Abstract
Head and neck squamous cell carcinomas (HNSCCs) are a heterogenous group of tumors and among the top 10 most common cancers and they arise from the epithelial tissues of the mucosal surfaces of the oral cavity, oropharynx, and larynx. Aberrant purinergic signaling has been associated with various cancer types. Here, we studied the role of the P2Y2 purinergic receptor (P2Y2R) in the context of oral cancer. We utilized bioinformatics analysis of deposited datasets to examine purinome gene expression in HNSCC tumors and cells lines and functionally characterized nucleotide-induced P2 receptor signaling in human FaDu and Cal27 and murine MOC2 oral cancer cell lines. Utilizing tumorigenesis assays with wild-type or P2ry2 knockout MOC2 cells we evaluated the role of P2Y2Rs in tumor growth and the host anti-tumor immune responses. Our data demonstrate that human and murine oral cancer cell lines express numerous P2 receptors, with the P2Y2R being highly expressed. Using syngeneic tumor grafts in wild-type mice, we observed that MOC2 tumors expressing P2Y2R were larger than P2Y2R−/− tumors. Wild-type MOC2 tumors contained a lower population of tumor-infiltrating CD11b+F4/80+ macrophages and CD3+ cells, which were revealed to be CD3+CD4+IFNγ+ T cells, compared to P2Y2R−/− tumors. These results were mirrored when utilizing P2Y2R−/− mice, indicating that the changes in MOC2 tumor growth and to the host anti-tumor immune response were independent of host derived P2Y2Rs. Results suggest that targeted suppression of the P2Y2R in HNSCC cells in vivo, rather than systemic P2Y2R antagonism, may be a more effective treatment strategy for HNSCCs.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
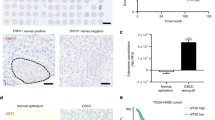
Head and neck cancers are among the top 10 most common cancers worldwide, and ~ 90% of diagnosed tumors are classified as head and neck squamous cell carcinomas (HNSCCs) that arise from the mucosal surfaces of the oral cavity, oropharynx, and larynx [1]. In the U.S., where overall cancer death rates declined during 2001–2018, HNSCC incidence and mortality continued to rise at an annual percentage change of 0.8 and 0.5, respectively [2]. The 5-year relative HNSCC survival among all races is 66.2% for females and 64.6% for males, below their respective national averages for 5-year relative survival for all cancers, i.e., 68.9% and 65.1%, respectively [3]. Heavy tobacco and alcohol use as well as infection with the human papillomavirus (HPV) are important risk factors for HNSCC.
Extracellular ATP (eATP) and extracellular adenosine (eADO) serve as biochemical signatures of cancer due to their significant concentrations in the tumor interstitium [4]. eATP hydrolysis is the putative major source of eADO, as eATP can be rapidly degraded by the CD39/CD73-axis of ectonucleotidases, and the ratio of these purines in the tumor microenvironment (TME) can be decisive for clinical outcomes [5]. Nucleotides in the TME serve as mediators of interactions between the tumor and host immune system, where eATP acts as a chemoattractant and eADO acts as an immunosuppressant [6]. Conventional cancer therapies, despite being able to damage tumor cells, also create an immunosuppressive microenvironment by inducing the release of intracellular nucleotides [7]. Additionally, extracellular nucleotides can act in an autocrine or paracrine manner and have been shown to regulate cell proliferation and migration, epithelial-to-mesenchymal transition, and angiogenesis. In addition, eATP is believed to promote tumor growth via activation of ERK1/2, which increases the activity of immunosuppressive regulatory CD4+ T cells [8,9,10].
On a cellular level, the effects of extracellular nucleotide signaling are due to the collective output of various purinoreceptors, ectoenzymes and transporters that form a receptosome known as the purinome [8, 11]. The main P2 receptors (P2Rs) of the purinome are P2 and P1 receptors for nucleotides and adenosine, respectively. P2 receptors are further divided on the basis of their agonist profile, and amino acid and 3-dimensional structure into P2Y G protein-coupled receptors (GPCRs) for extracellular purine and pyrimidine nucleotides and P2X ATP-gated ion channels [12]. This study focuses on the P2Y2 purinergic receptor (P2Y2R) subtype, a metabotropic GPCR that we demonstrated is equipotently activated by eATP and eUTP, which, when activated, leads to intracellular ERK1/2 activation [13]. P2Y2R is a cell surface nucleotide receptor in the purinome, whose upregulation has been associated with various cancers and inflammatory diseases [14, 15]. The purinome has been shown to regulate immune responses in part by inducing migration of immune cells to sites of inflammation, but also in the feedback regulation of immunosuppression through degradation of eATP to eADO [5]. P2Y2R has been studied in the context of multiple cancer types and has been shown to have diverse effects, from promoting epithelial-mesenchymal transition of gastric [16] and breast [17] cancer cells and enhancing glycolysis and cell growth in pancreatic ductal adenocarcinoma cells [18].
We have previously demonstrated that P2Y2R signaling influences the growth of xenograft tumors of human Cal27 and FaDu oral cancer cells in immunocompromised mice, which suggests that the P2Y2R could serve as a druggable target for HNSCC treatment [13]. However, the anti-tumor immune response cannot be discerned in immunocompromised mouse models. To circumvent this problem, we analyzed the expression in HNSCCs of molecules that comprise the purinome using published HNSCC datasets, and functionally characterized extracellular nucleotide responses of human and murine oral cancer cells with or without knockout of the P2Y2R. Subsequently, we evaluated tumor growth and the host anti-tumor response to wild-type and P2Y2R−/− murine MOC2 cells in a syngeneic, immunocompetent mouse model to investigate the role of purinergic signaling. Here, we report on the pleiotropic effects of the P2Y2R on tumor growth and the host anti-tumor response.
Materials and methods
All materials, reagents, and buffers used in this study are detailed and catalogued according to The Research Resource Identification Initiative.
Bioinformatic analysis of purinome gene expression
Data from The Cancer Genome Atlas (TCGA) Consortium’s PanCancer Atlas [19] or the Cancer Cell Line Encyclopaedia [20] were accessed through cBioPortal for Cancer Genomics [21, 22]. Purinome gene expression was queried in 515 available HNSCC patient tumors (72 HPV + , 415 HPV-, 28 not defined) and mRNA expression levels (calculated by cBioPortal using RNA-Seq by Expectation Maximization [23] and batch normalized from Illumina HiSeq RNA seq v2 were exported.
Cell culture and reagents
The mouse oral carcinoma (MOC2) (RRID:CVCL_ZD33) cell line was obtained from Dr. Ravindra Uppaluri (Dana Farber Cancer Institute, Boston, MA), verified using Mouse Cell STR Profiling (ATCC) and maintained in Iscove’s Modified Eagle’s Medium (IMEM)/F-12 (2:1) (Gibco) supplemented with 5% (v/v) FBS (Sigma), 100 units/ml penicillin (Gibco), 100 μg/ml streptomycin (Gibco), 400 ng/ml hydrocortisone (Sigma), 5 ng/ml EGF (Corning) and 5 μg/ml insulin (Gibco). MOC2 cells were confirmed to be free of mycoplasma contamination using the MycoAlert kit (Lonza). FaDu (RRID:CVCL_1218) human hypopharyngeal squamous cell carcinoma and Cal27 human tongue squamous cell carcinoma (RRID:CVCL_1107) cells were purchased from ATCC in 2019 and cultured in DMEM supplemented with 10% (v/v) FBS, 100 units/ml penicillin, 100 μg/ml streptomycin. Cells were cultured at 37°C with an atmosphere of 95% air and 5% CO2. All cell lines were used at a passage number ≤ 10 and under BSL-2 safety conditions. Nucleotides ATP, BzATP (2',3'-O-(4-benzoylbenzoyl ATP)), UTP, UDP, UDP-glucose, and 2-MeS-ADP (2-methylthioadenosine 5’-diphosphate) were purchased from Millipore Sigma. Fura-2 AM and the LV01 U6-gRNA:ef1a-puro-2A-Cas9-2A-tGFP plasmid containing guide RNA (gRNA) directed at the mouse P2ry2 (target ID MMPD0000028651) were purchased from Millipore Sigma. Specificity of gRNA was verified using CHOPCHOP [24]. AR-C118925 was purchased from Tocris. Rabbit anti-phospho-ERK1/2 (RRID:AB_331646), rabbit anti-total ERK1/2 (RRID:AB_390779), and horseradish peroxidase-linked goat anti-rabbit IgG (RRID:AB_2099233) antibodies were purchased from Cell Signaling Technology. Intracellular calcium ion flux assay buffer: 15mM HEPES, pH 7.4, 120mM NaCl, 5.4mM KCl, 1.2mM MgSO4, 1.2mM KH2PO4, 1mM CaCl2, 0.1% (w/v) albumin, and 0.18% (w/v) glucose. Digestion solution: 50µL of 50 mg/mL collagenase P (Roche), 50µL of 1 mg/mL DNase I (Invitrogen) in 5mL un-supplemented MOC media. Staining media: phenol red free RPMI-1640 (Gibco), 2% FBS, and 2mM EDTA. PEB: PBS, 0.5% (w/v) BSA (Sigma), and 2mM EDTA.
Real-Time PCR
RNA was isolated from MOC2 cells and MOC2 clones using the Qiagen RNeasy Mini kit and reverse transcribed using RNA to cDNA EcoDry Premix (Takara). Either 150ng, or 15ng of RNA was used per replicate for P2 receptors and 18S, respectively. RT-PCR was conducted using Applied Biosystems TaqMan probes for P2 receptors and 18S, where 18S was used as the internal control. Samples were run on an Applied Biosystems 7500 Real-Time PCR System according to the manufacturer’s protocol. The expression of targeted genes was normalized to 18S and quantified using the 2−ΔΔCt method.
Measurement of the intracellular Ca2+ concentration
Intracellular calcium ion flux measurements were performed on a Photon Technology International DeltaScan dual excitation spectrofluorometer controlled by the Felix software. Cells were incubated in assay buffer with 2µM Fura-2 AM at 37˚C for 45 minutes at 2 × 106 cells/mL. Pelleted cells were washed twice with 20mL of assay buffer then resuspended at 2 × 106 cells/mL in assay buffer and allowed to rest for 30 minutes. Prior to each assay, 1mL of Fura-2-loaded cells was spun down and the pellet was resuspended in 2mL of fresh 37˚C assay buffer. The cells were transferred to a pre-warmed cuvette with a magnetic stirrer and placed in the cuvette holder. After 90 seconds of equilibration, 20µL of agonist was added to a final concentration of 100µM. Agonist-induced changes in the 340/380nm fluorescence excitation ratio (505nm emission) were measured. Maximum and minimum fluorescence were measured 1 minute apart following the addition of 100µL of 2mM digitonin in ethanol and 100µL of 250mM EGTA in 2.5M Tris, pH 8, respectively. The intracellular Ca2+ concentration was calculated utilizing the Kd for the Ca2+/Fura-2 complex (224nM) and maximum and minimum fluorescence intensities at 340nm/505nm.
In vitro ERK1/2 phosphorylation assays
As previously described [13], cells were seeded in 12-well plates at 70,000 cells/well and allowed to adhere overnight. The culture media was then replaced with un-supplemented MOC media and cells were starved overnight. For assays involving AR-C118925, cells were pre-treated with 10µM AR-C118925 for 1 h. At time 0 minutes, UTP was added to each well at a final concentration of 100µM, except basal control wells. At each timepoint, the culture media was removed, and cells were lysed by addition of 300µL of warm 2X SDS Laemmli sample buffer.
SDS-PAGE and western blot analysis
As previously described [13], cell lysates were sonicated and boiled at 95˚C for 5 min and allowed to cool. Then, 25µL of lysate was loaded on to ExpressPlus 4–12% (Genscript) gradient gels alongside 10µL of Protein Plus Kaleidoscope marker (Bio-Rad). Gels were electrophoresed and transferred to nitrocellulose membranes utilizing the XCell II blotting system. Blots were rinsed in TBST (1X TBS, 0.1% (v/v) Tween-20) and blocked in 5% non-fat milk diluted in TBST for 1 hour at room temperature. Following blocking, primary antibody was added at 1:1,000 dilution to the blocking solution and incubated overnight at 4˚C. Blots were then washed 3 times with TBST for 10 minutes each and incubated with HRP-conjugated secondary antibody at 1:2,000 dilution in 1% non-fat milk/TBST for 1 hour at room temperature. Following secondary antibody incubation, blots were washed 3 times for 10 minutes each and developed using SuperSignal West Pico ECL (Pierce) and analyzed in LI-COR Image Studio.
Generation of P2Y2R-null cells using CRISPR/Cas9
MOC2 cells were seeded in a 6-well plate at 50,000 cells/well and the following day the media was removed and a solution of 4 µg plasmid DNA:10µL Lipofectamine 2000 in 500µL of serum-free MOC media was added per well. Following puromycin selection, LV01 GFP+ cells were sorted into 96-well plates on a MoFlo cell sorter (Beckman Coulter) at the University of Missouri Cell and Immunobiology Core. Single cell colonies are subsequently referred to as “MK” clones. The gDNA was extracted utilizing the Qiagen DNA Mini kit and a 331 bp portion of the P2ry2 gene was amplified utilizing NEB One Taq DNA polymerase with the following primers: LV01 Outer-F 5’-CTA CTT CGT CAC CAC CAG CGT GC-3' and LV01 Outer-R 5’-GTG GCA GCT GAG GTC AGG TGA TCG-3'. After confirming product size on a 1.5% (w/v) agarose gel, PCR product was prepared for Sanger sequencing analysis utilizing ExoSAP-IT (Invitrogen) and an internal set of oligonucleotide primers: LV01 Inner-F 5’-CCA TTT TGT GGC TTA CAG CTC CGT C-3' and LV01 Inner-R 5’-GCA GAG GGC GAA GAC GGC CAG-3'. Samples were run in half-reactions on a 3730xl 96-capillary DNA Analyzer with Applied Biosystems Big Dye Terminator cycle sequencing chemistry at the University of Missouri Genomics Technology Core. Sequences from both the forward and reverse reactions were utilized to confirm gene editing. MK mutants were functionally characterized utilizing assays for UTP-induced intracellular calcium ion flux and ERK1/2 phosphorylation, as described above. MK72 was selected based on a lack of UTP responses.
In vivo tumorigenesis assay
Male 8–10 week-old C57BL/6 J (RRID:IMSR_JAX:000664) and P2ry2 knockout (RRID:IMSR_JAX:009132) mice were purchased from The Jackson Laboratory and housed in the vivarium of The Christopher S. Bond Life Sciences Center at the University of Missouri on a 12 hour day/night cycle. At 10 weeks of age, mice were anesthetized intraperitoneally with Avertin (0.8 mg/g) dissolved in 1:40 tert-amyl alcohol:PBS. Fur was removed from both flanks and mice were injected with 100µL of 2 × 105 cells/mL in PBS (200,000 cells total) of both MOC2 and MK72 cells contralaterally. Tumor-bearing mice were inspected every other day for tumor growth and at 28 days post-inoculation, mice were euthanized by isoflurane overdose followed by cervical dislocation and tumors were resected. All experimental animal procedures were conducted in accordance with National Institutes of Health guidelines and were approved by the University of Missouri Animal Care and Use Committee.
Isolation and flow cytometry analysis of tumor-infiltrating leukocytes
Spleens were isolated and passed through a 40µM mesh filter. Isolated tumors were digested by mincing the tissue followed by incubation in 5 mL of digestion solution and shaken at 4˚C overnight. Digested tissue was then passed through a 70µm mesh filter followed by a 40µm mesh filter, which were washed each time with 5mL of staining media. Cells were then centrifuged at 500 × g for 5 minutes and CD45+ cells were isolated utilizing anti-CD45 magnetic beads (Miltenyi Biotec), as per the manufacturer’s instructions. Isolated leukocytes were then cultured for 6 hour in RPMI-1640 with 10% (v/v) FBS and 2µL/mL of Cell Activation Cocktail with Brefeldin A (BioLegend) at 37˚C in a 5% CO2 atmosphere. Following activation, cells were washed with staining media and stained for flow cytometry analysis.
Samples were resuspended in 100µL of PBS with 1:2,000 Zombie NIR viability dye (Biolegend) and 1:100 anti-mouse CD16/CD32 (BD Biosciences Cat# 553142, RRID: AB_394657) and incubated in the dark for 15 minutes at room temperature. Samples were washed with staining media and resuspended in 100µL of staining media with the extracellular marker antibodies against CD45 (Miltenyi Biotec Cat# 130–110-662, RRID:AB_2658231), CD3e (BioLegend Cat# 100328, RRID:AB_893318), CD4 (BioLegend Cat# 100467, RRID:AB_2734150), CD8a (BD Biosciences Cat# 563234, RRID:AB_2738084), CD11b (BioLegend Cat# 101216, RRID:AB_312799), F4/80 (Thermo Fisher Scientific Cat# 63–4801-82, RRID:AB_2723154), CD45R (Miltenyi Biotec Cat# 130–094-287, RRID:AB_10829022), Ly-6C (BD Biosciences Cat# 560592, RRID:AB_1727556), and Ly-6G (BioLegend Cat# 127624, RRID:AB_10640819) and incubated for 1 hour at 4˚C in the dark. The samples were washed with PEB followed by fixation and permeabilization with the True-Nuclear Transcription Factor Buffer Set (BioLegend) according to the manufacturer’s protocol. Following fixation and permeabilization, samples were incubated in 100µL of intracellular marker antibodies FoxP3 (Thermo Fisher Scientific Cat# 17–5773-82, RRID: AB_469457) and IFNγ (Thermo Fisher Scientific Cat# 11–7311-82, RRID: AB_465412) and incubated for 1 hour in the dark at room temperature. Samples were washed twice with 1mL of permeabilization buffer and resuspended in PEB for flow cytometry analysis. Unstained fixed samples were run as autofluorescence controls. Single stained compensation controls were generated using UltraComp eBeads (Invitrogen) and fluorescence minus one (FMO) controls were generated utilizing cells isolated from spleens for each fluorophore.
Data were acquired on a 4L Cytek Aurora spectral flow cytometer at the University of Missouri Cell and Immunobiology Core controlled by Cytek Spectroflo software. Spectral unmixing was performed in Spectroflo (v3.1) utilizing the single stained compensation spectra and the spectra of autofluorescence control samples from MOC2 tumors, MK72 tumors or spleens. Unmixed.fcs files were deposited at FLOWRepository under ID: FR-FCM-Z5H3.
Statistical analysis
Data were organized, statistical tests were performed, and figures were generated utilizing GraphPad (RRID:SCR_000306) (Prism). Western blot data were analyzed utilizing an unpaired t-test with Welch's correction. Tumorigenesis data normality was verified using a Shapiro–Wilk test and analyzed using either paired or unpaired parametric t-test, when appropriate. All statistical analyses were conducted against an alpha value of 0.05.
Results
Purinome gene expression and P2 receptor function in HNSCC tumors and cell lines
Analysis of RNA-Seq data from human HNSCC tumors compared to normal tissue using the TCGA PanCancer Atlas and cBioPortal informatics tools indicated that the expression of many purinergic mRNAs was elevated in HNSCC tumors, including mRNAs for hemichannels and gap junctions associated with eATP release (i.e., PANX1 (pannexin-1) and GJA1 (connexin 43)), P2X and P2Y receptors for extracellular ATP, UTP, ADP, UDP, UDP-glucose (i.e., P2RX1-7 and P2RY1,2,4,6,11–14), P1 receptors for eADO (i.e., ADORA1, ADORA2A, ADORA2B, ADORA3), ectonucleoside 5’-triphosphate diphosphohydrolase and ectonucleotide pyrophosphatase/phosphodiesterase that catalyze the stepwise breakdown of eATP to eADO (i.e., ENTPD1-8, ENPP1-4, NT5E), adenosine deaminases that catalyze conversion of adenosine to inosine (i.e., ADA, ADA2) and equilibrative nucleoside transporters that mediate cell reuptake of eADO (i.e., SLC29A1-3) (Fig. 1A and B). Genes involved in eATP release and catabolism, such as PANX1, ENTPD6 and NT5E, and the nucleoside transporter SLC29A1 (ENT1) were among the highest expressed purinome genes in HNSCC tumors. For P1 and P2 receptors, P2RY2 was the highest expressed, although P2RX4, P2RY1, P2RY6, and P2RY11 and ADORA2B were also expressed. Similarly, analysis of RNAseq data from the Cancer Cell Line Encyclopedia using cBioPortal informatics tools revealed that P2RY2 expression was highest among all P2X and P2Y receptors in human FaDu and Cal27 HNSCC cell lines (Figs. 2A and C). To functionally characterize P2R responses, we measured extracellular nucleotide-induced changes in the intracellular calcium ion concentration (Δ[Ca2+]i). As shown in Figs. 2B and D, both FaDu and Cal27 cells responded variably to distinct extracellular nucleotides. FaDu cells had the highest response to P2Y2R agonists ATP (240.8 nM peak) and UTP (269.4 nM peak), but also responded to UDP and BzATP, whereas Cal27 cells had the highest response to ATP (291.8 nM peak), but also responded to UTP, UDP, and BzATP. These data confirm that the P2Y2R is expressed in both human Cal27 and FaDu HNSCC cell lines, as in human HNSCC tumors (Fig. 1).
Purinergic landscape of HNSCC tumors. A Comparative analysis of purinome mRNA expression in human HNSCC tumors represented in TCGA Consortium’s PanCancer Atlas. B Extracellular purinome signal integration occurs through transport of nucleotides (e.g., ATP and UTP) via channels such as pannexins into the extracellular milieu leading to activation of P2XRs and P2YRs that produces intracellular second messengers via extracellular ion influx or G protein signaling, respectively. Extracellular nucleotides are degraded by ectoenzymes (i.e., ENTDPs, ENPPs) into nucleoside di- and mono-phosphates (e.g., ADP, AMP, UDP) products that can also activate P2YRs. Through the action of the ecto-5’-nucleotidase CD73, AMP is degraded to adenosine, which activates P1 adenosine receptors. Lastly, adenosine generated can enter cells via equilibrative nucleoside transporters (SLC29A1-3) and be broken down to inosine by adenosine deaminase (ADA)
P2Y receptor signalling in human head and neck cancer cell lines. Bioinformatic analysis of RNA-Seq data from the Cancer Cell Line Encyclopedia using cBioPortal reveals P2 receptor mRNA expression levels (calculated as reads per kilobase million; RPKM) in (A) FaDu and (C) Cal27 cell lines, where multiple P2R mRNAs are expressed, particularly P2RY2. Nucleotide-induced Δ[Ca2+]i in (B) FaDu and (D) Cal27 cells. Representative curves of Δ[Ca2+]i measured using the Ca2+-sensitive dye Fura-2 after adding 100 µM of the indicated nucleotide at t = 90 s and data were collected until a baseline was apparent
In mouse MOC2 oral carcinoma cells, we conducted real-time PCR to survey P2 receptor mRNA expression (Fig. 3A). Similar to FaDu and Cal27 cells, MOC2 cells highly expressed P2RY2, P2RX4, and P2RX7 receptor mRNA. Functional characterization of P2Rs in MOC2 cells showed similar activity profiles to Cal27 and FaDu cells, with maximum ATP- and UTP-induced changes in Δ[Ca2+]i being 429.3 nM and 504.4 nM, respectively (Fig. 3B). We also observed a Δ[Ca2+]i in response to the P2XR agonist BzATP (Fig. 3B). Treatment with the selective P2Y2R antagonist AR-C118925 inhibited UTP-induced Δ[Ca2+]i, demonstrating that P2Y2R mediates the UTP-induced Δ[Ca2+]i (Fig. 3C). To further characterize the kinetics of P2Y2R activation in MOC2 cells, since data for this cell line are lacking, we determined that the EC50 of P2Y2R-dependent, UTP-induced Δ[Ca2+]i was 3.68 µM (Fig. 3D) and the IC50 of AR-C118925 was 87.9 nM (Fig. 3E). Based on the well-established role of P2Y2Rs in eATP-induced ERK1/2 activation, we determined that UTP significantly increased ERK1/2 phosphorylation in MOC2 cells in a time-dependent manner with maximum phosphorylation occurring 5 min post-UTP addition (p = 0.0207) and basal levels returned within 60 min (Fig. 3F), similar to previous observations in human FaDu and Cal27 HNSCC cells [13]. In MOC2 cells, UTP-induced ERK1/2 phosphorylation was abolished by preincubation with 10 µM AR-C118925 (Fig. 3G). Taken together, these results indicate that both mouse and human HNSCC cell lines studied here express functional P2Y2Rs, and that overall P2R expression profiles are very similar in mouse MOC2 and human FaDu and Cal27 cells, suggesting that findings in mice may be directly relevant to HNSCCs in humans.
P2R mRNA profile and functional P2Y2R signaling in MOC2 cells. A Real-time PCR analysis of P2R expression in MOC2 cells. Data were normalized to 18S and relative to P2RY1. B Representative curve of nucleotide-induced Δ[Ca2+]i in MOC2 cells after addition of 100 µM nucleotide. C Representative curve of inhibition of UTP-induced Δ[Ca2+]i with 10 µM AR-C118925, a selective P2Y2R antagonist. D EC50 for UTP-induced Δ[Ca2+]i and E IC50 of AR-C118925-induced inhibition of UTP-induced Δ[Ca2+]i. F MOC2 cells were treated with or without 100 μM UTP for the indicated times, then subjected to immunoblot analysis using specific antibodies for phospho-ERK1/2 (Thr202/Tyr204) or total ERK1/2 as loading control. G MOC2 cells were pre-treated for 1 h with or without 10 μM AR-C118925 followed by 100 μM UTP for 5 min, then subjected to immunoblot analysis for phospho-ERK1/2 or total ERK1/2. Results are presented as means ± S.E.M. for n ≥ 3 independent experiments with n ≥ 3 technical replicates per experiment; where *, **, and **** indicate p < 0.05, p < 0.01, and p < 0.0001, respectively.
Genetic ablation of P2Y2R attenuates both UTP-induced Δ[Ca2+]i and ERK1/2 phosphorylation in MOC2 cells
To conclusively confirm the role of P2Y2R in UTP-induced MOC2 cell responses, we used MOC2 cells for CRISPR-Cas9-mediated generation of a P2Y2R-null clonal cell line named MK72 cells. Genomic DNA sequencing confirmed an 8-nucleotide deletion within the P2ry2 gene (Fig. 4A). Bioinformatic analysis using PSIPRED [25] and MemSATS [26] predicts that this deletion causes a premature stop codon, which truncates P2Y2R in intracellular loop 3 (Fig. 4B). In comparison to P2Y2R-expressing MOC2 cells from which MK72 cells were derived, these mutant P2Y2R−/− MK72 cells did not exhibit UTP-induced Δ[Ca2+]i (Fig. 4C), and UTP did not cause time-dependent changes in ERK1/2 phosphorylation (Fig. 4D), as seen in MOC2 cells (Fig. 3).
CRISPR-Cas9-mediated P2ry2 knockout prevents UTP-induced Δ[Ca2+]i and ERK1/2 phosphorylation in MK72 cells. A Chromatogram of Sanger sequencing and local alignment of P2ry2 nucleic acid sequences of wild-type (WT) P2ry2 in MOC2 cells and P2ry2 mutant generated in MK72 cells using CRISPR-Cas9; 8-nucleotide deletion highlighted in blue. B PSIPRED predicted P2Y2R amino acid residue sequence in WT MOC2 and MK72 cells with MemSAT predicted transmembrane domains in bold. Blue denotes first mismatched amino acid residue. C UTP-induced Δ[Ca2+]i in MOC2 (black trace) and MK72 (red trace) cells. D MK72 cells were treated with or without 100 μM UTP for the indicated times, then subjected to immunoblot analysis using specific antibodies for phospho-ERK1/2 (Thr202/Tyr204) or total ERK1/2 as loading control. Results are presented as means ± S.E.M. for n ≥ 3 independent experiments with n ≥ 3 technical replicates per experiment
Genetic ablation of P2Y2R reduces growth of syngeneic MOC2 tumor grafts
We have previously demonstrated that systemic administration of AR-C118925 slowed MOC2 tumor growth in vivo [13]. To determine whether P2Y2R ablation affects MOC2 tumor growth, we contralaterally implanted wild-type MOC2 or P2Y2R−/− MK72 cells into C57BL/6 J and P2Y2R−/− mice and at 28 days post-implantation tumors were excised and weighed. As shown in Fig. 5A, there was a statistically significant decrease in tumor weight of P2Y2R−/− MK72 tumors compared to the wild-type parental MOC2 tumors in both C57BL/6 J (p = 0.0104) or P2Y2R−/− mice (p = 0.0055). In Fig. 5B, we depict paired contralateral MOC2 and MK72 tumors generated in wild-type C57BL/6 J or P2Y2R−/− mice to visualize the size differences.
Tumoral P2Y2R deletion reduces tumor growth in wild-type and P2Y2R−/− mice. A Tumor weights of MOC2 and MK72 tumors isolated from wild-type C57BL/6 and P2Y2R−/− mice with accompanying representative images of in vivo tumor growths. Results are presented as means ± S.E.M. for n ≥ 3 more independent experiments with n ≥ 3 technical replicates per experiment. B Representative image of paired contralateral MOC2 and MK72 tumors grown in wild type C57BL/6 or P2Y2R−/− mice
Tumoral P2Y2R alters the host immune response
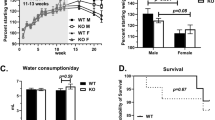
To study the host anti-tumor response, we isolated tumor-infiltrating immune cells as well as splenic immune cells and performed immunophenotyping using spectral flow cytometry (Fig. 6A). In both C57BL/6 J and P2Y2R−/− mice, MK72 tumors contained significantly increased levels of F4/80+ macrophages (Fig. 6B; wild-type p < 0.0001; P2Y2R−/− p = 0.0002) and CD3+ T cells (Fig. 6C; wild-type p = 0.0091; P2Y2R−/− p = 0.0021), as compared to MOC2 tumors. Within the T cell population, we observed statistically significant decreases in the percentage of CD3+CD4+ T helper cells in MOC2 tumors compared to MK72 tumors (Fig. 6D) (wild-type p = 0.0090; P2Y2R−/− p = 0.0290).
Genetic ablation of tumoral P2Y2R modulates the host anti-tumor response in C57BL/6 and P2Y2R−/− mice. A Representative gating strategy used for flow cytometry analysis to identify tumor-infiltrating immune cells. B-D Subpopulations of tumor-infiltrating immune cell types isolated from MOC2 and MK72 tumors of wild-type C57BL/6 and P2Y2R−/− mice. E MOC2 and MK72 tumors were isolated from C57BL/6 J mice 28 days after cell implantation. Tumor mRNA was extracted utilizing the standard TRIzol method followed by column purification utilizing the RNeasy mini kit from Qiagen. Real-time PCR was conducted as previously described. Data shown are relative quantifications normalized to 18S. Data represented are means ± S.E.M. for n ≥ 3 independent tumors, where * and ** indicate p < 0.05 and p < 0.01, respectively
Discussion
The tumor microenvironment (TME) is an, immune-privileged site of host immune and stromal tumor cell interactions that favors tumor growth. As a consequence of this TME, only ~ 20% of patients with solid head and neck tumors respond to immune checkpoint inhibitors [27]. Extracellular ATP (eATP) has been shown to have both immunosuppressive and immunostimulatory effects in the TME mainly due to its activation of both eATP- and eADO-binding purinergic receptors on both host and tumor cells [28, 29]. Additionally, eATP has been implicated in chemotherapeutic drug resistance and immunogenic cell death through alterations in the ratio of eATP and eADO levels catalyzed by CD39 and CD73 ecto-5’-nucleotidases [11]. In healthy tissues, eATP concentrations are maintained in the high nM to low µM range, while within the TME eATP concentrations have been reported to range from 0.1—1mM [4, 30]. Indeed, whereas ATP is normally thought of as an intracellular energy source, there exists a multicomponent, regulated system for eATP release and extracellular metabolism involving ion transporters and ecto-5’-nucleotidases in addition to P1 and P2 receptors that regulate tissue- and cell-specific responses mediated by binding of eADO and eATP, eUTP and other nucleotides, respectively (i.e., the purinome; Fig. 1). Previously, we observed that global pharmacological antagonism of the P2Y2R slowed MOC2 tumor growth in C57BL/6 J mice [13], which was hypothesized to result from blockade of tumoral rather than host P2Y2Rs. To address this hypothesis, the current study represents the first time that pleotropic effects of both wild-type and P2Y2R−/− tumors on the anti-tumor immune response have been investigated in both wild-type and P2Y2R−/− mice.
MOC2 cells were originally isolated from a 7,12-dimethylbenz(a)anthracene (DMBA)-induced oral tumor and have been previously described as a mouse model of oral squamous cell carcinoma (OSCC) [31]. MOC2 and MOC2-derived clonal cell lines MOC2-7 and MOC2-10 were demonstrated to possess a highly conserved genetic landscape and transcriptional signature comparable to human OSCCs [32]. Additionally, as highlighted by Kono et al. (2022) [33] MOC cells have been used extensively to study HNSCC for over 10 years. Using publicly available datasets accessed through cBioPortal, we determined the gene expression profiles of purinome components in 515 human HNSCC tumors (Fig. 1A) and found that genes involved in the initiation and termination of extracellular purinergic signaling (i.e., eATP release channels, ENTPDiphosphohydrolases, ecto-nucleotidases and ecto-nucleoside transporters) were generally more expressed than cell surface P1 and P2 purinergic receptors. However, the P2Y2R was the most expressed P2 receptor within the HSNCC dataset. Similarly, we determined that P2Y2R was the most expressed P2Y receptor in FaDu, Cal27, and MOC2 cell lines (Figs. 2 and 3), highlighting the utility of these cell lines for comparing HNSCC purinergic responses between mice and human cells in vitro and in vivo. Nonetheless, multiple P2 receptor subtypes are expressed in both HNSCC tumor samples and OSCC cell lines of human and murine origin indicating the complexity of P2R signaling in HNSCCs. Functional P2Y2R responses, such as increased Δ[Ca2+]i and ERK1/2 phosphorylation induced by the P2Y2R-selective agonist UTP, were observed in all 3 mouse and human HNSCC cell lines and were abolished by genetic ablation of P2Y2R expression or pharmacological inhibition with the selective P2Y2R antagonist AR-C118925 (Figs. 2, 3, and 4) [13]. As previously mentioned, purinergic signaling is the cumulative outcome of various purinome components (Fig. 1B) and here we show that a major part of the purinergic response in HNSCC is due to P2Y2R.
Hyperactivation of ERK1/2 is a common mechanism used by cancer cells to accelerate growth and has been demonstrated to be crucial for the aggressiveness and immune evasion of MOC2 cells [31]. To determine the contribution of tumoral P2Y2Rs to in vivo tumorigenesis, we used MOC2 cells to generate MK72 cells, a CRISPR-Cas9-mediated P2Y2R−/− clonal cell line (Fig. 4) and observed that loss of tumoral P2Y2R reduces tumor growth (Fig. 5) and leads to an altered host anti-tumor response independent of host P2Y2R (Fig. 6). P2Y2R has been shown to promote the growth of various tumors and cancer cell models via a plethora of mechanisms that range from a role in EGFR transactivation, ERK1/2 activation [13], calcium ion signaling to hyperactivation of glycolysis, and increased DNA damage [18, 34, 35]. Here, we show that P2Y2R−/− MK72 tumors have a greater infiltration of phagocytic CD11b+F4/80+ macrophages and CD3+CD4+ T cells than wild-type tumors. The combination of IFNγ secretion and macrophage infiltration are characteristic of an anti-tumoral Th1/M1 macrophage response [36]. This was confirmed by increased expression of monocyte chemotactic protein-1 (MCP-1/CCL2) and IFNγ in MK72 tumors from wild-type mice (Fig. 6E). Additionally, IFNγ secretion by CD4+ T cells is responsible for priming CD8+ cytotoxic T cells as part of the adaptive immune response [37, 38]. A significant increase in infiltration of CD11b+F4/80+ and CD4+ T cells was observed when comparing MOC2 tumors to the more indolent MOC1 clone tumors in wild-type mice [39]. This suggests that P2Y2R−/− MK72 tumors are more related to the MOC1 indolent tumor than the more aggressively growing MOC2 tumors, which may be due to the presence of immunosuppressive pro-tumor CD4+ regulatory T cells in MOC2 tumors. Taken together, these data indicate that modulation of the host anti-tumor immune response may represent a novel mechanism whereby P2Y2Rs regulate tumorigenesis.
Although much literature focuses on the role of P2Y2R expressed in host immune cells [40, 41], our data demonstrate that there is also a role for tumoral P2Y2R in the regulation of host immune cell responses in the TME, suggesting that targeted P2Y2R antagonism at the tumor site might be more effective than systemic P2Y2R antagonism in the treatment of HNSCC. Further research is needed to determine the contributions of other purinome components expressed in HNSCCs to the regulation of immune responses in the TME, and discrete roles of eATP and other extracellular nucleotides and their metabolites in host anti-tumor immune responses.
Data availability
The flow cytometry data underlying this article are available as .fcs files deposited at FLOWRepository under ID: FR-FCM-Z5H3. All other data underlying this article will be shared on reasonable request to the corresponding author.
References
Vigneswaran N, Williams MD (2014) Epidemiologic trends in head and neck cancer and aids in diagnosis. Oral Maxillofac Surg Clin North Am 26(2):123–141
Islami F et al (2021) Annual Report to the Nation on the Status of Cancer, Part 1: National Cancer Statistics. JNCI: J Natl Cancer Instit 113(12):1648–1669
U.S. Cancer Statistics Working Group (2023) U.S. Cancer Statistics Data Visualizations Tool, based on 2022 submission data (1999–2020): U.S. Department of Health and Human Services, Centers for Disease Control and Prevention and National Cancer Institute. https://www.cdc.gov/cancer/dataviz
Pellegatti P et al (2008) Increased Level of Extracellular ATP at Tumor Sites: In Vivo Imaging with Plasma Membrane Luciferase. PLoS ONE 3(7):e2599
de Andrade Mello P, Coutinho-Silva R, Savio LEB (2017) Multifaceted Effects of Extracellular Adenosine Triphosphate and Adenosine in the Tumor-Host Interaction and Therapeutic Perspectives. Front Immunol 8:1526
Feng LL et al (2020) The yin and yang functions of extracellular ATP and adenosine in tumor immunity. Cancer Cell Int 20:110
Mao C et al (2022) Delivery of an ectonucleotidase inhibitor with ROS-responsive nanoparticles overcomes adenosine-mediated cancer immunosuppression. Sci Trans Med 14(648):eabh1261
Woods LT et al (2021) P2Y receptors for extracellular nucleotides: Contributions to cancer progression and therapeutic implications. Biochem Pharmacol 187:114406
Jasmer KJ, Gilman KE, Muñoz Forti K, Weisman GA, Limesand KH (2020) Radiation-induced salivary gland dysfunction: mechanisms, therapeutics and future directions. J Clin Med 9(12):4095. https://doi.org/10.3390/jcm9124095
Alvarez CL, Troncoso MF, Espelt MV (2022) Extracellular ATP and adenosine in tumor microenvironment: Roles in epithelial-mesenchymal transition, cell migration, and invasion. J Cell Physiol 237(1):389–400
Volonte C, D’Ambrosi N (2009) Membrane compartments and purinergic signalling: the purinome, a complex interplay among ligands, degrading enzymes, receptors and transporters. FEBS J 276(2):318–329
Khalafalla MG, Woods LT, Jasmer KJ, Forti KM, Camden JM, Jensen JL, Limesand KH, Galtung HK, Weisman GA (2020) P2 receptors as therapeutic targets in the salivary gland: from physiology to dysfunction. Front Pharmacol 11:222. https://doi.org/10.3389/fphar.2020.00222
Woods LT et al (2020) P2Y2 receptors mediate nucleotide-induced EGFR phosphorylation and stimulate proliferation and tumorigenesis of head and neck squamous cell carcinoma cell lines. Oral Oncol 109:104808
Velázquez B et al (2000) Differential agonist-induced desensitization of P2Y2 nucleotide receptors by ATP and UTP. Mol Cell Biochem 206(1/2):75–89
Haystead TA (2006) The purinome, a complex mix of drug and toxicity targets. Curr Top Med Chem 6(11):1117–1127
Reyna-Jeldes M, De la Fuente-Ortega E, Cerda D, Velázquez-Miranda E, Pinto K, Vázquez-Cuevas FG, Coddou C (2021) Purinergic P2Y2 and P2X4 receptors are involved in the epithelial-mesenchymal transition and metastatic potential of gastric cancer derived cell lines. Pharmaceutics 13(8):1234. https://doi.org/10.3390/pharmaceutics13081234
Qiu Y et al (2018) P2Y2 receptor promotes the migration and invasion of breast cancer cells via EMT-related genes Snail and E-cadherin. Oncol Rep 39(1):138–150
Hu LP et al (2019) Targeting Purinergic Receptor P2Y2 Prevents the Growth of Pancreatic Ductal Adenocarcinoma by Inhibiting Cancer Cell Glycolysis. Clin Cancer Res 25(4):1318–1330
Hoadley KA et al (2018) Cell-of-Origin Patterns Dominate the Molecular Classification of 10,000 Tumors from 33 Types of Cancer. Cell 173(2):291-304 e6
Ghandi M et al (2019) Next-generation characterization of the Cancer Cell Line Encyclopedia. Nature 569(7757):503–508
Cerami E et al (2012) The cBio cancer genomics portal: an open platform for exploring multidimensional cancer genomics data. Cancer Discov 2(5):401–404
Gao J et al (2013) Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci Signal 6(269):pl1
Li B, Dewey CN (2011) RSEM: accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinformatics 12:323
Labun K et al (2019) CHOPCHOP v3: expanding the CRISPR web toolbox beyond genome editing. Nucleic Acids Res 47(W1):W171–W174
McGuffin LJ, Bryson K, Jones DT (2000) The PSIPRED protein structure prediction server. Bioinformatics 16(4):404–405
Aleksandrova AA, Sarti E, Forrest LR (2020) MemSTATS: A Benchmark Set of Membrane Protein Symmetries and Pseudosymmetries. J Mol Biol 432(2):597–604
Dogan V et al (2018) Current studies of immunotherapy in head and neck cancer. Clin Otolaryngol 43(1):13–21
Cekic C, Linden J (2016) Purinergic regulation of the immune system. Nat Rev Immunol 16(3):177–192
Draganov D, Lee PP (2021) Purinergic Signaling Within the Tumor Microenvironment. Adv Exp Med Biol 1270:73–87
Sorensen CE, Novak I (2001) Visualization of ATP release in pancreatic acini in response to cholinergic stimulus Use of fluorescent probes and confocal microscopy. J Biol Chem 276(35):32925–32
Judd NP et al (2012) ERK1/2 regulation of CD44 modulates oral cancer aggressiveness. Cancer Res 72(1):365–374
Onken MD et al (2014) A Surprising Cross-Species Conservation in the Genomic Landscape of Mouse and Human Oral Cancer Identifies a Transcriptional Signature Predicting Metastatic Disease. Clin Cancer Res 20(11):2873–2884
Kono M et al (2022) The mouse oral carcinoma (MOC) model: A 10-year retrospective on model development and head and neck cancer investigations. Oral Oncol 132:106012
Schulien I et al (2020) Extracellular ATP and Purinergic P2Y2 Receptor Signaling Promote Liver Tumorigenesis in Mice by Exacerbating DNA Damage. Cancer Res 80(4):699–708
Xie R et al (2014) The P2Y2 nucleotide receptor mediates the proliferation and migration of human hepatocellular carcinoma cells induced by ATP. J Biol Chem 289(27):19137–19149
Duan Z, Luo Y (2021) Targeting macrophages in cancer immunotherapy. Signal Transduct Target Ther 6(1):127
Green AM, Difazio R, Flynn JL (2013) IFN-gamma from CD4 T cells is essential for host survival and enhances CD8 T cell function during Mycobacterium tuberculosis infection. J Immunol 190(1):270–277
Nakiboneka R et al (2019) Interferon gamma (IFN-gamma) negative CD4+ and CD8+ T-cells can produce immune mediators in response to viral antigens. Vaccine 37(1):113–122
Judd NP et al (2012) Comparative analysis of tumor-infiltrating lymphocytes in a syngeneic mouse model of oral cancer. Otolaryngol Head Neck Surg 147(3):493–500
Campos-Contreras ADR, Díaz-Muñoz M, Vázquez-Cuevas FG (2020) Purinergic signaling in the hallmarks of cancer. Cells 9(7):1612. https://doi.org/10.3390/cells9071612
Ferrari D, Malavasi F, Antonioli L (2017) A Purinergic Trail for Metastases. Trends Pharmacol Sci 38(3):277–290
Acknowledgements
The authors would like to thank Mr. Daniel Jackson (University of Missouri, Cell and Immunobiology Core) and Dr. Joel Crespo (Cytek) for their assistance and consultation with spectral flow cytometry.
Funding
This work was supported by grant R01-DE007389 from the National Institute of Dental and Craniofacial Research and a TRIUMPH grant from the University of Missouri School of Medicine. KMF was supported by a Life Sciences Fellowship and The Wayne L. Ryan Fellowship from the University of Missouri and The Ryan Foundation, respectively.
Author information
Authors and Affiliations
Contributions
KMF, LTW, JMC, and KJJ were responsible for the experimental design. LTW was responsible for bioinformatic analysis. KMF was responsible for cell culture and animal experiments. KMF and KJJ were responsible for flow cytometry analysis. KMF and LTW were responsible for data curation and generating figures. GAW was responsible for securing funding and supervising data collection. All authors contributed to the writing and review of the manuscript.
Corresponding author
Ethics declarations
Ethical approval
All animal studies conducted in this work were reviewed and approved by the University of Missouri’s Animal Care and Use Committee (ACUC).
Competing interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Forti, K.M., Woods, L.T., Jasmer, K.J. et al. Tumoral P2Y2 receptor modulates tumor growth and host anti-tumor immune responses in a syngeneic murine model of oral cancer. Purinergic Signalling 20, 359–370 (2024). https://doi.org/10.1007/s11302-023-09960-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11302-023-09960-z