Abstract
Introduction
Trypanosoma brucei is the causative agent of human African trypanosomiasis, which is responsible for thousands of deaths every year. Current therapies are limited and there is an urgent need to develop new drugs. The anti-trypanosomal compound, 3-(oxazolo[4,5-b]pyridine-2-yl)anilide (OXPA), was initially identified in a phenotypic screen and subsequently optimized by structure–activity directed medicinal chemistry. It has been shown to be non-toxic and to be active against a number of trypanosomatid parasites. However, nothing is known about its mechanism of action.
Objective
Here, we have utilized an untargeted metabolomics approach to investigate the biochemical effects and potential mode of action of this compound in T. brucei.
Methods
Total metabolite extracts were analysed by HILIC-chromatography coupled to high resolution mass spectrometry.
Results
Significant accumulation of ceramides was observed in OXPA-treated T. brucei. To further understand drug-induced changes in lipid metabolism, a lipidomics method was developed which enables the measurement of hundreds of lipids with high throughput and precision. The application of this LC–MS based approach to cultured bloodstream-form T. brucei putatively identified over 500 lipids in the parasite including glycerophospholipids, sphingolipids and fatty acyls, and confirmed the OXPA-induced accumulation of ceramides. Labelling with BODIPY-ceramide further confirmed the ceramide accumulation following drug treatment.
Conclusion
These findings clearly demonstrate perturbation of ceramide metabolism by OXPA and indicate that the sphingolipid pathway is a promising drug target in T. brucei.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Human African trypanosomiasis (HAT), also referred to as sleeping sickness, is a vector-borne disease caused by the parasitic protozoa, Trypanosoma brucei (Brun et al. 2010). There are approximately 8000 new cases of HAT annually, and the disease has a significant socioeconomic impact on communities (Simarro et al. 2011). HAT progresses through two stages; stage one occurs while the parasites proliferate within the haemolymphatic system and is usually associated with flu-like symptoms (Brun et al. 2010). Stage two occurs when the parasites cross the blood–brain barrier (BBB) and invade the central nervous system (CNS), leading to severe sleep disturbances, neurological symptoms and eventual death (Brun et al. 2010).
Currently available drugs for HAT suffer several drawbacks including resistance, toxicity, cost and/or requirement for hospitalization. Pentamidine and suramin are used to treat first-stage HAT caused by T.b. gambiense and T.b. rhodesiense subspecies, respectively. Neither of these compounds cross the BBB, rendering them ineffective against stage-two HAT (Voogd et al. 1993; de Koning 2001). The use of pentamidine can lead to the development of diabetes mellitus and nephrotoxicity (Nok 2003), while suramin has been linked to exfoliative dermatitis and renal failure (Voogd et al. 1993). Melarsoprol is used to treat patients with stage-two disease and is effective against both subspecies of T. brucei (Nok 2003; Seebeck and Maser 2009). However, melarsoprol is extremely toxic and high failure rates have been reported, though resistance has not yet been proven (Seebeck and Maser 2009). Eflornithine is a safer alternative for the treatment of second stage HAT (Seebeck and Maser 2009). However, it is not effective against T.b. rhodesiense infection (Seebeck and Maser 2009) and administration requires four intravenous infusions daily for 14 days and this is impractical in many rural African facilities (Priotto et al. 2009). Recently, nifurtimox has been introduced as a combination therapy with eflornithine, commonly denoted as NECT (Priotto et al. 2009). NECT has the advantage of a shorter and simplified treatment regimen making it the current first-line treatment for second-stage HAT caused by T.b. gambiense (Priotto et al. 2009). Few new drugs are in clinical development for HAT. These include the nitroheterocycle, fexinidazole which has progressed through phase 1 clinical trials (Drugs for Neglected Diseases 2012). The orally active benzoxaborole, SCYX-7158 was selected to enter phase 1 clinical trials in 2012, though the progression of the study has been delayed due to a longer than expected half-life of the drug in human plasma (DNDi 2015). Notwithstanding these developments, there is still a great need for new trypanocidal compounds, particularly for the CNS-resident second stage of this disease.
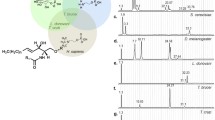
Recently, a high-throughput phenotypic screen of 87,000 compounds was undertaken against T.b. brucei leading to the identification of a novel lead inhibitor compound with a oxazolopyridine core (Sykes and Avery 2009; Sykes et al. 2012). Subsequent structure–activity relationship (SAR) investigations around this structure led to the development of 3-(oxazolo[4,5-b]pyridine-2-yl)anilide (OXPA; 1) as a potent inhibitor of T. brucei (Fig. 1a) (Ferrins et al. 2013). The same chemical scaffold has been identified in independent phenotypic screening campaigns (Schenkman et al. 1991; Chatterjee 2014). We have shown that these compounds demonstrate potent activity against T.b. brucei (IC50: 0.17 µM) and T.b. rhodesiense (IC50: 0.07 µM), and also against other kinetoplastid parasites, Trypanosoma cruzi and Leishmania donovani, the causative agents of Chagas disease and visceral Leishmaniasis, respectively (Ferrins et al. 2013). Minimal toxicity was observed in mammalian cells, suggesting that the oxazolopyridines are promising leads to discover new drugs for these neglected tropical diseases. The unique structure is unlike existing anti-kinetoplastid drugs and there is great interest in identifying the mechanism of action of this compound class to facilitate the optimisation of these broad spectrum anti-kinetoplastids.
a Structure of the novel anti-trypanosomal 3-(oxazolo[4,5-b]pyridine-2-yl)anilide (OXPA; 1). b Distribution of total metabolites putatively identified in treated and untreated bloodstream-form T.b. brucei. Heatmap depicts relative metabolite abundance of putative metabolites from each of the metabolite classes. C: Four replicates of untreated sample, T: Four replicates treated with 0.85 µM OXPA for 5 h. A total of 475 metabolites were detected. c Metabolites displaying significant differences between control and 0.85 µM OXPA-treated T.b. brucei samples by at least 50 %. All metabolites displayed were significantly perturbed according to unpaired Welch’s T test (α = 0.05) and fold change >1.5, <0.5 (n = 4). Metabolites categorized according to KEGG and Lipidmaps annotations. Succinate, 3-phospho-d-glycerate and o-acetylcarnitine changed by less than 50 % but are significantly perturbed metabolites from associated pathways
Untargeted metabolomics combined with in vitro cell culture methods provide a promising tool for pharmacological research to determine mechanisms of drug action (Creek et al. 2012; Drexler et al. 2011). This approach has been validated in T. brucei, with specific perturbation of polyamines detected following treatment with eflornithine, an inhibitor of ornithine decarboxylase (Vincent et al. 2012). Drug-specific metabolic perturbations in T. brucei were also observed for nifurtimox (Vincent et al. 2012), pentamidine (Creek et al. 2013) and halogenated pyrimidines (Ali et al. 2013), and in T. cruzi for benznidazole (Trochine et al. 2014). This unbiased approach is ideally suited to the de novo discovery of drug mechanisms for novel trypanocidal compounds identified by high-throughput phenotypic screening (Creek et al. 2013).
In this study, untargeted high resolution HILIC-MS metabolomics was applied to bloodstream-form T.b. brucei to elucidate the mechanism of action of OXPA. Treatment with this drug led to the selective accumulation of ceramides, which was confirmed using an optimized lipid profiling method and by measurement of the uptake and localization of fluorescently tagged ceramide. Collectively, these data suggest that sphingolipid metabolism is the major metabolic pathway targeted by OXPA.
2 Materials and methods
2.1 Parasite culturing for metabolic studies
T.b. brucei bloodstream forms were cultured in Creeks minimal media (CMM) containing additional 100 µM hypoxanthine (Creek et al. 2013) and 5 ml cultures maintained in a 25 ml vented flask (Corning) at 37 °C with 10 % CO2. The cultures were grown to a maximum density of 2 × 106 cells ml−1 and sub-cultured every 2 or 3 days. Cell density was measured with a Neubauer hemocytometer. Growth curves were obtained in the presence of five times the IC50 concentration of OXPA (0.85 µM) to confirm that this is a sub-lethal concentration for 5 h treatment (Creek et al. 2013).
For metabolomics and lipidomics studies, a confluent cell culture was sub-cultured into fresh medium at 1 × 105 cells ml−1, and OXPA (0.85 µM) or 4 μl DMSO (vehicle control) was added to flasks when cell density reached ~8 × 105 cells ml−1. Cultures were further incubated for 5 h (cell density reached 1 × 106 cells ml−1) and samples, including the controls, were quenched and extracted. Four independent biological replicates were prepared on separate days.
2.2 Untargeted metabolomic analysis of drug-treated Trypanosoma brucei
Parasites were metabolically quenched and extracted as previously described (Creek et al. 2013; Creek et al. 2011). A volume of 40 ml of cell culture was quenched by rapid cooling to 4 °C in a dry ice ethanol bath. Cell pellets were obtained by centrifugation at 1250×g for 10 min. The cell pellet was washed in 1 ml of phosphate-buffered-saline. The washed pellet, containing 4 × 107 cells was extracted with 100 μl chloroform:methanol:water (1:3:1 v/v) with periodic sonication and mixing for 1 h at 4 °C followed by centrifugation to remove the precipitate. The resulting metabolite solution was stored at −80 °C until analysis by liquid chromatography-mass spectrometry with a Dionex Ultimate 3000 UHPLC system (Thermo Fisher scientific) and high resolution mass spectrometry (Q-Exactive Orbitrap; Thermo Fisher scientific). Chromatography was performed using a ZIC-pHILIC (Merck Sequant) column with ammonium carbonate and acetonitrile in the mobile phase (Creek et al. 2013). The instrument was operated in both positive and negative ion mode. Parameters for the HPLC and MS analysis were applied as previously described (Creek et al. 2011).
2.3 Metabolomics data analysis
Metabolomics data analysis was performed with the freely available software packages mzMatch and IDEOM (http://mzmatch.sourceforge.net/ideom.php) as previously described (Trochine et al. 2014). Briefly, metabolite mixes containing 226 authentic metabolites were used to verify retention times and aid metabolite identification. Identification with those standards leads to a high confidence identification (MSI level 1) and these metabolites are highlighted yellow in the supplementary IDEOM file. Putative identification of all other metabolites was carried out by exact mass and predicted retention times from all metabolites from KEGG, Lipidmaps and MetaCyc databases (Creek et al. 2011). Metadata supporting the putative identification of each metabolite, and the associated metabolite identifiers are available in the supplementary IDEOM files which can be viewed in Excel. Raw data and experimental metadata are available in the Metabolights data repository (Accession number: MTBLS360). Relative quantification for the analysis is based on mean peak height and statistical analysis used unpaired Welch’s T test. The data was not normalized, and signal reproducibility was ensured by the analysis of four spiked internal standards (CHAPS, TRIS, PIPES and CAPS), total ion current chromatograms (TIC) and median peak heights. Identified metabolites were filtered to ensure a maximum relative standard deviation (RSD) of 50 % in the technical replicates of the pooled samples for the metabolomics study, and maximum RSD of 30 % for the lipidomics study. Blank extraction buffer was analysed to identify and remove contaminating chemical species and sample carryover.
2.4 Untargeted lipidomics analysis of drug treated Trypanosoma brucei
Parasites were grown as described above, and lipid extraction was carried out as described in the metabolomics methods. Extracts were analyzed utilizing a C8 reversed-phase column (Ascentis Express C8, 5 cm x2.1 mm, 2.7 μm, Supelco Anaytical) and Dionex Ultimate 3000 system (Thermo Fisher scientific) by applying the following settings: mobile phase A: 40 % isopropanol, 60 % H2O, 2 mM formic acid, 8 mM ammonium formate. Mobile phase B: 2 % H2O in isopropanol, 2 mM formic acid, 8 mM ammonium formate. The injector was washed before and after each injection with 3 ml 50:50 IPA:H2O. The column temperature was maintained at 40 °C. Gradient: 0 min 0 %B (100 %A) to 20 %B in 1.5 min linear, 1.5 min to 7 min linear to 28 %B, 7 min to 8 min linear to 35 %B, 8 min to 24 min linear to 65 %B, 24 min to 25 min linear to 100 %B, 25 min to 27 min 100 %B, 27 min to 29 min 100 %A. Flow rate: 0-24 min 200 μl/min, 24 min-29 min 500 μl/min. MS analysis on the Q Exactive Orbitrap was carried out with the following settings: positive and negative mode combined at 140 k resolution, AGC target 3e6, Maximum IT 200 ms, Scan range 140–2000 m/z. HESI source settings for flow rate 200 μl/min: Heater temperature 158 °C, S-lens RF level 50.00, Capillary temperature 300 °C, spray voltage 3.50 kV, sweep gas flow rate 2, aux gas flow rate 20, sheath gas flow rate 50. HESI source settings for flow rate 500 μl/min: Heater temperature 230 °C, S-lens RF level 50.00, Capillary temperature 350 °C, spray voltage 3.50 kV, sweep gas flow rate 5, aux gas flow rate 35, sheath gas flow rate 45. Data analysis was carried out as described above for metabolomics. The retention time prediction model was disabled. For the pooled sample (sample including a fraction of all samples) data-dependent MSMS was performed using the following settings: Chromatographic peak with 15 s, loop count 5, normalized collision energy (NCE) 25 %, positive and negative ionization ran separately, In source CID 0.0 eV, Microscans 1, resolution 17.5 k, AGC target 1e5, Maximum IT 50 ms, scan range from 200 to 2000 m/z, isolation window 4.0 m/z, underfill ratio 1 %, intensity threshold 2.0e4. The separation capability of the C8 reversed-phase column was validated by analysing 44 lipid standards prior to the drug treated T.b. brucei extract.
2.5 Accumulation of fluorescent ceramide
T.b. brucei bloodstream forms (4 × 105 ml−1) were incubated with BODIPY-FL C5-ceramide (2.5 µM, Life Technologies Molecular Probes), and 2 µM OXPA was added to 10 ml of the culture and incubated for 5 h and 24 h. An equivalent amount of DMSO (vehicle) was added to the control flasks. After incubation, cells were concentrated by centrifugation at 700×g, resuspended and fixed in 1 ml of 4 % paraformaldehyde in PBS and incubated for 5 min at 4 °C. Cells were centrifuged for 6 min at 700×g and washed with 1 ml of PBS. 1 µL of Rhodamine phalloidin (Sigma) solution and 0.4 µL of DAPI (Sigma) were added and incubated for 1 h. Cells were centrifuged at 700×g for 6 min and washed with 1 ml PBS and finally resuspended in 10 µL PBS for analysis by fluorescent microscopy (Leica TCS SP8 with a HC Plan APO 63x1.4 NA oil immersion objective, pictures captured in zoom 4). Wavelengths applied: DAPI Eex = 405 nm Eem = 415–450 nm, Phalloidin Eex = 561 nm Eem = 570–620 nm, BODIPY-FL-C5-Cer Eex = 488 nm Eem = 475–540 nm. Lipids were extracted and analysed as described above.
2.6 Determination of IC50 by resazurin growth inhibition assay
Compound activity against T.b. brucei was assessed in a resazurin viability assay as previously described by Sykes and Avery (Sykes and Avery 2009). Briefly, logarithmic phase T.b. brucei 427 bloodstream parasites at a concentration of 1200 cells/ml were added to 384-well microtiter plates (Greiner) in either 55 µL of HMI-9 medium + 10 % FCS or 55 µL of HMI-9 medium + 10 % FCS supplemented with 1 mM carnitine (Sigma). All assay plates were incubated for 24 h at 37 °C/5 % CO2. Serial drug concentrations of the test compound (OXPA) and carnitine were prepared in 100 % DMSO and subsequently diluted 1:21 in DMEM media. 5 µL of this dilution was added to assay plates to give final drug concentrations ranging from 41.67 to 0.04 µM. Plates were incubated for 48 h at 37 °C/5 % CO2. 10 µL of 0.49 mM of resazurin (Sigma Aldrich) prepared in HMI-9 media +10 % FCS was added to assay plates and plates incubated for a further 2 h at 37 °C/5 % CO2 followed by 22 h at room temperature. Assay plates were read at 535 nm excitation/590 nm emission on an Envision® multiplate reader (PerkinElmer, Massachusetts, USA). Data was analysed and IC50 values calculated using the software GraphPad Prism 5. Pentamidine, diminazene acetate and puromycin were included as controls and all experiments are a minimum of n = 2.
3 Results
3.1 Untargeted metabolomic analysis of drug treated T.b. brucei
Metabolites were extracted from T.b. brucei bloodstream forms after 5 h of incubation with either the test compound (OXPA) or DMSO and analysed with ZIC-pHILIC high resolution Orbitrap mass spectrometry. Signal extraction, artefact filtering and polarity merging of positive and negative ionization mode yielded a list of 475 putative metabolites which matched metabolite databases based on retention time and accurate mass. A list of putatively identified metabolites is supplied in Supporting Information File S1. According to the IDEOM software automated metabolite annotation, lipids were the most abundant metabolite class detected on this platform, representing 35 % of all putatively identified metabolites, followed by metabolites of amino acid metabolism (19 %). Putative metabolites that lack Lipidmaps or KEGG class/pathway annotations are listed as unmapped (23 %) (Fig. 1b). In order to detect metabolic changes induced by OXPA, metabolite abundances in untreated and treated cells were compared. Statistical analysis of the filtered data indicated that 11 (2.3 %) putative metabolites showed a significant change (α = 0.05) by at least ± 50 % associated with treatment by OXPA (Fig. 1c). Overall, 16 metabolites (3.4 %) changed significantly (Table 1). Interestingly, most of these putative metabolites were ceramides, all of which accumulated in the presence of drug. Metabolites putatively assigned as riboflavin and N-acetyllactosamine also showed drug-induced accumulation. In contrast, levels of L-carnitine decreased in the presence of drug. Levels of succinate, 6-phospho-D-gluconate and O-acetylcarnitine were also depleted by at least 30 % (Fig. 1c).
3.2 Untargeted lipidomics analysis of drug treated T.b. brucei
Further studies were undertaken to confirm the impact of OXPA on ceramide levels using an untargeted lipidomics approach. In order to detect molecular targets of OXPA, parasite cultures were incubated with drug and the cellular lipidome measured after 5 h of drug exposure. Lipids were extracted from bloodstream-form T.b. brucei and analyzed with a C8 reversed-phase column and high resolution mass spectra collected in both negative and positive ion mode. Several mobile phase conditions were compared and the isopropanol/ammonium formate buffer gradient described here provided optimal chromatographic separation and signal sensitivity, consistent with the findings of other recent lipidomics studies (Yamada et al. 2013; Hu et al. 2008; Chai 2014). Signal extraction, artefact filtering and polarity merging of positive and negative ion mode yielded a list of 517 lipids which matched metabolite databases based on accurate mass and retention times where available. A list of putatively identified metabolites is supplied in Supporting Information File S2. Major lipid classes detected by these analyses were molecular species of phosphatidylcholine (PC: 19 %), phosphatidylethanolamine (PE: 17 %), fatty acids (FA: 11 %), phosphatidylserine (PS: 11 %), phosphatidylinositol (PI: 6 %), sphingomyelin (SM: 6 %), neutral glycerolipid (GL: 6 %), ceramide (Cer: 5 %), glycerophosphoglycerol (PG: 4 %), sterol (ST: 4 %), ethanolamine phosphorylceramides (EPC: 2 %), glycerophosphates (PA: 2 %), sphingoid bases (1 %) and lipid metabolism intermediates (2 %). Additional lipids included prenols, flavonoids, gangliosides, phosphatidylinositol phosphates, phosphatidylethanolamine phosphates, glycerols and other hydrophobic metabolites which may be trypanosome-derived or may have been acquired from the serum in the growth medium (Fig. 2a). Our new lipidomics method shows low relative standard derivations (RSD < 30 %) for over 90 % of detected lipids (Fig. 2b), indicating excellent reproducibility and clear separation of authentic lipid standards (Fig. 2c) and biological extracts (Fig. 2d, e). Statistical analysis of automatically processed data indicated that 13 lipids (2.5 % of all putative lipids) showed a significant change (α < 0.05) induced by the treatment with OXPA by at least ± 30 % (Table 2) – a lower threshold was used for the lipidomic study due to the superior reproducibility demonstrated for this method (Fig. 2b). Consistent with the HILIC-based metabolomics experiment, OXPA primarily impacted on the levels of ceramides (85 % of significantly perturbed lipids), with a total of 11 out of 28 ceramide species exhibiting significant increases (Fig. 3). Data-dependent MS/MS allowed confirmation of the identity of the significantly perturbed ceramides (for the 8 ceramides where MS/MS spectra were available) by the signature ion at m/z 264 (or 266 for dihydroceramides; or 238 for C16-dihydroceramide) (Supporting Information, File S3). The corresponding SM species showed no significant changes except for the SM derived from Cer(32:0) and Cer(34:0), which showed a significant increase (20 %) according to an unpaired Welch’s T test (α = 0.05) (data in supplementary file 2). Eleven corresponding EPC species were detected with no significant perturbation caused by OXPA treatment. No IPC species were detected in any samples.
a Distribution of total lipids in bloodstream-form T.b. brucei sample. The pie chart depicts the number of putatively identified lipids from each lipid class, classified according to KEGG and Lipidmaps. A total of 517 lipids were analysed. b Relative standard deviation (RSD) for all lipids detected in C8 reversed phase analysis from bloodstream-form T.b. brucei. Lipids with RSD < 0.3: 92.5 %, RSD < 0.5: 97.5 %. c Separation of lipid standards by C8 reversed phase chromatography, RA relative abundance, RT retention time, 1 C14H25NO6:[Pimelylcarnitine], 2 C26H52NO7P [PC (18:1)], 3 C34H66NO10P [PS (14:0/14:0)], 4 C36H72NO8P [PC (14:0/14:0)], 5 C41H81N2O6P [SM(d18:1/18:1(9Z))], 6 C44H84NO8P [PC (18:1/18:1)], 7 C45H91N2O6P [SM(d18:1/22:0)], 8 C49H99N2O6P [SM(d18:1/26:0)], 9 C42H83NO3 [Cer(d18:1/24:0)], 10: C55H96O6 [TG(16:0/16:0/20:3)]. d Separation of T.b. brucei lipids by C8 reversed phase chromatography and mass spectrometry. e Separation of T.b. brucei sphingolipids by C8 reversed phase chromatography and mass spectrometry. Each dot represents a detected lipid species, coloured by lipid class. The size of each spot is proportional to the total number of double-bonds in the acyl chains. The general relationship between mass and retention time is observed within each class, with allowance for earlier elution of highly unsaturated species, and later elution of ether-linked phospholipids. GL neutral glycerolipids, PC phosphotidylcholines, PE phosphatidylethanolamines, PI phosphatidylinositols, PG phosphatidylglycerols, PS phosphatidylserines, PA phosphatidic acids, ST sterols, SE sterol esters, FA fatty acyls, SB sphingoid bases, SM sphingomyelins, EPC ethanolamine phosphorylceramides, Cer ceramides, Gan Gangliosides, –O indicates ether-lipids, OH indicates hydroxylated lipids
a The number of significantly perturbed lipids in each class after treatment with OXPA, expressed as a percentage of the total number of identified lipids per class. Significance determined by unpaired Welch’s T test (α = 0.05). Classification according to KEGG and Lipidmaps. b Volcano plot demonstrating that the most significantly perturbed lipids were ceramides. X-axis represents the relative abundance of each lipid in OXPA-treated parasites relative to untreated controls. Y-axis indicates significance determined by unpaired Welch’s T test (n = 4). Ceramides (red), ethanolamine phosphorylceramides (green) and sphingomyelins (blue) are shown in coloured spots. Black circles represent all other detected lipids. c Relative intensity of all detected ceramides in T.b. brucei treated with OXPA (dark columns) and untreated parasites (light columns). Ceramide abundance is shown as mean of LCMS peak intensity ± standard deviation (n = 4) with significant differences (asterisk) determined by Welch’s T test (α = 0.05). Note that four scales are included to allow for the three orders of dynamic range in detected ceramide levels
3.3 Localisation and metabolism of fluorescent ceramide
Bloodstream-form T.b. brucei were incubated with BODIPY-C5 ceramide to verify ceramide accumulation caused by incubation with OXPA, as observed in the metabolomics and lipidomics analyses. Parasites were incubated in the presence of BODIPY-FL-C5-cer for 5 h and 24 h with OXPA or DMSO control, and parasites were co-stained with Phalloidin and DAPI to monitor morphological changes to T.b. brucei due to treatment. Untreated (DMSO control 5 h, 24 h) T.b. brucei show a long slender form and the nuclear and mitochondrial DNA (kinetoplast) localization are consistent with healthy parasites (Matthews 2005). Treatment with OXPA for 5 h causes swelling of the parasite and a generally more rounded morphology. By 24 h, BODIPY ceramide accumulation was evident adjacent to the nucleus, consistent with staining of the Golgi apparatus where conversion of ceramide to more complex lipids (IPC, SM, EPC) occurs (Sutterwala et al. 2008; Fridberg et al. 2008; Sutterwala et al. 2007). A 24 h treatment caused further swelling and rounding of the parasite compared to the controls and 5 h treatment (Fig. 4). Lipids were extracted from 24 h treated bloodstream-form T.b. brucei and analyzed by the C8 reversed phase column and high resolution MS in order to confirm BODIPY-ceramide accumulation. Significant accumulation of BODIPY-ceramide was observed (54 % increase) when parasites were treated with OXPA for 24 h. Interestingly, the corresponding BODIPY-sphingomyelin (C39H66BF2N4O6P) and BODIPY-EPC (C36H60BF2N4O6P) species were also detected at significantly higher abundance in the drug-treated cells (Fig. 5), suggesting that accumulated BODIPY-ceramide is further processed and these steps are not inhibited by OXPA.
Ceramide accumulation in bloodstream-form T.b. brucei following treatment with OXPA. T.b. brucei bloodstream forms were incubated with BODIPY-FL-C5-ceramide in the presence or absence of OXPA and visualised by fluorescent microscopy. C 5 h and 24 h untreated, T treated for 5 h and 24 h with OXPA (two replicates shown). All cells were incubated with BODIPY-FL-C5-ceramide (green) and co-stained with DAPI (blue) and Rhodamine phalloidin (red). k kinetoplast, n nucleus, BF bright field
BODIPY-C5-FL-Ceramide accumulation after 24 h treatment with OXPA. a LC–MS peak intensity (mean ± standard deviation) for BODIPY-ceramide (C34H54BF2N3O3), b BODIPY-Sphingomyelin (C39H66BF2N4O6P) and c BODIPY-EPC (C36H60BF2N4O6P) from T.b. brucei incubated for 24 h with BODIPY-C5-FL-Ceramide and either OXPA or control (untreated). All comparisons show significant difference (asterisk) according to Welch’s unpaired T test (α = 0.05) (n = 2–3). d Acquired mass spectra for BODIPY-ceramide e BODIPY-Sphingomyelin and f BODIPY-EPC. Negative ion spectra demonstrating the accurate mass and boron-specific isotopic pattern
4 Discussion
In this study we have investigated the mode of action of OXPA, a lead anti-trypanosomatid drug candidate, using two complementary metabolomic profiling approaches involving HILIC and C8 reversed phase chromatography coupled to high resolution mass spectrometry. This study extends previous metabolic studies on trypanosomes that utilized the HILIC chromatography mass spectrometry platform (Vincent et al. 2012; Silva et al. 2011; t’Kindt et al. 2010; Kamleh et al. 2008; Trochine et al. 2014). Overall 475 metabolites were detected in the HILIC study, and the concentrations of 16 of those metabolites changed significantly when treated with the novel anti-trypanosomal compound OXPA. Strikingly, six of the significantly perturbed metabolites were ceramides, all of which accumulated by at least 50 % due to treatment. Ceramides contain a sphingolipid base and amide-linked fatty acid and can function as a key player in cell signaling as well as a precursor for complex sphingolipids (Futerman and Hannun 2004). In order to analyze effects of OXPA on the lipidome in more detail, we developed a new LC–MS method that employed a C8 reversed-phase chromatographic separation coupled to high resolution Orbitrap MS. This method can be used to screen total cellular lipids within 30 min in positive and negative ion mode with a mass resolution of 140 k. Analysis of drug treated bloodstream-form T.b. brucei with this lipidomic method shows excellent separation for nonpolar metabolites with low RSD’s and putative identification of 517 lipids. Glycerophospholipids were the most abundant lipid classes observed, which represented over 50 % of all detected lipids. All major phospholipid classes were found in bloodstream-form T.b. brucei in agreement with previous reports (Patnaik et al. 1993). Increased molecular species complexity was observed in the phosphatidylserines than reported in a previous study (Richmond et al. 2010), likely due to the different selectivity of the analytical platforms.
Analysis of the lipidome of bloodstream-form T.b. brucei after 5 h treatment with 0.85 µM OXPA, confirmed that all of the ceramides detected in the untargeted HILIC metabolite analysis were increased in the presence of OXPA. Overall, 11 ceramides showed a significant change (by at least 30 %) supporting the notion that OXPA directly impacts the ceramide metabolism and/or intracellular transport in the parasite. Furthermore, our imaging study with BODIPY-C5-Cer also clearly showed accumulation of ceramides in the treated parasites, most likely in the Golgi apparatus where SMs and EPCs are synthesized (Sutterwala et al. 2008), confirming our findings from metabolomics and lipidomics analyses.
Ceramide accumulation induced by OXPA could, in principle, arise as a result of decreased ceramide degradation or decreased conversion of ceramide to higher order sphingolipids. T. brucei lacks a known pathway for ceramide catabolism (consistent with the finding that neither sphingosine nor sphingosine 1-phosphate were detected in our analyses), suggesting that accumulation is due to altered metabolism. In T. brucei, ceramide is converted to complex sphingolipids such as SM, IPC and EPC, by the transfer of phosphocholine, phosphoinositol or phosphoethanolamine to ceramides, respectively (Becker and Lester 1980; Bromley et al. 2003; Denny et al. 2006; Sutterwala et al. 2008). Consistent with previous studies (Sutterwala et al. 2008; Sutterwala et al. 2007) we show that bloodstream forms predominantly synthesize EPCs and SMs, but not IPCs. Furthermore, BODIPY-labelled ceramide was efficiently incorporated into both SM and EPC. T. brucei possess a family of sphingolipid synthases, encoded by TbSLS1-4, that catalyse the synthesis of complex sphingolipids, and knockdown of these enzymes has been shown to lead to ceramide accumulation (Sutterwala et al. 2008; Serricchio and Butikofer 2011; Mina et al. 2009). However, the increased production of BODIPY-labelled SM and EPC in treated cells suggests that inhibition of sphingolipid synthase is not the primary mechanism of ceramide accumulation observed for OXPA. Interactions with alternative molecular targets in the ceramide uptake or sphingolipid metabolism pathways may be responsible for the observed ceramide accumulation. Alternative mechanisms, such as the activation of the endogenous neutral sphingomyelinase that degrades exogenous sphingolipids to ceramide (Young and Smith 2010) or the upregulation of de novo biosynthesis are unlikely, as up-regulation was also observed when exogenous BODIPY-Cer was added to parasite cultures. On the other hand, our data are consistent with a partial defect in the post-Golgi transport of sphingolipids, leading to an accumulation of ceramide and freshly synthesized complex sphingolipids. Future work will delineate the precise molecular target(s) involved in this mechanism.
The HILIC analysis of polar metabolites indicated that OXPA also led to depletion of the carnitines, O-acetylcarnitine (C9H17NO4) and L-carnitine (C7H15NO3). The potential role of carnitine acetylation/acylation in bloodstream-form T. brucei has not been fully defined. Unlike the insect stage, the mammalian infective bloodstream form is completely dependent on glycolysis for ATP production (Gilbert and Klein 1984). In other eukaryotes, conversion of L-carnitine to O-acetylcarnitine by carnitine acetyl transferase (CAT) (Friedman and Fraenkel 1955) results in depletion of acetyl-CoA and stimulation of pyruvate kinase activity directly or indirectly (Gilbert and Klein 1984). A decrease in L-carnitine levels in T. brucei could lead to a decrease of O-acetylcarnitine synthesis, with a concomitant increase in acetyl-CoA production and partial inhibition of pyruvate kinase activity, with subsequent effects on ATP synthesis and parasite survival. African trypanosomes cannot synthesize L-carnitine but scavenge carnitine from the medium/host via an active transporter, which has been reported as a validated drug target (Gilbert et al. 1983). These findings raised the possibility that carnitine uptake may be inhibited by OXPA. Interestingly, the glycolytic metabolite 3-phospho-D-glycerate (C3H7O7P) demonstrates a decrease in abundance by at least 30 % following treatment, consistent with a down-regulation of glycolysis. In order to test the hypothesis of carnitine uptake inhibition, the in vitro activity of OXPA was measured in the presence of high extracellular levels of L-carnitine (1 mM). No inhibition of activity was observed, and surprisingly our results indicate a twofold decrease in IC50 in the presence of excess L-carnitine (Supporting Information File S4). A similar trend was also observed with pentamidine, indicating some non-specific mild synergistic effect due to the very high concentration of L-carnitine. Overall, the data clearly rejects the hypothesis that excess carnitine would inhibit the activity of OXPA, and suggests that inhibition of carnitine uptake is not the primary mechanism of action of this compound.
The untargeted metabolomics analysis revealed few additional statistically significant perturbations to specific metabolites which may play a role in the activity, and/or potential toxicity of the oxazolopyridines. The metabolite annotated as N-acetyllactosamine (C14H25NO11) increased in abundance by 60 %, and depletion of succinate by 33 % and 6-phospho-d-gluconate by 56 %, may indicate some impact on the pathways of glycoconjugate salvage, central carbon metabolism and NAD(P) +/NAD(P)H balance (Hanau et al. 2004; Fairlamb and Opperdoes 1986; Tielens and Van Hellemond 1998; Besteiro et al. 2002). The mechanism of riboflavin accumulation is uncertain, and further investigations are necessary to confirm the relevance of these changes that were isolated to individual metabolites within diverse pathways. The perturbations to these individual metabolites may be relevant to a thorough understanding of the systems pharmacology of this drug class, but are unlikely to represent the trypanocidal target of OXPA, as other metabolites in these pathways were unaffected. In contrast, the reproducible accumulation observed for a large proportion of the observed ceramides suggests that sphingolipid metabolism is the primary target for the anti-trypanosomal activity of OXPA. It is important to note that most detected metabolites were not significantly impacted by treatment with OXPA. In particular, polyamine and sterol pathway metabolites were not perturbed, suggesting that this compound does not share targets with other common anti-trypanosomal drugs, such as ornithine decarboxylase (Vincent et al. 2012) and sterol 14α-demethylase (CYP51) (Choi et al. 2014). To confirm this, inhibition of CYP51 inhibition was measured in vitro using the purified enzyme, and no activity was observed for OXPA (IC50 > 10 µM), whereas the known CYP51 inhibitor posaconazole demonstrated an enzyme inhibition IC50 of 40 nM. The metabolic profile observed for OXPA was unique compared to those observed for several other anti-trypanosomal compounds tested using the same metabolomics methodology (Ali et al. 2013; Creek et al. 2013; Trochine et al. 2014; Vincent et al. 2012), confirming that the perturbations observed here are specific to this compound class, and do not represent a non-specific cell death phenotype.
Metabolomics and lipidomics enabled an unbiased assessment of the biochemical actions of OXPA in T. brucei. Mass spectrometry based metabolomics with our optimised C8 reversed-phase lipidomics method and fluorescence microscopy clearly demonstrated that the oxazolopyridine anti-trypanosomal compound (OXPA) caused significant accumulation of ceramides, indicating a mechanism of action targeting sphingolipid metabolism or trafficking. These findings provide chemical validation of sphingolipid metabolism and/or trafficking as an attractive pathway for future drug discovery in T. brucei. Specifically, this is the first description of the unique impact of oxazolopyridines on sphingolipid metabolism in T. brucei, providing the basis for future studies investigating the precise molecular target(s) and potential mechanism(s) of resistance for OXPA and related drug candidates. The mechanistic information described here will allow targeted analysis of the biochemical impact of oxazolopyridines in pre-clinical and clinical studies, thus facilitating the development of oxazolopyridines as drug candidates for HAT and other trypanosomatid diseases.
References
Ali, J. A., Creek, D. J., Burgess, K., Allison, H. C., Field, M. C., Maser, P., et al. (2013). Pyrimidine salvage in Trypanosoma brucei bloodstream forms and the trypanocidal action of halogenated pyrimidines. Molecular Pharmacology, 83(2), 439–453. doi:10.1124/mol.112.082321.
Becker, G. W., & Lester, R. L. (1980). Biosynthesis of phosphoinositol-containing sphingolipids from phosphatidylinositol by a membrane preparation from Saccharomyces cerevisiae. Journal of Bacteriology, 142(3), 747–754.
Besteiro, S., Biran, M., Biteau, N., Coustou, V., Baltz, T., Canioni, P., et al. (2002). Succinate secreted by Trypanosoma brucei is produced by a novel and unique glycosomal enzyme, NADH-dependent fumarate reductase. Journal of Biological Chemistry, 277(41), 38001–38012. doi:10.1074/jbc.M201759200.
Bromley, P. E., Li, Y. O., Murphy, S. M., Sumner, C. M., & Lynch, D. V. (2003). Complex sphingolipid synthesis in plants: characterization of inositolphosphorylceramide synthase activity in bean microsomes. Archives of Biochemistry and Biophysics, 417(2), 219–226.
Brun, R., Blum, J., Chappuis, F., & Burri, C. (2010). Human African trypanosomiasis. The Lancet, 375(9709), 148–159.
Chai, X. (2014). Untargeted lipidomic profiling of human plasma reveals differences due to race, gender and smoking status. Metabolomics, 4(131), 2153. doi:10.4172/2153-0769.1000131.
Chatterjee, A.K.N. A.S., Paraselli, P., Kondreddi, R.R., Leong, S.Y., Mishra, P.K., Moreau, R.J., Roland, J.T., Sim, W.L.S., Simon, O., Tan, L.J., Yeung, B.K., Zou, B., Bollu, V. (2014). Compounds and compositions for the treatment of parasitic diseases. US Patent 20140274926 A1. 18th September 2014.
Choi, J. Y., Podust, L. M., & Roush, W. R. (2014). Drug strategies targeting CYP51 in neglected tropical diseases. Chemical Reviews, 114(22), 11242–11271. doi:10.1021/cr5003134.
Creek, D. J., Anderson, J., McConville, M. J., & Barrett, M. P. (2012). Metabolomic analysis of trypanosomatid protozoa. Molecular and Biochemical Parasitology, 181(2), 73–84. doi:10.1016/j.molbiopara.2011.10.003.
Creek, D. J., Jankevics, A., Breitling, R., Watson, D. G., Barrett, M. P., & Burgess, K. E. V. (2011). Toward global metabolomics analysis with hydrophilic interaction liquid chromatography-mass spectrometry: Improved metabolite identification by retention time prediction. Analytical Chemistry, 83(22), 8703–8710. doi:10.1021/ac2021823.
Creek, D. J., Nijagal, B., Kim, D.-H., Rojas, F., Matthews, K. R., & Barrett, M. P. (2013). Metabolomics guides rational development of a simplified cell culture medium for drug screening against Trypanosoma brucei. Antimicrobial Agents and Chemotherapy, 57(6), 2768–2779. doi:10.1128/aac.00044-13.
de Koning, H. P. (2001). Transporters in African trypanosomes: Role in drug action and resistance. International Journal of Parasitology, 31(5–6), 512–522.
Denny, P. W., Shams-Eldin, H., Price, H. P., Smith, D. F., & Schwarz, R. T. (2006). The protozoan inositol phosphorylceramide synthase: A novel drug target that defines a new class of sphingolipid synthase. Journal of Biological Chemistry, 281(38), 28200–28209. doi:10.1074/jbc.M600796200.
DNDi Oxaborole SCYX-7158 (HAT). http://www.dndi.org/diseases-projects/portfolio/oxaborole-scyx-7158.html. Accessed 05 June 2015.
Drexler, D. M., Reily, M. D., & Shipkova, P. A. (2011). Advances in mass spectrometry applied to pharmaceutical metabolomics. Analytical and Bioanalytical Chemistry, 399(8), 2645–2653. doi:10.1007/s00216-010-4370-8.
Drugs for Neglected Diseases (2012). Pivotal Study of Fexinidazole for Human African Trypanosomiasis in Stage 2. http://www.clinicaltrials.gov/ct2/show/NCT01685827?term=fexinidazole&rank=3#wrapper. Accessed 23 Nov 2012.
Fairlamb, A., & Opperdoes, F. (1986). Carbohydrate metabolism in African trypanosomes, with special reference to the glycosome. In M. Morgan (Ed.), Carbohydrate Metabolism in Cultured Cells (pp. 183–224). US: Springer.
Ferrins, L., Rahmani, R., Sykes, M. L., Jones, A. J., Avery, V. M., Teston, E., et al. (2013). 3-(Oxazolo[4,5-b]pyridin-2-yl)anilides as a novel class of potent inhibitors for the kinetoplastid Trypanosoma brucei, the causative agent for human African trypanosomiasis. European Journal of Medicinal Chemistry, 66, 450–465. doi:10.1016/j.ejmech.2013.05.007.
Fridberg, A., Olson, C. L., Nakayasu, E. S., Tyler, K. M., Almeida, I. C., & Engman, D. M. (2008). Sphingolipid synthesis is necessary for kinetoplast segregation and cytokinesis in Trypanosoma brucei. Journal of Cell Science, 121(Pt 4), 522–535. doi:10.1242/jcs.016741.
Friedman, S., & Fraenkel, G. (1955). Reversible enzymatic acetylation of carnitine. Archives of Biochemistry and Biophysics, 59(2), 491–501.
Futerman, A. H., & Hannun, Y. A. (2004). The complex life of simple sphingolipids. EMBO Reports, 5(8), 777–782. doi:10.1038/sj.embor.7400208.
Gilbert, R. J., & Klein, R. A. (1984). Pyruvate kinase: A carnitine regulated site of ATP production in Trypanosoma brucei brucei. Comparative Biochemistry and Physiology Part B: Comparative Biochemistry, 78(3), 595–599. doi:10.1016/0305-0491(84)90104-4.
Gilbert, R. J., Klein, R. A., & Johnson, P. (1983). Bromoacetyl-l-carnitine: Biochemical and antitrypanosomal actions against Trypanosoma brucei brucei. Biochemical Pharmacology, 32(22), 3447–3451. doi:10.1016/0006-2952(83)90375-1.
Hanau, S., Rinaldi, E., Dallocchio, F., Gilbert, I. H., Dardonville, C., Adams, M. J., et al. (2004). 6-phosphogluconate dehydrogenase: a target for drugs in African trypanosomes. Current Medicinal Chemistry, 11(19), 2639–2650.
Hu, C., van Dommelen, J., van der Heijden, R., Spijksma, G., Reijmers, T. H., Wang, M., et al. (2008). RPLC-ion-trap-FTMS method for lipid profiling of plasma: method validation and application to p53 mutant mouse model. Journal of Proteome Research, 7(11), 4982–4991. doi:10.1021/pr800373m.
Kamleh, A., Barrett, M. P., Wildridge, D., Burchmore, R. J., Scheltema, R. A., & Watson, D. G. (2008). Metabolomic profiling using Orbitrap Fourier transform mass spectrometry with hydrophilic interaction chromatography: A method with wide applicability to analysis of biomolecules. Rapid Communications in Mass Spectrometry, 22(12), 1912–1918. doi:10.1002/rcm.3564.
Matthews, K. R. (2005). The developmental cell biology of Trypanosoma brucei. Journal of Cell Science, 118(2), 283–290. doi:10.1242/jcs.01649.
Mina, J. G., Pan, S.-Y., Wansadhipathi, N. K., Bruce, C. R., Shams-Eldin, H., Schwarz, R. T., et al. (2009). The Trypanosoma brucei sphingolipid synthase, an essential enzyme and drug target. Molecular and Biochemical Parasitology, 168(1), 16–23. doi:10.1016/j.molbiopara.2009.06.002.
Nok, A. J. (2003). Arsenicals (melarsoprol), pentamidine and suramin in the treatment of human African trypanosomiasis. Parasitology Research, 90(1), 71–79. doi:10.1007/s00436-002-0799-9.
Patnaik, P. K., Field, M. C., Menon, A. K., Cross, G. A. M., Yee, M. C., & Bütikofer, P. (1993). Molecular species analysis of phospholipids from Trypanosoma brucei bloodstream and procyclic forms. Molecular and Biochemical Parasitology, 58(1), 97–105. doi:10.1016/0166-6851(93)90094-E.
Priotto, G., Kasparian, S., Mutombo, W., Ngouama, D., Ghorashian, S., Arnold, U., et al. (2009). Nifurtimox-eflornithine combination therapy for second-stage African Trypanosoma brucei gambiense trypanosomiasis: a multicentre, randomised, phase III, non-inferiority trial. The Lancet, 374(9683), 56–64.
Richmond, G. S., Gibellini, F., Young, S. A., Major, L., Denton, H., Lilley, A., et al. (2010). Lipidomic analysis of bloodstream and procyclic form Trypanosoma brucei. Parasitology, 137(9), 1357–1392. doi:10.1017/s0031182010000715.
Schenkman, S., Jiang, M. S., Hart, G. W., & Nussenzweig, V. (1991). A novel cell surface trans-sialidase of Trypanosoma cruzi generates a stage-specific epitope required for invasion of mammalian cells. Cell, 65(7), 1117–1125.
Seebeck, T., & Maser, P. (2009). Drug resistance in African Trypanosomiasis. In D. L. Mayers (Ed.), Antimicrobial drug resistance (pp. 589–604). Newyork: Humana Press.
Serricchio, M., & Butikofer, P. (2011). Trypanosoma brucei: a model micro-organism to study eukaryotic phospholipid biosynthesis. FEBS Journal, 278(7), 1035–1046. doi:10.1111/j.1742-4658.2011.08012.x.
Silva, A. M., Cordeiro-da-Silva, A., & Coombs, G. H. (2011). Metabolic variation during development in culture of Leishmania donovani promastigotes. PLoS Neglected Tropical Diseases, 5(12), e1451. doi:10.1371/journal.pntd.0001451.
Simarro, P. P., Diarra, A., Ruiz Postigo, J. A., Franco, J. R., & Jannin, J. G. (2011). The human African trypanosomiasis control and surveillance programme of the World Health Organization 2000–2009: The way forward. PLoS Neglected Tropical Diseases, 5(2), e1007. doi:10.1371/journal.pntd.0001007.
Sutterwala, S. S., Creswell, C. H., Sanyal, S., Menon, A. K., & Bangs, J. D. (2007). De novo sphingolipid synthesis is essential for viability, but not for transport of glycosylphosphatidylinositol-anchored proteins, in African trypanosomes. Eukaryotic Cell, 6(3), 454–464. doi:10.1128/EC.00283-06.
Sutterwala, S. S., Hsu, F. F., Sevova, E. S., Schwartz, K. J., Zhang, K., Key, P., et al. (2008). Developmentally regulated sphingolipid synthesis in African trypanosomes. Molecular Microbiology, 70(2), 281–296. doi:10.1111/j.1365-2958.2008.06393.x.
Sykes, M. L., & Avery, V. M. (2009). Development of an Alamar Blue viability assay in 384-well format for high throughput whole cell screening of Trypanosoma brucei brucei bloodstream form strain 427. American Journal of Tropical Medicine and Hygeine, 81(4), 665–674. doi:10.4269/ajtmh.2009.09-0015.
Sykes, M. L., Baell, J. B., Kaiser, M., Chatelain, E., Moawad, S. R., Ganame, D., et al. (2012). Identification of compounds with anti-proliferative activity against Trypanosoma brucei brucei strain 427 by a whole cell viability based HTS campaign. PLoS Neglected Tropical Diseases, 6(11), e1896. doi:10.1371/journal.pntd.0001896.
Tielens, A. G. M., & Van Hellemond, J. J. (1998). Differences in energy metabolism between Trypanosomatidae. Parasitology Today, 14(7), 265–272. doi:10.1016/S0169-4758(98)01263-0.
t’Kindt, R., Scheltema, R. A., Jankevics, A., Brunker, K., Rijal, S., Dujardin, J.-C., Breitling, R., et al. (2010). Metabolomics to unveil and understand phenotypic diversity between pathogen populations. PLoS Neglected Tropical Diseases. 4(11), e904. doi:10.1371/journal.pntd.0000904.
Trochine, A., Creek, D. J., Faral-Tello, P., Barrett, M. P., & Robello, C. (2014). Benznidazole biotransformation and multiple targets in Trypanosoma cruzi revealed by metabolomics. PLoS Neglected Tropical Diseases, 8(5), e2844. doi:10.1371/journal.pntd.0002844.
Vincent, I. M., Creek, D. J., Burgess, K., Woods, D. J., Burchmore, R. J., & Barrett, M. P. (2012). Untargeted metabolomics reveals a lack of synergy between nifurtimox and eflornithine against Trypanosoma brucei. PLoS Neglected Tropical Diseases, 6(5), e1618. doi:10.1371/journal.pntd.0001618.
Voogd, T. E., Vansterkenburg, E. L., Wilting, J., & Janssen, L. H. (1993). Recent research on the biological activity of suramin. Pharmacological Reviews, 45(2), 177–203.
Yamada, T., Uchikata, T., Sakamoto, S., Yokoi, Y., Fukusaki, E., & Bamba, T. (2013). Development of a lipid profiling system using reverse-phase liquid chromatography coupled to high-resolution mass spectrometry with rapid polarity switching and an automated lipid identification software. Journal of Chromatography A, 1292, 211–218.
Young, S., & Smith, T. K. (2010). The essential neutral sphingomyelinase is involved in the trafficking of the variant surface glycoprotein in the bloodstream form of Trypanosoma brucei. Molecular Microbiology, 76(6), 1461–1482. doi:10.1111/j.1365-2958.2010.07151.x.
Funding
DJC acknowledges support from a NHMRC training fellowship. LF acknowledges an Australian Postgraduate Award. MJM is an NHMRC Principal Research Fellow. Financial support was received from NHMRC project Grants APP1025581 and APP1067728.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interests
All authors declare that they have no conflict of interest.
Human and animal rights
No human participants or animals were involved in this study.
Electronic supplementary material
Below is the link to the electronic supplementary material.
S1
IDEOM metabolite list and metadata from HILIC metabolomics study (XLSB 31541 kb)
S2
IDEOM metabolite list and metadata from lipidomics study (XLSB 23466 kb)
S3
MSMS spectra of significant ceramides and acetylcarnitine (PDF 375 kb)
S4
Results from IC50 analysis in presence of Carnitine (PDF 38 kb)
Rights and permissions
About this article
Cite this article
Stoessel, D., Nowell, C.J., Jones, A.J. et al. Metabolomics and lipidomics reveal perturbation of sphingolipid metabolism by a novel anti-trypanosomal 3-(oxazolo[4,5-b]pyridine-2-yl)anilide. Metabolomics 12, 126 (2016). https://doi.org/10.1007/s11306-016-1062-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11306-016-1062-1