Abstract
Purpose
Attenuated Salmonella typhimurium is a potential biotherapeutic antitumor agent because it can colonize tumors and inhibit their growth. The present study aimed to develop a doxycycline (Doxy)-inducible gene switch system in attenuated S. typhimurium and assess its therapeutic efficacy in various tumor-bearing mice models.
Procedures
A Doxy-inducible gene switch system comprising two plasmids was engineered to trigger the expression of cargo genes (Rluc8 and clyA). Attenuated S. typhimurium carrying Rluc8 were injected intravenously into BALB/c mice bearing CT26 tumors, and bioluminescence images were captured at specified intervals post-administration of doxycycline. The tumor-suppressive effects of bacteria carrying clyA were evaluated in BALB/c mice bearing CT26 tumors and in C57BL/6 mice bearing MC38 tumors.
Results
Expression of the fimE gene, induced only in the presence of Doxy, triggered a unidirectional switch of the POXB20 promoter to induce expression of the cargo genes. The switch event was maintained over a long period of bacterial culture. After intravenous injection of transformed Salmonella into mice bearing CT26 tumors, the bacteria transformed with the Doxy-inducible gene switch system for Rluc8 targeted only tumor tissues and expressed the payloads 2 days after Doxy treatment. Notably, bacteria carrying the Doxy-inducible gene switch system for clyA effectively suppressed tumor growth and prolonged survival, even after just one Doxy induction.
Conclusions
These results suggest that attenuated S. typhimurium carrying this novel gene switch system elicited significant therapeutic effects through a single induction triggering and were a potential biotherapeutic agent for tumor therapy.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Various strategies have been studied to overcome the limitations of conventional cancer therapies such as surgery, chemotherapy, and radiotherapy [1,2,3,4]. Since Coley’s toxin was introduced in the late nineteenth century, immunotherapy using live bacteria has gained much attention [5]. Attenuated bacteria, including Clostridium [6], Bifidobacterium [7], Listeria [8], Escherichia coli [9], and Salmonella [10], have demonstrated antitumor efficacy with low systemic toxicity. These bacteria selectively colonize and multiply in tumors, leading to oxygen deprivation, excessive nutrient leakage, and an antitumor immune response [11, 12].
Bacteria can be genetically engineered to express therapeutic payload genes that enhance their antitumor efficacy [13, 14]. When delivering therapeutic payloads, such as cytotoxic molecules and cytokines, it is crucial to minimize off-target effects on normal host cells. Bacteria, when administered via the tail vein, initially and temporarily concentrate in reticuloendothelial (RE) organs. To ensure efficacy, the release of cytotoxic payloads should be timed to occur after the bacteria have been cleared from the RE organs and have accumulated in the targeted tumor tissue, typically taking place 1–3 days post-administration of the bacteria. Consequently, gene-triggering strategies have been explored to mitigate or prevent off-target effects by regulating the timing and location of payload expression, ultimately improving safety [15].
Gene expression can be triggered by signals within the tumor microenvironment. Bacteria localize mainly in the hypoxic region of tumor tissues, where changes in the partial pressure of oxygen (pO2), pH, and metabolites occur during tumor development and bacterial colonization [16, 17]. Some bacterial genes can be induced in response to such stresses, thereby serving as internal signals for gene expression [18,19,20]. Externally supplied chemicals can also be used to induce gene expression. Inducible promoters such as a bidirectional tetracycline-inducible (Ptet) [21, 22], arabinose-inducible araBAD (PBAD) [10, 23], and lactose-inducible (Ptac) [24] promoters have been used to express therapeutic cargos in bacterial cancer therapy. Among them, the Ptet system stands out as an ideal tool for bacterial-mediated cancer therapy (BCT). This is primarily due to its ability to regulate gene expression at low concentrations of doxycycline (Doxy), which readily penetrates the bacterial membrane and possesses an appropriate half-life (18 h) for in vivo applications [25]. Notwithstanding these notable advantages, the Ptet system does exhibit certain limitations. For instance, repetitive administration of Doxy could induce side effects such as alterations in the microbiome and mitochondrial dysfunction [26]. Consequently, it became imperative to develop a gene switch system capable of constitutive expression of drug payloads, thereby eliciting therapeutic effects through a single induction with Doxy.
The advent of synthetic biology technology has led to the design of gene circuits that are specific for biological activity; the aim is to exert fine control over the timing and strength of cargo gene expression in response to a specific input signal(s) [27,28,29]. Similar to the design of silicon-based electronic circuitry, genetic circuits combine various input signals in the form of AND (the output is high only when all inputs are high), NAND (the output is low only when all inputs are high), and NOR (the output is low when either input is high) logic gates [30]. Bacterial gene circuits can be designed to induce gene expression by external signals or to control the direction of gene expression through site-specific recombination, a phenomenon known as a gene switch [31]. The gene switch is mediated by two types of enzymes: tyrosine recombinases such as Cre, Flp, FimB, and FimE and serine integrases such as phiC31 and Bxb1 [32,33,34]. Generally, genes encoding enzymes inducing unidirectional recombination have been used to construct circuits that stably maintain gene expression after the switch event [35].
The fim operon encodes a set of structural genes for type 1 fimbria of uropathogenic E. coli; these genes are virulence factors that encode adhesins that bind to host epithelial cells [36]. The expression state of this operon (ON and OFF) is controlled by a 314 bp fimS fragment that is located upstream of the structural genes. This fragment is bounded by 9 bp inverted repeats (IRL, inverted repeat left; and IRR, inverted repeat right) and includes a fimA promoter (PfimA) [37, 38]. The tyrosine recombinases FimB and FimE catalyze a fimS switch through recombination, including DNA cleavage, strand exchange, and relegation against repeats. Even though the two enzymes show almost equal efficiency, FimE catalyzes a fimS switch only unidirectionally in the ON-to-OFF direction, whereas FimB does so bidirectionally (i.e., both ON-to-OFF and OFF-to-ON) [38]. Due to the unidirectional inverting property of FimE, this enzyme has been used to design recombinase-based gene circuits in bacteria [32].
In the present study, we employed synthetic biology techniques to fabricate a Doxy-inducible gene switch system consisting of two plasmids. One plasmid harbors the recombinase fimE gene downstream of the Ptet promoter, while another one contains payload genes downstream of a constitutive promoter bound by two inverted repeats. Within this gene switch system, a bioluminescence reporter Renilla luciferase variant 8 (Rluc8) [22] and a bacterial toxin cytolysin A (ClyA) [10, 14] were loaded as cargo genes, respectively. These systems were then introduced into S. typhimurium CNC018 (ΔppGpp, ΔSPI-1, ΔSPI-2), which was newly established through genetic disruption of two gene clusters called Salmonella pathogenicity islands (SPI-1 and SPI-2) in a ΔppGpp strain previously used in BCT [15]. The gene clusters encode genes related to host cell invasion and intracellular survival/proliferation of bacteria, respectively [39,40,41]. We subsequently evaluated the precise and sustained control of the gene switch system in the transformed CNC018 and assessed its antitumor efficacy in CT26 and MC38 tumor–bearing mice.
Materials and Methods
Bacterial Strains and Tumor Cell Lines
E. coli DH10-beta (New England Biolabs, USA) was used to clone and amplify plasmids. The plasmids were constructed using T4 DNA ligase (Thermo Fisher Scientific, USA) after digestion with restriction enzymes (New England Biolabs, USA) or using the Gibson assembly kit (New England Biolabs, USA). Speed pfu DNA polymerase (NanoHelix, Korea) was used for polymerase chain reaction (PCR) amplification. The primers used in the study were chemically synthesized by Macrogen, Korea, and listed in Table S1.
The attenuated S. typhimurium strain ΔppGpp was deficient in ppGpp biosynthesis through disruption of relA and spoT genes [22]. S. typhimurium CNC018 (ΔppGpp, ΔSPI-1, ΔSPI-2) was created from ΔppGpp bacterium via complete deletion of two Salmonella pathogenicity island gene clusters, SPI-1 and SPI-2, which are responsible for bacterial invasion into host cells (39 genes) and intracellular survival and replication (31 genes), respectively [39,40,41]. Bacterial transformation with plasmids was performed using an electroporator set at 2.5 kV (Bio-Rad, USA). All transformed bacteria were grown at 37 °C in Luria–Bertani (LB) broth or on LB agar plates supplemented with ampicillin (100 μg/mL) and/or chloramphenicol (34 μg/mL). Bacteria were stored at − 80 °C in 25% glycerol before use.
The murine colon carcinoma cell lines CT26 (CRL-2638, American Type Culture Collection, USA) and MC38 (ENH204-FP, Kerafast, USA) were purchased and cultured at 37 °C/5% CO2 atmosphere in high-glucose Dulbecco’s Modified Eagle Medium (DMEM) (WELGENE, Korea) containing 10% fetal bovine serum (FBS) and 1% penicillin–streptomycin.
Plasmid Construction of Doxy-Inducible Gene Switch System
NIA3, a plasmid containing the fimE recombinase gene, was a kind gift from Dr. M. Shapiro (California Institute of Technology, USA). pTU2S-a (#74088, Addgene, USA), a plasmid containing the p15A replicon (ori), was a kind gift from Dr. P. Freemont (Imperial College London, UK). pSF-OXB20, a plasmid containing the OXB20 promoter (POXB20), was purchased (#OG50R1, Oxford Genetics, UK). pBAD-Rluc8 and pBAD-ClyA, plasmids containing the Rluc8 and the bacterial cytolysin ClyA (clyA) genes, respectively, were described previously [10].
To construct plasmids expressing FimE, the fimE gene fragment with (FimE-Flag) or without (FimE-NoTag) a Flag tag sequence was amplified by PCR using NIA3 as a template. The primer sets for FimE-Flag were FimE-Flag-F and FimE-Flag-R, and those for FimE-NoTag were FimE-NoTag-F and FimE-NoTag-R. These two fragments did not contain a ribosome binding site (RBS). Using the Gibson assembly kit, the purified PCR fragments were digested with SpeI and StuI and ligated into the same sites of the pJH18 plasmid harboring an ampicillin resistance (AmpR) gene [22]. The resulting plasmids were named pJHFimTag and pJHFimNoTag, respectively, in which fimE genes with or without Flag tag were placed under the control of PtetA in Ptet dual promoter. As a target for the switch, the DNA sequence of an artificial strong constitutive promoter, POXB20, with inverted repeats at its left and right ends (IRL (36 bp) and IRR (39 bp)), was designed. The POXB20 fragment was amplified by PCR using the primers OXB-F1 and OXB-R1, with pSF-OXB20 as the template. Using the amplified fragment as a template, a second PCR reaction was performed with primers OXB-F2 and OXB-R2. The amplicon from this reaction was then used as a template for a third PCR reaction with primers OXB-F2 and OXB-R3. The final PCR product, which contained the POXB20 fragment with IRL and IRR at the ends, was named the OXB20 switch block.
The DNA fragment of the Rluc8 gene containing a highly efficient RBS sequence (5′-AAAGAGGAGAAA) was amplified using primers Rluc8-F and Rluc8-R1, with the pBAD-Rluc8 plasmid as a template. The amplified fragment was then used as a template for a second PCR using primers Rluc8-F and Rluc8-R2 to obtain the Rluc8 gene fragment carrying a His tag (5′-CATCACCATCACCATCAC) at the 3′-end. The OXB20 switch block and the His-tagged Rluc8 gene fragment were triply ligated into the pTU2S-a plasmid (cut with NdeI and EcoRI) using the Gibson assembly kit to construct a plasmid containing the Rluc8 gene with a switched promoter. The resulting plasmid, named p15AOXBR, contained a chloramphenicol resistance (CmR) gene.
The DNA fragment of the clyA gene was amplified using primers ClyA-F and ClyA-R1, with the pBAD-ClyA plasmid as a template. The amplicon was then used as a template in a second PCR with primers ClyA-F and ClyA-R2 to obtain the clyA gene with a Myc tag (5′-GAACAAAAACTCATCTCAGAAGAGGATCTG). The Myc-tagged clyA gene fragment was ligated into the p15AOXBR plasmid (cut with NdeI and PmeI) using the Gibson assembly kit to construct a plasmid containing the clyA gene with a switched promoter. The resulting plasmid was named p15AOXBC. All plasmids constructed in this study were sequenced by Macrogen, Korea.
Each plasmid used for FimE expression (i.e., pJHFimTag and pJHFimNoTag) was co-transformed into CNC018 bacteria along with p15AOXBR, resulting in CNC018::pFimTagOXBR and CNC018::pFimNoTagOXBR, respectively. The co-transformant bacteria harboring pJHFimTag and p15AOXBC was named CNC018::pFimTagOXBC. To generate controls without the Doxy-inducible gene switch system, CNC018 bacteria were transformed with pJH18-Rluc8(AP), pJH18-Rluc8(RP), or pJH18-ClyA(AP). In these constructs, the Rluc8 gene is driven by the PtetA and PtetR promoters, while the clyA gene is driven by the PtetA promoter, respectively [22]. The resulting bacteria were named CNC018::pJH18-Rluc8(AP), CNC018::pJH18-Rluc8(RP), and CNC018::pJH18-ClyA(AP), respectively.
The CNC018 transformants used in the study are listed in Table S2.
Western Blot Analysis and Luciferase Activity Assay
The transformed Salmonella CNC018::pFimTagOXBR and CNC018::pFimNoTagOXBR were cultured at 37 °C overnight (with shaking at 200 rpm) in LB broth supplemented with ampicillin (100 μg/mL) and chloramphenicol (34 μg/mL). These cultures were diluted 100-fold into 3 mL of fresh LB broth and cultured until mid-log phase (optical density at 600 nm (OD600) 0.5–0.7), at which point various concentrations of Doxy (0–500 ng/mL) (Sigma-Aldrich, Germany) were added. The cultures were then incubated for a further 22 h. During culture, OD600 values were measured at different time points (from 6 to 22 h) by spectrometry, and the bacterial pellet at each time point was collected by centrifugation at 4000 rpm for 5 min prior to further analysis.
For Western blot analysis, the bacterial pellets were resuspended in sodium dodecyl sulfate (SDS) sample buffer containing 0.2% β-mercaptoethanol (Sigma-Aldrich, Germany), sonicated on ice, and heat-treated for 10 min at 95 °C. The dissolved pellets (0.1 OD600 equivalents per lane) were separated by SDS–polyacrylamide gel electrophoresis (PAGE) on a 15% gel. The proteins in the gel were transferred to a polyvinylidene fluoride (PVDF) membrane (GE Healthcare Life Sciences, USA), which was then incubated in blocking buffer (5% (w/v) skim milk in Tris-buffered saline buffer containing 0.1% Tween 20 (TBS-T; Sigma-Aldrich, Germany)) at room temperature for 1 h. After decanting the blocking buffer, the membranes were treated with primary antibodies diluted in the blocking buffer. After incubation overnight at 4 °C, the membranes were washed three times with TBS-T and probed for 1 h with horseradish peroxidase (HRP)-conjugated secondary antibodies diluted in blocking buffer. After three washes with TBS-T, a chemiluminescent HRP substrate (Merck Millipore, MA, USA) was added, and images were captured using a ChemiDoc™ XRS + system imager (Bio-Rad, USA). The band intensity of a specific protein was quantified using the ImageJ program (National Institutes of Health and Laboratory for Optical and Computational Instrumentation, USA). All antibodies used in this study, and their dilution factors, are listed in Table S3.
To measure Rluc8 activity, the bacterial pellets (0.1 OD600) were resuspended in 100 μL of phosphate-buffered saline (PBS) and transferred to the wells of a 96-well plate. To measure bioluminescence generated by Rluc8 activity, 5 μL of 0.2 μg/μL coelenterazine (Biotium, USA) was added to each well, and bioluminescence was measured immediately at room temperature using the Orion L Microplate luminometer (Titertek Berthold, Germany). The value of the signal for each well was normalized to OD600.
PCR Analysis to Detect a Gene Switch Event
CNC018::pFimTagOXBR or CNC018::pFimNoTagOXBR bacteria were treated for 18 h with the indicated concentrations of Doxy. After measuring the OD600 value, bacteria were collected by centrifugation at 12,000 rpm for 1 min. The bacterial pellet (0.2 OD600) was resuspended in 200 μL of PBS and boiled at 95 °C for 5 min. Next, 5 μL of heat-killed bacteria was mixed with 20 μL of AccuPower PCR PreMix (Bioneer, Korea) containing the p15A-F and pOXB-R primers, and PCR was performed for 25 cycles. The PCR samples were separated by electrophoresis on a 1% agarose gel. A fragment with a size of 616 bp was expected after a switch event. The PCR fragment appearing on the gel was purified using the Zymoclean Gel DNA Recovery kit (Zymo Research, USA) and sequenced to confirm the switch event [42].
Analysis of Switch Event Maintenance by the Doxy-Inducible Gene Switch System
CNC018::pFimTagOXBR was cultured at 37 °C in the presence of various concentrations of Doxy. CNC018::pJH18-Rluc8(AP) and CNC018::pJH18-Rluc8(RP) were cultured as negative controls in the presence of 300 ng/mL Doxy. After 18 h, each bacterium was diluted 100-fold in 3 mL of fresh medium containing appropriate antibiotics in the absence of Doxy and then cultured for a further 24 h. This sub-culture was repeated two more times. Finally, the bacteria were precipitated by centrifugation at 10,000 rpm for 1 min, and Rluc8 protein levels were determined by Western blotting with an anti-His antibody as described above.
Bacterial Distribution in CT26 Tumor–Bearing Mice
All animal experiments and euthanasia procedures were performed in accordance with protocols approved by the Animal Research Committee of Chonnam National University, Korea, and NIH Guidelines for the Care and Use of Laboratory Animals [43]. Female BALB/c mice at 6 weeks of age were purchased from Orient, Korea. After anesthetization with 2% isoflurane (Hana Pharm, Korea), CT26 tumor cells (5 × 105 in 50 μL PBS) were implanted subcutaneously (s.c.) into the right flank of each mouse. Once the tumor volume reached 80–120 mm3, CNC018 or ΔppGpp bacteria (2 × 107 colony-forming unit (CFU) in 100 μL of PBS) were injected intravenously (i.v.). At the indicated days after bacterial injection, tumors and other organs were collected from the mice as described [44]. After measuring the weights, they were homogenized in PBS using the IKAT 10 Basic ULTRA-TURRAX homogenizer (IKA Dispersers, Germany). Homogenates were serially diluted in PBS and spread onto an LB agar plate. After an overnight incubation at 37 °C, the colonies were enumerated and computed as CFU per gram of tissue. Detection limit was over 103 CFU/g [44].
Bioluminescence Imaging of CT26 Tumor–Bearing Mice Treated with Bacteria Transformed with the Doxy-Inducible Rluc8 Switch System
Once the tumor volume reached approximately 150 mm3, CNC018::pFimTagOXBR or CNC018::pJH18-Rluc8(AP) bacteria (2 × 107 CFU in 100 μL of PBS) were injected intravenously. The mice were grouped following the timing of oral Doxy administration. Days 1, 2, or 3 after bacteria injection corresponded to the treatment-induction interval (TII) 1, 2, and 3 groups, respectively. Doxy (1.7 mg/kg body weight) was orally administered once. Bioluminescence imaging was performed using an in vivo imaging system (IVIS) (Lumina S5; Perkin Elmer, Waltham, MA, USA) starting from the day after Doxy administration. Coelenterazine (0.7 mg/kg body weight) was i.v. injected to observe bioluminescence. The bioluminescence intensity was described as the photon signal in the gated tumor region.
Hemolytic Assay of Bacteria with the Doxy-Inducible clyA Switch System
CNC018::pFimTagOXBC or CNC018::pJH18-ClyA(AP) bacteria were freshly cultured, collected by centrifugation, and resuspended in 100 μL of PBS at a concentration of 0.1 OD600. The bacterial suspension was then grown on blood agar plates that were pre-spread with various concentrations of Doxy. Following overnight incubation at 37 °C, hemolysis was judged as the appearance of hollow zones around the bacterial colonies.
Evaluation of Antitumor Effects in Tumor-Bearing Mice by Bacteria with the Doxy-Inducible clyA Switch System
CT26 or MC38 tumor cells (5 × 105 in 100 μL of PBS) were s.c. implanted into the right flank of female BALB/c or C57BL/6 mice at 6 weeks of age (Orient, Korea). Once the tumor volume reached 80–120 mm3, CNC018::pFimTagOXBC bacteria (2 × 107 CFU in 100 μL PBS) were i.v. injected. Doxy were orally administered to the mice of the TII 1, 2, and 3 groups. As a control, CNC018::pJH18-ClyA(AP) bacteria (2 × 107 CFU in 100 μL PBS) were i.v. injected into CT26-bearing mice, and Doxy were orally administered into the mice like the TII 1 group. Tumor size was measured with a caliper every 3 days until day 90 after bacteria injection. Tumor volume (cubic millimeter) was calculated using the formula (L × H × W)/2 formula, where L, W, and H represented the length, width, and height of the tumor, respectively. Mice with a tumor volume ≥ 1500 mm3 were euthanized.
Serum Biochemistry Parameter Analysis
After treatment of CNC018::pFimTagOXBC bacteria, venous blood from CT26 tumor–bearing mice was collected from the retro-orbital venous plexus using a micro-hematocrit capillary tube at day 5 from the TII 0, 1, 2, and 3 groups. These samples were then placed into BD Vacutainer serum tubes (Becton–Dickinson, NJ, USA) and centrifuged with 9300 × g at 4 °C for 30 min. Subsequently, the obtained serums were immediately utilized for biochemistry analyses. Levels of aspartate aminotransferase (AST) and concentration of blood urea nitrogen (BUN), creatinine, C-reactive protein (CRP), and procalcitonin (PCT) were determined using Cobas 8000 c702 and e801 automatic biochemical analysis machines (Roche Diagnostics system, USA) following the manufactures’ instructions. Standard controls were run before each determination.
Statistical Analysis
Statistical analysis was performed using Prism 9.0 software (GraphPad, USA). Student’s t-test or one-way ANOVA was used to compare single variables, while two-way ANOVA with Tukey’s correction was used for multiple comparisons. Survival rate analysis was conducted by constructing the Kaplan–Meier curves, and a long-rank (Mantel–Cox) test was used for comparison. The detailed results of statistical tests are shown in the figures. P-values < 0.05 were considered significant. All data represent the mean ± standard error of the mean (SEM).
Results
Study Design and Construction of the Doxy-Inducible Gene Switch System
The Doxy-inducible gene switch system used in this study comprised two plasmids (Fig. 1a). One, pJHFimTag, was developed from the high-copy plasmid pJH18, which contains the ColE1 ori and AmpR genes [22]. pJHFimTag encodes the TetR repressor under the weak constitutive promoter POXB1 and the Flag-tagged FimE under the PtetA promoter. Because only a low level of FimE is necessary for the rapid inversion reaction [32], the typical RBS was deleted upstream of the fimE open reading frame (ORF). The other plasmid, containing the switching target (either p15AOXBR for Rluc8 or p15AOXBC for clyA), is a low-copy number plasmid containing p15A ori and CmR and carrying the OXB20 switch block, which contains a strong constitutive promoter, POXB20, with recombinase target sequences (IRL and IRR) at either end. The gene of interest (GOI), either Rluc8 or clyA, was directed opposite to POXB20.
Schematic diagram of the Doxy-inducible gene switch system. a Plasmids comprising the Doxy-inducible gene switch system. The system consists of two plasmids p15AOXBR or p15AOXBC, as well as pJHFimTag. pJHFimTag, derived from the pJH18 plasmid [22], carries an RBS-less fimE gene driven by PtetA of the tet dual promoter and a tetR gene driven by a weak constitutive promoter, POXB1. p15AOXBR or p15AOXBC carries the OXB20 switch block (IRL-POXB20-IRR), in which the direction of POXB20 is opposite to that of the gene of interest (GOI), i.e., Rluc8 or clyA. b Mechanism of action of the Doxy-inducible gene switch system. In the absence of Doxy, the TetR repressor is expressed constitutively by a weak promoter, POXB1. It then binds to the Ptet promoter, resulting in repression of fimE gene expression. The GOI, located in the reverse direction to POXB20, is not expressed (i.e., the OFF state). Doxy binds to TetR, releasing it from the Ptet promoter and allowing expression of FimE. Consequently, FimE catalyzes inversion of the OXB20 switch block, leading to expression of the GOI (i.e., the ON state)
Expression of GOIs in the pJHFimTag system was induced by Doxy (Fig. 1b). In the absence of Doxy, TetR was expressed constitutively, which repressed fimE gene expression (the OFF state). Thus, the GOI was not expressed in either p15AOXBR or p15AOXBC. In the presence of Doxy, TetR binds to Doxy and is released from Ptet, thereby inducing the expression of fimE by PtetA (the ON state). Subsequently, FimE induces switching of the OXB20 switch block containing the POXB20 promoter to direct it in the direction of the GOIs.
Characterization of the Doxy-Inducible Gene Switch System in Bacteria
Next, we evaluated the functionality of the Doxy-inducible gene switch system in bacteria by measuring Rluc8 gene switching through fimE expression (Fig. 2a). We transformed CNC018 bacteria with the system and then measured bioluminescence generated by Rluc8. The bioluminescence signal in both CNC018::pFimTagOXBR and CNC018::pFimNoTagOXBR bacteria, which carry a fimE gene with (pJHFimNoTag) or without (pJHFimTag) a Flag tag, increased in a Doxy-dependent manner up until 18 h, before decreasing slightly hereafter. However, the bioluminescence signal generated by CNC018::pFimTagOXBR was significantly higher than that generated by CNC018::pFimNoTagOXBR at the same Doxy concentration, indicating that the fimE gene with a Flag tag is better expressed by the system.
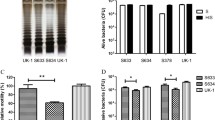
Expression of the switch target gene Rluc8 via the Doxy-inducible gene switch system. CNC018 bacteria were co-transformed with pJH18FimTag and p15AOXBR (CNC018::pFimTagOXBR), or with pJH18FimNoTag and p15AOXBR (CNC018::pFimNoTagOXBR). Plasmids pJH18FimTag and pJH18FimNoTag express FimE with and without a Flag tag, respectively, and plasmid p15AOXBR contains the Rluc8 gene as the switch target. The bacteria were diluted 100-fold, inoculated into fresh medium (0 h), and cultured at 37 °C. At an OD600 0.5–0.7 (2 h), the bacteria were treated with the indicated concentrations of Doxy and further cultured. Rluc8 activity at each time point was measured in the presence of coelenterazine and normalized to bacteria cell number (OD600). The switch events at p15AOXBR, as well as Rluc8 expression, were analyzed by PCR and Western blotting, respectively, at 18 h. a Target gene switching by FimE recombinase. CNC018::pFimTagOXBR (left) and CNC018::pFimNoTagOXBR (right) were cultured and induced by Doxy. Bioluminescence generated by Rluc8 activity at each time point was measured in the presence of coelenterazine and normalized to bacteria cell number (OD600). b PCR analysis to detect a gene switch. Positions and directions that primers p15A-F and pOXB-R bind with p15AOXBR are marked in schematic illustration (upper panels). PCR was done against heat-killed bacteria containing CNC018::pFimTagOXBR (lower left panel) and CNC018::pFimNoTagOXBR (lower right panel) using as the templates at 18 h. Subsequently, PCR products were separated in agarose gel electrophoresis. Red arrowheads, amplified PCR fragments. c Western blot analysis of proteins expressed by CNC018::pFimTagOXBR bacteria after Doxy induction. After induction with the indicated concentrations of Doxy, the proteins in the bacterial pellet (obtained at 18 h post-induction) were separated by SDS-PAGE and blotted onto a PVDF membrane. The membrane was then probed with antibodies specific for the His tag of Rluc8 (upper panel) or the Flag tag of FimE (middle panel). The stripped membrane was re-stained with an antibody specific for DnaK, an endogenous Salmonella protein (bottom panel). d Growth curve of CNC018::pFimTagOXBR bacteria. Bacteria were cultured in the presence of various concentrations of Doxy, and OD600 values were measured
PCR analysis was also performed to measure gene switching (Fig. 2b). To detect a POXB20 switch event in p15AOXBR, PCR amplification using primers p15A-F and pOXB-R should occur only in the ON state (Fig. 2b, upper panels). The expected PCR fragments were not detected in either CNC018::pFimTagOXBR (left) or CNC018::pFimNoTagOXBR (right) bacteria until treatment with Doxy at a concentration of 5 ng/mL (Fig. 2b, lower panels), indicating that leaky expression of FimE was negligible in both bacteria. The fragments were significantly detected in CNC018::pFimTagOXBR exposed to 20 ng/mL Doxy and in CNC018::pFimNoTagOXBR exposed to 100 ng/mL Doxy and appeared more strongly in those exposed to higher Doxy concentrations. The band intensity for CNC018::pFimTagOXBR was much stronger than that for CNC018::pFimNoTagOXBR in the presence of 20–500 ng/mL Doxy. The results of Western blot analysis of CNC018::pFimTagOXBR were consistent with those of bioluminescence and PCR (Fig. 2c). The FimE and Rluc8 bands were faint at 20 ng/mL Doxy, but the intensity increased gradually in a Doxy-dependent manner. Growth of CNC018::pFimTagOXBR was not affected by Rluc8 expression (Fig. 2d). These results indicate that fimE expression is tightly controlled by Doxy in our system and that the fimE gene downstream of PtetA is more efficiently and strongly expressed with a Flag tag than without.
Genetic Stability of a Switch Event Induced by the Doxy-Inducible Gene Switch System
FimE induces unidirectional inversion of a gene surrounded by inverted repeats; this is because this bacterial recombinase recognizes IRR more than IRL [32, 38]. Thus, a gene switch event induced by FimE is maintained permanently and is inheritable [20, 45].
We assessed whether a gene switch event induced by the Doxy-inducible gene switch system was stably maintained in bacteria (Fig. 3a). CNC018::pFimTagOXBR bacteria were induced for 18 h with various concentrations of Doxy and then cultured without Doxy for 3 days. Expression of Rluc8 was evaluated by Western blotting (Fig. 3b). The Rluc8 band in bacteria cultured with Doxy for 18 h was faint after induction with 20 ng/mL but gradually became more pronounced as the concentration of Doxy increased to 500 ng/mL (Fig. 3b, left panel). After sub-culture without Doxy for 3 days, the Rluc8 band in cells cultured initially with 100 ng/mL Doxy was faint, but again, it became gradually more pronounced as the initial concentration increased to 500 ng/mL. This indicated that a switch event induced by the Doxy-inducible gene switch system at this concentration was stably maintained for at least 3 days. However, at concentrations below 300 ng/mL, the level of expression after culture with Doxy for 18 h was higher than that after sub-culture without Doxy for 3 days (Fig. 3b, right panel). This may be due to incomplete gene switching at these Doxy concentrations.
Permanent inheritance of a gene switch event in Doxy-inducible switch system. a Experimental design. CNC018::pFimTagOXBR bacteria were cultured in the presence of the indicated concentrations of Doxy. After 18 h, the bacteria were diluted 100-fold and sub-cultured for 1 day in fresh medium supplemented with appropriate antibiotics but without Doxy. This sub-culture process was repeated three times. CNC018::pJH18-Rluc8(AP) and CNC018::pJH18-Rluc8(RP) bacteria, treated with 300 ng/mL Doxy, were used as negative controls. b Expression of Rluc8 in CNC018::pFimTagOXBR bacteria cultured with Doxy for 18 h and then sub-cultured without Doxy for 3 days. Rluc8 protein in bacterial pellets was analyzed by Western blotting with an anti-His tag antibody (left panel). The intensity of the Rluc8 bands was quantified and depicted as a graph (right panel). c Rluc8 expression in CNC018 bacteria without Doxy-inducible switch system in cultured with Doxy for 18 h and sub-culture without Doxy for 3 days. Rluc8 proteins in CNC018::pJH18-Rluc8(AP) and CNC018::pJH18-Rluc8(RP) were analyzed in Western blot analysis with anti-His tag antibody (left panel). The Rluc8 band intensities were depicted as a quantitation graph (right panel)
Next, we conducted the same experiment using CNC018::pJH18-Rluc8(AP) and CNC018::pJH18-Rluc8(RP) and found that Rluc8 expression was induced by 300 ng/mL Doxy (Fig. 3c, left panel). These bacteria expressed significant amounts of Rluc8 upon culture with these inducers for 18 h, but not after sub-culture in their absence for 3 days. This was confirmed in the quantitation data (Fig. 3c, right panel). These results indicate that a switch event in the Doxy-inducible system occurs in response to Doxy and is permanently maintained during bacterial division and plasmid replication.
Biodistribution of CNC018 Bacteria in CT26 Tumor–Bearing Mice
We investigated the biodistribution of CNC018 and ΔppGpp strains within the tumor, liver, spleen, and blood by quantifying viable bacterial populations at 1, 2, 3, and 5 days following bacterial injection. Notably, both CNC018 and the ΔppGpp strain demonstrated a substantial presence within the tumor, persisting from day 1 through day 5. While a significant number of ΔppGpp strains were detected in the liver and spleen on days 1 and 2, no cultivable CNC018 were recovered from these organs above the limit of detection (approximately 1 × 103 colony-forming units (CFU)). After 1 day following injection, neither CNC018 nor the ΔppGpp strain were detectable in the bloodstream (Fig. S1). These findings underscore that CNC018 bacteria were effectively cleared from the bloodstream and normal organs while concurrently establishing colonization and proliferation within the tumor. Consequently, the Doxy-inducible system in the CNC018 strain was found to be inducible from day 1 following bacterial injection.
Tumor Targeting of Bacteria Transformed with the Doxy-Inducible Rluc8 Gene Switch System
To evaluate the tumor-targeting efficacy of bacteria transformed with the Doxy-inducible gene switch system, we injected CNC018::pFimTagOXBR i.v. into CT26 tumor–bearing mice (Fig. 4a). Bioluminescence signals were not detectable for 3 days after bacterial treatment of mice not administered Doxy (pre-treatment) (Fig. 4b, c). When the mice received Doxy administration at days 1, 2, or 3 after bacteria injection (treatment-induction interval (TII) 1, 2, and 3 groups), signals were detected in the tumor region (days 2, 3, and 4 in the TII 1, 2, and 3 groups, respectively) and reached their maximum level in all groups at 2–3 days after Doxy induction (days 3, 4, and 6 in the TII 1, 2, and 3 groups, respectively). Furthermore, the maximum bioluminescence signal intensity was significantly higher in the TII 1 and 2 groups than in the TII 3 group. In addition, in a control group of mice injected with CNC018::pJH18-Rluc8(AP) and subjected to a single dose of Doxy on day 1 following bacteria injection (TII 1 group), signals were observed within the tumor region as early as the day following Doxy induction (day 2). However, these signals subsequently diminished and eventually disappeared (Fig. 4b). Furthermore, it is noteworthy that the maximum signal intensity of bioluminescence was significantly greater in the CNC018::pFimTagOXBR TII 1 group when compared to CNC018::pJH18-Rluc8(AP) TII 1 group (Fig. 4b, c). These results indicate that bacteria carrying the Doxy-inducible gene switch system specifically target tumor tissues within 2 days after intravenous injection and that Doxy induction at 1 or 2 days after bacterial injection induces optimal gene switching.
Targeting of bacteria carrying Doxy-inducible switch system of Rluc8 gene in tumor xenograft mice. a Experimental scheme. BALB/c (n = 3) were subcutaneously (s.c.) implanted with CT26 cells (5 × 105) on the hind flank. At about 150 mm3 tumors volume, the mice were intravenously (i.v.) injected with CNC018::pFimTagOXBR or CNC018::pJH18-Rluc8(AP) (2 × 107 CFU) (day 0). Doxy (1.7 mg/kg body weight) was orally administrated once at days 1, 2, and 3 (treatment-induction interval (TII) 1, 2, and 3 groups, respectively). Mice treated with CNC018::pJH18-Rluc8(AP) received a single oral dose of Doxy on day 1 following bacteria injection (TII 1 group). Bioluminescence images of whole body were obtained at the indicated days soon after coelenterazine injection (0.7 mg/kg body weight) via tail vein. b Bioluminescence images of whole body. The mice treated with coelenterazine just before Doxy injection were taken as imaging controls (pre-treatment, red boxes). c Quantification of bioluminescence signals in the tumor sites. Arrows, days of Doxy treatment. Statistical significance was calculated with two-way analysis of variance (ANOVA) with Sidák’s multiple comparison test. CNC018::pFimTagOXBR: *P = 0.0233 (TII 2 group vs. TII 3 group at day 4); *P = 0.0123 (TII 1 group vs. TII 3 group at day 6); **P = 0.0029 (TII 2 group vs. TII 3 group at day 10); *P = 0.0301 (TII 1 group vs. TII 3 group at day 10). CNC018::pFimTagOXBR TII 1 group vs. CNC018::pJH18-Rluc8(AP) TII 1 group: **P = 0.0018 (at day 3); **P = 0.0022 (at day 6). All data are shown as mean ± SEM
Antitumor Effects of Bacteria Transformed with the Doxy-Inducible clyA Gene Switch System
Finally, we characterized the in vivo antitumor effects of bacterium CNC018::pFimTagOXBC, which contains both pJHFimTag for Flag-tagged FimE and p15AOXBC for ClyA with a switch block (Fig. 1a and S2A–C). Upon in vitro culture, ClyA (34 kDa) was not detected in the absence of Doxy treatment (Fig. S2A). The protein was detected faintly at 5 ng/mL Doxy, and maximum levels were observed in the pellet at > 20 ng/mL Doxy. A significant amount of ClyA was detected in the culture medium in the presence of 20 ng/mL Doxy, with the maximum level being detected at 300 ng/mL Doxy (Fig. S2A). This result indicates that ClyA expression is specifically induced by Doxy, and the protein is secreted by the bacteria. The hemolytic activity of this bacterium was assessed on blood agar plates pre-spread with various concentrations of Doxy (Fig. S2B). After culture for 1 day in the absence of Doxy, no hemolytic zones appeared around colonies. Colonies with hemolytic zones started to appear in plates containing 5 ng/mL Doxy, and the numbers increased at higher Doxy concentrations. This suggests that ClyA is functionally expressed only after gene switching induced by Doxy induction. It is notable that colonies with and without hemolysis appeared on all Doxy plates, indicating that gene switching did not occur completely, even at higher Doxy concentrations. There was no difference in ClyA activity between CNC018::pFimTagOXBC and CNC018::pJH18-ClyA(RP) (Fig. S3). The growth of this bacterium was not affected by any of the concentrations of Doxy (Fig. S2C).
The antitumor effect of CNC018::pFimTagOXBC was assessed in BALB/c mice bearing CT26 tumors (Fig. 5a–c). As in the experiment shown in Fig. 4, Doxy was administrated orally 1, 2, or 3 days after bacteria injection (the TII 1, 2, and 3 groups, respectively). Mice treated with bacteria in the absence of Doxy (no induction) showed significant tumor suppression compared with non-treated mice (PBS) (Fig. 5a, b). Thus, suppressive effect was enhanced in all Doxy-induced groups but was most prominent in the TII 1 group (Fig. 5b). Additionally, complete tumor eradication on day 51 was observed only in the Doxy-induced groups (10 of 12 (83%) in TII 1, 9 of 12 (75%) in TII 2, and 7 of 12 (58%) in TII 3). In the CNC018::pJH18-ClyA(AP) TII 1 group, 1 of 5 (20%) complete tumor eradication. The survival rates were consistent with these results (Fig. 5b, c and Fig. S4).
Tumor-suppressive effect of bacteria carrying the Doxy-inducible switch system for the clyA gene. The tumor-suppressive effects were assessed in BALB/c mice bearing CT26 tumors. The CNC018::pFimTagOXBC or CNC018::pJH18-ClyA(AP) bacteria (2 × 107 CFU) were injected intravenously into BALB/c mice bearing CT26 tumors on day 0. Control mice were treated with PBS (n = 5). Mice treated with CNC018::pFimTagOXBC (n = 12 each group) were orally administered a single dose once with Doxy (1.7 mg/kg body weight) on days 1, 2, or 3 after bacterial treatment (treatment-induction interval (TII) 1, 2, and 3 groups, respectively); mice treated with CNC018::pJH18-ClyA(AP) (n = 5) were received a single dose once with Doxy (1.7 mg/kg body weight) on day 1 (TII 1 group). Other bacteria-treated mice were not treated with Doxy (no induction group) (n = 5). a Representative images of BALB/c mice bearing CT26 tumors. b Average tumor growth curves for mice bearing CT26 tumors. Statistical significance was calculated using two-way analysis of variance (ANOVA) with Sidák’s multiple comparison test. *P = 0.0427 on day 12 and ****P < 0.0001 on the other days (PBS vs. no induction group); *P = 0.012 on day 15 and ****P < 0.0001 on days 18, 21, and 24 (TII 1 group vs. no induction group); **P = 0.014 on day 18 and ****P < 0.0001 on days 21 and 24 (TII 2 group vs. no induction group); *P = 0.0197 on day 18, ***P = 0.0001 on day 21, and ****P < 0.0001 on day 24 (TII 3 group vs. no induction group); **P = 0.01 on day 24 (CNC018::pJH18-ClyA(AP) TII 1 group vs. no induction group). c The Kaplan–Meier survival curves for BALB/c mice bearing CT26 tumors. Statistical significance was calculated using the log-rank (Mantel–Cox) test (induction groups vs. the no induction group). The number of mice in which tumors were eradicated is shown for each group. ****P < 0.0001 for the TII 1 and 2 groups; ***P = 0.0002 for the TII 3 group; P = 0.0993 for the CNC018::pJH18-ClyA(AP) TII 1 group. All data represent the mean ± SEM
The antitumor effect of CNC018::pFimTagOXBC was similarly observed in C57BL/6 mice bearing MC38 tumors (Fig. S5A-C). Compared with the group without Doxy induction, all Doxy-induced groups showed significant tumor suppression (Fig. S5A). The antitumor effect was most notable in the TII 1 group. Complete tumor eradication was observed only in this group (i.e., 5 of 5 (100%) in TII 1, 4 of 5 (80%) in TII 2, and 3 of 5 (60%) in TII 3), and survival rates were consistent with this result (Fig. S5B, S5C and Fig. S6). The results demonstrated that bacteria carrying the Doxy-inducible clyA gene switch system exhibited robust antitumor efficacy, a phenomenon observed solely in the presence of Doxy.
Safety Analysis of Bacteria Transformed with the Doxy-Inducible clyA Gene Switch System
The body weight of mice in all bacteria-treated groups fell by approximately 10% by day 3 post-bacteria injection, regardless of Doxy induction. However, by day 12, body weight had returned to normal levels (Fig. S7). This observation is consistent with previous reports showing that bacterial infection of the bloodstream can trigger an immune response, leading to temporary weight loss. However, clearance of bacteria within hours to days after treatment allows for recovery of body weight [10, 12, 22].
Finally, to further assess biosafety, biochemistry parameters were evaluated in the serum of mice subjected to treatment with CNC018::pFimTagOXBC. These parameters encompassed AST (54–298 IU/L), BUN (8–33 mg/dl), creatinine (0.2–0.9 mg/dl), CRP (< 0.5 mg/dL), and PCT (< 0.5 ng/mL). The results revealed that there were no significant alterations in AST, BUN, creatinine, CRP, and PCT levels in the treatment groups when compared to the control group (Fig. S8). Consequently, our system exhibited no discernible evidence of inducing severe systemic toxicity and demonstrated a high degree of biosafety when employed for cancer immunotherapy.
Discussion
The present study reports the design, construction, and characterization of a Doxy-inducible gene switch system for the expression of therapeutic payload genes by cancer-targeting bacteria. The system comprises two plasmids: the pJHFimTag plasmid encodes the TetR repressor and the Flag-tagged FimE recombinase under the control of constitutive and inducible promoters, respectively, while the p15AOXBR and p15AOXBC plasmids contain the GOI, either Rluc8 or clyA, respectively, under the control of the POXB20 promoter harboring recombinase target sequences. This system enabled tight and sustainable control of target gene expression after a single administration of the Doxy. The system was evaluated both in vitro and in vivo, and it was effective at suppressing tumor growth in mice.
A single administration of Doxy to induce expression of therapeutic payloads could serve as an advanced approach because multiple administrations might cause inconvenience and side effects to patients, as well as make more work for busy medical professionals. In our system, a single Doxy treatment induced gene switching and allowed for the sufficient expression of a target cargo after a single injection. The activities of Rluc8 and ClyA were detectable in vitro and clearly measurable in tumor-bearing mice. The unidirectional switch event in bacteria also increased the stable expression of the payloads. Furthermore, the gene expression level of the Doxy-inducible gene switch system was comparable to that of the Doxy-inducible tet system, highlighting its potential as an approach for in vivo cancer treatment.
Doxy has been shown to be an immunomodulator with wide-spectrum antitumor effects [46]. Sun et al. [46] explored the antitumor effect by injecting Doxy (150 mg/kg/day) into the peritoneal cavity of mice bearing B16/F10 melanoma for 14 days, resulting in a 35.6% reduction in tumor growth. In the present study, we employed a much lower dose of Doxy (1.7 mg/kg/day) for 1 day. Given the differences in dosage and treatment duration, it appears that the Doxy regimen we employed may not have exerted a significant impact on tumor growth.
The CNC018::pFimTagOXBR and CNC018::pFimTagOXBC studies suggest that the optimal time of Doxy administration might be 1 or 2 days after bacterial treatment; indeed, we noted higher bioluminescence signals and better therapeutic effects in mice from the TII 1 and 2 groups than in those from the TII 3 group. This is consistent with previous reports indicating that 2 or 3 days are required for S. typhimurium to colonize tumors in mice [20]. Notably, even when induced at an early time point (TII 1 or 2), there was no evidence of systemic toxicity.
The Flag tag sequence encodes an octapeptide (DYKDDDDK) widely used for identification and purification of various recombinant proteins [47]. We found accidentally that the PtetA promoter expressed the fimE gene with a Flag tag at its 3′-end more efficiently than the gene without this tag, even though the gene did not contain a canonical RBS upstream. This may be linked to mRNA stability. Several reports show that the addition of an extra-nucleotide at the 3′-end can increase mRNA stability [48, 49]. Another possibility is the solubility of Flag-tagged FimE. The amino acid residues in the tag are highly hydrophilic. E. coli sometimes express recombinant proteins conjugated to soluble tags in higher amounts than untagged proteins [50].
Regarding the translational potential of the engineered bacteria expressing ClyA (CNC018::pFimTagOXBC), our study has shown that a single dose of Doxy induction leads to significant tumor suppression across various murine tumor models. However, it is essential to note that our present study did not delve into the mechanism by which ClyA-expressing bacteria induce tumor cell death. We postulate that ClyA, secreted by tumor-targeting bacteria, selectively binds to tumor cells via cholesterol, potentially inducing tumor cells death. The process may, in turn, facilitate the recruitment of immune cells. Nevertheless, a further in-depth investigation into this mechanism is currently underway.
In conclusion, the Doxy-inducible gene switch system enables precise and sustainable control of target gene expression after a single Doxy administration. The tumor-targeting bacteria carrying this system have the potential for use as a highly effective and safe treatment for cancer.
References
Chau CH, Steeg PS, Figg WD (2019) Antibody–drug conjugates for cancer. Lancet 394:793–804
Zhong L, Li Y, Xiong L et al (2021) Small molecules in targeted cancer therapy: advances, challenges, and future perspectives. Signal Transduct Target Ther 6:201
Mitchell MJ, Billingsley MM, Haley RM, Wechsler ME, Peppas NA, Langer R (2021) Engineering precision nanoparticles for drug delivery. Nat Rev Drug Discovery 20:101–124
Sgouros G, Bodei L, McDevitt MR, Nedrow JR (2020) Radiopharmaceutical therapy in cancer: clinical advances and challenges. Nat Rev Drug Discovery 19:589–608
Loughlin KR (2020) William B. Coley: his hypothesis, his toxin, and the birth of immunotherapy. Urol Clin 47:413–417
Roberts NJ, Zhang L, Janku F et al (2014) Intratumoral injection of Clostridium novyi-NT spores induces antitumor responses. Sci Transl Med 6:249ra111
Longhi G, Van Sinderen D, Ventura M, Turroni F (2020) Microbiota and cancer: the emerging beneficial role of bifidobacteria in cancer immunotherapy. Front Microbiol 11:575072
Vitiello M, Evangelista M, Di Lascio N et al (2019) Antitumoral effects of attenuated Listeria monocytogenes in a genetically engineered mouse model of melanoma. Oncogene 38:3756–3762
Kang S-R, Jo EJ, Nguyen VH et al (2020) Imaging of tumor colonization by Escherichia coli using 18 F-FDS PET. Theranostics 10:4958
Nguyen VH, Kim H-S, Ha J-M, Hong Y, Choy HE, Min J-J (2010) Genetically engineered Salmonella typhimurium as an imageable therapeutic probe for cancer. Can Res 70:18–23
Lou X, Chen Z, He Z, Sun M, Sun J (2021) Bacteria-mediated synergistic cancer therapy: small microbiome has a big hope. Nano-Micro Lett 13:1–26
Zhou S, Gravekamp C, Bermudes D, Liu K (2018) Tumour-targeting bacteria engineered to fight cancer. Nat Rev Cancer 18:727–743
Sieow BF-L, Wun KS, Yong WP, Hwang IY, Chang MW (2021) Tweak to treat: reprograming bacteria for cancer treatment. Trends Cancer 7:447–464
Nguyen D-H, Chong A, Hong Y, Min J-J (2023) Bioengineering of bacteria for cancer immunotherapy. Nat Commun 14:3553
Kang S-R, Nguyen D-H, Yoo SW, Min J-J (2022) Bacteria and bacterial derivatives as delivery carriers for immunotherapy. Adv Drug Deliv Rev 181:114085
Chen W, Wang Y, Qin M et al (2018) Bacteria-driven hypoxia targeting for combined biotherapy and photothermal therapy. ACS Nano 12:5995–6005
Chien T, Harimoto T, Kepecs B et al (2022) Enhancing the tropism of bacteria via genetically programmed biosensors. Nat Biomed Eng 6:94–104
Dharanishanthi V, Orgad A, Rotem N et al (2021) Bacterial-induced pH shifts link individual cell physiology to macroscale collective behavior. Proc Natl Acad Sci 118:e2014346118
George SE, Hrubesch J, Breuing I et al (2019) Oxidative stress drives the selection of quorum sensing mutants in the Staphylococcus aureus population. Proc Natl Acad Sci 116:19145–19154
Qin Y, You S-H, Zhang Y, Venu A, Hong Y, Min J-J (2023) Genetic programming by nitric oxide-sensing gene switch system in tumor-targeting bacteria. Biosensors 13:266
Leventhal DS, Sokolovska A, Li N et al (2020) Immunotherapy with engineered bacteria by targeting the STING pathway for anti-tumor immunity. Nat Commun 11:2739
Nguyen D-H, You S-H, Vo A-TN et al (2022) Optimized doxycycline-inducible gene expression system for genetic programming of tumor-targeting bacteria. Mol Imag Biol 24:82–92
Zheng JH, Nguyen VH, Jiang S-N et al (2017) Two-step enhanced cancer immunotherapy with engineered Salmonella typhimurium secreting heterologous flagellin. Sci Transl Med 9:eaak9537
Cronin CA, Gluba W, Scrable H (2001) The lac operator-repressor system is functional in the mouse. Genes Dev 15:1506–1517
Jiang S-N, Park S-H, Lee HJ et al (2013) Engineering of bacteria for the visualization of targeted delivery of a cytolytic anticancer agent. Mol Ther 21:1985–1995
Wüst RC, Houtkooper RH, Auwerx J (2020) Confounding factors from inducible systems for spatiotemporal gene expression regulation. J Cell Biol 219 (7):e202003031
Cubillos-Ruiz A, Guo T, Sokolovska A et al (2021) Engineering living therapeutics with synthetic biology. Nat Rev Drug Discovery 20:941–960
Sedlmayer F, Aubel D, Fussenegger M (2018) Synthetic gene circuits for the detection, elimination and prevention of disease. Nat Biomed Eng 2:399–415
Ruder WC, Lu T, Collins JJ (2011) Synthetic biology moving into the clinic. Science 333:1248–1252
Lim WA (2010) Designing customized cell signalling circuits. Nat Rev Mol Cell Biol 11:393–403
Brophy JA, Voigt CA (2014) Principles of genetic circuit design. Nat Methods 11:508–520
Ham TS, Lee SK, Keasling JD, Arkin AP (2006) A tightly regulated inducible expression system utilizing the fim inversion recombination switch. Biotechnol Bioeng 94:1–4
Mohaisen MR, McCarthy AJ, Adriaenssens EM, Allison HE (2020) The site-specific recombination system of the Escherichia coli bacteriophage Φ24B. Front Microbiol 11:578056
Yao S, Yuan P, Ouellette B et al (2020) RecV recombinase system for in vivo targeted optogenomic modifications of single cells or cell populations. Nat Methods 17:422–429
Bonnet J, Subsoontorn P, Endy D (2012) Rewritable digital data storage in live cells via engineered control of recombination directionality. Proc Natl Acad Sci 109:8884–8889
Schwan WR (2011) Regulation of fim genes in uropathogenic Escherichia coli. World J Clin Infect Dis 1:17
Klemm P (1986) Two regulatory fim genes, fimB and fimE, control the phase variation of type 1 fimbriae in Escherichia coli. EMBO J 5:1389–1393
McCusker MP, Turner EC, Dorman CJ (2008) DNA sequence heterogeneity in Fim tyrosine-integrase recombinase-binding elements and functional motif asymmetries determine the directionality of the fim genetic switch in Escherichia coli K-12. Mol Microbiol 67:171–187
Lawrence A-LE, Abuaita BH, Berger RP et al (2021) Salmonella enterica serovar Typhimurium SPI-1 and SPI-2 shape the global transcriptional landscape in a human intestinal organoid model system. MBio 12:e00399-e321
Liss V, Swart AL, Kehl A et al (2017) Salmonella enterica remodels the host cell endosomal system for efficient intravacuolar nutrition. Cell Host Microbe 21:390–402
Jennings E, Thurston TL, Holden DW (2017) Salmonella SPI-2 type III secretion system effectors: molecular mechanisms and physiological consequences. Cell Host Microbe 22:217–231
Yang L, Nielsen AA, Fernandez-Rodriguez J et al (2014) Permanent genetic memory with> 1-byte capacity. Nat Methods 11:1261–1266
National Research Council (2011) Guide for the care and use of laboratory animals: eight edition. National Academies Press, Washington, DC. Available from: https://www.ncbi.nlm.nih.gov/books/NBK54050/, https://doi.org/10.17226/12910
Chowdhury S, Castro S, Coker C, Hinchliffe TE, Arpaia N, Danino T (2019) Programmable bacteria induce durable tumor regression and systemic antitumor immunity. Nat Med 25:1057–1063
Harbaugh SV, Goodson MS, Dillon K, Zabarnick S, Kelley-Loughnane N (2017) Riboswitch-based reversible dual color sensor. ACS Synth Biol 6:766–781
Sun B, Zhang S, Zhang D, Yin X, Wang S, Gu Y, Wang Y (2007) Doxycycline influences microcirculation patterns in B16 melanoma. Exp Biol Med 232:1300–1307
Kosobokova E, Skrypnik K, Kosorukov V (2016) Overview of fusion tags for recombinant proteins. Biochem Mosc 81:187–200
Hajnsdorf E, Kaberdin VR (2018) RNA polyadenylation and its consequences in prokaryotes. Philos Trans R Soc B: Biol Sci 373:20180166
Frederick MI, Heinemann IU (2021) Regulation of RNA stability at the 3′ end. Biol Chem 402:425–431
Ki M-R, Pack SP (2020) Fusion tags to enhance heterologous protein expression. Appl Microbiol Biotechnol 104:2411–2425
Funding
This work was supported by the Bio & Medical Technology Development Program of National Research Foundation of Korea (NRF) (No. NRF-2020M3A9G3080282), NRF grants (Nos. 2020R1A5A2031185, 2021M3C1C3097637, 2021C300) funded by Ministry of Science and IT (MSIT), and by the Korea Drug Development Fund funded by Ministry of Science and ICT, Ministry of Trade, Industry, and Energy, and Ministry of Health and Welfare (HN22C0637, RS-2022-00167101). Y.H. was supported by the Bio & Medical Technology Development Program of NRF (No. NRF-2020M3A9G3080330).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of Interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Ngo, H.TT., Nguyen, DH., You, SH. et al. Reprogramming a Doxycycline-Inducible Gene Switch System for Bacteria-Mediated Cancer Therapy. Mol Imaging Biol 26, 148–161 (2024). https://doi.org/10.1007/s11307-023-01879-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11307-023-01879-6