Abstract
Purpose
The purpose of this study was to evaluate magnetic particle imaging (MPI) as a method for the in vivo tracking of dendritic cells (DC). DC are used in cancer immunotherapy and must migrate from the site of implantation to lymph nodes to be effective. The magnitude of the ensuing T cell response is proportional to the number of lymph node-migrated DC. With current protocols, less than 10% of DC are expected to reach target nodes. Therefore, imaging techniques for studying DC migration must be sensitive and quantitative. Here, we describe the first study using MPI to detect and track DC injected into the footpads of C57BL/6 mice migrating to the popliteal lymph nodes (pLNs).
Procedures
DC were labelled with Synomag-D™ and injected into each hind footpad of C57BL/6 mice (n = 6). In vivo MPI was conducted immediately and repeated 48 h later. The MPI signal was measured from images and related to the signal from a known number of cells to calculate iron content. DC numbers were estimated by dividing iron content in the image by the iron per cell measured from a separate cell sample. The presence of SPIO-labeled DC in nodes was validated by ex vivo MPI, histology, and fluorescence microscopy.
Results
Day 2 imaging showed a decrease in MPI signal in the footpads and an increase in signal at the pLNs, indicating DC migration. MPI signal was detected in the left pLN in four of the six mice and two of the six mice showed MPI signal in the right pLN. Ex vivo imaging detected signal in 11/12 nodes. We report a sensitivity of approximately 4000 cells (0.015 µg Fe) in vivo and 2000 cells (0.007 µg Fe) ex vivo.
Conclusions
Here, we describe the first study to use MPI to detect and track DC in a migration model with immunotherapeutic applications. We also bring attention to the issue of resolving unequal signals within close proximity, a challenge for any pre-clinical study using a highly concentrated tracer bolus that shadows nearby lower signals.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Cell therapy involves the therapeutic administration of immune cells (such as T cells or dendritic cells (DC)) and stem cells (such as mesenchymal stem cells or pluripotent stem cells). Significant progress in cell therapy has occurred over the past decade, used to treat cancers [1], autoimmune [2], degenerative [3], genetic [4], and infectious diseases [5].
Despite its immense potential, clinical results from DC cancer vaccine trials have been variable and discordant because of disparities in cell source, preparation, route of administration, and implantation methodologies [6]. Many critical questions about the delivery, anatomical location/distribution, numbers, and persistence of DC remain unanswered. For example, in DC cancer immunotherapy, the magnitude of an anti-tumour response is proportional to the quantity of antigen presenting cells that reach a target lymph node [7,8,9], and therefore, it is crucial to know whether the injected cells have migrated to the target and in what magnitude to establish correlates with vaccine efficacy. Unfortunately, it remains difficult to spatially and quantitatively track the migration of cells and the ensuing immune response that is elicited.
Magnetic resonance imaging (MRI) has been used to track DC migration. This involves ex vivo labelling of DCs with superparamagnetic iron oxide nanoparticles (SPIO) prior to administration and imaging with pulse sequences sensitive to iron. In MRI, SPIOs produce negative contrast (signal loss) that is indirectly detected through its effect on proton relaxation. SPIO-based MRI cell tracking has very high sensitivity (tens to hundreds of cells), albeit with low specificity, and it is difficult to accurately quantify the local tissue concentration of SPIO particles [10].
Magnetic particle imaging (MPI) is a tracer-based imaging modality that directly detects the response of SPIO to an applied magnetic field [11, 12]. MPI has potential for overcoming the challenges of MRI-based cell tracking because it has high specificity [13], and the MPI signal is linearly related to iron mass allowing for quantitation of the amount of iron from images, and with knowledge of the amount of iron/cell after SPIO labelling, the cell number can be calculated [14]. A current challenge with MPI is resolution. The shape of the point spread function from the acquired signal can limit the ability to quantify two unequal sources of MPI signal in close proximity. This manifests as large signals concealing nearby smaller signals. This becomes an issue when several samples with different concentrations are being imaged [15].
In this study, we evaluate the use of MPI for quantitative tracking of DC in a pre-clinical mouse model. DCs are subcutaneously administered into the hind footpad and migrate to the draining popliteal node (pLN) located on the back of the knee. The ability to detect and resolve the high signal in the footpad from the low signal in the lymph node is studied using samples of SPIO and injections of SPIO-labeled DC in mice.
Methods
Cell Labelling
Animals
C57BL/6 mice were sourced from the breeding operation at the West Valley Barrier Facility at Western University. All applicable animal protocols were approved by the University of Western Ontario Animal Care and Use Subcommittee.
Mouse Bone Marrow–Derived Dendritic Cell Culture
Mouse bone marrow–derived DCs were isolated from the femurs and tibias of C57BL/6 mice and cultured as previously described [16,17,18]. Briefly, bone marrow progenitor cells were cultured in complete RPMI media supplemented with interleukin-4 (IL-4, 10 ng/ml, PeproTech, Canada) and granulocyte–macrophage colony-stimulating factor (GM-CSF, 4 ng/ml, PeproTech, Canada) for 4 days. On day 4, immature DC were enriched from culture using Histodenz™ (Millipore Sigma, Canada) gradient centrifugation.
SPIO Labelling of DC
Following gradient enrichment of immature DC on day 4, DC were labelled with Synomag-D (Micromod GmbH) (200 µg Fe/ml) via simple co-incubation in complete RPMI (+ IL-4 and GM-CSF). To increase Synomag-D uptake by DC, transfection agents (TAs) were employed. Protamine sulfate (0.24 mg/ml) and heparin (8 USP units/ml) were individually diluted in OPTI-MEM in Eppendorf tubes. Following Synomag-D addition to OPTI-MEM containing heparin, the contents of both Eppendorf tubes were combined, vortexed, and added to immature DC. Synomag-D was added such that DC were cultured in a final concentration of 200 µg Fe/ml for 4 h, after which time complete RPMI (+ IL-4 and GM-CSF) was added to ensure DC were cultured overnight at the same cell concentration as DC labelled via simple co-incubation. On day 5 of culture, regardless of which SPIO labelling method was used on day 4, a previously defined maturation cocktail [19] was added to the labelled DC which were then cultured for an additional 24 h.
Magnetic Selection of SPIO + DC
To ensure that only Synomag-D + DC were administered, magnetic column enrichment was performed as previously described [20]. Briefly, on day 6, mature DC were collected, washed in PBS and then resuspended in 2 ml PBS per DC culture condition in a 12- × 75-mm sterile tube. DC suspensions were incubated in an EasySep™ magnet (Stemcell Technologies, Vancouver, CAN) at room temperature. After 5 min, both tube and magnet were inverted to collect flow through containing unlabeled DC, while Synomag-D + DC remained in the tube and were collected, counted, and prepared for subsequent adoptive cell transfer. An aliquot of Synomag-D + DC was collected to histologically confirm Synomag-D uptake through Perls Prussian Blue (PPB) staining. Unlabeled DC were used for viability and immunophenotyping analysis.
Flow Cytometry
DC were phenotyped and assessed for viability upon completion of a 6-day culture using a previously outlined protocol [21]. Briefly, Zombie NIR™ fixable viability dye (Biolegend, San Diego, USA) was employed to assess DC viability and preceded surface immunofluorescence staining. In the presence of TruStain FcX™ anti-mouse CD16/32 block (clone: 93), DC were stained with PE CD45 (30-F11) and APC CD11c (N418) for 25 min at 4 °C (all Biolegend). Excess antibodies were removed by washing with HBSS + 0.1% BSA. Upon resuspension in HBSS + 0.1% BSA, DC were fixed with 4% paraformaldehyde and kept at 4 °C until acquisition using a LSRII analytical flow cytometer (BD Biosciences, San Jose, USA). FlowJo software (v10, Tree Star, Inc., Ashland, USA) was used for all analyses.
In Vitro MPI
MPI Relaxometry
Synomag-D was compared VivoTrax™ (Magnetic Insight Inc.) for MPI signal and resolution. The active ingredient of VivoTrax™ is carboxydextran and is magnetically identical to ferucarbotran, which was adopted early for MPI and is one of the most commonly used tracers for MPI.
Using the Relax™ module equipped on the Momentum™ scanner (Magnetic Insight Inc.), measurements were collected in triplicate for (i) VivoTrax™, (ii) Synomag-D, (iii) Synomag-D with TAs, and (iv) Synomag-D + DC with TAs. For (i) and (ii), 1 µL of the agent was used. For (iii), 1.25% of the volumes used for the labelling procedure were prepared (1.125 µL Synomag-D, 62.5 µL serum free media, 0.0375 µL protamine sulfate, 0.0125 µL heparin). For (iv), 500 K (K = 1000) SPIO-labelled DC suspended in 40 µL PBS were collected in a PCR tube. Point spread functions (PSF) from MPI relaxometry were analysed for signal and resolution, normalized to iron content. To compare resolutions, full width half maximum (FWHM) values in milli-Tesla (mT) were converted to millimetres (mm) by dividing by the 3.0 T per meter (T/m) gradient strength. Ordinary one-way ANOVA tests were performed to determine differences in sensitivity and resolution for relaxometry measurements.
DC Detection with MPI
Triplicate sets of cell samples containing 250 K, 100 K, 50 K, 25 K, 12 K, and 6 K Synomag-D + DC labelled with (n = 18) and without (n = 18) TAs in 40 µL PBS were imaged with MPI. All images were acquired on a Momentum™ MPI scanner (Magnetic Insight Inc.). Projection images were acquired in 2D with a 3.0-T/m selection field gradient and drive field strengths of 20 mT and 23 mT in the X and Z axes, respectively. These 2D images took ~ 1.5 min to acquire for a 6 × 6-cm field of view (FOV). A simple linear regression was performed to determine the relationship between iron content and cell number.
DC Migration Model
To model DC migration, a 250K cell sample (high signal) was placed 2 cm from a 25K cell sample (low signal) to approximate the distance and signal strength difference between the footpad and the pLN. The 25K cell sample was then moved incrementally 1 cm away, up to 5 cm, from the 250K cell sample to determine the minimum distance at which two distinct signals could be resolved. Projection images were acquired in 2D with a 3.0-T/m selection field gradient and drive field strengths of 20 mT and 23 mT in the X and Z axes, respectively. These 2D images took ~ 2 min to acquire for a 12 × 6-cm FOV. The same image parameters were used for 3D imaging, which combines 35 projections (~ 30 min).
In Vivo MPI
In Vivo Modelling of DC Migration
25 K and 250 K Synomag-D + DC suspended in 40 µL PBS were subcutaneously injected into the upper back of anesthetized C57BL/6 mice (n = 4) at distances of 2 cm (n = 2) or 5 cm (n = 2) apart. MPI was performed immediately post injection using the same image parameters described above.
In Vivo DC Migration
300 K (n = 3) or 500 K (n = 3) Synomag-D + DC were intradermally injected into each hind footpad immediately following Tag-It Violet™ (Violet +) cell tracking dye incorporation (Biolegend, San Diego, USA). MPI was performed immediately following injection (day 0) and repeated 48 h later (day 2). Mice were fasted overnight for 12 h prior to imaging with only water, a laxative, and corn bedding. Following imaging, mice were returned to cages with food ad libitum. This was done to reduce gastrointestinal signal in MPI images, following approved animal protocols by the University of Western Ontario Animal Care and Use Subcommittee. On day 3, whole mouse body proton MRI was acquired for the two mice that had the highest detected pLN signal from each injection condition using a clinical 3 T MRI (Discovery MR750, General Electric) system and a 4.3 × 4.3cm surface coil (Clinical MR Solutions, Wisconsin) as described [22]. Left and right pLNs from all mice were excised and imaged ex vivo with MPI on day 3. pLNs were prepared and sectioned as previously described [17]. Qualitative imaging of pLNs was performed to identify Violet + Synomag-D + DC using an Olympus IX50 phase contrast inverted microscope (Richmond Hill, CA) and Infinity3-3URF camera (Lumenera, Ottawa, CA). Iron mass was quantified from measured MPI signal, as described below. Additionally, two cell pellet samples each containing 1000 K DC were imaged using 2D and 3D high sensitivity isotropic (3.0 T/m gradient) parameters to measure the amount of iron per cell and subsequently estimate cell number.
Mice were anesthetized initially with 2% isoflurane and maintained with 1% isoflurane during imaging. All in vivo images and ex vivo pLN images were acquired in 2D and 3D using a 3.0 T/m selection field gradient and drive field strengths of 20 mT and 23 mT in the X and Z axes, respectively, and a 12 × 6 cm FOV.
Quantification
Calibration lines (2D and 3D) for Synomag-D were made to determine the relationship between iron content and MPI signal using previously established methods [22]. To verify calibrations, manufacturer reported iron concentrations were compared to iron concentrations calculated using MPI, with no statistically significant difference. All MPI images were imported and viewed with a custom MPI colour look-up table (CLUT) using the open-source Horos™ image analysis software (version 3.3.6, Annapolis, MD USA). MPI signal was measured within a specific region of interest (ROI) using a semi-automatic segmentation tool for both 2D and 3D images. Total MPI signal for an ROI was calculated by multiplying the ROI area (2D) or volume (3D) by the mean signal. The signal-to-noise ratio (SNR) was calculated by dividing the mean signal for a ROI by the standard deviation of the background noise. The SNR had to be greater than 5 for the MPI signal to be considered detectable and for images to be further quantified. Iron content was calculated by dividing the total MPI signal by the slope of the calibration line. All MPI images (calibrations, in vitro DC pellets, and in vivo experiments) were delineated and analysed in the same way to ensure consistency. Iron content per cell was calculated by dividing the measured iron content from the signal ROI by the number of cells in the cell pellet sample (1000 K). Cell numbers for the true migration experiment were then estimated by dividing the iron quantified within a given ROI by the respective iron/cell measurements.
Results
In Vitro MPI
MPI Relaxometry
MPI relaxometry compared the performance of Synomag-D and VivoTrax™ and measured differences in signal after Synomag-D internalization by DC. Relative sensitivities (Fig. 1a) and resolutions (Fig. 1b) are shown by the point spread functions (PSF) for free SPIOs, Synomag-D mixed with TAs, and Synomag-D-labelled DC. Synomag-D had approximately 3.46 times higher sensitivity and improved resolution (2.49 mm versus 3.29 mm) compared to VivoTrax™ (Fig. 1c). This agrees with Vogel et al. [23]. Sensitivity was reduced when Synomag-D was mixed with heparin and protamine sulfate and was further reduced when internalized into DC (Fig. 1d). There was no statistically significant difference in resolution between Synomag-D and Synomag-D with TAs; however, the resolution was significantly lower for intracellular Synomag-D (Fig. 1e). The reduced MPI signal which occurs either upon combination with TAs and/or internalization has been shown previously, resulting from aggregation and slower MPI relaxation [24].
Comparing MPI sensitivity and resolution of VivoTrax™, Synomag-D, Synomag-D with transfection agents (TAs), and intracellular Synomag-D. a Relative sensitivities are compared by signal peak intensities, normalized to the iron content of the sample. b Relative resolutions are compared by full width half maximum (FWHM) values with MPI signal normalized to the iron content and signal peak intensities of the sample. c Numerical values for the graphs (a) and (b) are shown. There were significant differences in both sensitivity and resolution with intracellular Synomag-D. d There was a significant difference in sensitivity and e a non-significant difference in resolution between Synomag-D and Synomag-D with TAs. Data shown as mean ± SD (ns — no statistical significance (p > 0.05), *p ≤ 0.05, ***p ≤ 0.001, ****p ≤ 0.0001, ordinary one-way ANOVA).
DC Detection with MPI
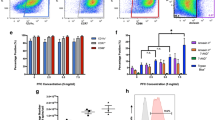
PPB staining showed the presence of iron in DC labelled with and without TAs (Fig. 2a, b). TAs improved DC labelling with a higher amount of iron/cell (5.5 pg Fe/cell vs 1.4 pg Fe/cells) and more iron positive cells (76.6% vs 51.6%). Cell viability was not affected by cell labelling (Fig. 2c). Both labelling strategies show strong linear correlations (R2 = 0.99) between measured iron content and cell number (Fig. 2d). DC labelled with TAs had a higher cellular sensitivity as indicated by a steeper slope (5.73 × 10−6 vs 1.24 × 10−6). MPI of Synomag-D + DC samples without TAs was limited to a detection of 25 K cells due to lower labelling efficiency (Fig. 2e). With more iron/cell from enhanced labelling with TAs, 6 K cells are detected with MPI, the lowest number tested (Fig. 2f). The addition of TAs enhanced cell labelling which was confirmed by PPB stains, MPI, and measured iron content. This labelling strategy was used for in vivo experiments.
DC labelling and MPI cellular detection with Synomag-D. PPB staining shows the presence of iron (Synomag-D) in DC labeled a with and b without TAs. The addition of TAs had a higher labelling efficiency (5.5 pg/cell vs 1.4 pg/cell) with more Synomag-D + cells (76.6% vs 51.8%). c CD11c + DC viability is unaffected by Synomag-D labelling or the use of TAs. Flow cytometry was used to identify singlet, CD45 + CD11c + DC (gating not shown) and subsequently assessed for viability using Zombie NIR™ fixable viability dye. CD11c + DC viability was unchanged (grey bar) following Synomag-D labelling (200 μg/ml) in the presence of TAs (protamine sulfate and heparin) when compared to unlabeled CD11c + DC from the same culture (white bar). Data shown as mean ± SD (n = 4, ns — no statistical significance, p > 0.05, paired t-test). d DC labelled with TAs have a higher cellular sensitivity than without TAs, indicated by a steeper slope (5.729 × 10-6 vs 1.237 × 10-6). Iron content was quantified from MPI images of cells. e Detection was limited to 25 K cells without using TAs, below which signal was not detected, indicated by the red X. f With TAs, 6 K cells are detected, the lowest number tested in this experiment.
In Vitro Modelling of DC Migration
Using 2D imaging, two samples of 25 K cells (Fig. 3a) and 250 K cells (Fig. 3b) placed 2 cm apart were detected with two distinct peaks shown in the adjacent corresponding signal intensity profile. When the 25K cell phantom is imaged with the 250K cell phantom, creating unequal signals, the signal from 25 K cells (less iron) was not detected next to the higher signal from 250 K cells. The corresponding signal intensity profile shows the 25K cell sample as a small shoulder off the tail of the higher peak from the 250K cell phantom placed 2 cm away (Fig. 3c). To fully separate the unequal signals into two distinct peaks, a minimum distance of 5 cm was required for 2D imaging (Fig. 3d). Windowing the image to the minimum and maximum signal from the 25K cell phantom oversaturated MPI signal from the 250K cell phantom, expanding into the region of the lower signal, and prevented detection of the 25K cell phantom. Distinct signals were detected using 3D imaging at 2 cm; however, window levelling to the 25K cell phantom signal still oversaturated signal from the 250K cell phantom (Fig. 3e).
Imaging two proximal sources of unequal MPI signal. Two equal samples of either a 25 K or b 250 K Synomag-D + DC were detected and can be resolved at 2 cm separation (images shown in full dynamic range). c Signal from 25 K cells was partially concealed when placed 2 cm from 250 K cells. d At a distance of 5 cm, both signals were fully resolved with 2D imaging. Window levels for c and d were set to the minimum and maximum signal of the 25 K cell sample. Corresponding signal intensity profiles are shown adjacent to images. e With 3D imaging (as in the samples and position in c), both signals are resolved at 2 cm. Data is displayed in landscape mode for 2D images and portrait mode for 3D.
In Vivo MPI
In Vivo Modelling of DC Migration
The in vitro DC migration model was repeated to compare signal detection and resolution between two unequal signal sources in an in vivo mouse model. Subcutaneous injections of 25 K and 250 K cells 2 cm apart (Fig. 4a) could not be resolved with 2D imaging (Fig. 4b) but were resolved with 3D imaging (Fig. 4c). To clearly detect the 25K cell injection site, the contrast and intensity of the images had to be adjusted (Fig. 4d). When imaging mice with cell injections 5 cm apart (Fig. 4e), a gastrointestinal (GI) signal was detected in all mice due to iron contained within mouse feed and ingested cage bedding as shown in the Suppl. Figure 1 (See ESM). Three distinct signal regions were detected with both 2D and 3D imaging (1) 250 K cells, (2) GI, and (3) 25 K cells. Signal from the lower source (25 K) was partially concealed within signal from the GI using 2D imaging (Fig. 4f). With 3D imaging, the signal from 25 K is 10 times weaker than the 250K signal and not perceptible at the full dynamic range (Fig. 4g). The dynamic range was lowered to show the signal detected from the 25K sample (Fig. 4h).
In vivo cellular sensitivity and resolution after 25 K and 250 K Synomag-D + DC were injected subcutaneously into the upper back of anesthetized C57BL/6 mice (n = 4) separated by either 2 cm (n = 2) or 5 cm (n = 2). a At a distance of 2 cm, in 2D images, the signal from 25 K cells was obscured by the signal from 250 K cells, b with unresolved peaks in the corresponding signal intensity profile. c With 3D imaging the two regions of signal were resolved after (d) window levelling to the lower signal. e When separated by 5 cm, the signal from the 25K cells was partially obscured by f stronger signal from the GI tract, shown in the corresponding signal intensity profile. g With 3D imaging, signal from 25 K cells is resolved after h window levelling to the 25K cell sample. Data is displayed in landscape mode for 2D images and portrait mode for 3D.
In Vivo DC Migration
Mice were injected with either 300 K (n = 3) or 500 K (n = 3) Violet + Synomag-D + DC. In this experiment, mice were prepared with bowel prep to remove GI signals that are similar in magnitude to the pLN signals. Table 1 summarizes the data from all 6 mice, showing cell number and iron estimates for the footpads and pLNs on day 0, day 2, and ex vivo. Footpad signal (left and right) was detected and quantified on both day 0 and day 2 imaging. On day 2, MPI signal was detected in the left pLN in four of the six mice and in the right pLN in 2 of the 6 mice. The number of DC injected did not affect the ability to detect MPI signal in the pLNs; for each cell number, two of the three mice had signal in the left pLN, and one of the three mice had signal in the right pLN. Ex vivo imaging of the pLNs improved the dynamic range of MPI signal; signal was detected in 11/12 nodes with 3D imaging.
Figure 5 shows images from one representative mouse (mouse 4) that underwent both MPI and MRI following 300 K cells injected into the hind footpads. Day 0 MPI detected signal in the left and right hind footpads, as expected (Fig. 5a). Day 2 MPI detected signal in the left and right pLNs, with quantification (see Table 1) suggesting a 3% and 2% migration rate, respectively (Fig. 5b). Day 3 MRI indirectly detected regions of signal loss in both the left and right pLNs (Fig. 5c). This was also true for the second mouse imaged with both MPI and MRI (mouse 3, not shown). The signal void volumes were 0.31 mm3 (left) and 0.35 mm3 (right) for mouse 3 and 0.43 mm3 (left) and 0.32 mm3 (right) for mouse 4. Ex vivo MPI of the left and right pLNs detected signal in both pLNs (Fig. 5d). After imaging with MPI and MRI, left and right pLNs were removed from all mice and sectioned for fluorescence microscopy. The presence of migration-competent Violet + DC is shown in both left (Fig. 6a) and right (Fig. 6b) pLN sections, confirming that the source of signal from MPI and MRI was from originally injected Synomag-D + DC.
DC migration experiment showing in vivo and ex vivo MPI of C57BL/6 mouse DC migration following injection of 300 K Synomag-D + DC into the hind footpads. All images are slices selected from 3D high sensitivity isotropic (3.0 T/m gradient) images. MPI signal values are indicated by the scale bars. a Day 0 imaging detected two regions of MPI signal from the hind footpads (left — orange bracket, right — orange dashed bracket). b Day 2 imaging detected signal in both pLNs (left — blue bracket, right — blue dashed bracket). Oversaturation of footpad signal is apparent with extreme window levelling to the lower signal intensities of the pLNs. c On day 3, coronal MRI indirectly detected signal in the pLNs, shown by signal voids indicated by the white arrows (left — white box, right — white dashed box). d Ex vivo MPI of excised pLNs detected signal in the left and right pLNs, also confirming DC migration. Iron quantification and cell number estimates are shown below images.
Discussion
This study demonstrates the use of MPI for detecting the migration of DC to lymph nodes; a critical step required for DC cancer vaccines to generate effective and long-lasting immune responses. The success of DC immunotherapy is strongly related to the number of administered DC accumulating in the nodes [25, 26]. However, the best route and timing for DC administration and the ideal strategies for maturation and activation of DC to maximize migration to lymphoid organs are not clear. In vivo imaging can be used to evaluate migration efficacy.
The main advantage of MPI for DC tracking is the ability to directly quantify iron from acquired images. Quantification of the MPI signal is directly related to the amount of iron present in the ROI. This can subsequently be used, alongside a measurement of the amount of iron/cell, to estimate cell number. In this study, MPI allowed for the estimation of the number of migrated DC in lymph nodes. To the best of our knowledge, MPI has not yet been applied to imaging of DC. In our in vivo migration experiment (Table 1), six mice were injected with either 300 K (n = 3) or 500 K (n = 3) Synomag-D + DC. In vivo pLN signal was detected, and cell number was estimated in half of the target nodes. There was variability in the iron content in the nodes and the migrated cell numbers estimated from the MPI values. This is expected given the biological variability in the mouse lymphatic system and in migration and injection efficiency. Ex vivo MPI was used to verify that the signal was due to iron in the pLNs. In 4/6 cases, the MPI signal was reduced ex vivo. This reduction in signal is likely due to sampling error. It is difficult to dissect the popliteal lymph nodes, and, in some cases, tissue may have been left behind. It is also possible that some of the in vivo signal results from iron labelled DC that are in the fat pad; this would not be removed. For the other two mice where the MPI signal in nodes was compared in vivo and ex vivo, the values were higher ex vivo. Since in vivo imaging was done on day 2 post-injection and ex vivo imaging was done with pLNs removed on day 3, there is a possibility that in some cases, migration continued after day 2 increasing the presence of iron labelled DC on day 3.
MPI signal was not detected in the pLNs of all mice. This may be explained by cell migration being below our detection threshold. Only 3–5% of injected DC migrate to a target lymph node post-administration, and this may be lower since varying numbers of DC are known to remain at the injection site, lose viability, and be cleared by infiltrating macrophages within 2 days [27]. The inability to separate and resolve pLN signal from the larger footpad signal likely also contributes to our detection limits. Overall, this investigation provides the proof-of-principle that MPI can be used as a modality to track and potentially quantify DC migration.
Iron-based MRI cell tracking has been the most widely used imaging modality for pre-clinical studies of DC migration [16, 19, 28, 29], explored in reviews by Dekaban et al. and Bulte et al. [30, 31]. Experimental studies in mice have demonstrated that SPIO-labelled DC can be detected in MRI as regions of signal loss in T2- or T2*-weighted images after their intradermal or intranodal administration. These studies used signal void volume, number of black pixels, signal intensity, or fractional signal loss as a measure to relate to the number of cells injected, histological analysis of DC in nodes, the iron content in lymph nodes measured ex vivo, or tumour size [16, 19, 20, 29, 32,33,34,35,36]. For example, De Chickera et al. demonstrated that MRI could detect differences in the migratory abilities of two different DC preparations, ex vivo untreated, resting DC versus ex vivo matured DC [19]. Despite the evidence that pre-clinical MRI of SPIO-labelled DC can be used to assess the efficacy of DC immunotherapy, it is challenging to absolutely quantify iron content and cell number. Quantifying signal loss is not straightforward because the relationship between signal loss and iron concentration is not linear and because the presence of iron produces a localized disruption to the magnetic field homogeneity, resulting in a region of signal loss that extends far beyond the actual boundary of the cells.
The main challenge for quantification of our MPI data was the isolation of neighbouring strong and weak MPI signals as displayed in Fig. 3. Detection of pLN signal was challenging in vivo where the larger footpad signal from remaining cells partially obscured the weaker signal from pLNs, preventing quantification in some mice. For this type of study, a wide dynamic range of signal is expected with high iron concentrations at the injection sites and low iron concentrations at migratory sites. The lower gradient (3.0 T/m) was chosen for this study to give the sensitivity needed to detect these low quantities of iron in the pLNs, at the cost of resolution. These varying signal intensities contained in a small FOV result in the lower signal being concealed by the higher signal in additional experiments as shown in the Suppl. Figure 2 and Suppl. Figure 3 (see ESM). Resolving these varying signal intensities was improved with 3D imaging, which provides higher SNR and higher resolution compared to 2D imaging.
GI signal was another challenge as a source of high signal, although we were able to mitigate GI signal by overnight fasting of animals with laxatives. Surprisingly, iron in mouse feed showed significant MPI signal and after ingestion, translating to gastrointestinal (GI) signal observed in in vivo images. We have since investigated methods to mitigate unwanted GI signal including fasting, choice of bedding, and using laxatives. Without taking steps to mitigate food-related iron in the GI tract, detecting MPI signal in the surrounding area will be challenging, for example detecting MPI signal in the mesenteric lymph nodes.
Several studies have reported on the issue of resolving a wide range of differing iron concentrations [37,38,39]. Most recently, Boberg et al. discussed the idealized dynamic range of single sample and single iron concentration scenarios versus the effective dynamic range when there are multiple samples of different iron concentrations within the FOV [15]. They found that the presence of high particle concentrations within the FOV can significantly reduce the dynamic range that the system is able to resolve. Herz et al. discuss the issue as a function of the limited dynamic range of the analog to digital converter (ADC), with additional effects from reconstruction, discretization of the PSF, and the ADC not being used in the entire dynamic range [39]. It should be noted that although the MPI systems differ across these studies, including the present one, the issue of resolving proximal, varying iron concentrations persists and solutions are being investigated. In our study, increasing the gradient strength could improve resolution of signals; however, this comes at the cost of sensitivity and longer imaging times [40, 41]. Changing the FOV to exclude the high signal source is another potential solution. This is challenging in cases where the exact location of signal is unknown, since image artefacts are caused by iron contained on or near the edge of the FOV, resulting in a large negative signal region [42, 43].
MPI does have some other limitations for tracking cells in vivo. Compared to iron-based MRI cell tracking, MPI has much lower resolution. This can make it challenging to pinpoint where the signal is located, especially without prior knowledge. It is expected that the ongoing development of MPI-tailored SPIOs will substantially improve image resolution [23, 41, 44,45,46,47,48]. In addition, quantification of cell number can only be determined in experiments where SPIO is used to pre-label cells prior to their administration and the mass of iron per cell is known.
Conclusions
MPI enables the sensitive detection of DCs migrating to the pLNs with better specificity than MRI. Future work will be performed to study and validate the quantification of DC. Future developments to refine MPI for cell tracking applications include increasing sensitivity without compromising resolution, improving the dynamic range, or creating unique reconstruction algorithms. These can all be aided by the development of new nanoparticles. Ideally, non-invasive imaging will be used as a biomarker for predicting the efficacy of DC immunotherapy. Quantification of the number of DC in lymph nodes by MPI could serve as an in vivo migration assay which could be used to determine whether more, or less, DC reach the target lymph nodes under different test conditions. These include but are not limited to injection route, site of injection, timing of injection, maturation cocktails, cell injection number and antigen loading. This new approach to measuring the migration and accumulation of DC will provide valuable insight into the further development of cell-based cancer vaccines.
References
Met Ö, Jensen KM, Chamberlain CA et al (2019) Principles of adoptive T cell therapy in cancer. Semin Immunopathol 41:49–58. https://doi.org/10.1007/s00281-018-0703-z
Munir H, McGettrick HM (2015) Mesenchymal stem cell therapy for autoimmune disease: risks and rewards. Stem Cells Dev 24:2091–2100. https://doi.org/10.1089/scd.2015.0008
Chang Y-H, Wu K-C, Harn H-J et al (2018) Exosomes and stem cells in degenerative disease diagnosis and therapy. Cell Transplant 27:349–363. https://doi.org/10.1177/0963689717723636
Sun JM, Kurtzberg J (2018) Cell therapy for diverse central nervous system disorders: inherited metabolic diseases and autism. Pediatr Res 83:364–371. https://doi.org/10.1038/pr.2017.254
Hamdan H, Hashmi SK, Lazarus H et al (2021) Promising role for mesenchymal stromal cells in coronavirus infectious disease-19 (COVID-19)-related severe acute respiratory syndrome? Blood Rev 46:100742. https://doi.org/10.1016/j.blre.2020.100742
Gilboa E, Nair SK, Lyerly HK (1998) Immunotherapy of cancer with dendritic-cell-based vaccines. Cancer Immunol Immunother 46:82–87. https://doi.org/10.1007/s002620050465
Förster R, Braun A, Worbs T (2012) Lymph node homing of T cells and dendritic cells via afferent lymphatics. Trends Immunol 33:271–280. https://doi.org/10.1016/j.it.2012.02.007
Martin-Fontecha A, Sebastiani S, Höpken UE et al (2003) Regulation of dendritic cell migration to the draining lymph node: impact on T lymphocyte traffic and priming. J Exp Med 198:615–621. https://doi.org/10.1084/jem.20030448
Wang B, Sun C, Wang S et al (2018) Mouse dendritic cell migration in abdominal lymph nodes by intraperitoneal administration. Am J Transl Res 10:2859–2867
Liu W, Frank JA (2009) Detection and quantification of magnetically labeled cells by cellular MRI. Eur J Radiol 70:258–264. https://doi.org/10.1016/j.ejrad.2008.09.021
Zheng B, Yu E, Orendorff R et al (2017) Seeing SPIOs directly in vivo with magnetic particle imaging. Mol Imaging Biol 19:385–390. https://doi.org/10.1007/s11307-017-1081-y
Bulte JWM (2019) Superparamagnetic iron oxides as MPI tracers: a primer and review of early applications. Adv Drug Deliv Rev 138:293–301. https://doi.org/10.1016/j.addr.2018.12.007
Song G, Chen M, Zhang Y et al (2018) Janus iron oxides @ semiconducting polymer nanoparticle tracer for cell tracking by magnetic particle imaging. Nano Lett 18:182–189. https://doi.org/10.1021/acs.nanolett.7b03829
Makela AV, Gaudet JM, Schott MA et al (2020) Magnetic particle imaging of macrophages associated with cancer: filling the voids left by iron-based magnetic resonance imaging. Mol Imaging Biol. https://doi.org/10.1007/s11307-020-01473-0
Boberg M, Gdaniec N, Szwargulski P et al (2021) Simultaneous imaging of widely differing particle concentrations in MPI: problem statement and algorithmic proposal for improvement. Phys Med Biol 66:095004. https://doi.org/10.1088/1361-6560/abf202
Dekaban GA, Snir J, Shrum B et al (2009) Semiquantitation of mouse dendritic cell migration in vivo using cellular MRI. J Immunother 32:240–251. https://doi.org/10.1097/CJI.0b013e318197b2a0
Fink C, Smith M, Gaudet JM et al (2020) Fluorine-19 cellular MRI detection of in vivo dendritic cell migration and subsequent induction of tumor antigen-specific immunotherapeutic response. Mol Imaging Biol 22:549–561. https://doi.org/10.1007/s11307-019-01393-8
Inaba K, Inaba M, Romani N et al (1992) Generation of large numbers of dendritic cells from mouse bone marrow cultures supplemented with granulocyte/macrophage colony-stimulating factor. J Exp Med 176:1693–1702
de Chickera S, Willert C, Mallet C et al (2012) Cellular MRI as a suitable, sensitive non-invasive modality for correlating in vivo migratory efficiencies of different dendritic cell populations with subsequent immunological outcomes. Int Immunol 24:29–41. https://doi.org/10.1093/intimm/dxr095
de Chickera SN, Snir J, Willert C et al (2011) Labelling dendritic cells with SPIO has implications for their subsequent in vivo migration as assessed with cellular MRI. Contrast Media Mol Imaging 6:314–327. https://doi.org/10.1002/cmmi.433
Fink C, Gaudet JM, Fox MS et al (2018) 19F-perfluorocarbon-labeled human peripheral blood mononuclear cells can be detected in vivo using clinical MRI parameters in a therapeutic cell setting. Sci Rep 8:590. https://doi.org/10.1038/s41598-017-19031-0
Sehl O, Makela A, Hamilton A, Foster P (2019) Trimodal cell tracking in vivo: combining iron- and fluorine-based magnetic resonance imaging with magnetic particle imaging to monitor the delivery of mesenchymal stem cells and the ensuing inflammation. Tomography 5:367–376. https://doi.org/10.18383/j.tom.2019.00020
Vogel P, Kampf T, Rückert M et al (2021) Synomag®: the new high-performance tracer for magnetic particle imaging. Int J Magn Part Imaging 7. https://doi.org/10.18416/IJMPI.2021.2103003
Suzuka H, Mimura A, Inaoka Y, Murase K (2019) Magnetic nanoparticles in macrophages and cancer cells exhibit different signal behavior on magnetic particle imaging. J Nanosci Nanotechnol 19:6857–6865. https://doi.org/10.1166/jnn.2019.16619
Fong L, Brockstedt D, Benike C et al (2001) Dendritic cells injected via different routes induce immunity in cancer patients. J Immunol 166:4254–4259. https://doi.org/10.4049/jimmunol.166.6.4254
Eggert AAO, Schreurs MWJ, Boerman OC et al (1999) Biodistribution and vaccine efficiency of murine dendritic cells are dependent on the route of administration. Cancer Res 59:3340–3345
Verdijk P, Aarntzen EHJG, Lesterhuis WJ et al (2009) Limited amounts of dendritic cells migrate into the T-cell area of lymph nodes but have high immune activating potential in melanoma patients. Clin Cancer Res 15:2531–2540. https://doi.org/10.1158/1078-0432.CCR-08-2729
Baumjohann D, Hess A, Budinsky L et al (2006) In vivo magnetic resonance imaging of dendritic cell migration into the draining lymph nodes of mice. Eur J Immunol 36:2544–2555. https://doi.org/10.1002/eji.200535742
Rohani R, de Chickera SN, Willert C et al (2011) In vivo cellular MRI of dendritic cell migration using micrometer-sized iron oxide (MPIO) particles. Mol Imaging Biol 13:679–694. https://doi.org/10.1007/s11307-010-0403-0
Dekaban GA, Hamilton AM, Fink CA et al (2013) Tracking and evaluation of dendritic cell migration by cellular magnetic resonance imaging. WIREs Nanomed Nanobiotechnol 5:469–483. https://doi.org/10.1002/wnan.1227
Ahrens ET, Bulte JWM (2013) Tracking immune cells in vivo using magnetic resonance imaging. Nat Rev Immunol 13:755–763. https://doi.org/10.1038/nri3531
Zhang X, de Chickera SN, Willert C et al (2011) Cellular magnetic resonance imaging of monocyte-derived dendritic cell migration from healthy donors and cancer patients as assessed in a scid mouse model. Cytotherapy 13:1234–1248. https://doi.org/10.3109/14653249.2011.605349
Ferguson PM, Slocombe A, Tilley RD, Hermans IF (2013) Using magnetic resonance imaging to evaluate dendritic cell-based vaccination. PLoS ONE 8:e65318. https://doi.org/10.1371/journal.pone.0065318
Townson JL, Ramadan SS, Simedrea C et al (2009) Three-dimensional imaging and quantification of both solitary cells and metastases in whole mouse liver by magnetic resonance imaging. Cancer Res 69:8326–8331. https://doi.org/10.1158/0008-5472.CAN-09-1496
Long CM, van Laarhoven HWM, Bulte JWM, Levitsky HI (2009) Magnetovaccination as a novel method to assess and quantify dendritic cell tumor antigen capture and delivery to lymph nodes. Cancer Res 69:3180–3187. https://doi.org/10.1158/0008-5472.CAN-08-3691
Mills PH, Hitchens TK, Foley LM et al (2012) Automated detection and characterization of SPIO-labeled cells and capsules using magnetic field perturbations. Magn Reson Med 67:278–289. https://doi.org/10.1002/mrm.22998
Franke J, Heinen U, Lehr H et al (2016) System characterization of a highly integrated preclinical hybrid MPI-MRI scanner. IEEE Trans Med Imaging 35:1993–2004. https://doi.org/10.1109/TMI.2016.2542041
Graeser M, Knopp T, Sattel TF et al (2012) Signal separation in magnetic particle imaging. IEEE Nuclear Science Symposium and Medical Imaging Conference Record (NSS/MIC) 2483–2485. https://doi.org/10.1109/NSSMIC.2012.6551566
Herz S, Vogel P, Kampf T et al (2017) Selective signal suppression in traveling wave MPI: focusing on areas with low concentration of magnetic particles. Int J Magn Part Imaging 3(2):1709001. https://doi.org/10.18416/ijmpi.2017.1709001
Croft LR, Goodwill PW, Konkle JJ et al (2016) Low drive field amplitude for improved image resolution in magnetic particle imaging. Med Phys 43:424–435. https://doi.org/10.1118/1.4938097
Tay ZW, Hensley DW, Vreeland EC et al (2017) The relaxation wall: experimental limits to improving MPI spatial resolution by increasing nanoparticle core size. Biomed Phys Eng Express 3:035003–035003. https://doi.org/10.1088/2057-1976/aa6ab6
Weizenecker J, Gleich B, Rahmer J et al (2009) Three-dimensional real-time in vivo magnetic particle imaging. Phys Med Biol 54:L1–L10. https://doi.org/10.1088/0031-9155/54/5/L01
Weber A, Werner F, Weizenecker J et al (2016) Artifact free reconstruction with the system matrix approach by overscanning the field-free-point trajectory in magnetic particle imaging. Phys Med Biol 61:475. https://doi.org/10.1088/0031-9155/61/2/475
Bulte JWM (2018) Superparamagnetic iron oxides as MPI tracers: a primer and review of early applications. Adv Drug Deliv Rev. https://doi.org/10.1016/J.ADDR.2018.12.007
Du Y, Lai PT, Leung CH, Pong PWT (2013) Design of superparamagnetic nanoparticles for magnetic particle imaging (MPI). Int J Mol Sci 14:18682–18710. https://doi.org/10.3390/ijms140918682
Ferguson RM, Khandhar AP, Kemp SJ et al (2015) Magnetic particle imaging with tailored iron oxide nanoparticle Tracers. IEEE Trans Med Imaging 34:1077–1084. https://doi.org/10.1109/TMI.2014.2375065
Khandhar AP, Ferguson RM, Arami H et al (2015) Tuning surface coatings of optimized magnetite nanoparticle tracers for in vivo magnetic particle imaging. IEEE Trans Magn 51:1–4. https://doi.org/10.1109/TMAG.2014.2321096
Khandhar AP, Ferguson RM, Arami H, Krishnan KM (2013) Monodisperse magnetite nanoparticle tracers for in vivo magnetic particle imaging. Biomaterials 34:3837–3845. https://doi.org/10.1016/j.biomaterials.2013.01.087
Funding
This work was supported by the Canadian Institutes of Health Research (CIHR) grant no. OPG 363209 and the National Sciences and Engineering Research Council (NSERC) of Canada, the Molecular Imaging Graduate Program (Western University), Translational Breast Cancer Research Unit, Ontario Graduate Scholarship, and NSERC post-graduate scholarship.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Gevaert, J.J., Fink, C., Dikeakos, J.D. et al. Magnetic Particle Imaging Is a Sensitive In Vivo Imaging Modality for the Detection of Dendritic Cell Migration. Mol Imaging Biol 24, 886–897 (2022). https://doi.org/10.1007/s11307-022-01738-w
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11307-022-01738-w