Abstract
In estimating the number of species per quadrat with a given area, we usually need much time and labor, because we have no simple ways to easily count it using hardware. We devised here a software method to estimate the mean number of species by counting them only partially in the survey, and estimating it mathematically. We classify quadrats into three classes composed of k, k + 1 and more than k + 1 species; or classify quadrats into four classes composed of k, k + 1, k + 2, and more than k + 2 (referred to as “censored sampling”), where k is the minimum number of species per quadrat. We do not need to count more than k + 2 or k + 3 species, respectively. Using only these 3 or 4 numerals we can estimate the mean of number of species per quadrat, the 95% confidence intervals for the mean and know the spatial pattern value such as aggregated, random or uniform pattern of the number of species per quadrat. Using this method, 10–20% time and labor in counting will be saved compared to the full census. The estimates are obtained through the maximum likelihood calculation. Computer programs to estimate the mean number of species per quadrat are attached as ESMs.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
To understand the nature of vegetation, we measure various traits of plants and their distributions. We set at random points or arrange systematically many quadrats in the vegetation, each of which has a given area, on line transects, and plants in each quadrats are measured and recorded. The traits of vegetation often measured by field plant scientists are cover, number of occurrences, biomass and number of individuals/density per quadrat. Based on these measurements, traits of vegetation such as species richness, and relations working among species composing the community and interactions such as their spatial pattern could be understood (Pielou 1977).
To measure these vegetation traits, various rational methods have been developed; for example, in measuring cover, the line-intercept method (Bonham 2013) and point-count method (Chen et al. 2008a). Although we can obtain clear results from these surveys based on statistical manipulations, extensive physical labor and a long time are usually required in field survey. For this reason, various time-saving methods have been proposed: one is to measure using such as machineries or photo-sensors (hardware-dependent methods) (e.g., for small-scale quadrats Shibayama 2001, 2003; Shibayama and Akita 2002; for large-scale areas Gould 2000; Yasuda 2018), and another is to measure based on software-dependent methods such as statistical/mathematical model (e.g., Nicholas et al. 2011). We consider hereafter the software-dependent method. In measuring cover through line-intercept and point-count methods, there are methods to measure only a few objects (e.g., plant individuals) instead of the entire objects visually or using simple tools, and mathematically estimated the parameter values (software-dependent method) (Chen et al. 2006, 2008c). The similar software-dependent methods have also been developed for estimating herbaceous biomass (Shiyomi 1991, 1992) and estimating the number of insect individuals (Shiyomi 1974, 1977, 1978). The method for estimating the number of insects has a possibility applicable to measure the number of plant individuals/density per quadrat. These methods can be used based on some probability distributions.
In this study, apart from these traits such as cover and biomass, we consider “the number of species (richness) per quadrat”, which is another important trait in vegetation study of grasslands. Species richness influences to the primary plant production (e.g., Tilman et al. 2001; Pallett et al. 2016), contributes for protecting and maintaining vegetation from natural or artificial disturbances, and recovering the vegetation destroyed once (e.g., Lavorel 1999; Katharina et al. 2001; Nicholas et al. 2011; Oliver et al. 2015). In analyzing vegetation composing of many species, interesting findings for the niche rule have been obtained (van der Maarel et al. 1995; Zobel et al. 2000; Chen et al. 2008b; Chytrý et al. 2015). Chen et al. (2016) proposed a model of Poisson-like probability distribution in which the number of species per quadrat follows, referred to as “extended Poisson distribution” in their paper. In the followings, we introduce a time-saving method in vegetation survey based on this model, we count rapidly the number of species per quadrat, and estimate mathematically the mean, the variance among quadrats, and the spatial pattern value.
Fundamental model, materials and methods
The Poisson-like frequency distribution model of per-quadrat number of species
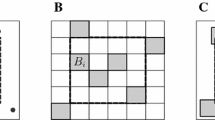
Now, suppose a situation where we set small quadrats at N random points on a herbaceous plant community such as grassland and riverside meadow. Each quadrat contains k species at least (referred to as “the fundamental number of species per quadrat” or FNS hereafter) (e.g., k = 4 in Fig. 1a), and the number of species more than k, i follows a Poisson distribution with mean μ (i = 0, 1, 2, …). Then, the number of species per quadrat, j, is composed of two values, k and i, where k is a constant through all quadrats as defined above, and i is a variate and indicating that i species are allocated at random quadrats. Therefore, j = k + i, where both constant k and variate i are non-negative integers. The probability that j species occur in a quadrat (Chen et al. 2016) is as follows:
where letting λ be the mean of j, and μ be the mean of i; therefore,
The number of species per quadrat and a model of the frequency distribution: an example observed in a semi-arid natural grassland in Shenmu, Shaanxi Province, China (after Chen et al. 2016). a Original data: Ninety 50 × 50-cm quadrats were set along a line on the grassland, and the number of species was counted per quadrat. Since each of the quadrats contains four or more species, i.e., k = 4, letting the number of species in excess of k (= 4) be i, the number of species per quadrat, j, equals k + i. The mean of j is λ, and λ = k + μ, where μ is the mean of i. b Observed frequency distribution: The number of species in each quadrat shown in Fig. 1a, is summarized as a frequency distribution, where the mean of j is λ = 6.76. c Fitted frequency distribution: The frequency distribution of black bars indicates the distribution (Eq. 1) fitted to the frequency distribution of the number of species counted per quadrat. The shaded bars indicate that the frequency distribution moved 4 (= k) units to the left, which is a Poisson distribution with μ = 2.76
The variance for Eq. (1) is given as same as for the Poisson distribution of e−μμi/i! (i = 0, 1, 2,…): μ. Therefore, the mean, λ, is always larger than the variance for k ≥ 1. Equation (1) indicates the equation moved the Poisson distribution with the mean of μ to just k units rightward along the axis of abscissa (Poisson-like distribution), as shown in Fig. 1c by an example obtained at 90 quadrats set in the Shenmu grassland, in which k = 4, μ = 2.9181 and λ = k + μ = 4 + 2.9181 = 6.9181. The variance for the example, v, is 2.2317, which is close to 2.9181.
A probable reason why we have arrived at the present model, especially the introduction of k, after long-term, trial-and-error considerations is as follows:
-
1.
The theoretical frequency distribution of the number of species per quadrat can be assumed to be constructed of a positive, infinite series of integer variates, such as 0, 1, 2, …
-
2.
According to the experimental results reported in Chen et al. (2016), the mean number of species per quadrat in a community is always larger than the variance among quadrats.
-
3.
All grassland communities are a mixture of species with low to high occurrence.
Our model was synthesized so as to satisfy these three concepts.
According to our idea, each species appears in random quadrats, depending on their number of occurrence. Since several species with a high occurrence fill most quadrats up to k species, a uniform frequency distribution of the number of species per quadrat, ranged between 0 and k, is approximately formed by these species. Indeed, the 11 survey results reported in Chen et al. (2016) suggest that we can determine k if we know the number of species that occur in more than 50% of the quadrats (i.e., high occurrence) (Fig. 2). The other i species with a low occurrence add at random quadrats according to a Poisson distribution. Then, the total number of species at an arbitrary quadrat, j, is k + i, which is the Poisson-like distribution shown in Fig. 1c (black bars). This distribution is an infinite series constructed of the variate of non-negative integers as shown in Eq. (1). If k > 0, the mean number of species is larger than the variance, but if k = 0, the mean number of species is equal to the variance (i.e., usual Poisson distribution).
Regression of the FNS, k, on the number of species occurring in more than 50% of the quadrats, x. The approximately 85% variation among the FNS, k, is explained by the variation in terms of the number of species in more than 50% of the quadrats, x: the regression is k = 0.450 + 0.953 X and the contribution, R2, is 0.849. These data are cited from Chen et al. (2016), where Shenmu, Banska Bystrica, and Nishinasuno are grasslands in the Shaanxi region of China, Slovakia, and Japan, respectively
We have already shown that most frequency distributions of the number of species per quadrat fit the model defined by Eq. (1). Chen et al. (2016) studied 55 grassland communities in a variety of locations including semiarid areas in China and temperate areas in Japan and central Europe. Of these communities, 54 fit the model (based on the goodness-of-fit test using χ2). In every case tested, the mean number of species was larger than the variance between the quadrats. Therefore, Eq. (1) is a reasonable frequency distribution model for the number of species per quadrat.
A time-saving counting and estimation of the mean
When counting the number of species quadrat to quadrat, we classify quadrats into n + 1 classes, that is, quadrats with k, k + 1, …, and (k + n − 1) species, and more than (k + n − 1) species. Then, we count the number of species k to (k + n − 1) species at all the N quadrats, but we do not count the number more than (k + n − 1) species for saving the surveying time (referred to as “censored” sampling). In the other words, we divide the probability distribution that number of species follows into k + n + 1 terms: (1) the k terms of \( P(j = 0) = P(j = 1) = \cdots = P(j = k - 1) = 0 \) for the per-quadrat number of species, (2) the n terms of P(j = k), P(j = k + 1), …, P(j = k + n − 1): e−μ, e−μμ, …, e−μμn−1/(n − 1)!, respectively, and (3) one term of P(j ≥ k + n): \( \sum\nolimits_{j = k + n}^{\infty } {P(j)} = \sum\nolimits_{j = k + n}^{\infty } {\frac{{e^{ - \upmu } \upmu^{j - k} }}{(j - k)!}} \),where j is a variate, k is a constant and μ is an unknown parameter in all terms.
For example, suppose that each quadrat is composed of 4 or more species as shown in Fig. 1a, i.e., k = 4. If setting as n = 2, we count the number of species at each quadrat, and classify quadrats into three classes in which the number of quadrats composed of 4, 5 and more than 5 species. In Fig. 1a, b, for example, the numbers of quadrats composed of 4, 5 and more than 5 species are 5, 14 and 90 − (5 + 14) = 71, respectively, where 90 is the total number of quadrats observed. Thus, we do not need to count the number of species at quadrats composed of species more than 5.
If setting n = 3 in Fig. 1a, we count the number of quadrats composed of 4–6 species separately, and the pooled number of quadrats containing 7 or more species, where we classify quadrats into four classes. In this case, the numbers of quadrats composed of 4, 5, 6 and more than 6 species are 5, 14, 23 and 90 − (5 + 14 + 23) = 48, respectively.
Let N quadrats be observed in a vegetation survey, and the numbers of quadrats of containing j species be expressed by x j . Furthermore, let the numbers of quadrats of containing less than k species be 0 (i.e., x0, x1,…, xk−1 = 0), the number of quadrats of containing k, k + 1, k + 2,…,k + i,…,k + n − 1 species be x k , xk+1, xk+2,…,xk+i,…,xk+n−1, and the number of quadrats containing k + n or more species be xk+n+. Then, a logarithmic likelihood equation, l, based on Eq. (1), can be expressed as follows:
Differentiated Eq. (3) by μ and putting 0, we solve the equation for μ as shown in Eq. (4).
The mean of number of species per quadrat is obtained by \( k + \hat{\upmu } \), which is \( \hat{\lambda } \), where the estimate of μ is \( \hat{\upmu } \), and that of λ is \( \hat{\lambda } \).
Next, we calculate the Fisher’s information for estimating the large sample variance as follows:
where omitted ^ of \( \hat{\upmu } \). Then, the estimate of large sample variance for \( \hat{\lambda } \) and \( \hat{\upmu } \) (i.e., \( \hat{\upsigma }_{{\hat{\lambda }}}^{2} \) and \( \upsigma_{{\hat{\upmu }}}^{2} \)), is given by:
where \( \upmu = \hat{\upmu } \) indicates to substitute \( \hat{\upmu } \) for μ. Then, the 95% confidence intervals for μ and λ are given as:
where 1.96 is the 95%-point value of the standard normal distribution.
Vegetation survey in grasslands
Although there are many cases which can be shown as examples of vegetation survey, we will adopt again the 11 examples already shown in Chen et al. (2016). (1) In a semiarid grassland at Shenmu of Shaanxi Province, China (a) 360 25 cm × 25 cm quadrats were surveyed, and the number of species at each quadrat was counted. (b) Next, two adjacent quadrats were incorporated into a 25 cm × 50 cm quadrats, and produced 180 quadrats. (c) Third, two adjacent 25 cm × 50 cm quadrats were incorporated into a 50 cm × 50 cm quadrats, and produced 90 quadrats. (d) Lastly, two adjacent 50 cm × 50 cm quadrats were incorporated into a 50 cm × 100 cm, produced 45 large quadrats. (2) In Banská Bystrica, Slovakia, we selected three different grasslands of (a) a sown grassland (b) a natural grassland, and (c) a grassland over-sown improved herbage seeds as the survey areas. Each survey area with 5 m × 5 m was divided into 100 50 cm × 50 cm quadrats, 50 alternate quadrats like chessboard were selected for surveys, and the number of species at each quadrat was counted. (3) In the National Grassland Research Institute, Nishinasuno, Japan, two semi-natural grasslands different in the grazing rate were selected for vegetation surveys. In each grassland, 100 10 cm × 10 cm quadrats were set, and the number of species at each quadrat was counted in May and August.
According to the plan of the present study, first we determined k-value, constructed censored frequency distributions for n = 2 and n = 3, then estimated μ and λ (= k + μ) based on Eq. (4). The actual calculation will be described in the following sections, and computer programs written in R and Excel-Basic for estimating the mean number of species per quadrat are provided in Supplements ESM 1 to ESM 4.
Results
Here, we show an example of estimating the mean of number of species based on data obtained at the Shenmu grassland, China. In the survey area, 90 50 cm × 50 cm, steel-made quadrats were set, and the number of species within each quadrat was recorded (see Fig. 1a). The result obtained from the complete survey data (i.e., not censored survey) were as follows: k = 4, \( \hat{\upmu } \) = 2.76, \( \hat{\lambda } \) = 4 + 2.76 = 6.76, and v = 2.2317. The frequency distribution of the number of species per quadrat obtained based on the survey data is shown in Fig. 1b. In the complete survey data, it is easy to find the k-value in Fig. 1a.
Next, applying these data to Eq. (4), we estimate the mean number of species per quadrat, \( \hat{\upmu } \). The calculations were conducted on the Excel-BASIC program (see Supplement ESM3 and ESM4). We first set k = 4 and n = 2, that is, the 90 quadrats are classified into three classes such as 5 quadrats containing 4 species, 14 quadrats containing 5 species, and 71 quadrats containing 6 or more species (“censored frequency distribution”; cf. Table 1a). Then, we estimated the mean number of species per quadrat, \( \hat{\upmu } \), based on Eq. (4): \( \hat{\upmu } \) = 2.9181 and \( \hat{\lambda } \) = k + \( \hat{\upmu } \) = 4 + 2.9181 = 6.9181. The values of \( \hat{\upmu } \) and \( \hat{\lambda } \) were comparatively near to the values estimated based on the complete survey data (Fig. 3a).
The number of species per quadrat and model of the frequency distribution: an example observed in a Shenmu semiarid natural grassland of Shaanxi Province, China. a The frequency distributions of observed and estimated numbers of species in each quadrat for n = 2, where the estimated mean based on the censored sample is λ = 6.9181 for k = 4 and μ = 2.9181. b The frequency distributions of observed and estimated numbers of species in each quadrat for n = 3, where the estimated mean based on the censored sample is λ = 6.8431 for k = 4 and μ = 2.8431
Next, let us try to calculate for the case of k = 4 and n = 3. In this case, the number of quadrats where 6 species were found was 23, and the number of quadrats in which 7 or more species found was also 48 (see Table 1). The calculated results based on the censored sample are: \( \hat{\upmu } \) = 2.8431, \( \hat{\lambda } \) = k + \( \hat{\upmu } \) = 4 + 2.8 431 = 6.8431, which are also considerably near to those estimated based on the complete survey data. We constructed the complete frequency distribution for the number of species using these k and μ estimated based on the censored sample, as shown in Fig. 3b. For k = 4 and n = 1, based on 2 classes of the numbers at only 5 quadrats with four species and residual 85 quadrats with more than four species, we can, of course, calculate the mean number of species per quadrat. However, we cannot expect sufficiently high precision for \( \hat{\upmu } \) and \( \hat{\lambda } \); actually, we obtained \( \hat{\upmu } \) = 2.30 and \( \hat{\lambda } \) = 6.30, which are much different from those of the complete survey in this example.
Next, we estimated the large sample variance of \( \hat{\upmu } \) (and \( \hat{\lambda } \)) based on the Fisher’s information, and then estimated the 95% confidence intervals of λ: 6.84–6.99 for n = 2, and 6.74–6.94 for n = 3.
Based on our experiences, n = 2 or 3 is generally adequate when considering labor and time for surveys (in the above example, we count by 6 or 7 species, respectively). Although we can also estimate based on other larger n-values than 2 or 3, it will need more survey time and labor though we can attain a higher precision.
Whether can per-quadrat means of the number of species estimated based on the time-saving method approximate the estimates based on the complete samples? An empirical answer to this question, which is calculated for the 11 survey results shown in Chen et al. (2016), is as Fig. 4, which indicates the relation between the estimates calculated using the complete survey data, and estimates using the censored method. Figure 4a, b are for n = 2 and 3, respectively, in estimating μ. The correlation coefficients between the estimates calculated using the complete survey data, \( \hat{\upmu } \) com, and the estimates using the censored method,\( \hat{\upmu }_{{}} \) rap, are 0.86 for n = 2, and 0.97 for n = 3, which indicate there is a high linear relationship, especially in the case of n = 3. Next, we show the relation between the estimates calculated using the complete survey data, \( \hat{\lambda } \) com, and the estimates using the censored method, \( \hat{\lambda } \) rap, in Fig. 4c, d. The correlation coefficients between \( \hat{\lambda } \) com and \( \hat{\lambda } \) rap reached almost 1.
Comparisons of means of the per-quadrat number of species estimated based on the entire samples (x) and the censored samples (y). a Relation in μ for the case of n = 2; b Relations in µ for the case of n = 3; c Relation in λ for the case of n = 2; d Relations in λ for the case of n = 3. Correlation coefficient is denoted by r. Although the relation of \( \hat{\lambda } \) = k + \( \hat{\upmu } \) is held between \( \hat{\upmu } \) and \( \hat{\lambda } \), k-values are different among different samples
Discussion
In discussion of species richness/diversity of vegetation, the argument is generally made grounded on vegetation traits of each species such as cover or biomass, which relates to productivity, and resilience and resistance to environmental disturbances (Lavorel 1999; Katharina et al. 2001; Tilman et al. 2001; Oliver et al. 2015; Pallett et al. 2016).
Why is the censored sampling necessary?
To estimate accurate mean of number of species, we have no way except for visual count, since we have not developed mechanical or optical sensors to easily count the number of species or easily and accurately distinguish different species at present. However, visual count of the number of species in general requires much time and labor. The censored method stated here will save 10–20% of time and labor compared to those in the complete survey.
Except for the number of species, methods easily measuring cover, biomass and the number of individuals based on the censored sampling have been already established. The prerequisite is that the theoretical frequency distributions for these traits of vegetation are known. For example, the frequency distribution of cover per quadrat follows the beta distribution (Chen et al. 2006, 2008a; Bonham 2013); the frequency distribution of biomass per quadrat follows the gamma distribution (Chen et al. 2008d; Chen and Shiyomi 2014), and the frequency distribution of individuals follows the negative binomial distribution (Chen et al. 2008d). In such cases, we can easily apply simple, censored methods as same as for the number of individuals (Shiyomi 1974, 1977), for cover (Chen et al. 2006, 2008c), and biomass (Shiyomi 1991, 1992).
In Eq. (1), for k = 0 the mean must be equal to the variance. If expressing the spatial heterogeneity value for the number of species by I = μ/λ (David and Moore 1954), I = 1 indicates “random pattern in spatial heterogeneity”, I < 1 indicates “lower heterogeneity than random pattern (uniform pattern)”, and I > 1 indicates “higher heterogeneity than random pattern (aggregated pattern; according to our experience I > 1 does not occur in the number of species per quadrat in a community)”. In our examples shown in Chen et al. (2016), three grasslands in Slovakia exhibited small I, four grasslands in Nishinasuno exhibited I near 1 although I were a little smaller than 1, grasslands in Shenmu is inbetween. A large k causes a low spatial heterogeneity (i.e., small I) because I = μ/(k + μ) → small for large k.
Measuring procedures and conclusions
The procedure is as follows: (1) In the herbaceous community of survey target, set N quadrats, each of which has a given area, where N was 50–100, and quadrat size was 0.01–0.25 m2 in our case. (2) Based on a preliminary or the past surveys, determine the FNS, k (usually an integer). (3) Count the number of species quadrat to quadrat, and classify quadrats into 3 classes: x k quadrats with k species, xk + 1 quadrats with k + 1 species, and xk+1+ quadrats with more than k + 1species; or in case of 4 classes: x k quadrats with k species, xk+1 quadrats with k + 1 species, xk+2 quadrats with k + 2 species and xk+2+ quadrats with more than k + 2 species. In an actual survey, we have to determine, in advance, which of n = 2 or n = 3 will be used. If k is previously known, it will be easy to determine the number of classes to be counted the number of species per quadrat. (4) Substitute x k to xk+1+ or x k to xk+2+ into Eq. (4), estimate μ and k + μ = λ (using computer programs written in Excel-BASIC or R by us: see Supplements ESM1 to ESM4). (5) Using Eqs. (5) and (6), estimate the 95% confidence intervals of μ and λ using the same program.
We explain this procedure using the survey in Shenmu (see Fig. 1 and Table 1), where actual counting and estimating were made to the number of species per quadrat. (1) We set 90 quadrats, each of which had 50 cm × 50 cm. (2) The minimum of number of species per quadrat was four, i.e., k = 4. (3) n must be ≥ 1. In our example, if we decide to survey based on n = 2, we classify quadrats into three classes of quadrats with 4, 5 or more than 5 species in counting; each class was composed of 5, 14 and 71 quadrats, respectively. If we decide to survey based on n = 3, we classify quadrats into four classes of quadrats with 4, 5, 6 and more than 6 species when counting in the grassland; each class was composed of 5, 14, 23 and 48, respectively (see Table 1). If k = 4 is previously known, it will be useful to determine the number of classes, n. (4) For n = 2, the following estimates were obtained using Eq. (4): \( \hat{\upmu } = \) 2.9181 and \( \hat{\lambda } \) = 6.9181; and for n = 3, \( \hat{\upmu } = \) 2.8431 and \( \hat{\lambda } \) = 6.8431 (see Table 1). Both the values for n = 2 and 3 are not very different. (5) The 95% confidence intervals for λ: for n = 2, 6.85–6.99, and for n = 3, 6.75–6.94 (Table 1). If we use n = 3, we need more time and labor for the survey than using n = 2, but we can obtain a narrower range for the confidence intervals than using n = 2, and usually the estimate for n = 3 will be near to one obtained by the complete survey compared to the estimate for n = 2. (5) The estimates of spatial heterogeneity values, I, for the number of species were 0.4218 for n = 2, and 0.4155 for n = 3. Both the values for n = 2 and 3 are not very different, here too.
References
Bonham DC (2013) Measurements for terrestrial vegetation, 2nd edn. Blackwell-Wiley, New York
Chen J, Shiyomi M (2014) Spatial pattern model of herbaceous plant mass as species level. Ecol Inform 24:124–131. https://doi.org/10.1016/j.ecoinf.2014.08.001
Chen J, Shiyomi M, Yamamura Y, Hori Y (2006) Distribution model and spatial variation of cover in grassland vegetation. Grassl Sci 52:167–173. https://doi.org/10.1111/j.1744-697X.2006.00065.x
Chen J, Okumura K, Takada H (2008a) Estimation of clover biomass and percentage in a performance trial of white clover-timothy binary mixtures: use of multiple regression equations incorporating plant cover measured with the grid-point plate. Grassl Sci 56:127–133. https://doi.org/10.1111/j.1744-697X.2010.00184.x
Chen J, Yamamura Y, Hori Y, Shiyomi M, Yasuda T, Zhou H-K, Li Y-N, Tang Y-H (2008b) Small-scale species richness and its spatial variation in an alpine meadow on the Qinghai-Tibet Plateau. Ecol Res 23:657–663. https://doi.org/10.1007/s11284-007-0423-7
Chen J, Shiyomi M, Bonham CD, Yasuda T, Hori Y, Yamamura Y (2008c) Plant cover estimation based on the beta distribution in grassland vegetation. Ecol Res 23:813–819. https://doi.org/10.1007/s11284-007-0443-3
Chen J, Shiyomi M, Hori Y, Yamamura Y (2008d) Frequency distribution models for spatial patterns of vegetation abundance. Ecol Model 211:403–410. https://doi.org/10.1016/j.ecolmodel.2007.09.017
Chen J, Gaborčík N, Shiyomi M (2016) A probability distribution model of small-scale species richness in plant communities. Ecol Inform 33:101–108. https://doi.org/10.1016/j.ecoinf.2016.04.003
Chytrý M, Dražil T, Hájek M et al (2015) The most species-rich plant communities in the Czech Republic and Slovakia (with new world records). Preslia 87:217–278
David F, Moore PG (1954) Notes on contagious distributions in plant populations. Ann Bot 18:47–53. https://doi.org/10.1093/oxfordjournals.aob.a083381
Gould W (2000) Remote sensing of vegetation, plant species richness and regional biodiversity hotspots. Ecol Appl 10:1861–1870. https://doi.org/10.2307/2641244
Katharina A, Engelhardt M, Kadlec JA (2001) Species traits, species richness and the resilience of wetlands after disturbance. J Aquat Plant Manag 39:36–39
Lavorel S (1999) Ecological diversity and resilience of Mediterranean vegetation to disturbance. Divers Distrib 5:3–13. https://doi.org/10.1046/j.1472-4642.1999.00033.x
Nicholas J, Gotelli NJ, Colwell RK (2011) Estimating species richness. In: Magurran AE, MacGill BJ (eds) Frontiers in measurement and assessment. Oxford University Press, Oxford
Oliver TH, Heard MS, Isaac NJB et al (2015) Biodiversity and resilience of ecosystem functions. Trends Ecol Evol 30:673–684. https://doi.org/10.1016/j.tree.2015.08.009
Pallett DW, Pescott OL, Schäfer SM (2016) Changes in plant species richness and productivity in response to decreased nitrogen inputs in grassland in southern England. Ecol Indic 68:73–81. https://doi.org/10.1016/j.ecolind.2015.12.024
Pielou EC (1977) Mathematical ecology. Wiley, New York
Shibayama M (2001) Estimation of leaf area and leaf inclination distributions of perennial ryegrass, tall fescue, and white clover canopies using an electromagnetic 3-D digitizer. Grassl Sci 47:303–306
Shibayama M (2003) Prediction of ratio of legumes in a mixed seeding pasture by polarized reflected light. Grassl Sci 49:229–237
Shibayama M, Akita S (2002) A portable spectropolarimeter for field crop canopies: distinguishing species and cultivars of fully developed canopies by polarized light. Plant Prod Sci 5:311–319. https://doi.org/10.1626/pps.5.311
Shiyomi M (1974) A method for estimating the number of individuals on the assumption of a Poisson or negative binomial distribution. Jpn J Appl Stat 18:106–114 (in Japanese with English summary)
Shiyomi M (1977) A graphic method of quick estimation of animal population density using a censored sample. Appl Entomol Zool 12:18–26. https://doi.org/10.1303/aez.12.18
Shiyomi M (1978) A rapid graphic estimation of density by a qusi-sequential method. Bull Natl Inst Agric Sci Ser A 25:33–57
Shiyomi M (1991) Method for the estimation of herbaceous biomass in grazed pasture by visual observation. Jpn J Grassl Sci 37:231–239
Shiyomi M (1992) Maximum likelihood estimation of herbaceous biomass using data obtained by visual observation in grazing pasture. Jpn J Grassl Sci 38:36–43
Tilman D, Reich PB, Knops J, Wedin D, Mielke T, Lehman C (2001) Diversity and productivity in a long-term grassland experiment. Science 294:843–845. https://doi.org/10.1126/science.1060391
van der Maarel E, Noest V, Palmer MW (1995) Variation in species richness on small grassland quadrats: niche structure or small-scale plant mobility? J Veg Sci 6:741–752. https://doi.org/10.2307/3236445
Yasuda T (2018) Semi-natural grassland vegetation mapping utilizing unmanned aerial vehicles and image analysis. Jpn J Grassl Sci (in press) (in Japanese)
Zobel M, Otsus M, Liina J, Moora M, Mőls T (2000) Is small-scale species richness limited by seed availability or micro-site availability? Ecology 81:3274–3282. https://doi.org/10.1890/0012-9658(2000)081[3274:ISSSRL]2.0.CO;2
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
About this article
Cite this article
Chen, J., Shiyomi, M. & Bai, H. A timesaving estimation of per-quadrat species number in grassland communities based on a Poisson-like model. Ecol Res 33, 427–434 (2018). https://doi.org/10.1007/s11284-017-1544-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11284-017-1544-2