Abstract
Recently, there is an increased interest in the function of polyamine (PA) catabolism during fruit ripening, but little is known about its role during the ripening of peach (Prunus persica L.). In this study, the contents of both free and conjugated PAs markedly decreased as the peach fruit ripened. However, RNA-seq analysis showed that the transcript levels of PA synthesis–related genes increased, suggesting that the decreasing PA content during peach ripening was associated with PA catabolism. To better understanding the function of PA catabolism in peach ripening, genes encoding potential polyamine oxidases (PAOs), which are involved in PA catabolism, were identified across the peach genome. Four putative PAO genes (PpePAO1-PpePAO4) were identified in peach using the recently released genome database. The mRNA level of PpePAO1 was significantly increased during peach fruit ripening. Detection of free PAs in tobacco leaves transiently over-expressing PpePAO1 suggested that this gene is probably involved in terminal catabolism of PA. Treatment with the PAO inhibitor guazatine significantly reduced ethylene production and flesh softening of peach fruit, decreased the expression levels of fruit ripening-related genes, and significantly decreased the expression level of PpePAO1 and PAO activity, while PA contents were dramatically higher compared to control treatments. Our results suggests that PpePAO1-mediated PA catabolism is associated with decreased PA content during peach fruit ripening. These data provide valuable knowledge for better understanding the roles of PA catabolism in peach development and ripening.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Peach (Prunus persica L.) is one of the important fruits worldwide. However, peach fruits are highly perishable and deteriorate after ripening. This fruit ripening is primarily regulated by the phytohormone ethylene. The auxin IAA helps to stimulate the synthesis of ethylene in peach (Tatsuki et al. 2013; Pan et al. 2015). In addition, some transcription factors (TFs) also play important roles in the regulation of fruit ripening, including NAC transcription factors that affect fruit ripening through ethylene and other hormone pathways. The ethylene signaling pathway includes ethylene-induced NAC TFs that regulate fruit ripening, NAC TFs that regulate TFs upstream in the ethylene signaling network, and TFs that directly regulate the main ethylene synthesis genes to control fruit maturity (Ma et al. 2014; Zhu et al. 2014; Gao et al. 2018). In peach, the NAC gene ppa008301m (PpeNAC72) co-segregated with the maturity date locus, suggesting that this gene may control ripening time in peach (Pirona et al. 2013). Recently, the ethylene response factor (ERF) was found to regulate ABA biosynthesis by activating or repressing transcription of the NAC transcription factor PpeNCED2/3 during fruit ripening in peach (Wang et al. 2019).
Recent studies have shown that PAs serve as endogenous anti-senescence agents. PAs are small molecules such as putrescine (Put), spermidine (Spd), and spermine (Spm) that carry at least two amino groups (more details below). PAs may interact with plant hormones, including ethylene, auxin, and ABA, to regulate fruit development and ripening (Alcazar et al. 2010; Cui et al. 2010; Parra-Lobato and Gomez-Jimenez 2011). Bregoli et al. (2002) reported that exogenous PAs affect ethylene emission and flesh firmness in peach. Liu et al. (2007) reported the possible role of PAs in peach development and discussed the relationship between PAs and ethylene biosynthesis. It has been more recently reported that exogenous application of PAs affect peach fruit development and ripening (Torrigiani et al. 2012). Generally, PAs are usually found at abundant levels during early stages of fruit development, but at relatively low levels in mature fruits (Liu et al., 2006; Gomez-Jimenez et al. 2010; Agudelo-Romero et al. 2013).
Polyamines (PAs) are low molecular weight aliphatic cations that are found in almost all living organisms (Cona et al. 2006). PAs are involved in various physiological and developmental processes, including plant growth, fruit development and ripening, and abiotic and biotic stress responses (Bregoli et al. 2002; Torrigiani et al. 2012; Guo et al. 2018; Alcazar et al. 2010; Hatmi et al. 2013). Putrescine (Put), spermidine (Spd), and spermine (Spm) are the most common PAs in plants. In addition, some less abundant PAs, including cadaverine (Cad), thermospermine (Therm-Spm), norspermidine (Nor-Spd), and norspermine (Nor-Spm), have been reported in plants. The intercellular concentrations of PAs are decided by the dynamic balance between PA biosynthesis and PA catabolism. In plants, PA biosynthesis starts with the formation of Put from ornithine, which is mediated by arginine decarboxylase (ADC) (Alcazar et al. 2010). The transitions from Put to Spd and Spd to Spm are mediated by Spd synthase (SPDS) and Spm synthase (SPMS), respectively. The Spd and Spm biosynthesis reactions require one or two aminopropyl moieties, which are produced through decarboxylation of S-adenosylmethionine (SAM) by S-adenosylmethionine decarboxylase (SAMDC). SAM is also a precursor of ethylene production, suggesting a competitive relationship between PA and ethylene biosynthesis.
PA catabolism is mediated by two kinds of amine oxidases, diamine oxidase (DAO) and polyamine oxidase (PAO). DAOs are copper-containing enzymes and usually named copper-containing amine oxidases (CuAOs). Generally speaking, CuAOs catalyze the degradation of Put and Cad, producing 4-aminobutanal, H2O2, and ammonia (Alcazar et al. 2010). However, some studies reported that some CuAOs in Arabidopsis thaliana are also involved in the catabolism of Spd (Planas-Portell et al. 2013). The most studied CuAOs are the Arabidopsis thaliana CuAOs. Five AtCuAOs have been identified at the protein level in Arabidopsis (Møller and McPherson 1998; Planas-Portell et al. 2013; Ghuge et al. 2015; Groβ et al. 2017). CuAOs have also been reported in pea (Tipping and McPherson 1995), chickpea (Rea et al. 1998), tobacco (Naconsie et al. 2014), apple (Zarei et al. 2015), and sweet orange (Wang et al. 2017). In contrast to CuAOs, PAOs are divided into two groups, according to whether they function in PA terminal catabolism or PA back-conversion reactions. The first group of PAOs catalyze the breakdown of Spd and Spm, with the generation of 4-aminobutanal, 1,3-diaminopropane (Dap), and H2O2 (Cona et al. 2006). The second group of PAOs catalyze the back-conversion of Spm to Spd or Spd to Put with the concomitant generation of H2O2 (Moschou et al. 2012). So far, PAOs have been identified in many plant species, such as maize (Cervelli et al. 2000), barley (Smith 1972; Cervelli et al. 2001), tobacco (Yoda et al. 2006), poplar (Tuskan et al. 2006), Arabidopsis (Fincato et al. 2011), rice (Ono et al. 2012), sweet orange (Wang and Liu 2015, 2016), Brachypodium distachyon (Takahashi et al. 2018), upland cotton (Cheng et al. 2017), and tomato (Hao et al. 2018). Although the first PAO genes identified to be involved in PA terminal catabolism is from barley (HvPAO1 and HvPAO2) (Smith 1972), the best-studied gene of the first group is the maize PAO gene (ZmPAO) (Cona et al. 2006). Subsequently, the PAO genes from rice (OsPAO7), sweet orange (CsPAO4), and B. distachyon (BdPAO2) were also found to be involved in PA terminal catabolism (Liu et al. 2014; Wangand Liu 2016; Takahashi et al. 2018). In contrast, most PAO genes that have been reported belong to the second group and catalyze PA back-conversion reactions. There are five PAO genes in Arabidopsis (AtPAO1-AtPAO5) and all of them catalyze the back-conversion reactions of PAs (Tavladoraki et al. 2006; Moschou et al. 2008; Kamada-Nobusada et al. 2008; Fincato et al. 2011; Ahou et al. 2014). So far, seven PAO genes have been reported in the rice genome, and four of them (OsPAO1, OsPAO3, OsPAO4, and OsPAO5) have been shown to catalyze back-conversion reactions of PAs (Ono et al. 2012; Liu et al. 2014). More recently, some PAO genes in sweet orange (Wang and Liu 2015) and upland cotton (Cheng et al. 2017) were reported to be involved in the PA back-conversion pathway.
The role of PA catabolism in fruit ripening has been poorly studied. Agudelo-Romero et al. (2013) reported that the PA content dramatically decreased during grape ripening, while the levels of CuAO and PAO transcripts were markedly upregulated, suggesting that the increased oxidation of PA leads to the decreased PA levels. Furthermore, perturbation of PA catabolism by treatment with a PAO inhibitor affects cell expansion and aroma development during grape fruit ripening (Agudelo-Romero et al. 2014). Recently, the genes involved in PA biosynthesis and catabolism are found highly expressed during the fast growing and ripening stages of tomato (Tsaniklidis et al. 2016). These studies suggest that PA catabolism plays important roles in fruit ripening.
Understanding how the peach fruit ripens can lead to better control of the deterioration common during storage. It is just coming to light how PAs interact with plant hormone signaling, especially influencing how ethylene, during fruit ripening in peach. This study was undertaken to learn more about the function of PA catabolism in peach ripening. RNA-seq analysis suggested that the decrease in the levels of free PAs was associated with PA catabolism during peach fruit ripening. The genome-wide identification of PAO genes identified four putative PAO genes (PpePAO1-PpePAO4) in the peach genome. Analysis of free PAs in transgenic tobacco leaves suggested that PpePAO1 and PpePAO4 were involved in PA terminal catabolism, while PpePAO2 was involved in PA back-conversion. The function of PAO-mediated PA catabolism during peach fruit ripening was also investigated using guazatine treatment, a potent inhibitor of PAO. The result showed that guazatine treatment significantly reduced ethylene production and flesh softening of peach fruit. Our results suggested that PpePAO1-mediated PA catabolism was significantly increased during peach fruit ripening, which probably led to the decreased PA content. Furthermore, PAO-mediated PA catabolism affected the expression of fruit ripening–related genes. These data provide valuable knowledge for better understanding the roles of PAO-mediated PA catabolism during peach ripening.
Materials and methods
Plant materials and treatments
The experiment was carried out on 5-year-old Prunus persica (L.) Batch cv. ‘Huangshuimi’ and 6-year-old Prunus persica (L.) Batch cv. ‘Okubo’ trees that were grown at the experimental farm of Henan Agricultural University, China. Fruit diameter was measured at weekly intervals on 200 fruits from 20 days after full bloom (dAFB) to establish the fruit growth curve in cv. Huangshuimi. The growth pattern was evaluated using the first derivative of the diameter. Sixty fruits with homogeneous diameter, size, color, and position on the plant were selected for treatments. For treatment with a PAO inhibitor, guazatine acetate (J & K Chemical, China) containing a surfactant was diluted in distilled water and applied at the concentration of 1 mM and 5 mM. Blank control was performed by applying Tween 20 (0.5 mL/L), while positive control was performed with mixture of 1 mM Spd and Tween 20. The guazatine acetate and Spd were applied at 68 days (the second half of S3 stage) and 82 days (S3/S4 stage) after full bloom (dAFB) in cv. ‘Huangshuimi,’ respectively. For cv. ‘Okubo,’ the guazatine acetate and Spd were applied at 130 days after full bloom (20 days before commercial harvest). The harvest index was determined as follows: twenty control and treated fruits of ‘Huangshuimi’ peach were collected at 75 (late S3), 82 (S3/S4), and 89 dAFB (S4, commercial harvest), while twenty control and treated fruits of ‘Okubo’ peach were collected at 150 dAFB (commercial harvest), respectively. Ethylene production and fruit quality traits were determined using the whole fresh fruit. For polyamine content analysis and gene expression analysis, ten untreated fruits of ‘Huangshuimi’ peach of similar diameter, size, color, and position on the plant were collected at 28 (S1), 43 (S2), 53 (S2/S3), 68 (S3), 82 (S3/S4), and 89 dAFB (S4, commercial harvest). All the samples were quickly frozen using liquid nitrogen and stored at − 80 °C until use. For each experiment, at least three biological samples were collected for further analysis.
Identification and sequence analysis of peach PAOs
To identify the putative PAO genes in peach, Arabidopsis PAOs (AtPAO1-AtPAO5) were used to search against the peach genome database (https://www.rosaceae.org/). Multiple sequence alignment of the peach PAO genes was performed with Genedoc program (version 2.7). Gene structure of the peach PAO genes was analyzed by the Gene Structure Display Server (http://gsds.cbi.pku. edu.cn/). The molecular weights (MVs) and isoelectric points (pIs) of the peach PAO proteins were identified by ExPasy (http://web.expasy.org/compute_pi/), while the conserved and potential motifs of peach PAOs were identified by MEME tool (http://meme-suite.org/tools/meme). Phylogenetic relationships of plant PAOs were analyzed using MEGA 4.1 software by the neighbor-joining (NJ) method (Saitou and Nei 1987). The cis-elements of the promoter regions of each PpePAO were predicted by the Plant Care program (http://bioinformatics.psb.ugent.be/webtools/plantcare/ html/).
RNA-seq analysis
For RNA-seq analysis, the mesocarp of ‘Huangshuimi’ peach was collected at 68 (S3), 82 (S3/S4), and 89 dAFB and quickly frozen in liquid nitrogen. Total RNA was extracted using the Spin Column Plant Total RNA Purification Kit (Sangon, Shanghai, China) according to the manufacturer’s instructions. RNA quantity and quality (purity and integrity) were analyzed using a NanoPhotometer spectrophotometer (IMPLEN, Westlake Village, CA, USA) and an Agilent Bioanalyzer 2100 system (Agilent Technologies, CA, USA), respectively. The mRNA was purified from the total RNA using poly-T oligo-attached magnetic beads and then was broken into short fragments of approximately 200 bp. The fragments were used to synthesize first-strand cDNA using random oligonucleotides and SuperScript II. The library fragments were purified with the AMPure XP system (Beckman Coulter, USA). Eligible cDNAs were selected for PCR amplification, which was performed with Phusion® High-Fidelity DNA polymerase, Universal PCR primers, and Index (X) Primer. Raw reads (primary cDNA) were produced using the Illumina HiSeq 2500 platform by BGI Tech (Shenzhen, China). A total of 138.60 GB data were generated. The reads containing the adapter and poly-N as well as low-quality reads were removed to obtain clean reads. In addition, Q20, Q30, GC-content, and sequence duplication values of the clean reads were calculated. The summary statistics of assembly are shown in Suppl. Table 1. The trimmed reads were aligned to the Prunus persica genome (https://www.rosaceae.org/organism/24333). Quantification of gene expression levels was estimated as fragments per kilobase of transcript per million fragments mapped (FPKM).
Real-time qRT-PCR analysis
To investigate the tissue-specific expression profiles of the PpePAO genes, flowers (including pistil, stamen, petal, and calyx), tender leaves (10 dAFB), old leaves (40 dAFB), and young fruits (20 dAFB) were collected, and total RNA was extracted as previously described. First-strand cDNA synthesis was performed by reverse transcription (TaKaRa, PrimeScript® 1st Strand cDNA Synthesis Kit, Dalian, China) according to the manufacturer’s descriptions. The primers used for real-time qRT-PCR (qPCR) were designed using Primer Premier 5. The qPCR was performed with a QuantStudio 7 Flex system (Applied Biosystems, USA) and SYBR Premix Ex Taq II (TaKaRa, Dalian, China). The qPCR reaction system contained 1 μL of diluted cDNA, 5 μL of SYBR® Premix Ex Taq™ II, 0.25 μL of each primer, and 3.5 μL of RNase-free H2O. For each sample, at least three technical replicates were performed. The peach EF2 gene (GeneBank ID: XM_007213735.2) was used as an internal control. The expression levels of the samples were calculated with 2−∆∆CT method (Livak and Schmittgen 2001). The expression levels of previously reported fruit ripening–related genes were also analyzed by qRT-PCR. These genes included ethylene synthesis (ACS1 and ACO1), ethylene perception and signal transduction (ETR1/ETR2 and ERF2), cell wall metabolism (PG and PMEI), ABA and gibberellins synthesis (NCED2 and GA2-OX), auxin conjugation, perception and transport (GH3, TIR1, and PIN1), and three NAC transcription factors (NAC1, NAC4, and NAC72) (Trainotti et al. 2003; Paponov et al. 2005; Staswick et al. 2005; Zhang et al. 2009; Torrigiani et al. 2012; Pirona et al. 2013). These genes or their homologous genes in peach genome were selected for gene expression analysis. The primers for expression analysis of the PpePAOs and fruit ripening–related genes are listed in Suppl. Table 2 and Suppl. Table 3.
Vector construction and transient expression assay
Full-length cDNA of PpePAO1, PpePAO2, and PpePAO4 were PCR-amplified with primers (Suppl. Table 1) containing SalI and EcoRI restriction sites and cloned into the pRI-101AN binary vector, under the control of the CaMV 35S promoter producing recombinant constructs. The three recombinant constructs were introduced into Agrobacterium tumefaciens strain GV3101, respectively, as was the empty pRI-101AN vector (EV) as control. An Agrobacterium-mediated transient transformation system was used to generate transgenic tobacco as described by Voinnet et al. (2003). The transgenic plants were kept at 25 °C for 3 days before the transgenic samples were collected for further analysis.
Detection of PAs, ethylene, and fruit quality traits
The extraction, derivation, and benzoylation of free and conjugated PAs (soluble and insoluble) were performed as described by Fu et al. (2011). The benzoylated PAs were separated and detected by HPLC (Waters, Milford MA, US) with a reverse-phase C18 column and an UV detector (230 nm) according to the description of Wang and Liu (2016).
Dry weight (DW) was obtained after drying mesocarp slices in an oven at 60 °C for 48 h. Ethylene production was detected by placing the whole detached fruit in a 500-mL box sealed with an air-tight lid equipped with a rubber stopper at room temperature. One hour later, 1 mL of gas was taken and measured by gas chromatography (GC 2010PLUS, Shimadzu, Japan) as described by Bregoli et al. (2002). Flesh firmness (FF) was measured using a pressure tester (GY-4, Hengmei, China). Soluble solids content (SSC) was measured with a digital refractometer (PAL-1, Atago, Japan). Each measurement was repeated at least three times.
Statistical analysis
Values are the means ± SE. Data on polyamine and fruit quality parameters were analyzed by the two-way ANOVA test (n=4).
Results
Detection of PA concentrations during peach fruit development and ripening
The diameter of the fruits from peach cultivar ‘Huangshuimi’ was measured and plotted over time. The first derivative of the growth curve exhibited a classic double sigmoidal pattern over the four stages of growth (S1 to S4; Fig. 1a). The concentrations of free and conjugated PA (soluble and insoluble) were detected during peach fruit development and ripening. The concentration of free Put slightly increased at S2 and then dramatically decreased from S2 to S4 (Fig. 1b). The concentrations of both free Spd and Spm were significantly increased at S2 and then dramatically decreased from S2 to S4 (Fig. 1b). Meanwhile, the concentrations of both soluble and insoluble conjugated PAs dramatically decreased during peach fruit development and ripening (Fig. 1c, d). However, it is unclear whether the decrease in free PA was associated with the decrease in conjugated PA.
Fruit growth curve and free polyamines content in the peach cultivar Huangshuimi during fruit development and ripening. a Peach fruit growth curve based on fruit diameter (red line) and its first derivative (green line) from 28 to 90 dAFB. S1–S4 are the four stages of growth up to harvest. b The free polyamine concentration during fruit development and ripening. c The content of soluble conjugated polyamines during peach fruit development and ripening. d The content of insoluble conjugated polyamines during peach fruit development and ripening. All measurements used 5-year-old orchard-grown trees of peach cultivar Huangshuimi. Error bars represent standard deviations for three replicates. Asterisks indicate *P <0.05, **P <0.01, ***P <0.001 (Student’s t test)
RNA-seq analysis of the expression levels of PA synthesis and catabolism-related genes
In order to investigate the molecular mechanisms behind the decrease in PA content during peach fruit ripening, RNA-seq was performed to analyze the expression of PA synthesis and catabolism-related genes. Across the later stages, the transcription levels increased or only slightly changed for PA synthesis–related genes such as Prupe.8G083100 (an arginine decarboxylase gene) and Prupe.8G048200 (an ornithine decarboxylase gene) (Fig. 2). Meanwhile, the expression of some PA catabolism–related genes significantly increased, such as Prupe.2G020100 (a polyamine oxidase gene) and Prupe.1G011900 (a copper amine oxidase gene). These results suggested that the decrease of PA content during peach fruit ripening was probably associated with PA catabolism.
Transcriptome analysis of the expression levels of PA synthesis and catabolism-related genes. Prupe.8G083100, arginine decarboxylase; Prupe.8G048000, Prupe.8G044900, and Prupe.8G048200, ornithine decarboxylase; Prupe.2G289700, Prupe.1G405900, and Prupe.1G011700, thermospermine synthase; Prupe.3G243800 and Prupe.1G299600, S-adenosylmethionine decarboxylase; Prupe.1G255300 and Prupe.5G176100, spermidine synthase; Prupe.1G012100, Prupe.5G078900, Prupe.1G011900, and Prupe.1G243500, copper amine oxidase; Prupe.2G020100, Prupe.3G179900, Prupe.3G013800, and Prupe.7G209800, polyamine oxidase
Identification of PAO genes in peach
Four putative PpePAO genes were identified by searching against the peach genome database using Arabidopsis AtPAO amino acid sequences (Table 1). After confirmation of the gene sequences by PCR amplification and sequencing, the peach PAO genes were designated as PpePAO1-4, respectively. The lengths of the coding sequences (CDS) of the four PpePAO genes ranged from 1485 bp (PpePAO2) to 1563 bp (PpePAO4), with corresponding polypeptides ranging from 494 to 520 amino acids (aa). The theoretical isoelectric points (pIs) of the PpePAOs ranged from 5.17 (PpePAO1) to 5.40 (PpePAO4), while the molecular weights (MWs) ranged from 54.5 kDa (PpePAO2) to 56.8 kDa (PpePAO4) (Table 1).
Sequence alignment, conserved motif identification, and phylogenetic analysis
Amino acid sequence alignment was performed with the four PpePAOs and one typical Zea mays PAO member (ZmPAO) (Fig. 3). The peach PAOs displayed a great divergence from each other. PpePAO2 and PpePAO3 were the most identical (61.0% identity), followed by ZmPAO and PpePAO1 (39.2%). PpePAO3 exhibited the lowest sequence similarity, of 13.7%, to PpePAO4, while the identities among other PpePAOs ranged from 14.0 to 23.4%.
Alignment of the amino acid sequences of peach PAO and maize PAO amino acid sequences. Identical and similar residues are shaded in black and gray background, respectively. Black lines above indicate the signal peptide and transmembrane domain (SP+TD). Peroxisomal targeting signals of PpePAO2 and PpePAO3 are indicated in red
The peach PAOs were analyzed for conserved and potential motifs. A peroxisomal targeting signal (PTS) was found in two of the PpePAO proteins. A PTS is a C-terminal tripeptide consisting of the consensus sequence (S/P/A/C) (R/K/N/M/S/L/H/)(L/M/I/V/Y) (Lingner et al. 2011). In plants, the preferred targeting signal is SKL (Reumann et al. 2007). The amino acid sequences of PpePAO2 and PpePAO3 contained a putative PTS1 (Fig. 3). PpePAO2 contained an SRI sequence, whereas PpePAO3 contained an SRM sequence, indicating that both of them are probably targeted to the peroxisome. All of the peach PAOs contained a transmembrane domain (TD), while only PpePAO2 and PpePAO3 possessed a signal peptide (SP) (Fig. 3).
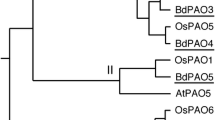
A phylogenetic analysis of the PAO proteins from different plants (including maize, grape, and apple) divided the protein family into four major groups (I, II, III, and IV) (Fig. 4). Consistent with the amino acid sequence alignment, PpePAO2 and PpePAO3 belonged to group I, PpePAO1 to group III, and PpePAO4 to group II. No proteins from peach were found in group IV, which contained genes from both monocots and other dicots.
Phylogenetic tree showing sequence relationships among polyamine oxidase proteins (PAOs) from peach (Ppe), sweet orange (Cs), apple (Md), Arabidopsis (At), tobacco (Ni), grape (Vv), humans, maize (Zm), barley (Hv), Brassica juncea (Bj), and rice (Os). The accession numbers of the genes are listed in Supplemental Table 1
Gene structure analysis and chromosomal distribution of the PpePAOs
Gene structure analysis revealed that PpePAO2 and PpePAO3 possessed similar gene structures, with each having ten exons and nine introns (Fig. 5a). PpePAO4 contained only 2 exons and 1 introns (Fig. 5a). In contrast, PpePAO1 lacked introns. The chromosomal distribution analysis suggested that the four PpePAOs were anchored on three chromosomes (Fig. 5b). Both PpePAO2 and PpePAO3 were located on chromosome 3, while PpePAO1 and PpePAO4 were located on chromosomes 2 and 7, respectively (Fig. 5b).
Genomic sequence and location analysis of four PpePAO genes. a Gene structures of the peach PAO genes generated using GSDS (http://gsds.cbi.pku.edu.cn/chinese.php). b Distribution of the PAO genes on peach chromosomes. The triangles indicate the transcription direction. The chromosome numbers are indicated at the top of each bar
Expressions of PpePAOs in different tissues
To investigate the expression profiles of the PAO genes in different tissues, transcript levels of the PpePAOs in flowers (including pistil, stamen, petal, and calyx), young leaves, old leaves, and young fruits were analyzed (Fig. 6). The expression level of each PpePAO gene in the flower sample was set at 1. The results showed that the PpePAO genes exhibited tissue-specific expression patterns. Specifically, PpePAO1 had its highest expression level in old leaves, and much lower expression levels in tender leaf and young fruit (Fig. 6a). In the flower, the mRNA level of PpePAO1 was much higher in stamen than in other tested tissues (Fig. 6a). PpePAO2 exhibited nearly equivalent expression levels in flower, young fruit, and old leaves, but presented a much lower expression level in tender leaves (Fig. 6b). The expression level of PpePAO2 was much higher in pistil than in other tested tissues of the flower (Fig. 6a). Similarly, PpePAO3 exhibited nearly equivalent expression levels in flower, tender leaves, and old leaves, but presented a much lower expression level in young fruit (Fig. 6c). In contrast, the mRNA level of PpePAO4 was highest in tender leaves, followed by old leaves, young fruit, and flower (Fig. 6d). Interestingly, the transcript levels of PpePAO4 in the petal were markedly higher than those in the pistil, stamen, and calyx (Fig. 6d).
Expression patterns of PpePAOs during peach fruit development and ripening
The expression patterns of the PpePAO genes in developing and ripening peach fruit were also investigated over six stages. The mRNA level of PpePAO1 gradually increased during fruit development and ripening (S1 to S3), peaking at 14-fold higher than the level of S1 at S3/S4 (Fig. 7b), then slightly decreasing at S4, although remaining higher compared to S1 (Fig. 7b). The expression level of PpePAO2 was stable at stages 1 and 2, sharply decreased to its minimum value (only 1/6 of its initial value) at S2/S3, dramatically increased to its peak value (2-fold higher than its initial value) at S3, and slightly decreased from S3/S4 to S4 (Fig. 7b). PpePAO3 exhibited an expression pattern similar to that of PpePAO2, of low expression at stages 1 and 2, followed by a rapid decrease at S2/S3, and a subsequent increase to the initial level from S3 to S4 (Fig. 7b). Compared to PpePAO1, PpePAO4 exhibited a converse expression pattern, gradually decreasing from S1 to S4 with its minimal levels during the S3/S4 and S4 (Fig. 7b).
Expression patterns of peach PAO genes during peach fruit development and ripening. a The phenotype of developing fruit from cv. Huangshuimi from S1 to S4 stage. b The expression patterns of PpePAO1-4 during ‘Huangshuimi’ peach development and ripening stage. Error bars represent standard deviations for four replicates
Detection of free PA concentration in plants transiently expressing peach PpePAOs
To further investigate the function of the PpePAOs in PA catabolism, PpePAO1, PpePAO2, and PpePAO4 were transiently expressed in tobacco. Genomic PCR and RT-PCR analyses demonstrated that these three genes were successfully expressed in the tested leaves (Supplementary Data Fig. 3). The content of free PAs was detected in PpePAO1, PpePAO2, and PpePAO4 transgenic leaves. The levels of Put in all transgenic lines were decreased compared to the level in the EV control line (Fig. 8). The levels of both Spd and Spm in PpePAO1 transgenic leaves were significantly decreased, by 55.9% and 57.8%, respectively, in comparison with the control (Fig. 8). Interestingly, in the PpePAO2 transgenic leaves, the level of Spd was increased by 33.7%, while the level of Spm was dramatically decreased by 34.2% compared to the EV line. In the PpePAO4 transgenic leaves, the concentration of Spd was similar to EV, while the level of Spm decreased by 55.1% compared to the control level (Fig. 8).
Inhibiting PAO-mediated PA catabolism delayed peach fruit ripening
The function of PAO-mediated PA catabolism during peach fruit ripening was explored by treating ‘Huangshuimi’ and ‘Okubo’ peach fruits at different days AFB with either Spd (as a positive control) or the PAO inhibitor guazatine. When fruit were treated with Spd or guazatine at S3 (68 dAFB), fruit ripening of ‘Huangshuimi’ was delayed (Fig. 9a). Ethylene production was significantly reduced after both Spd and guazatine treatments compared to control at S3/S4 and S4, except the ethylene content under 1 mM guazatine treatment at S3/S4 stage exhibited no difference compared to control (Fig. 9b). During normal ripening, the fruit firmness decreases, especially at S3 and S4. Spd and guazatine treatments prevented this decrease from S3 to S4 (Fig. 9c). Spd and guazatine treatments did not alter the content of soluble solids compared to control (Fig. 9d). Most interestingly, Spd and guazatine treatments did cause decreases in PpePAO1 transcript level and PAO activity (Fig. 9e, f), while guazatine treatment increased the content of free PAs compared to control (Fig. 10).
The phenotype (a), ethylene production (b), flesh firmness (c), soluble solids content (d), and expression levels of PpePAO1 (e), and PAO activity (f) of Huangshuimi peach fruit after treatment with 1 mM Spd or 1 mM or 5 mM guazatine (Gu) treatment at 68 dAFB (after full bloom). Error bars represent standard deviations for four replicates. Asterisks indicate *P <0.05, **P <0.01, ***P <0.001 (Student’s t test)
To comparing the timing of PAO inhibitor treatment, ‘Huangshuimi’ peach fruit were treated with 5 mM guazatine treatment at later stage of development, at S3/S4 (82 dAFB) (Supplementary Data Fig. 1B). Later treatment with guazatine produced a smaller effect on delaying fruit ripening compared to treatment at S3. Ethylene production was significantly decreased only on the first after guazatine treatment, and exhibited no obvious difference between control and guazatine treatment from day 2 to day 7 after treatment (Supplementary Data Fig. 1B). Later inhibitor treatment produced differences in fruit firmness only in days 3 and 5 after treatment (Supplementary Data Fig. 1C). The soluble solids content exhibited no obvious changes with later guazatine treatment compare to control (Supplementary data Fig. 1D).
To further verify the effect of guazatine treatment on peach fruit ripening, another peach cultivar, ‘Okubo,’ was treated with 1 mM Spd or 5 mM guazatine at 120 dAFB and harvested at 150 dAFB (commercial harvest). Guazatine treatment also inhibited ethylene production and firmness reduction in this cultivar (Supplementary Data Fig. 2B and C). These data suggested that peach fruit ripening was delayed by guazatine treatment and the effect was dependent on growth stage.
To further investigate the molecular mechanism underlying how PA catabolism can delay peach fruit ripening, the transcript levels of fruit ripening-related genes in ‘Huangshuimi’ peach were analyzed by qRT-PCR. Spd and guazatine treatments decreased the transcript levels of ethylene biosynthetic (PpACS and PpACO) and signaling genes (PpeETR1, PpeETR2, and PpeERF2) during fruit ripening (Fig. 11(A–E)). The transcript levels of the cell wall–related genes PpePG and PpePMEI were also downregulated by Spd and guazatine treatments (Fig. 11(F, G)). Spd and guazatine treatments also decreased the mRNA levels of ABA, GA, and IAA biosynthetic and signaling genes, such as PpeNCED, PpeGA2OX, PpeAux/IAA, PpePIN1, and PpeGH3, compared to the control levels (Fig. 11(H–L)). Furthermore, the expression levels of the previously reported NAC gene (PpeNAC72) as well as PpeNAC1 and PpeNAC4 (genes homologous with SlNAC1 and SlNAC4 in tomato) were also decreased after guazatine treatment. These results suggested that PA catabolism functions in peach fruit ripening by affecting the expression levels of fruit ripening-related genes.
Expression analysis by qRT-PCR of ripening-related genes in S3 to S4 peach fruit under 1 mM Spd or 5 mM guazatine treatment. (A–E) Transcript levels of ethylene biosynthetic and signaling genes during peach fruit ripening. (F, G) Transcript levels of cell wall–related genes during peach fruit ripening. (H) Transcript level of ABA biosynthetic gene PpeNCED during peach fruit ripening. (I) Transcript level of GA biosynthetic gene PpeGA2OX during peach fruit ripening. (J–L) Transcript level of IAA signaling genes during peach fruit ripening. (M–O) Transcript levels of NAC transcription factors during peach fruit ripening. The relative expression value was set to 1 for control at S3 stage
Discussion
Higher concentrations of PAs have been detected at the early stages of fruit development, followed by a gradual decrease as fruit mature. Similar phenomena have been reported in both non-climacteric and climacteric fruits, such as peach, apple, grape, and orange (Liu and Moriguchi 2007; Agudelo-Romero et al. 2013). Here, our results also showed that the concentrations of both free and conjugated PAs significantly decreased during peach ripening, in accordance with previous reports (Torrigiani et al. 2012). The decrease in free and conjugated PAs likely reflects the balance of PA in the plant cells; however, we are not sure whether the decrease of free PA was associated with conjugated PA. RNA-seq analysis showed that the expression levels of most PA synthesis–related genes were dramatically increased or unchanged, while some PA catabolism–related genes were markedly upregulated. These results suggested that the reduction of PA accumulation was probably associated with increased PA catabolism, especially PAO-mediated PA oxidation, which was consistent with a previous report in grape (Agudelo-Romero et al. 2013).
Therefore, the PAO genes in the peach genome were identified, and their temporal and spatial expression patterns were analyzed. Four putative PAO genes (PpePAO1-PpePAO4) were identified in the peach genome, and these genes exhibited tissue-specific expression. Most of the PpePAOs were more highly expressed in old leaves, indicating that PA oxidation and breakdown may be greater in old leaves. In the flower, PpePAO1 and PpePAO3 were most highly expressed in the stamen, while PpePAO2 and PpePAO4 were most highly expressed in pistil and petal, respectively. It has been reported that Spd-derived H2O2 signals Ca2+ influx and regulates pollen tube growth (Wu et al. 2010).
In order to further investigate the functions of the PpePAOs in PA catabolism, PpePAO1, PpePAO2, and PpePAO4 were transient expressed in tobacco leaves, and the concentration of free PAs was detected in transgenic lines. PpePAO3 was not tested because it exhibited the similar expression pattern with PpePAO2 during fruit development and ripening suggesting they probably play approximate functions during these process. Our results showed that PpePAO1 and PpePAO4 were probably involved in the terminal catabolism of Spd and Spm, while PpePAO2 was involved in the back-conversion of Spm to Spd. These data suggested PpePAO1 was probably involved in pollen tube growth.
In addition, many studies suggest that PA catabolism is involved in hormone and biotic stress responses (Alcazar et al. 2010). Here, PpePAO4 is reported to be significantly induced by ABA treatment, suggesting that this gene is involved in ABA response. In addition, most of the PpePAOs were induced by cold stress, indicating that PA oxidation probably takes part in cold stress response. In contrast, most of the PpePAOs were downregulated by heat, salt, and oxidative stress treatments, suggesting that PA oxidation exhibited different roles during different stress responses.
The transcript level of PpePAO1 was significantly upregulated during peach fruit development and ripening, indicating that PA catabolism was increased during these processes. However, the function of PA catabolism in peach fruit ripening remains largely unknown. After treatment with the PAO inhibitor guazatine, the expression level of PpePAO1 and PAO activity was markedly decreased, while the content of free PAs was much higher than control. These data suggested that PAO-mediated PA catabolism probably leads to the decrease of PA content during peach fruit ripening. In addition, peach fruit ripening and softening was delayed and ethylene production was significantly reduced after guazatine treatment. Furthermore, the expression levels of ethylene, ABA, GA, and IAA biosynthetic, signaling genes, and cell wall–related genes as well as some NAC genes were decreased after guazatine treatment. These results suggested that PA catabolism functions in peach fruit ripening by affecting the expression of genes related to hormone biosynthesis and signaling, and cell wall breakdown, as well as NAC genes. Our study provided strong evidence showing that PAO-mediated PA catabolism plays important roles in peach fruit ripening. However, the specific roles of PpePAOs, especially PpePAO1, in peach fruit ripening needs to be further investigated.
Abbreviations
- PAs:
-
Polyamines
- PAO:
-
Polyamine oxidase
- CuAO:
-
Copper-containing amine oxidase
- FAD:
-
Flavin adenine dinucleotide
- Put:
-
Putrescine
- Spd:
-
Spermidine
- Spm:
-
Spermine
- Dap:
-
1,3-Diaminopropane
- PpePAO :
-
Peach polyamine oxidase
- qPCR:
-
Real-time quantitative PCR
- PCD:
-
Programmed cell death
- PTSl:
-
Peroxisomal targeting signals
References
Agudelo-Romero P, Bortolloti C, Pais MS (2013) Study of polyamines during grape ripening indicate an important role of polyamine catabolism. Plant Physiol Biochem 67:105–119
Agudelo-Romero P, Ali K, Choi YH, Sousa L, Verpoorte R, Tiburcio AF, Fortes AM (2014) Perturbation of polyamine catabolism affects grape ripening of Vitis vinifera cv. Trincadeira. Plant Physiol Biochem 74:141–155
Ahou A, Martignago D, Alabdallah O, Tavazza R, Stano P, Macone A, Pivato M, Masi A, Rambla JL, Vera-Sirera F, Angelini R, Federico R, Tavladoraki P (2014) A plant spermine oxidase/dehydrogenase regulated by the proteasome and polyamine. J Exp Bot 65:1585–1603
Alcazar R, Altabella T, Marco F, Bortolotti C, Reymond M, Koncz C, Carrasco P, Tiburcio AF (2010) Polyamines: molecules with regulatory functions in plant abiotic stress tolerance. Planta 231:1237–1249
Bregoli AM, Scaramagli S, Costa G, Sabatini E, Ziosi V, Biondi S (2002) Peach (Prunus persica) fruit ripening: aminoethoxyvinylglycine (AVG) and exogenous polyamines affect ethylene emission and flesh firmness. Physiol Plant 114:472–481
Cervelli M,Tavladoraki P,Di Agostino S,Angelini R,Federico R,Mariottini P (2000) Isolation and characterization of three polyamine oxidase genes from Zea mays. Plant Physiol Biochem 38(9):667–677
Cervelli M, Cona A, Angelini R, Polticelli F, Federico R, Mariottini P (2001) A barley polyamine oxidase isoform with distinct structural features and subcellular localization. Eur J Biochem 268:3816–3830
Cheng X-Q, Zhu X-F, Tian W-G, Cheng W-H, Sun HJ, Jin S-X, Zhu H-G (2017) Genome-wide identification and expression analysis of polyamine oxidase genes in upland cotton (Gossypium hirsutum L.). Plant Cell Tissue Organ Culture (PCTOC) 129(2):237–249
Cona A, Rea G, Angelini R, Federico R, Tavladoraki P (2006) Function of amine oxidases in plant development and defence. Trends Plant Sci 11:80–88
Cui X, Ge CM, Wang RX, Wang R, Wang H, Chen W, Fu Z, Jiang X, Li J, Wang Y (2010) The BUD2 mutation affects plant architecture through altering cytokinin and auxin responses in Arabidopsis. Cell Res 20:576–586
Fincato P, Moschou PN, Spedaletti V, Tavazza R, Angelini R, Federico R, Roubelakis-Angelakis KA, Tavladoraki P (2011) Functional diversity inside the Arabidopsis polyamine oxidase gene family. J Exp Bot 62:1155–1168
Fu XZ, Chen CW, Wang Y, Liu JH, Moriguchi T (2011) Ectopic expression of MdSPDS1 in sweet orange (Citrus sinensis Osbeck) reduces canker susceptibility: involvement of H2O2 production and transcriptional alteration. BMC Plant Biol 11:55
Gao Y, Wei W, Zhao X, Tian XL, Fan ZQ, Zhang YP, Jing Y, Meng LH et al (2018) A NAC transcription factor, NOR-like1, is a new positive regulator of tomato fruit ripening. Hortic Res 5:75
Ghuge SA, Carucci A, Rodrigues-Pousada RA, Tisi A, Franchi S, Tavladoraki P, Angelini R, Cona A (2015) The apoplastic copper AMINE OXIDASE1 mediates jasmonic acid-induced protoxylem differentiation in Arabidopsis roots. Plant Physiol 168:690–707
Gomez-Jimenez MC, Paredes MA, Gallardo M (2010) Tissue-specific expression of olive S-adenosyl methionine decarboxylase and spermidine synthase genes and polyamine metabolism during flower opening and early fruit development. Planta 232:629–647
Groβ F, Rudolf EE, Thiele B, Durner J, Astier J (2017) Copper amine oxidase 8 regulates arginine-dependent nitric oxide production in Arabidopsis thaliana. J Exp Bot 68:2149–2162
Guo J, Wang S, Yu X, Dong R, Li Y, Mei X, Shen Y (2018) Polyamines regulate strawberry fruit ripening by ABA, IAA, and ethylene. Plant Physiol 177:339–351
Hao Y, Huang B, Jia D, Mann T, Jiang X, Qiu Y, Niitsu M, Berberich T, Kusano T, Liu T (2018) Identification of seven polyamine oxidase genes in tomato (Solanum lycopersicum L.) and their expression profiles under physiological and various stress conditions. J Plant Physiol 228:1–11
Hatmi S, Trotel-Aziz P, Villaume S, Couderchet M, Clément C, Aziz A (2013) Osmotic stress-induced polyamine oxidati-on mediates defence responses and reduces stress-enhanced grapevine susceptibility to Botrytis cinerea. J Exp Bot 65:75–88
Kamada-Nobusada T, Hayashi M, Fukazawa M, Sakakibara H, Nishimura M (2008) A putative peroxisomal polyamine oxidase, AtPAO4, is involved in polyamine catabolism in Arabidopsis thaliana. Plant Cell Physiol 49:1272–1282
Lingner T, Kataya AR, Antonicelli GE, Benichou A, Nilssen K, Chen XY, Siemsen T, Morgenstern B, Meinicke P, Reumann S (2011) Identification of novel plant peroxisomal targeting signals by a combination of machine learning methods and in vivo subcellular targeting analyses. Plant Cell 23:1556–1572
Liu JH, Moriguchi T (2007) Changes in free polyamine titers and expression of polyamine biosynthetic genes during growth of peach in vitro callus. Plant Cell Rep 26:125–131
Liu T, Kim DW, Niitsu M, Maeda S, Watanabe M, Kamio Y, Berberich T, Kusano T (2014) Polyamine oxidase 7 is a terminal catabolism-type enzyme in Oryza sativa and is specifically expressed in anthers. Plant Cell Physiol 55:1110–1122
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2 −ΔΔCT method. Methods 25:402–408
Ma NN, Feng HL, Meng X, Li D, Yang DY, Wu CG, Meng QW (2014) Overexpression of tomato SlNAC1 transcription factor alters fruit pigmentation and softening. BMC Plant Biol 14:351
Møller SG, McPherson MJ (1998) Developmental expression and biochemical analysis of the Arabidopsis atao1 gene encoding an H2O2-generating diamine oxidase. Plant J 13:781–791
Moschou PN, Sanmartin M, Andriopoulou AH, Rojo E, Sanchez-Serrano JJ, Roubelakis-Angelakis KA (2008) Bridging the gap between plant and mammalian polyamine catabolism: a novel peroxisomal polyamine oxidase responsible for a full back-conversion pathway in Arabidopsis. Plant Physiol 147:1845–1857
Moschou PN, Wu J, Cona A, Tavladoraki P, Angelini R, Roubelakis-Angelakis KA (2012) The polyamines and their catabolic products are significant players in the turnover of nitrogenous molecules in plants. J Exp Bot 63:5003–5015
Naconsie M, Kato K, Shoji T, Hashimoto T (2014) Molecular evolution of N-methylputrescine oxidase in tobacco. Plant Cell Physiol 55:436–444
Ono Y, Kim DW, Watanabe K, Sasaki A, Niitsu M, Berberich T, Kusano T, Takahashi Y (2012) Constitutively and highly expressed Oryza sativa polyamine oxidases localize in peroxisomes and catalyze polyamine back conversion. Amino Acids 42:867–876
Pan L, Zeng WF, Niu L, Lu ZH, Liu H, Cui GC, Zhu YQ, Chu JF, Li WP, Fang WC, Cai ZG, Li GH, Wang ZQ (2015) PpYUC11, a strong candidate gene for the stony hard phenotype in peach (Prunus persica L. Batsch), participates in IAA biosynthesis during fruit ripening. J Exp Bot 66:7031–7044
Paponov IA, Teale WD, Trebar M, Blilou I, Palme K (2005) The PIN auxin efflux facilitators: evolutionary and functional perspectives. Trends Plant Sci 10:170–177
Parra-Lobato MC, Gomez-Jimenez MC (2011) Polyamine-induced modulation of genes involved in ethylene biosynthesis and signalling pathways and nitric oxide production during olive mature fruit abscission. J Exp Bot 62:4447–4465
Pirona R, Eduardo I, Pacheco I, Linge CDS, Miculan M, Verde I, Tartarini S, Dondini L, Pea G, Bassi D, Rossini L (2013) Fine mapping and identification of a candidate gene for a major locus controlling maturity date in peach. BMC Plant Biol 13:166
Planas-Portell, J., Gallart, M., Tiburcio, A.F., Altabella,T., 2013. Copper-containing amine oxidases contribute to terminal polyamine oxidation in peroxisomes and apoplast of Arabidopsis thaliana. BMC Plant Biol 13, 109.
Rea G, Laurenzi M, Tranquilli E, D’Ovidio R, Federico R, Angelini R (1998) Developmentally and wound-regulated expression of the gene encoding a cell wall copper amine oxidase in chickpea seedlings. FEBS Lett 437:177–182
Reumann S, Babujee L,Ma C, Wienkoop S, Siemsen T, Antonicelli GE, Rasche N,Franziska L, Weckwerth W, Jahn O (2007) Proteome Analysis ofLeaf Peroxisomes Reveals Novel Targeting Peptides, Metabolic Pathways, and Defense Mechanisms. Plant Cell 19(10):3170–3193
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Smith TA (1972) Purification and properties of the polyamine oxidase of barley plants. Phytochemistry 11:899–910
Staswick PE, Serban B, Rowe M, Tiryaki I, Maldonado MT, Maldonado MC, Suza W (2005) Characterization of an Arabidopsis enzyme family that conjugates amino acids to indole-3-acetic acid. Plant Cell 17:616–627
Takahashi Y, Ono K, Akamine Y, Asano T, Ezaki M, Mouri I (2018) Highly-expressed polyamine oxidases catalyze polyamine back conversion in Brachypodium distachyon. J Plant Res 131:341–348
Tatsuki M, Nakajim N, Fujii H, Shimada T, Nakano M, Hayashi K, Hayama H, Yoshioka H, Nakamura Y (2013) Increased levels of IAA are required for system 2 ethylene synthesis causing fruit softening in peach (Prunus persica L. Batsch). J Exp Bot 64:1049–1059
Tavladoraki P, Rossi MN, Saccuti G, Perez-Amador MA, Polticelli F, Angelini R, Ferderico R (2006) Heterologous expression and biochemical characterization of a polyamine oxidase from Arabidopsis involved in polyamine back conversion. Plant Physiol 141:1519–1532
Tipping AJ, McPherson MJ (1995) Cloning and molecular analysis of the pea seedling copper amine oxidase. J Biol Chem 270:16939–16946
Torrigiani P, Bressanin D, Beatriz Ruiz K, Tadiello A, Trainotti L, Bonghi C, Ziosi V, Costa G (2012) Spermidine application to young developing peach fruits leads to a slowing down of ripening by impairing ripening-related ethylene and auxin metabolism and signaling. Physiol Plant 146:86–98
Trainotti L, Zanin D, Casadoro G (2003) A cell wall-oriented genomic approach reveals a new and unexpected complexity of the softening in peaches. J Exp Bot 54:1821–1832
Tsaniklidis G, Kotsiras A, Tsafouros A (2016) Spatial and temporal distribution of genes involved in polyamine metabolism during tomato fruit development. Plant Physiol Biochem 100:27–36
Tuskan GA, DiFazio S, Jansson S, Bohlmann J, Grigoriev I, Hellsten U, Putnam N (2006) The genome of black cottonwood, Populus trichocarpa (Torr. & Gray). Science 313:1596–1604
Voinnet O, Rivas S, Mestre P, Baulcombe D (2003) An enhanced transient expression system in plants based on suppression of gene silencing by the p19 protein of tomato bushy stunt virus. Plant J 33:949–956
Wang W, Liu JH (2015) Genome-wide identification and expression analysis of the polyamine oxidase gene family in sweet orange (Citrus sinensis). Gene 555:421–429
Wang W, Liu JH (2016) CsPAO4 of Citrus sinensis functions in polyamine terminal catabolism and inhibits plant growth under salt stress. Sci Rep 6:31384
Wang W, Wu H, Liu JH (2017) Genome-wide identification and expression profiling of copper-containing amine oxidase genes in sweet orange (Citrus sinensis). Tree Genet Genomes 13:31
Wang X, Zeng W, Ding YF, Wang Y, Niu L, Yao JL, Pan L, Lu ZH, Cui GC, Li GH, Wang ZQ (2019) Peach ethylene response factor PpeERF2 represses the expression of ABA biosynthesis and cell wall degradation genes during fruit ripening. Plant Sci 283:116–126
Wu J, Shang Z, Wu J, Jiang X, Moschou PN, Sun W, Roubelakis-Angelakis KA (2010) Spermidine oxidase-derived H2O2 regulates pollen plasma membrane hyperpolarization-activated Ca2+-permeable channels and pollen tube growth. Plant J 63:1042–1053
Yoda H, Hiroi Y, Sano H (2006) Polyamine oxidase is one of the key elements for oxidative burst to induce programmed cell death in tobacco cultured cells. Plant Physiol 142:193–206
Zarei A, Trobacher CP, Cooke AR, Meyers AJ, Hall JC, Shelp BJ (2015) Apple fruit copper amine oxidase isoforms: peroxisomal MdAO1 prefers diamines as substrates whereas extracellular MdAO2 exclusively utilizes monoamines. Plant Cell Physiol 56:137–147
Zhang M, Ping L, Zhang G, Li X (2009) Cloning and functional analysis of 9-cis-epoxycarotenoid dioxygenase (NCED) genes encoding a key enzyme during abscisic acid biosynthesis from peach and grape fruits. J Plant Physiol 166:1241–1252
Zhu MK, Chen GP, Zhou S, Tu Y, Wang Y, Dong TT, Hu ZL (2014) A new tomato NAC(NAM/ATAF1/2/CUC2)transcription factor, SlNAC4, functions as a positive regulator of fruit ripening and carotenoid accumulation. Plant Cell Physiol 55:119–135
Acknowledgements
The author thanks Anita K. Snyder, M. Sc., for the critical reading and language editing of the manuscript.
Data archiving statement
The transcriptome data have been deposited in NCBI database (with the following link: https://submit.ncbi.nlm.nih.gov/subs/sra/SUB9021282/files). Genomic sequences and information of genes related to polyamine metabolism and fruit ripening were downloaded from the JGI database (https://phytozome.jgi.doe.gov/pz/portal.html).
Funding
This work was supported by the National Key Research and Development Program of China (2018YFD1000300), the Major Science and Technology Project in Henan Province (151100110900), the Science and Technology Creative Team in Henan Province (19IRTSTHN009), and the Innovation and Technology Fund of Henan Agricultural University (KJCX2018A15).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Communicated by L.A. Meisel
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Wang, W., Zheng, X., Liu, S. et al. Polyamine oxidase (PAO)–mediated polyamine catabolism plays potential roles in peach (Prunus persica L.) fruit development and ripening. Tree Genetics & Genomes 17, 10 (2021). https://doi.org/10.1007/s11295-021-01495-x
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11295-021-01495-x