Abstract
Background
Oral cancer is one of the most frequently occurring cancers. Metabolic reprogramming is an important hallmark of cancer. Metabolomics characterizes all the small molecules in a biological sample, and a complete set of small molecules in such sample is referred as metabolome. Nuclear magnetic resonance spectroscopy and mass spectrometry are two widely used techniques in metabolomics studies. Increasing evidence demonstrates that metabolomics techniques can be used to explore the metabolic signatures in oral cancer. Elucidation of metabolic alterations in oral cancer is also important for the understanding of its pathological mechanisms.
Aim of review
In this paper, we summarize the latest progress of metabolomics study in oral cancer and provide the suggestions for the future studies.
Key scientific concepts of review
The metabolomics studies in saliva, serum, and tumor tissues revealed the existence of metabolic signatures in bio-fluids and tissues of oral cancer, and several tumor-specific metabolites identified in individual study could discriminate oral cancer from healthy controls or precancerous lesions, which are potential biomarkers for the screening or early diagnosis of oral cancer. Metabolomics study of oral cancers in the future should aim to establish a routine procedure with high sensitivity, profile intracellular metabolites to find out the metabolic characteristics of tumor cells, and investigate the mechanism behind metabolomic alterations and the metabolic response of cancer cells to chemotherapy.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Oral cancer, a type of head and neck cancer, is the sixth most common cancer in the world, and over 90% of them are oral squamous cell carcinoma (OSCC) (Rai et al. 2018). Several oral lesions, such as lichen planus, leukoplakia, erythroplakia, and oral sub-mucous fibrosis are considered as oral potentially malignant disorders (Chen and Zhao 2017). The pathological mechanisms of oral cancer are not well understood, and thus the early diagnosis is critical for the improvement of patient’s survival rates. Metabolic reprogramming has been demonstrated to be an important hallmark of cancer (Yu et al. 2017). Cancer cells are more likely to use glycolysis, even when sufficient oxygen is available. This phenomenon is known as aerobic glycolysis or the Warburg effect (Warburg 1956; Yu et al. 2017), which promotes tumorigenesis and cancer progression. In addition to the Warburg effect, cancer undergoes complex metabolic changes, including metabolic reprogramming of lipids and amino acids, such as glutaminolysis (Yang et al. 2017; Nakagawa et al. 2018; Sun et al. 2018). Whether oral cancer has special metabolic characteristics has not been well studied. Elucidation of metabolic alterations in oral cancer is of great importance for identifying novel biomarkers and understanding the development and progression of oral cancer.
Metabolomics aims to characterize all the small molecules in a sample, and a complete set of small molecule chemicals found within a such biological sample is referred as metabolome (Nicholson and Lindon 2008). Nuclear magnetic resonance (NMR) spectroscopy and mass spectrometry (MS) are two common analytical techniques used for metabolomics (Markley et al. 2017; Rai et al. 2018). Nuclear magnetic resonance detects hydrogen atoms in metabolites of a biological sample, and all hydrogen-containing molecules will produce an 1H NMR spectrum. The preparation of NMR samples is simple, and some bio-fluids, such as serum, may require no preparation (Bag et al. 2015; Gupta et al. 2015). Plasma containing proteins and lipids that interfere with NMR spectral quality can be treated with methanol to remove lipoproteins (Markley et al. 2017). Nuclear magnetic resonance provides excellent analytical precision in identification of cancer metabolomic signature. Mass spectrometry is an analytical technique that ionizes chemical species and sorts the ions based on their mass to charge ratio, producing a mass spectrum. Mass spectrometry is usually coupled with chromatography and other separation techniques to improve its mass-resolving and mass-determining capabilities (Ye et al. 2014; Ohshima et al. 2017). Increasing evidence has demonstrated that metabolomics techniques can be used to explore the metabolic signatures in bio-fluids and tissues of oral cancer (Table 1) (Shin et al. 2016; Rai et al. 2018). In the current paper, PubMed search engine and various combinations of keywords such as oral, cancer (tumor), metabolomics (metabolomic), and HNSCC were used for literature retrieval. A total of 78 articles were obtained, of which 22 were used for preparation of the review after omitting reviews and repetitive articles. We summarize the latest progress of metabolomics study in oral cancer and propose the suggestions for the future studies.
2 Salivary metabolome of oral cancer
Saliva is an informative biological fluid and saliva-based diagnostics has been widely used in clinical practice (Yakob et al. 2014). Metabolomics of saliva, the measurement of all metabolites in saliva, has become a new powerful tool to characterize the association between salivary metabolites and the diseases, particularly cancer (Shankar et al. 2014; Mikkonen et al. 2016). When saliva collected from oral cancers and other cancers was analyzed using capillary electrophoresis and time-of-flight mass spectrometry (CE-TOF-MS), the 57 principal metabolites were identified to be associated with these cancers (Sugimoto et al. 2010), suggesting that cancer-specific signatures are embedded in saliva metabolites. Twenty-eight differential metabolites, such as pyrroline hydroxycarboxylic acid, choline, and amino acids, were revealed between oral cancers and healthy controls (Sugimoto et al. 2010). Compared to controls the significantly higher level of salivary polyamines was detected in patients with oral cancer, and taurine and piperidine were identified to be oral cancer-specific metabolites that provide promising markers for oral cancer screening (Sugimoto et al. 2010). Up-regulation of polyamine has been found to increase cell proliferation, decrease apoptosis, and promote tumor invasion and metastasis (Gerner and Meyskens 2004). For example, putrescine was used to monitor the effect of chemotherapy on oral cancer (Okamura et al. 2007). In another study the metabolome of saliva from OSCC patients was analyzed, and a total of 25 discriminant metabolites, including seven metabolites detected in the previous study (Sugimoto et al. 2010), were identified between OSCC and healthy controls (Table 1) (Ohshima et al. 2017).
2.1 Metabolic stratification of oral epithelial lesions
Saliva metabolomics was applied for the early diagnosis and monitoring of OSCC (Wang et al. 2014b, c). Metabolomics analysis of saliva revealed a higher level of choline, betaine, and pipecolinic acid, and a lower level of l-carnitine in OSCC (Wang et al. 2014c). Concentrations of these metabolites can distinguish OSCC I-II from healthy controls, suggesting that salivary biomarkers are valuable for OSCC early diagnosis (Wang et al. 2014c). The researchers in the same group further modified the experimental procedure to overcome the limitations of a single chromatographic method due to different polarity of metabolites and identified 14 potential discriminant metabolites in the saliva of OSCC patients (Table 1) (Wang et al. 2014b). Combination of 5 salivary metabolites, propionylcholine, N-acetyl-l-phenylalanine, sphinganine, phytosphingosine, and S-carboxymethyl-l-cysteine can precisely distinguish early stage of OSCC from the controls (Wang et al. 2014b). The results are exciting, but more studies are needed to validate them.
Salivary metabolomics was explored as a diagnostic and stratification tool for oral cancer. In an early study, saliva from OSCC and precancerous lesions oral lichen planus (OLP) and oral leukoplakia (OLK) was analyzed with HPLC-MS. The diagnostic models established based on metabolic profiles could precisely diagnose and discriminate OSCC, OLP, and OLK (Yan et al. 2008). Salivary metabolites in OSCC, OLK and controls were profiled in another similar study, and each group had characteristic salivary metabolic signatures (Wei et al. 2011). A total of 41, 61, and 27 discriminant metabolites were identified between OSCC and control, OSCC and OLK, and OLK and control, respectively. The five most discriminant metabolites are γ-aminobutyric acid, phenylalanine, valine, n-eicosanoic acid, and lactate (Wei et al. 2011). The potential of these metabolites in detection of OSCC was evaluated, and a combination of valine, lactate, and phenylalanine was found to be a powerful predictive marker in distinguishing OSCC from the controls or OLK, suggesting a supplementary role of salivary metabolome diagnostics in the clinical detection of OSCC and stratification of the different oral epithelial lesions (Wei et al. 2011).
In order to explore the application of salivary metabolite biomarkers in oral cancer screening, the hydrophilic metabolites in saliva and tumor tissues of patients with oral cancer were analyzed by CE-TOF-MS (Ishikawa et al. 2016). In total, 85 metabolites in tumor and 45 metabolites in saliva were identified to be significantly different between oral cancer and controls, and 17 metabolites among them shared a similar tendency in both saliva and tissue. Of 17 metabolites, the combination of S-adenosylmethionine and pipecolate can discriminate oral cancers from controls. Results suggest that integrating metabolomics of saliva and tumor tissue is useful for the identification of salivary metabolite biomarkers for non-invasive oral cancer screening (Ishikawa et al. 2016).
2.2 Factors impacting metabolomic profiles of saliva
Although several studies revealed the existence of salivary metabolic signatures of oral cancer, it is still hard to get a set of common salivary metabolite biomarkers for the OSCC screening or diagnosis. Multiple factors may affect metabolomic profiles, including biological and technical aspects. The differences of salivary metabolome between male and female, stimulated and unstimulated, and smoking status were observed (Takeda et al. 2009). Almost all metabolites are higher in the unstimulated saliva than in the stimulated saliva (Takeda et al. 2009). Smokers had higher concentrations of citrate, lactate, pyruvate, and sucrose and lower level of formate than non-smokers. Concentrations of several metabolites such as acetate, formate, glycine and lactate in male saliva were significantly higher than those in female saliva (Takeda et al. 2009). Diurnal variation was also observed in salivary metabolomic profiles (Cooke et al. 2003; Kawanishi et al. 2018). Putrescine was the most abundant amine studied, followed by cadaverine and indole (Cooke et al. 2003). When people first woke up, the concentration of amines was found to be the highest, dropping rapidly after breakfast and brushing their teeth (Cooke et al. 2003). These metabolites were known as oral malodour (Goldberg et al. 1994) and may be affected by age. Salivary amino acids showed age-dependent changes (Tanaka et al. 2010). Glycine was the most abundant amino acid in saliva. Glycine and lysine levels increased significantly with age, but not with gender differences (Tanaka et al. 2010). Therefore, these factors must be fully taken into account when interpreting the results of salivary metabolome (Sugimoto et al. 2010).
The stability of metabolites in saliva may depend on the characteristics of metabolites, the time and temperature of sample storage, and the processing conditions of samples. For example, the time point at which saliva is collected after meals affects the level of salivary metabolites, and the 12 h fasting time point after dinner is the best (Ishikawa et al. 2017). Some metabolites, such as choline, betaine, pipecolinic acid and L-carnitine, are relatively stable in saliva at room temperature for 24 h or at least one month at − 35 °C (Wang et al. 2014c). The effect of saliva storage at − 18 °C on polyamines and amino acids was the smallest (Tomita et al. 2018). Polyamines were more stable than amino acids at a higher temperature, and addition of ethanol significantly stabilized polyamine (Tomita et al. 2018). The charged metabolites in plasma showed better stability than those in serum (Hirayama et al. 2015). Understanding the stability of metabolites under various saliva storage conditions is helpful to establish a protocol for the clinical usage.
In addition to biological aspects, there is also a need for improvement in detection methods. It is necessary to establish a standard protocol by taking the advantages of various techniques or pay attention to other new methods (Yuvaraj et al. 2014). Moreover, the source and mechanism of tumor-related metabolites in saliva need further study. It is not clear how oral and even distant cancers release their metabolites into saliva (Wang et al. 2017). It is possible to release exosomes through cancer cells (Lau et al. 2013).
2.3 Salivary metabolomics in other cancers or periodontal diseases
Not only salivary metabolomics of oral cancer has been studied. In addition, the metabolite profiles of saliva in other cancers or diseases were analyzed (Takayama et al. 2016; Zhong et al. 2016). Sugimoto et al. conducted a comprehensive metabolite analysis of saliva samples from patients with oral cancer, pancreatic cancer, breast cancer and periodontal disease, and identified 57 metabolites that were significantly different between patients and healthy controls, including 28 metabolites for oral cancer, 28 for breast cancer and 48 for pancreatic cancer (Table 1) (Sugimoto et al. 2010). Although salivary metabolites contain cancer-specific characteristics, some of them in the three cancers are overlapping. For example, ornithine and putrescine levels in breast cancer or pancreatic cancer patients were higher than those in healthy controls. However, the levels of these metabolites in oral cancer patients were markedly higher than those in healthy controls, suggesting that polyamines in saliva were affected by cancer types (Sugimoto et al. 2010). The spermine and acetyl-spermidine in saliva of the patients with breast cancer (Takayama et al. 2016) or pancreatic cancer (Asai et al. 2018) were significantly different from those in healthy controls. Since changes in polyamines have been observed in many types of cancer, the specificity of these changes in oral cancer needs to be verified.
Salivary metabolomics is used not only in the diagnosis of oral cancer, but also in periodontal diseases (Mikkonen et al. 2016; Sakanaka et al. 2017; Romano et al. 2018). Salivary metabolites cadaverine and hydrocinnamate were associated with periodontal inflamed surface area (PISA), which is an indicator of periodontal inflammation, while uric acid and ethanolamine were associated with the decline of PISA (Sakanaka et al. 2017). In another study, a combination of cadaverine, 5-oxoproline, and histidine in saliva was found to have satisfactory accuracy in the diagnosis of periodontitis (Kuboniwa et al. 2016). Compared with healthy periodontal individuals, the salivary levels of pyruvate, N-acetyl groups and lactate in patients with generalized chronic periodontitis and generalized aggressive periodontitis (GAgP) were significantly lower, while the levels of proline, phenylalanine, and tyrosine were significantly higher (Romano et al. 2018). We note that some metabolites such as cadaverine, histidine, and phenylalanine are associated with oral cancer and periodontitis (Sugimoto et al. 2010; Kuboniwa et al. 2016; Sakanaka et al. 2017; Romano et al. 2018), and their importance in oral cancer needs to be carefully evaluated.
3 Serum metabolome of oral cancer
Serum metabolome assay of oral cancer is another application of metabolomics. In an early study when blood samples from oral cancer patients were analyzed using 1H NMR-based metabolomics, a distinct signature of altered energy metabolism was observed, which presents as accumulation of ketone bodies, suppression of tricarboxylic acid (TCA) cycle, and abnormal amino acid catabolism (Tiziani et al. 2009). The metabolic profile alteration occurs in early stage of oral cancer and distinguishes from the finding in the controls, suggesting existence of a systemic metabolic response to cancer and a potential value for early diagnosis (Tiziani et al. 2009). In another study, 1H and 13C NMR was used to analyze OSCC serum, and down-regulation of choline was detected in OSCC, with concomitant up-regulation of its break-down product of trimethylamine N-oxide (Bag et al. 2015). Plasma metabolite profiles also revealed that phospholipid metabolism plays a critical role in the oncogenesis of esophageal squamous cell carcinoma (Liu et al. 2013). Malonate was found to be up-regulated, suggesting the existence of an alternative glucose metabolism pathway (Bag et al. 2015).
The serum metabolomics was used to discriminate OSCC, oral potentially malignant disorder, and healthy controls (Zhou et al. 2009; Gupta et al. 2015). It was found that four metabolites, glutamine, propionate, acetone, and choline, were able to accurately distinguish oral cancer from healthy controls, and a combination of glutamine, acetone, acetate, and choline were able to precisely distinguish OLK from OSCC (Gupta et al. 2015). The results demonstrated that serum metabolite signature exists in oral cancer, and even in OLK stage. In order to explore time-dependent metabolic changes during the oral carcinogenesis, the lesions of OLK and OSCC were induced in Sprague–Dawley rats by the administration of 4-nitroquinoline-1-oxide and NMR-based blood plasma metabolome was analyzed (Kong et al. 2015). Up-regulation of lactate, choline and glucose and down-regulation of proline, valine, isoleucine, aspartate and 2-hydroxybutyric acid may contribute to the oral cancer development, suggesting that plasma metabolites are potential metabolic biomarkers for oral carcinogenesis, which are probably applied in the early diagnosis and prevention of oral cancer (Kong et al. 2015).
When the metabolomic changes of serum and tissue samples of head and neck squamous cell carcinoma (HNSCC) patients were comparatively analyzed using GC-MS, it was found that the serum levels of several amino acids were down-regulated in HNSCC patients, and levels of a few of metabolites involved in the glycolytic pathway were up-regulated (Yonezawa et al. 2013). In contrast to sera, levels of some amino acids, such as valine, tyrosine, serine and methionine, were increased and the levels of many glycolysis-related metabolites were decreased in tumor tissues of HNSCC compared with non-tumorous tissues (Yonezawa et al. 2013). The different directional changes of certain metabolites in cancer tissue and serum may not be an inconsistency. The metabolite increased in cancer tissue but decreased in serum, which may indicate an increased uptake of that metabolite in cancer. This needs to be verified.
As discussed above, serum metabolomics was also applied to the other cancers or diseases. For example, taurine, glutamate and ethylmalonic acid in serum were identified as important metabolites for detecting human breast cancer (Wang et al. 2018). Compared with healthy controls, serum urea and allo-inositol levels in GAgP patients increased significantly, while the levels of glutathione, 2,5-dihydroxybenzaldehyde, adipic acid and 2-deoxyguanosine decreased (Chen et al. 2018). Although these metabolites are not consistent with those in the serum of oral cancer patients, some of them, such as taurine and glutamate, increase in saliva (Sugimoto et al. 2010; Ohshima et al. 2017) or tumor tissue (Somashekar et al. 2011) of oral cancer patients. Therefore, the source, mechanism and specificity of serum cancer-related metabolites in patients with oral cancer need further study.
4 Metabolomics study of oral cancer tissues and cells
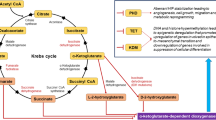
Metabolomics study of oral cancer or HNSCC is valuable for the understanding of the pathophysiological processes and underlying mechanisms of these cancers. In early studies of metabolomics, certain amino acids, glutathione, and polyamine were found more likely to be detected in HNSCC than in normal tissues (Mukherji et al. 1997), and taurine, choline, glutamate, lactate, and lipid were found to have diagnostic potential (El-Sayed et al. 2002). When metabolic profiles of matched HNSCC, adjacent normal tissues, and associated lymph-node metastatic tissues were explored, both primary and metastatic HNSCC tissues were found to have the increased levels of amino acids, choline-containing compounds, creatine, taurine, and glutathione, and decreased levels of triglycerides (Somashekar et al. 2011). Metabolomics study in OSCC supported the findings above (Srivastava et al. 2011). A recent study revealed an enhancement of glucose and glutamine consumption in OSCC tissues (Table 1) (Ogawa et al. 2014). Glutamate is the highest prominent amino acid in OSCC tissues, whereas the most abundant amino acid in normal tissues is glutamine. The enhanced glutamine consumption and lactate production represent the existence of glutaminolysis and the Warburg effect (Ogawa et al. 2014). The 2-hydroxyglutarate, a TCA cycle analog, was detected in HNSCC tumor and cell lines but not in adjacent normal tissues, suggesting its potential as non-invasive biomarker (Mukherjee et al. 2017).
Considering the heterogeneous property of tumor tissues, metabolome of HNSCC cell lines and primary cultures of normal human oral keratinocytes was analyzed (Tripathi et al. 2012). More than 35 metabolites were identified in cell extracts, indicating that HNSCC cells have extensive metabolic reprogramming. Glutaminolysis was found to be a major carbon source in HNSCC cells (Tripathi et al. 2012). The altered levels of choline-containing metabolites and elevated arachidonic acid in HNSCC cells suggested that there is a change in membrane choline phospholipid metabolism. This was further supported by the enhanced cytosolic activity of phospholipase A2 (PLA2) (Tripathi et al. 2012). The extract of HNSCC cells showed similar metabolomic profile to that of HNSCC tissues (Tripathi et al. 2012), and the tumor tissues of different anatomical parts of head and neck also showed similar metabolomic profile (Somashekar et al. 2011).
Metabolomics analysis of tumor tissues or cells has also been used to identify the metabolites associated with metastatic potential, stemness, and precancerous lesion of oral cancer. SCC-9, a human tongue squamous cell carcinoma cell, was inoculated into the footpads of nude mice to increase metastatic potential (Sant’Anna-Silva et al. 2018), and the metabolites of cell extract were analyzed by NMR. Compared with tongue fibroblasts, malonate, methyl malonic acid, n-acetyl, and unsaturated fatty acids were found to accumulate along with the metastatic potential progression, suggesting that lipogenesis is related to the increased invasiveness (Sant’Anna-Silva et al. 2018). The sugar monophosphates, in particular, d-fructose 6-phosphate and α-d-glucose 6-phosphate, were found to be significantly higher in stem-like cancer cells of HNSCC compared with nonstem cancer cells, implying that glycolysis pathway may be activated in the stem-like cancer cells (Wang et al. 2014a). Metabolome of OSCC, oral sub-mucous fibrosis, and controls was analyzed with GC-MS, and 31 differential compounds were identified among samples, suggesting potential roles of tissue metabolic profiles in detecting oral cancer (Musharraf et al. 2016). Metabolomics approach is also applied to analyze the effect of key metabolic genes on the metabolism in oral cancer. The knockdown of metabolic enzyme adenylate kinase 2 or phosphorylate glycerol kinase 1 gene resulted in distinct changes in metabolic phenotypes in oral cancer cells (Ji et al. 2017).
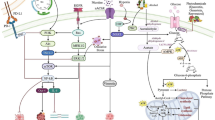
Multiple metabolic pathway alterations were observed in oral cancer, which include highly active glycolysis, increased influx of amino acids, glutaminolysis, lipolysis, TCA cycle, membrane choline phospholipid metabolism, and anti-oxidant mechanism (Somashekar et al. 2011; Tripathi et al. 2012; Wang et al. 2014a). Increased levels of choline-containing molecules and alteration of membrane biogenesis were observed in OSCC tissues in numerous studies. Choline to creatine ratio was found to be increased in HNSCC compared to healthy controls (Mukherji et al. 1997; Star-Lack et al. 2000; El-Sayed et al. 2002), predicting a poor treatment response and bad prognosis (Bezabeh et al. 2005). Choline is an important constituent of cellular membrane phospholipid, and abnormal choline metabolism is regarded as a metabolic hallmark for tumor development and progression (Glunde et al. 2011). Metabolomics analysis identified a significant change of lipid metabolism intermediates in oral sub-mucous fibrosis and cancer, revealing the alteration of membrane biogenesis in oral cancer and precancerous lesion (Bag et al. 2016).
5 Metabolomics of chemo-resistant oral cancer cells
Understanding of cancer type-specific metabolic pathways and the effects of chemotherapeutic drugs on these pathways is valuable for the development of new strategies for diagnosing specific cancers and more effective anticancer agents (Urakami et al. 2013). Metabolomics analysis can reflect the unique chemical fingerprints of specific cell processes, which can be used to evaluate the response of chemotherapy and identify novel therapeutic biomarkers (Zhang et al. 2016; Zaal et al. 2017; Cardoso et al. 2018). It has been applied to identify the serum metabolic signatures of OSCC patients in response to TPF (docetaxel, cisplation, and fluorouracil) induction chemotherapy, which uses chemotherapeutic drugs as initial treatment prior to radiotherapy or surgery of cancer (Ye et al. 2014). Chemotherapy leads to up-regulation of fatty acids, steroids, and antioxidant substances in all patients. The metabolic response of amino acids and carbohydrates were significantly different in patients with significant or no significant efficacy of chemotherapy, and metabolites related to glycolysis and amino acid metabolism were reversely regulated (Ye et al. 2014). The lactate, glucose, glutamate, aspartate, leucine, and glycerol were remarkably associated with efficacy of induction chemotherapy. The lactate, glutamate, and aspartate were defined as potential biomarkers, which can precisely predict the suitability and efficacy of induction chemotherapy. These results suggest the potential application of metabolomics in personalized induction chemotherapy (Ye et al. 2014).
The development of multidrug resistance (MDR) is a major obstacle in the cancer chemotherapy. Carboplatin and pingyangmycin were used to induce MDR in oral squamous cell carcinoma cell line Tca8113 in vitro, and extracellular metabolome was analyzed using NMR (Wang et al. 2015). The results indicated that the amount of glutamate, glycerophosphoethanol amine, and glucose was significantly altered in the drug-induced Tca8113 cells compared with the parental Tca8113 cell line (Wang et al. 2015). A relatively higher level of acetate and lower level of lactate were observed in drug-resistant cells, suggesting their important role in the drug resistance of cells. Metabolomic analysis may have potential for monitoring the formation of MDR during chemotherapy (Wang et al. 2015).
Metabolomics studies of chemo-resistance in oral cancer are relatively limited, and the metabolic changes caused by chemotherapeutic resistance need to be verified. For example, patients with oral cancer have higher levels of glutamate and lactate in tumor tissues (Srivastava et al. 2011; Ogawa et al. 2014) and higher level of glutamate in saliva (Sugimoto et al. 2010; Ohshima et al. 2017), indicating enhanced glutamine consumption and lactate production. The levels of serum glutamate and lactate significantly decreased after successful induction chemotherapy (Ye et al. 2014). However, lactate level also decreased in drug-resistant cell culture (Wang et al. 2015). These metabolites come from different sources, making them difficult to compare. We note that only extracellular metabolites have been profiled in the two studies described. Further studies may focus on intracellular metabolome of chemo-resistant cells in order to understand the cellular metabolic responses to chemotherapy and identify the potential targets for tumor therapy.
6 Conclusion
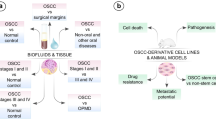
The metabolomics has been applied in many aspects of cancer study including cancer pathophysiology, biomarker discovery, and treatment response (Backshall et al. 2011; Roodhart et al. 2011; Tan et al. 2012; Weaver et al. 2012; Ye et al. 2012). Majority of metabolomics studies in oral cancer focus on the metabolic profiles of saliva, serum, and tumor tissues in order to identify potential biomarkers for the screening or early diagnosis (Fig. 1), although a few of studies involve other samples or applications. For example, a set of metabolites identified through urine metabolomics can accurately predict OSCC (Xie et al. 2012), and metabolomics was applied to screen the novel structural chemicals with potential activity against OSCC (Suzuki et al. 2014). While a panel of tumor-specific metabolites are usually identified in individual study, it seems difficult to get consensus biomarkers derived from multiple studies. Some studies even obtained discrepant results, such as whether the Warburg effect exists or not in OSCC (Ogawa et al. 2014; Bag et al. 2015). Therefore, it is necessary to perform comprehensive studies to evaluate the various discriminant metabolites in different bio-fluids or tissues in order to identify several genuine biomarkers for the clinical application. In addition, few metabolite biomarkers in oral cancer are validated. This is why these metabolites could not yet be used as clinical markers for the screening or diagnosis of oral cancer.
Applications of metabolomics studies in oral cancer. The saliva, serum and tissue obtained from oral cancer are subject to metabolomics analysis and discriminant metabolites or biomarkers are identified. A panel of discriminant metabolites will be used for the screening or early diagnosis of oral cancer, evaluation of the responses of cancer cells to chemotherapy, and investigation of the association of metabolic characteristics of oral cancer with clinic-pathophysiology
Some suggestions for future research are put forward. Methodologically, the recently used techniques and methods should be optimized to establish a highly sensitive routine procedure, which can be suitable for different polarities of metabolites in various samples. Secondly, most of the available studies focus on the metabolomics of bio-fluids or extracellular metabolites. It is critical to profile intracellular metabolites and find the metabolic characteristics of cancer cells. Thirdly, and most importantly, the mechanism behind metabolomic alterations should be clarified. This is probably accomplished by integrating transcriptomics and/or proteomics analysis to find out the genes or proteins resulted in or related to the metabolomic changes, which could be potential targets for the therapy of oral cancer. The metabolic changes of cancer cells in response to drug should be analyzed to uncover the metabolic mechanisms of drug-resistance in oral cancer, which may provide opportunity to overcome the chemo-resistance of cancer or revert the sensitivity of cancer by the regulation of metabolism. The metabolomics offers novel insights into cancer metabolism and will be widely applied in the diagnosis and therapy of oral cancer in the future.
References
Asai, Y., et al. (2018). Elevated polyamines in saliva of pancreatic cancer. Cancers, 10, E43. https://doi.org/10.3390/cancers10020043.
Backshall, A., Sharma, R., Clarke, S. J., & Keun, H. C. (2011). Pharmacometabonomic profiling as a predictor of toxicity in patients with inoperable colorectal cancer treated with capecitabine. Clinical Cancer Research, 17, 3019–3028. https://doi.org/10.1158/1078-0432.CCR-10-2474.
Bag, S., et al. (2015). NMR ((1)H and (13)C) based signatures of abnormal choline metabolism in oral squamous cell carcinoma with no prominent Warburg effect. Biochemical and Biophysical Research Communications, 459, 574–578. https://doi.org/10.1016/j.bbrc.2015.02.149.
Bag, S., et al. (2016). NanoLC MALDI MS/MS based quantitative metabolomics reveals the alteration of membrane biogenesis in oral cancer. Rsc Advances, 6, 62420–62433. https://doi.org/10.1039/c6ra07001a.
Bezabeh, T., et al. (2005). Prediction of treatment response in head and neck cancer by magnetic resonance spectroscopy. AJNR American Journal of Neuroradiology, 26, 2108–2113.
Cardoso, M. R., Santos, J. C., Ribeiro, M. L., Talarico, M. C. R., Viana, L. R., & Derchain, S. F. M. (2018). A metabolomic approach to predict breast cancer behavior and chemotherapy response. International Journal of Molecular Sciences, 19, E617. https://doi.org/10.3390/ijms19020617.
Chen, H. W., Zhou, W., Liao, Y., Hu, S. C., Chen, T. L., & Song, Z. C. (2018). Analysis of metabolic profiles of generalized aggressive periodontitis. Journal of Periodontal Research, 53, 894–901. https://doi.org/10.1111/jre.12579.
Chen, X., & Zhao, Y. (2017). Human papillomavirus infection in oral potentially malignant disorders and cancer. Archives of Oral Biology, 83, 334–339. https://doi.org/10.1016/j.archoralbio.2017.08.011.
Cooke, M., Leeves, N., & White, C. (2003). Time profile of putrescine, cadaverine, indole and skatole in human saliva. Archives of Oral Biology, 48, 323–327.
El-Sayed, S., et al. (2002). An ex vivo study exploring the diagnostic potential of 1H magnetic resonance spectroscopy in squamous cell carcinoma of the head and neck region. Head and Neck, 24, 766–772. https://doi.org/10.1002/hed.10125.
Gerner, E. W., & Meyskens, F. L. Jr. (2004). Polyamines and cancer: old molecules, new understanding. Nature Reviews Cancer, 4, 781–792. https://doi.org/10.1038/nrc1454.
Glunde, K., Bhujwalla, Z. M., & Ronen, S. M. (2011). Choline metabolism in malignant transformation. Nature Reviews Cancer, 11, 835–848. https://doi.org/10.1038/nrc3162.
Goldberg, S., Kozlovsky, A., Gordon, D., Gelernter, I., Sintov, A., & Rosenberg, M. (1994). Cadaverine as a putative component of oral malodor. Journal of Dental Research, 73, 1168–1172. https://doi.org/10.1177/00220345940730060701.
Gupta, A., Gupta, S., & Mahdi, A. A. (2015). (1)H NMR-derived serum metabolomics of leukoplakia and squamous cell carcinoma. Clinica Chimica Acta, 441, 47–55. https://doi.org/10.1016/j.cca.2014.12.003.
Hirayama, A., et al. (2015). Effects of processing and storage conditions on charged metabolomic profiles in blood. Electrophoresis, 36, 2148–2155. https://doi.org/10.1002/elps.201400600.
Ishikawa, S., et al. (2016). Identification of salivary metabolomic biomarkers for oral cancer screening. Scientific Reports, 6, 31520. https://doi.org/10.1038/srep31520.
Ishikawa, S., et al. (2017). Effect of timing of collection of salivary metabolomic biomarkers on oral cancer detection. Amino Acids, 49, 761–770. https://doi.org/10.1007/s00726-017-2378-5.
Ji, E. H., et al. (2017). Metabolomic analysis of human oral cancer cells with adenylate kinase 2 or phosphorylate glycerol kinase 1 inhibition. Journal of Cancer, 8, 298–304. https://doi.org/10.7150/jca.17521.
Kawanishi, N., et al. (2018). Effects of inter-day and intra-day variation on salivary metabolomic profiles. Clinica Chimica Acta, 489, 41–48. https://doi.org/10.1016/j.cca.2018.11.030.
Kong, X., et al. (2015). Analysis of plasma metabolic biomarkers in the development of 4-nitroquinoline-1-oxide-induced oral carcinogenesis in rats. Oncology Letters, 9, 283–289. https://doi.org/10.3892/ol.2014.2619.
Kuboniwa, M., Sakanaka, A., Hashino, E., Bamba, T., Fukusaki, E., & Amano, A. (2016). Prediction of periodontal inflammation via metabolic profiling of saliva. Journal of Dental Research, 95, 1381–1386. https://doi.org/10.1177/0022034516661142.
Lau, C., et al. (2013). Role of pancreatic cancer-derived exosomes in salivary biomarker development. Journal of Biological Chemistry, 288, 26888–26897. https://doi.org/10.1074/jbc.M113.452458.
Liu, R., et al. (2013). Identification of plasma metabolomic profiling for diagnosis of esophageal squamous-cell carcinoma using an UPLC/TOF/MS platform. International Journal of Molecular Sciences, 14, 8899–8911. https://doi.org/10.3390/ijms14058899.
Markley, J. L., et al. (2017). The future of NMR-based metabolomics. Current Opinion in Biotechnology, 43, 34–40. https://doi.org/10.1016/j.copbio.2016.08.001.
Mikkonen, J. J., Singh, S. P., Herrala, M., Lappalainen, R., Myllymaa, S., & Kullaa, A. M. (2016). Salivary metabolomics in the diagnosis of oral cancer and periodontal diseases. Journal of Periodontal Research, 51, 431–437. https://doi.org/10.1111/jre.12327.
Mukherjee, P. K., et al. (2017). Metabolomic analysis identifies differentially produced oral metabolites, including the oncometabolite 2-hydroxyglutarate, in patients with head and neck squamous cell carcinoma. BBA Clinical, 7, 8–15. https://doi.org/10.1016/j.bbacli.2016.12.001.
Mukherji, S. K., Schiro, S., Castillo, M., Kwock, L., Muller, K. E., & Blackstock, W. (1997). Proton MR spectroscopy of squamous cell carcinoma of the extracranial head and neck: in vitro and in vivo studies. AJNR American Journal of Neuroradiology, 18, 1057–1072.
Musharraf, S. G., Shahid, N., Naqvi, S. M., Saleem, M., Siddiqui, A. J., & Ali, A. (2016). Metabolite profiling of preneoplastic and neoplastic lesions of oral cavity tissue samples revealed a biomarker pattern. Science Reports, 6, 38985. https://doi.org/10.1038/srep38985.
Nakagawa, H., Hayata, Y., Kawamura, S., Yamada, T., Fujiwara, N., & Koike, K. (2018). Lipid metabolic reprogramming in hepatocellular carcinoma. Cancers, 10, E447. https://doi.org/10.3390/cancers10110447.
Nicholson, J. K., & Lindon, J. C. (2008). Systems biology: metabonomics. Nature, 455, 1054–1056. https://doi.org/10.1038/4551054a.
Ogawa, T., Washio, J., Takahashi, T., Echigo, S., & Takahashi, N. (2014). Glucose and glutamine metabolism in oral squamous cell carcinoma: insight from a quantitative metabolomic approach. Oral Surgery, Oral Medicine, Oral Pathology, Oral Radiology, 118, 218–225. https://doi.org/10.1016/j.oooo.2014.04.003.
Ohshima, M., Sugahara, K., Kasahara, K., & Katakura, A. (2017). Metabolomic analysis of the saliva of Japanese patients with oral squamous cell carcinoma. Oncology Reports, 37, 2727–2734. https://doi.org/10.3892/or.2017.5561.
Okamura, M., Kobayashi, M., Suzuki, F., Shimada, J., & Sakagami, H. (2007). Induction of cell death by combination treatment with cisplatin and 5-fluorouracil in a human oral squamous cell carcinoma cell line. Anticancer Research, 27, 3331–3337.
Rai, V., Mukherjee, R., Ghosh, A. K., Routray, A., & Chakraborty, C. (2018). “Omics” in oral cancer: new approaches for biomarker discovery. Archives of Oral Biology, 87, 15–34. https://doi.org/10.1016/j.archoralbio.2017.12.003.
Romano, F., et al. (2018). Analysis of salivary phenotypes of generalized aggressive and chronic periodontitis through nuclear magnetic resonance-based metabolomics. Journal of Periodontology, 89, 1452–1460. https://doi.org/10.1002/Jper.18-0097.
Roodhart, J. M., et al. (2011). Mesenchymal stem cells induce resistance to chemotherapy through the release of platinum-induced fatty acids. Cancer Cell, 20, 370–383. https://doi.org/10.1016/j.ccr.2011.08.010.
Sakanaka, A., Kuboniwa, M., Hashino, E., Bamba, T., Fukusaki, E., & Amano, A. (2017). Distinct signatures of dental plaque metabolic byproducts dictated by periodontal inflammatory status. Science Reports, 7, 42818. https://doi.org/10.1038/srep42818.
Sant’Anna-Silva, A. C. B., Santos, G. C., Campos, S. P. C., Oliveira Gomes, A. M., Perez-Valencia, J. A., & Rumjanek, F. D. (2018). Metabolic profile of oral squamous carcinoma cell lines relies on a higher demand of lipid metabolism in metastatic cells. Frontiers Oncology, 8, 13. https://doi.org/10.3389/fonc.2018.00013.
Shankar, A. A., Alex, S., & Routray, S. (2014). Incorporation of salivary metabolomics in oral cancer diagnostics. Oral Oncology, 50, e53–e54. https://doi.org/10.1016/j.oraloncology.2014.07.013.
Shin, J. M., Kamarajan, P., Fenno, J. C., Rickard, A. H., & Kapila, Y. L. (2016). Metabolomics of head and neck cancer: a mini-review. Frontiers in Physiology, 7, 526. https://doi.org/10.3389/fphys.2016.00526.
Somashekar, B. S., et al. (2011). Magic angle spinning NMR-based metabolic profiling of head and neck squamous cell carcinoma tissues. Journal of Proteome Research, 10, 5232–5241. https://doi.org/10.1021/pr200800w.
Srivastava, S., Roy, R., Gupta, V., Tiwari, A., Srivastava, A. N., & Sonkar, A. (2011). Proton HR-MAS MR spectroscopy of oral squamous cell carcinoma tissues: an ex vivo study to identify malignancy induced metabolic fingerprints. Metabolomics, 7, 278–288. https://doi.org/10.1007/s11306-010-0253-4.
Star-Lack, J. M., et al. (2000). In vivo 1H MR spectroscopy of human head and neck lymph node metastasis and comparison with oxygen tension measurements. AJNR American Journal of Neuroradiology, 21, 183–193.
Sugimoto, M., Wong, D. T., Hirayama, A., Soga, T., & Tomita, M. (2010). Capillary electrophoresis mass spectrometry-based saliva metabolomics identified oral, breast and pancreatic cancer-specific profiles. Metabolomics, 6, 78–95. https://doi.org/10.1007/s11306-009-0178-y.
Sun, L. C., Suo, C. X., Li, S. T., Zhang, H. F., & Gao, P. (2018). Metabolic reprogramming for cancer cells and their microenvironment: beyond the Warburg effect. Biochimica et Biophysica Acta, 1870, 51–66. https://doi.org/10.1016/j.bbcan.2018.06.005.
Suzuki, R., Matsuno, S., Sakagami, H., Okada, Y., & Shirataki, Y. (2014). Search of new cytotoxic crude materials against human oral squamous cell carcinoma using 1H NMR-based metabolomics. Anticancer Research, 34, 4117–4120.
Takayama, T., et al. (2016). Diagnostic approach to breast cancer patients based on target metabolomics in saliva by liquid chromatography with tandem mass spectrometry. Clinica Chimica Acta, 452, 18–26. https://doi.org/10.1016/j.cca.2015.10.032.
Takeda, I., et al. (2009). Understanding the human salivary metabolome. NMR in Biomedicine, 22, 577–584. https://doi.org/10.1002/nbm.1369.
Tan, Y., et al. (2012). Metabolomics study of stepwise hepatocarcinogenesis from the model rats to patients: potential biomarkers effective for small hepatocellular carcinoma diagnosis. Molecular and Cell Proteomics, 11, M111 010694. https://doi.org/10.1074/mcp.M111.010694.
Tanaka, S., Machino, M., Akita, S., Yokote, Y., & Sakagami, H. (2010). Changes in salivary amino acid composition during aging. In Vivo, 24, 853–856.
Tiziani, S., Lopes, V., & Gunther, U. L. (2009). Early stage diagnosis of oral cancer using 1H NMR-based metabolomics. Neoplasia, 11, 269–276, 4p following 269.
Tomita, A., et al. (2018). Effect of storage conditions on salivary polyamines quantified via liquid chromatography-mass spectrometry. Science Reports, 8, 12075. https://doi.org/10.1038/s41598-018-30482-x.
Tripathi, P., et al. (2012). Delineating metabolic signatures of head and neck squamous cell carcinoma: phospholipase A2, a potential therapeutic target. International Journal of Biochemistry and Cell Biology, 44, 1852–1861. https://doi.org/10.1016/j.biocel.2012.06.025.
Urakami, K., Zangiacomi, V., Yamaguchi, K., & Kusuhara, M. (2013). Impact of 2-deoxy-D-glucose on the target metabolome profile of a human endometrial cancer cell line. Biomedical Research, 34, 221–229.
Wang, H., et al. (2015). (1)H nuclear magnetic resonance-based extracellular metabolomic analysis of multidrug resistant Tca8113 oral squamous carcinoma cells. Oncology Letters, 9, 2551–2559. https://doi.org/10.3892/ol.2015.3128.
Wang, J., et al. (2014a). Metabolomic profiling of anionic metabolites in head and neck cancer cells by capillary ion chromatography with Orbitrap mass spectrometry. Analytical Chemistry, 86, 5116–5124. https://doi.org/10.1021/ac500951v.
Wang, Q., Gao, P., Wang, X., & Duan, Y. (2014b). The early diagnosis and monitoring of squamous cell carcinoma via saliva metabolomics. Science Reports, 4, 6802. https://doi.org/10.1038/srep06802.
Wang, Q., Gao, P., Wang, X., & Duan, Y. (2014c). Investigation and identification of potential biomarkers in human saliva for the early diagnosis of oral squamous cell carcinoma. Clinica Chimica Acta, 427, 79–85. https://doi.org/10.1016/j.cca.2013.10.004.
Wang, X., et al. (2018). Taurine, glutamic acid and ethylmalonic acid as important metabolites for detecting human breast cancer based on the targeted metabolomics. Cancer Biomarkers, 23, 255–268. https://doi.org/10.3233/CBM-181500.
Wang, X., Kaczor-Urbanowicz, K. E., & Wong, D. T. (2017). Salivary biomarkers in cancer detection. Medical Oncology, 34, 7. https://doi.org/10.1007/s12032-016-0863-4.
Warburg, O. (1956). On the origin of cancer cells. Science, 123, 309–314.
Weaver, Z., et al. (2012). Temporal molecular and biological assessment of an erlotinib-resistant lung adenocarcinoma model reveals markers of tumor progression and treatment response. Cancer Research, 72, 5921–5933. https://doi.org/10.1158/0008-5472.CAN-12-0736.
Wei, J., et al. (2011). Salivary metabolite signatures of oral cancer and leukoplakia. International Journal of Cancer, 129, 2207–2217. https://doi.org/10.1002/ijc.25881.
Xie, G. X., et al. (2012). Urine metabolite profiling offers potential early diagnosis of oral cancer. Metabolomics, 8, 220–231. https://doi.org/10.1007/s11306-011-0302-7.
Yakob, M., Fuentes, L., Wang, M. B., Abemayor, E., & Wong, D. T. (2014). Salivary biomarkers for detection of oral squamous cell carcinoma - current state and recent advances. Current Oral Health Reports, 1, 133–141. https://doi.org/10.1007/s40496-014-0014-y.
Yan, S. K., Wei, B. J., Lin, Z. Y., Yang, Y., Zhou, Z. T., & Zhang, W. D. (2008). A metabonomic approach to the diagnosis of oral squamous cell carcinoma, oral lichen planus and oral leukoplakia. Oral Oncology, 44, 477–483. https://doi.org/10.1016/j.oraloncology.2007.06.007.
Yang, L. F., Venneti, S., & Nagrath, D. (2017). Glutaminolysis: A hallmark of cancer metabolism. Annual Review of Biomedical Engineering, 19, 163–194. https://doi.org/10.1146/annurev-bioeng-071516044546.
Ye, G., et al. (2012). Analysis of urinary metabolic signatures of early hepatocellular carcinoma recurrence after surgical removal using gas chromatography-mass spectrometry. Journal of Proteome Research, 11, 4361–4372. https://doi.org/10.1021/pr300502v.
Ye, G., et al. (2014). Study of induction chemotherapy efficacy in oral squamous cell carcinoma using pseudotargeted metabolomics. Journal of Proteome Research, 13, 1994–2004. https://doi.org/10.1021/pr4011298.
Yonezawa, K., et al. (2013). Serum and tissue metabolomics of head and neck cancer. Cancer Genomics & Proteomics, 10, 233–238.
Yu, L., Chen, X., Sun, X., Wang, L., & Chen, S. (2017). The glycolytic switch in tumors: How many players are involved? Journal of Cancer, 8, 3430–3440. https://doi.org/10.7150/jca.21125.
Yuvaraj, M., et al. (2014). Fluorescence spectroscopic characterization of salivary metabolites of oral cancer patients. Journal of Photochemistry and Photobiology B, 130, 153–160. https://doi.org/10.1016/j.jphotobiol.2013.11.006.
Zaal, E. A., Wu, W., Jansen, G., Zweegman, S., Cloos, J., & Berkers, C. R. (2017). Bortezomib resistance in multiple myeloma is associated with increased serine synthesis. Cancer & Metabolism, 5, 7. https://doi.org/10.1186/s40170-017-0169-9.
Zhang, R. X., Zhuang, X. Y., Zong, L., Liu, S., Liu, Z. Q., & Song, F. R. (2016). Investigations on the cell metabolomics basis of multidrug resistance from tumor cells by ultra-performance liquid chromatography-mass spectrometry. Analytical and Bioanalytical Chemistry, 408, 5843–5854. https://doi.org/10.1007/s00216-016-9696-4.
Zhong, L. P., Cheng, F., Lu, X. Y., Duan, Y. X., & Wang, X. D. (2016). Untargeted saliva metabonomics study of breast cancer based on ultra performance liquid chromatography coupled to mass spectrometry with HILIC and RPLC separations. Talanta, 158, 351–360. https://doi.org/10.1016/j.talanta.2016.04.049.
Zhou, J., et al. (2009). 1H NMR-based metabonomic and pattern recognition analysis for detection of oral squamous cell carcinoma. Clinica Chimica Acta, 401, 8–13. https://doi.org/10.1016/j.cca.2008.10.030.
Acknowledgements
This study was funded by the National Natural Science Foundation of China (No. 81873711 and No. 31670788) and Open Fund of Guangdong Key Laboratory of Pharmaceutical Functional Genes (No.2014B030301028 and No.2017B030314021).
Author information
Authors and Affiliations
Contributions
DY conceived and designed review. XC and DY wrote, read and approved the manuscript. XC drew figure.
Corresponding author
Ethics declarations
Conflict of interest
The authors confirm that there are no conflicts of interest.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Chen, X., Yu, D. Metabolomics study of oral cancers. Metabolomics 15, 22 (2019). https://doi.org/10.1007/s11306-019-1483-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11306-019-1483-8