimicrobial Susceptibility Testing, EUCAST (2015) Breakpoint tables for interpretation of MICs and zone diamteres; Version 5.0 valid from 2015
Fernández M, Fernández M, Laca A, Laca A, Díaz M (2014) Seasonal occurrence of removal of pharmaceutical products in municipal wastewaters. J Environ Chem Eng 2:495–502
Feuerpfeil I, Sterzel W (1992) Presence of antibiotic-resistant coliform bacteria in the human intestinal flora. Bundesgesudheitsblatt 35:61–65
Fraia D et al (2018) A novel energy assessment of urban wastewater treatment plants. Energ Convers Manage 163:304–313
Garrett JH (2015) An era of antimicrobial resistance: imminent threats of superbugs to vascular access clinicians and patients. JAVA 20(3):133–136. https://doi.org/10.1016/j.java.2015.06.002
Hamiwe T, Kock MM, Magwira CA, Antiabong JF, Ehlers MM (2019) Occurrence of enterococci harbouring clinically important antibiotic resistance genes in the aquatic environment in Gauteng, South Africa. Environ Pollut 245:1041–1049
Hrenovic et al (2017) The fate of carbapenem-resistant bacteria in wastewater treatment plant. Water Res 126:232–239
Huang JJ, Xi J, Hu HY, Li Y, Lu SQ, Tang F, Pang YC (2016) UV light tolerance and reactivation potential of tetracycline-resistant bacteria from secondary effluents of a wastewater treatment plant. J Environ Sci 41:146–153
Iweriebor BCH, Gaqavu S, Obi L, Nwodo U, Okoh A (2015) Antibiotic susceptibilities of Enterococcus species isolated from hospital and domestic wastewater effluents in Alice, eastern Cape Province of South Africa. Int J Environ Res Public Health 12:4231–4246. https://doi.org/10.3390/ijerph120404231
Kay D, Crowther J, Stapleton CM, Wyer MD, Fewtrell L, Edwards A, Francis CA, McDonald AT, Watkins J, Wilkinson J (2008) Faecal indicator organism concentrations in sewage and treated effluents. Water Res 42(1):442–454
Khan HA, Ahmad A, Mehboob R (2015) Nosocomial infections and their control strategies. Asian Pac J Trop Biomed 5(7):509–514
Kumar A, Pal D (2018) Antibiotic resistance and wastewater: correlation, impact and critical human health challenges. JECE 6:52–58
Laishram S, Sahni RD, Anandan S, Balaji V (2014) Evaluation of the antimicrobial activity of daptomycin and linezolid against vancomycin-resistant Enterococcus spp. isolates in south India. J Glob Antimicrob Re 2:194–197
Lamba M, Ahammad SZ (2017) Sewage treatment effluent in Delhi: a key contributor of β-lactam resistant bacteria and genes to the environment. Chemosphere 188:249–256
Lépesová K, Kraková L, Pangallo D, Medveďová A, Olejníková P, Mackuľak T, Tichý J, Grabic R, Birošová L (2018) Prevalence of antibiotic-resistant coliform bacteria, Enterococcus spp. and Staphylococcus spp. in wastewater sewerage biofilm. J Glob Antimicrob Res 14:145–151
Maal-Bared R, Bartlett KH, Bowie WR, Hall ER (2013) Phenotypic antibiotic resistance of Escherichia coli and E. coli O157 isolated from water, sediment and biofilms in an agricultural watershed in British Columbia. Sci Total Environ 443:315–323
Mackuľak T, Škubák J, Grabic R, Ryba J, Birošová L, Fedorova G, Špalková V, Bodík I (2014) National study of illicit drug use in Slovakia based on wastewater analysis. Sci Total Environ 494-495:158–165
Mackuľak T, Nagyová K, Faberová M, Grabic R, Koba O, Gál M, Birošová L (2015) Utilization of Fenton-like reaction for antibiotics and resistant bacteria elimination in different parts of WWTP. Environ Toxicol Pharmacol 40:492–497
Magiorakos AP, Srinivasan A, Carey RB, Carmeli Y, Falagas ME, Giske CG, Harbarth S, Hindler JF, Kahlmeter G, Olsson-Liljequist B, Paterson DL, Rice LB, Stelling J, Struelens MJ, Vatopoulos A, Weber JT, Monnet DL (2012) Multidrug-resistant, extensively drug-resistant and pandrug-resistant bacteria: an international expert proposal for interim standard definitions for acquired resistance. Clin Microbiol Infect 18:268–281
Maheshwari M, Ahmad I, Althubiani AS (2016) Multidrug resistance and transferability of bla CTX-M among extended-spectrum β-lactamase-producing enteric bacteria in biofilm. J Glob Antimicrob Res 6:142–149
Metri BC, Jyothi P, Peerapur BV (2013) Antibiotic resistance in Citrobacter spp. isolated from urinary tract infection. Urol Ann 5(4):312–313
Naidoo S, Olaniran AO (2014) Treated effluent wastewater as a source of microbial pollution of surface water resources. Int J Environ Res Public Health 11:249–270. https://doi.org/10.3390/ijerph110100249
Neudorf KD, Huang YN, Ragush CM, Yost CK, Jamieson RC, Truelstrup Hansen L (2017) Antibiotic resistance genes in municipal wastewater treatment systems and receiving waters in Arctic Canada. Sci Total Environ 598:1085–1094
Novo A, André S, Viana P, Nunes OC, Manaia CM (2013) Antibiotic resistance, antimicrobial residues and bacterial community composition in urban wastewater. Water Res 47:1875–1887
Olejníková P, Kurucová M, Švorc L, Marchalín Š (2012) Induction of resistance in Mycobacterium smegmatis. Can J Microbiol 59(2):126–129
Oravcová V et al (2017) Vancomycin-resistant enterococci with vanA gene in treated municipal wastewater and their association with human hospital strains. Sci Total Environ 609:633–643
Ory J, Bricheux G, Togola A, Bonnet JL, Donnadieu-Bernard F, Nakusi L, Forestier C, Traore O (2016) Ciprofloxacin residue and antibiotic-resistant biofilm bacteria in hospital effluent. Environ Pollut 214:635–645
Ou D, Chen B, Bai R, Song P, Lin H (2015) Contamination of sulfonamide antibiotics and sulfamethazine-resistant bacteria in the downstream and estuarine areas of Jiulong River in Southeast China. Environ Sci Pollut Res Int 22:12104–12113
Picot-Guéraud R, Batailler P, Caspar Y, Hennebique A, Mallaret MR (2015) Bacteremia caused by multidrug-resistant bacteria in a French university hospital center: 3 years of collection. Am J Infect Control 43:960–964
Qiao M, Ying GG, Singer AC, Zhu YG (2018) Review of antibiotic resistance in China and its environment. Environ Int 110:160–172
Ramos S, Chafsey I, Silva N, Hébraud M, Santos H, Capelo-Martinez JL, Poeta P, Igrejas G (2015) Effect of vancomycin on the proteome of the multiresistant Enterococcus faecium SU18 strain. J Proteome 113:378–387
Ranotkar S, Kumar P, Zutshi S, Prashanth KS, Bezbaruah B, Anand J, Lahkar M (2014) Vancomycin-resistant enterococci: troublemaker of the 21st century. J Glob Antimicrob Res 2:205–212
Rizzo L, Manaia C, Merlin C, Schwartz T, Dagot C, Ploy MC, Michael I, Fatta-Kassinos D (2013) Urban wastewater treatment plants as hotspots for antibiotic resistant bacteria and genes spread into the environment: a review. Sci Total Environ 447:345–360
Rodriguez-Mozaz S, Chamorro S, Marti E, Huerta B, Gros M, Sànchez-Melsió A, Borrego CM, Barceló D, Balcázar JL (2015) Occurrence of antibiotics and antibiotic resistance genes in hospital and urban wastewaters and their impact on the receiving river. Water Res 69:234–242
Ruelle V, Moualij BE, Zorzi W, Ledent P, Pauw ED (2004) Rapid identification of environmental bacterial strains by matrix-assisted laser desorption/ionization time-of-flight mass spectrometry. Rapid Commun Mass Spectrom 18:2013–2019
Santos T, Capelo JL, Santos HM, Oliveira I, Marinho C, Gonçalves A, Araújo JE, Poeta P, Igrejas G (2015) Use of MALDI-TOF mass spectrometry fingerprinting to characterize Enterococcus spp. and Escherichia coli isolates. J Proteome 127:321–331
Schwartz T, Kohnen W, Jansen B, Obst U (2003) Detection of antibiotic-resistant bacteria and their resistance genes in wastewater, surface water, and drinking water biofilms. FEMS Microbiol Ecol 43:325–335
Sharma VK et al (2015) A review of the influence of treatment strategies on antibiotic resistant bacteria and antibiotic resistance genes. Chemosphere 150:702–714
Smanthong N, Tavichakorntrakool R, Saisud P, Prasongwatana V, Sribenjalux P, Lulitanond A, Tunkamnerdthai O, Wongkham C, Boonsiri P (2015) Biofilm formation in trimethoprime/sulfamethoxazole-susceptible and trimethoprim/sulfamethoxazole-resistant uropathogenic Escherichia coli. Asian Pac J Trop Biomed 5(6):485–487
Sukmawinata E, Sato W, Uemura R, Sueyoshi M (2018) Antimicrobial resistant Enterococcus faecium, Enterococcus faecalis, and other Enterococcus species isolated from foal feces in Japan. J Equine Vet Sci 63:51–54
Tahrani L, Soufi L, Mehri I, Najjari A, Hassan A, van Loco J, Reyns T, Cherif A, Mansour HB (2015) Isolation and characterization of antibiotic-resistant bacteria from pharmaceutical industrial wastewaters. Microb Pathog 89:54–61
Taniguchi L, de Fátima Faria B, Rosa RT, de Paula e Carvalho A, Gursky LC, Elifio-Esposito SL, Parahitiyawa N, Samaranayake LP, Rosa EAR (2009) Proposal of a low-cost protocol for colorimetric semi-quantification of secretory phospholipase by Candida albicans grown in planktonic and biofilm phases. J Microbiol Methods 78(2):171–174. https://doi.org/10.1016/j.mimet.2009.05.012
Taučer-Kapteijn M, Hoogenboezem W, Heiliegers L, de Bolster D, Medema G (2016) Screening municipal wastewater effluent and surface water used for drinking water production for the presence of ampicillin and vancomycin resistant enterococci. Int J Hyg Environ Health 219:437–442
Tenaillon O, Skurnik D, Picard B, Denamur E (2010) The population genetics of commensal Escherichia coli. Nat Rev Microbiol 8:207–217. https://doi.org/10.1038/nrmicro2298
Thenmozhi S et al (2014) Antibiotic resistance mechanisms of ESBL producing Enterobacteriaceae in clinical field: a review. Int J Pure Appl Biosci 2:207–224
Tolouei S, Burnet JB, Autixier L, Taghipour M, Bonsteel J, Duy SV, Sauvé S, Prévost M, Dorner S (2019) Temporal variability of parasities, bacterial indicators, and wastewater micropollutants in a water resource recovery facility under various weather conditions. Water Res 148:446–458
Turolla A, Cattaneo M, Marazzi F, Mezzanotte V, Antonelli M (2018) Antibiotic resistant bacteria in urban sewage: role of full-scale wastewater treatment plants on environmental spreading. Chemosphere 191:761–769
Wan MT, Chou CHCH (2014) Spreading of β-lactam resistance gene (mecA) and methicillin-resistant Staphylococcus aureus through municipal and swine slaughterhouse wastewaters. Water Res 64:288–295
Wang Q, Wang P, Yang Q (2018) Occurrence and diversity of antibiotic resistance in untreated hospital wastewater. Sci Total Environ 621:990–999
Welker M, Moore ER (2011) Applications of whole-cell matrix-assisted laser-desorption/ionization time-of-flight mass spectrometry in systematic microbiology. Syst Appl Microbiol 34:2–11
World Health Organization WHO (2016) Antibiotic resistance. (Online in 5.2.2016). http://www.who.int/news-room/fact-sheets/detail/antibiotic-resistance
Yang YX et al (2015) Molecular characterization of resistance, virulence and clonality in vancomycin-resistant Enterococcus faecium and Enterococcus faecalis: a hospital-based study in Biejing, China. Infect Genet Evol 33:253–260
Zhang S, Han B, Gu J, Wang C, Wang P, Ma Y, Cao J, He Z (2015) Fate of antibiotic resistant cultivable heterotrophic bacteria and antibiotic resistance genes in wastewater treatment processes. Chemosphere 135:138–145
Funding
This work was financially supported by the Scientific Grant Agency VEGA (Grant number VEGA 1/0096/17), by the Slovak Research and Development Agency (Grant number APVV-16-0171) and by a project for the building of infrastructure for the modern research of civilization diseases (ITMS 26230120006).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Responsible editor: Diane Purchase
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Highlights
• First annual monitoring of resistant bacteria prevalence in wastewaters in Slovakia.
• Of all monthly sampling, the prevalence of antibiotic-resistant E. coli in influent water was the highest in October.
• Samples of effluent water taken in March, May, and December contained a high number of resistant E. coli.
• Effluent water in March and October contained vancomycin-resistant enterococci.
• Coliform bacteria resistant to antibiotics were identified as Morganella ssp. and Citrobacter ssp.
Rights and permissions
About this article
Cite this article
Lépesová, K., Olejníková, P., Mackuľak, T. et al. Annual changes in the occurrence of antibiotic-resistant coliform bacteria and enterococci in municipal wastewater. Environ Sci Pollut Res 26, 18470–18483 (2019). https://doi.org/10.1007/s11356-019-05240-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11356-019-05240-9