Abstract
Nucleic acid demethylases of α-ketoglutarate-dependent dioxygenase (AlkB) family can reversibly erase methyl adducts from nucleobases, thus dynamically regulating the methylation status of DNA/RNA and playing critical roles in multiple cellular processes. But little is known about AlkB demethylases in filamentous fungi so far. The present study reports that Monascus purpureus genomes contain a total of five MpAlkB genes. The MpAlkB1 gene was disrupted and complemented through homologous recombination strategy to analyze its biological functions in M. purpureus. MpAlkB1 knockout significantly accelerated the growth of strain, increased biomass, promoted sporulation and cleistothecia development, reduced the content of Monascus pigments (Mps), and strongly inhibited citrinin biosynthesis. The downregulated expression of the global regulator gene LaeA, and genes of Mps biosynthesis gene cluster (BGC) or citrinin BGC in MpAlkB1 disruption strain supported the pleiotropic trait changes caused by MpAlkB1 deletion. These results indicate that MpAlkB1-mediated demethylation of nucleic acid plays important roles in regulating the growth and development, and secondary metabolism in Monascus spp.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Monascus spp. are traditional medicinal and edible filamentous fungi and widely used in food processing, brewing, medicine and health care and other fields in China and Southeast Asian countries (Shi et al. 2011). Monascus can produce a variety of valuable secondary metabolites, such as Monascus pigments (Mps), Monacolins, ergosterol, γ-aminobutyric acid, and so on (Cheng et al. 2010; Patakova 2013). Mps is one of the main beneficial secondary metabolites, which can be divided into yellow pigment, orange pigment and red pigment based on the color or the difference of maximum absorption wavelength. Besides as edible natural colorants, Mps has multiple functions such as lowering blood pressure, anti-inflammatory, antibacterial, inhibiting cancer and antioxidant (Cheng et al. 2011; Hsu et al. 2013; Tan et al. 2018; Choe et al. 2020). However, during the fermentation process, Monascus also produce a harmful secondary metabolite known as citrinin, which is a mycotoxin and has strong kidney and liver toxicity to mammals (Zhang et al. 2016). Food safety issues caused by citrinin has greatly restricted the application of Monascus-fermented products. Therefore, Mps and citrinin are two concerned secondary metabolites in Monascus spp. Although the biosynthesis gene clusters (BGCs) of some secondary metabolites have been revealed, especially Mps and citrinin, it is still unclear whether the epigenetics of nucleic acid methylation regulate the formation of these secondary metabolites.
It is generally acknowledged that the methylation of nucleic acid is the most common forms of epigenetic modification and plays crucial roles in multiple cellular processes through regulating gene expression, DNA replication, DNA repair and so on. Methyl adducts can be removed from the targets by the corresponding nucleic acid demethylase. The methylation levels and its dynamic changes controlled by the reversibility between methylation and demethylation are vital to regulate life processes (Luo et al. 2018). To date, the best-identified nucleic acid demethylases are α-ketoglutarate-dependent dioxygenases (AlkB), which belong to Fe2+/2-oxoglutarate-dependent (Fe2OG) dioxygenase domain-containing protein subfamily. AlkBs acting as nucleic acid repair enzymes can remove alkyl adducts, predominately methyl substituent, to regenerate the natural nucleotides (Delaney et al. 2005; Fu et al. 2013). AlkB was first identified as a DNA repair enzyme to reverse DNA damage through oxidative demethylation in E. coli (Falnes et al. 2002; Trewick et al. 2002). Subsequently, an increasing number of studies have confirmed that AlkB homologies are prevalent in eukaryote and prokaryote, and can catalyze the demethylation of both DNA and RNA substrates (Michalak et al. 2019).
Functional studies on AlkB demethylases are an important way to investigate the physiological processes regulated by epigenetics of nucleic acid methylation in given organism. In recent years, multiple roles of AlkB family in regulating physiological and pathological processes have been discovered in mammals and plants (Xie et al. 2020). However, little is known about the functions of AlkB demethylase in filamentous fungi. In the present study, a demethylase AlkB gene MpAlkB1 was identified from M. purpureus. To analyze the biological functions of MpAlkB1, its disruption strain (ΔMpAlkB1) and complementation strain (ΔMpAlkB1::MpAlkB1) were successfully obtained by Agrobacterium-mediated homologous recombination technology. The results revealed that MpAlkB1 is involved in regulating the growth and development of Monascus and the formation of secondary metabolites. This work provides a sound basis for exploring the physiological functions and regulatory mechanism of nucleic acid methylation epigenetics in Monascus filamentous fungi.
Materials and methods
Strains, medium, and culture conditions
M. purpureus CGMCC 3.15548 (China General Microbiological Culture Collection Center) was used for performing the MpAlkB1 gene deletion. Potato dextrose agar medium (PDA) and malt extract agar medium (MA) were used for morphological characterization. All Monascus strains were maintained on PDA slants at 30 °C for sporulation, producing Mps and citrinin, and total DNA extraction. Escherichia coli DH5α was the host of recombinant DNA manipulation. Agrobacterium tumefaciens AGL-1 was used for fungal transformation.
Gene acquisition and phylogenetic analysis
The genome of M. purpureus was re-annotated based on genome sequencing data (GenBank assembly accession: GCA_011319195.1) by the fungal genome annotation software (Funannotate v1.8.13). Then the sequences of Fe2+ and 2-oxoglutarate (2OG)-dependent dioxygenase domain-containing proteins were extracted from the annotation results, and the enzymes with 2OG-FeII_Oxy_2 domain (Pfam ID: PF13532) ware classified to AlkB subfamily. The 3D structure of MpAlkB1 was modeled by online tool (https://swissmodel.expasy.org/) using the real protein structure of human HpALKBH1 (PDB ID: 6IE2.4) as a homology template. Protein sequence was aligned performed by ClustalX 2.1 and visualized by ESPript 3.0 (https://espript.ibcp.fr/ESPript/ESPript/). The phylogenetic tree was constructed using Neighbor-Joining (NJ) method of MEGA X software and visualized by iTOL 6.8.2 (https://itol.embl.de/).
DNA extraction
Fungal genomic DNA was isolated from mycelia grown on PDA plates covered with cellophane membranes using the cetyltrimethylammonium bromide (CTAB) method.
MpAlkB1 disruption and complementation
MpAlkB1 was deleted from genome through homologous recombination strategy. The 5ʹ and 3ʹ flanking regions (985 bp and 989 bp) of MpAlkB1 were respectively cloned into the upstream and downstream of the hygromycin phosphotransferase gene hph cassette in pAg1-H3 vector through Kpn I/Xho I and Spe I/Hind III sites to generate the disruption vector pAg1-H3-AlkB1K (Fig. 1A). Subsequently, the vector was delivered into wild-type (WT) M. purpureus via Agrobacterium tumefaciens-mediated transformation (ATMT) method to generate the MpAlkB1 gene deletion strain (ΔMpAlkB1) as previously described (Shao et al. 2009). The transformants were selected on PDA medium containing 50 µg/mL hygromycin B. ΔMpAlkB1 stains were screened by PCR and DNA blotting for MpAlkB1 gene replacement by the hph cassette. Probe preparation, membrane hybridization and visualization were performed according to the manufacturer’s instructions of the DIG-High Prime DNA Labeling & Detection Starter Kit I (Roche, Switzerland).
MpAlkB1 belongs to AlkB subfamily. (A) Phylogenetic tree of Fe2+/2-oxoglutarate-dependent dioxygenases from A. nidulans and M. purpureus. The light blue shading indicates the AlkB subfamily. The names beginning with FUN00 indicates the 2OGD proteins of M. purpureus obtained by genome reannotation. The conserved domain of each protein is exhibited in the corresponding position on the periphery of the phylogenetic tree, the blue box represents the 2OG-FeII_Oxy_2 domain, and the gray box represents the 2OG-FeII_Oxy domain. (B) Sequence alignments of AlkB proteins. Gray shading indicates conserved domain sequences. The spatial structure sites of MpAlkB1 and HsALKBH1 are shown above the corresponding sequences. The UniProt identifiers for the proteins: AnAlkB1 (Q5B672), SpAlkB1 (O60066), HsALKBH1 (Q13686), and EcAlkB (P05050). (C) Three-dimensional structures of MpAlkB1 and human HsALKBH1 (6IE2) proteins. Superposition of structures indicates the structural similarity between MpAlkB1 (blue) and HsALKBH1 (red) compared by TM-align online (https://zhanggroup.org/TM-align/)
To complement ΔMpAlkB1, the complete MpAlkB1 gene including promoter, coding sequence and terminator, neomycin phosphotransferase resistance gene neo cassette, and downstream region of MpAlkB1 were recombined with the pAg1-H3 vector by In-Fusion cloning technology. The resulting construct was transformed into the ΔMpAlkB1 strain via ATMT. The complement stains (ΔMpAlkB1::MpAlkB1) were selected by 15 µg/mL G418 sulfate and confirmed by PCR and DNA blotting. The expression levels of MpAlkB1 were detected by quantitative real-time PCR (qRT-qPCR). Primers used in construction of vectors and verification of engineered strains were listed in Table S1. Primers used for preparing the probes were listed in Table S2.
Morphology observation and growth rate analysis
To prepare spore suspensions, the WT, ΔMpAlkB1 and complemented strains were incubated on PDA agar plate at 30 °C. After 10 days culture, fresh spores were harvested from each plate with 7 mL of sterilized distilled water and filtered through lens wiping paper to remove hyphae. Then, spores were collected by centrifugation and resuspended to obtain a final concentration of 1 × 105 spores/mL. To analyses colony morphology, 5 µL spore suspension for each strain were pipetted onto PDA and MA plates, which were incubated at 30 °C and observed the phenotypes for 14 days. For the growth rate analysis, the colony diameters of each strain cultured on PDA plates were measured on the 4th, 6th, 8th, 10th, 12th and 14th days. To measure the biomass, 100 µL of spore suspension for each strain was inoculated into 30 mL PDB liquid medium and incubated at 30 °C with shaking at 150 rpm. The culture mediums were taken every other day and filtered to obtain mycelium, which were dried and weighed. All assays above were repeated in triplicate at least.
For micro-morphological analysis, 5 µL of spore suspension of each strain was inoculated on PDA medium covered with cellophane membrane and cultured at 30 °C. The morphologies of mycelia, conidia and cleistothecia were observed using M150 upright microscope (Motic, China). Meanwhile, spores were isolated from mycelia and their numbers were counted by hemacytometer at least three times for each sample. The yields of spores were calculated, and the values of different strains were compared.
Measurement of mps and citrinin
100 µL spore suspension of each strain were coated on PDA plates covered with cellophane membranes and cultured at 30 °C. The mycelia and media were taken every 2 days, beginning at 4th day. The collected samples were dried by vacuum freeze drying (−56 °C, 3 Pa, 24 h; SCIENTZ-10 N, Scientz, China) and ground into powder. Mps yields were detected according to the method described in the Chinese standard (GB1886.15-2015) and Xu et al. (2022) with a slight modification. Appropriate amount of dry mycelium or 0.1 g media accurately weighed was treated with 1 mL 70% ethanol (v/v) and incubated at 60 °C for 1 h. After centrifugation, the supernatant was filtrated by 0.22 μm organic filter membrane. The intracellular and extracellular Mps extracts were diluted with 70% ethanol, and their absorbance values of the red, orange, and yellow pigments were measured at the specific wavelengths 510, 470, and 380 nm, respectively. The color value was calculated using the formula [X (U/g) = OD (nm) × dilution ratio/Monascus powder quality (g)].
To evaluate whether MpAlkB1 knockout affected the formation of citrinin, LC-MS 8060 (Shimadzu, Japan) was employed to measure the citrinin content in mycelium. 0.01 g of mycelium powder was treated by 1.5 mL of methanol and shaken for 30 min. After centrifugation, the supernatant was filtered through 0.22 μm membrane, and put into liquid phase sampling bottle for LC-MS detection. Shim-pack™ GISS C18 (2.1 × 100 mm, 1.9 μm) column was used. The LC-MS parameters were as follows: mobile phase, 75% acetonitrile/25% water (v/v) (pH 2.5, adjusted by orthophosphoric acid); flow rate, 0.2 mL/min; column temperature, 35 °C. The elution was monitored using a fluorescence detector at an emission wavelength of 500 nm and an excitation wavelength of 331 nm. A citrinin standard (Yuanye, China) was used to verify the LC-MS analysis.
Gene expression analysis by quantitative real-time PCR (qRT-PCR)
Strains were grown on PDA plates at 30 °C, and total RNA of mycelia was extracted using Fungal RNA Extraction Kit.E.Z.N.A.® (Omega, USA). Removal of residual genomic DNA and synthesis of first-strand cDNA was performed by PrimeScript™ RT reagent Kit (Takara, Japan). The qRT-PCR was carried out using SYBR green master mix (Takara, Japan) on a CFX96 Touch™ Real-Time PCR Detection System (Bio-Rad, USA) with the following steps: 95 °C for 2 min, followed by 40 cycles (95 °C for 15 s and 58 °C for 40 s) and one cycle (95 °C for 15 s and 60 °C for 15 s). Samples were analyzed in triplicate, and three technical replicates were run for each sample. The average Ct values was counted and the data were analyzed using the 2−ΔΔCt method. Relative gene expression levels were normalized against the expression of the housekeeping gene GAPDH. The primers used in qRT-PCR were listed in Table S3.
Statistics analysis
For the significant difference analysis between ΔMpAlkB1 and WT control strains, the corresponding experimental data were analyzed using a student’s t test. Statistical differences were considered as significant when * P < 0.05 or ** P < 0.01.
Results
MpAlkB1 belongs to AlkB subfamily of Fe2+/2-oxoglutarate-dependent dioxygenase protein family
Based on the re-annotation of genome, a total of 24 Fe2+/2-oxoglutarate-dependent dioxygenase (Fe2OGDio) domain-containing proteins were found in M. purpureus. Among them, five members contain 2OG-FeII_Oxy_2 (Pfam number: PF13532) domain, which is the characteristic conserved domain of nucleic acid demethylase AlkB subfamily. However, the other Fe2OGDio members all contain the conserved domain of 2OG-FeII_Oxy (Pfam number: PF03171). The phylogenetic tree of Fe2OGDio proteins from M. purpureus and filamentous fungi model organism Aspergillus nidulans showed that MpAlkB1 belongs AlkB subfamily (Fig. 1A). A 1,635 bp full-length sequence of MpAlkB1 consists of 2 exons and 1 intron, and encodes a deduced protein of 353 amino acids. The well-conserved 2OG-FeII_Oxy_2 domain made of 240 amino acids accounts for most of the whole sequence of MpAlkB1 and is closer to the C-terminus, consistent with the corresponding domains of Aspergillus nidulans AnAlkB1, Schizosaccharomyces pombe SpAlkB1, Homo sapiens HsALKBH1, and Escherichia coli EcAlkB (Fig. 1B). The result of homology modeling and structure comparison showed that the spatial structure of MpAlkB1 is very similar to that of human HsALKHB1 (PDB ID: 6IE2) (Fig. 1C). Especially, the conserved domain regions of them have consistent structural characteristics, that is, 11 β-folds form a double-stranded β-helix domain (DSBH, also referred to as a ‘jelly-roll fold’) (Fig. 1B, C). The key catalytic residues interacting with Fe2+ and 2-oxoglutarate are located inside of the DSBH domain and surrounded by multiple α-helix structures on both sides of the domain (Shen et al. 2014). It is further proved that MpAlkB1 is a member of the nucleic acid demethylase AlkB subfamily, so it is likely to have the corresponding demethylase activity and functions in M. purpureus.
Verification of the MpAlkB1 deletion and complementation strains
To analyze the function of MpAlkB1 in M. purpureus, a homologous recombination strategy was employed to generate MpAlkB1 disruption strain (ΔMpAlkB1) and its complementation strain (ΔMpAlkB1::MpAlkB1) (Fig. 2A). The successfully constructed strains were confirmed by PCR and southern blot. As indicated in Fig. 2B, the expected PCR products of MpAlkB1 were obtained from WT and ΔMpAlkB1::MpAlkB1, but not from ΔMpAlkB1; the target fragments of resistant gene hph and neo could be amplified from ΔMpAlkB1 and ΔMpAlkB1::MpAlkB1, respectively; moreover, amplicons of 2.5 kb, 3.7 kb and 4.8 kb were respectively obtained from WT, ΔMpAlkB1 and ΔMpAlkB1::MpAlkB1 using primer pair LAlkB1-5 F/LAlkB1-3R annealed to homologous arms. To further demonstrate the occurrence of the homologous recombination events, the genomic DNA digested by Hind III was subjected to Southern blot. As showed in Fig. 2C, a single hybridization band was detected not only in WT but also in ΔMpAlkB1::MpAlkB1 using probe 1 targeting the coding region of MpAlkB1, while no fragment was obtained in ΔMpAlkB1; in addition, the target band was detected only in ΔMpAlkB1 with probe 2 (probe hph), and only in ΔMpAlkB1::MpAlkB1 with probe 3 (probe neo). Both the results of PCR and Southern blot showed that the disruption mutant and complementary strain were successfully constructed.
Construction and confirmation of the MpAlkB1 disruption mutant and complementation. (A) Construction strategy of homologous recombination. The short arrows indicate the primer pairs for transformant detection. Hind III and EcoR I indicate the corresponding restriction sites. hph, hygromycin phosphotransferase gene; neo, neomycin phosphotransferase resistance gene. (B) Verification of homologous recombination events by PCR. The primer pairs used are shown at the top of the map. (C) Southern blot analysis using MpAlkB1 probe (probe 1), hph probe (probe 2) and neo probe (probe 3). Genomic DNA was digested by Hind III for hybridization with probe 1 or probe 3, and digested by EcoR I for hybridization with probe 2. M, DNA marker; lane 1, wild-type (WT); lane 2, ΔMpAlkB1; lane 3, ΔMpAlkB1::MpAlkB1. (D) Expression analysis of MpAlkB1 gene by qRT-PCR. Each value represents the mean ± SE (standard error) of three biological replicates. Asterisks represent significant differences between the transformed strain and WT strain at same growth stage (* P < 0.05, ** P < 0.01, Student’s t-test)
Expression pattern of MpAlkB1 in wild-type strain, ΔMpAlkB1, and ΔMpAlkB1::MpAlkB1
The transcript abundance of MpAlkB1 in WT, ΔMpAlkB1, and ΔMpAlkB1::MpAlkB1 were analyzed by qRT-PCR. In the WT strain, the expression of MpAlkB1 gradually weakens with the growth and development, while keeps steady level at the later growth stage of 12 to 14 days (Fig. 2D), which indicated that MpAlkB1 may be involved in regulating growth and development of M. purpureus. No transcript of MpAlkB1 was detected in ΔMpAlkB1, while the expression level of MpAlkB1 in ΔMpAlkB1::MpAlkB1 was almost consistent with that in WT (Fig. 2D), which further proved the success of MpAlkB1 gene knockout and complementation.
MpAlkB1 knockout promoted the growth and development of M. Purpureus
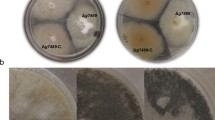
To investigate the influences of MpAlkB1 deletion on developmental processes, the colony phenotypes of ΔMpAlkB1, ΔMpAlkB1::MpAlkB1 and WT strains grown on PDA and MA media were observed. Among the three strains, the most readily visible morphological difference was that the growth of ΔMpAlkB1 strain was obviously accelerated and its colony was the largest (Fig. 3A). In the same growth period of 4 to 14 days, the colony diameter of ΔMpAlkB1 was about 1.4–1.8 times that of WT strain, and the final colony size of ΔMpAlkB1 was also significantly larger than that of WT control (Fig. 3B). In addition, the biomass of the three strains cultured in PDB liquid medium was measured. The results showed that the biomass of ΔMpAlkB1 increased by about 40–65% compared with that of WT strain in the same growth period (Fig. 3C). No significant phenotypic difference between the complementation strain (ΔMpAlkB1::MpAlkB1) and the WT strain indicated that the enhancement of growth and development in ΔMpAlkB1 strain was caused by the deletion of MpAlkB1 gene.
Comparison of MpAlkB1 transformants and WT strains in colony phenotype and biomass. (A) Colony morphology of ΔMpAlkB1, ΔMpAlkB1::MpAlkB1 and WT strains on potato dextrose agar (PDA) and malt extract ager (MA) media after cultivating at 30 °C for 14 days. (B) The colony diameter of strains grown on PDA plates. (C) Biomass of strains incubated in PDB liquid medium. The error bar represents the standard deviations of three biological replicates. Significant analysis was performed between MpAlkB1 transformant and WT strain at the same growth stage (** P < 0.01, Student’s t-test)
To investigate the potential physiological reasons of MpAlkB1 knockout promoting growth and development, the microstructures of ΔMpAlkB1, ΔMpAlkB1::MpAlkB1 and wild-type strains were observed microscopically. There were no visible difference in mycelium thickness, branching ability and spore morphology among the three strains, while the spores of ΔMpAlkB1 strain were obviously denser than those of WT strain or ΔMpAlkB1::MpAlkB1 strain (Fig. 4A). The statistical results showed that the spore yield of ΔMpAlkB1 was 20 times higher than that of WT or ΔMpAlkB1::MpAlkB1 (Fig. 4B). The cleistothecia of ΔMpAlkB1 was larger, contained more ascospores and matured earlier than that of WT strain (Fig. 4C), which implied that MpAlkB1 may play a role in regulating sexual development. These results indicated that MpAlkB1 knockout promoted the growth and development of M. purpureus through enhancing the sexual growth and sporulation.
Microscopic morphology of spores and cleistothecia. (A) Micromorphology of hypha and conidiospores. The arrows indicate the conidiospores. (B) The spore yields. The error bars represent the standard deviations of three biological replicates. The spore yield of each transformed strain was compared with WT for significance analysis (** P < 0.01, Student’s t-test). (C) Microscopic structure of cleistothecia and ascospores. The arrows indicate the cleistothecia. All strains analyzed were cultivated on PDA at 30 °C for 10 days
MpAlkB1 deletion reduced the production of MPs and citrinin
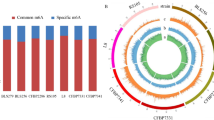
To investigate whether MpAlkB1 deletion affected the formation of secondary metabolites, the contents of Mps and citrinin, the two most representative secondary metabolites of Monascus spp., were determined at different growth stages of WT and transformed strains. As shown in Fig. 5, both intracellular and extracellular pigment accumulations of ΔMpAlkB1 strain increased with the growth time before the 12th day, which was consistent with the change trend of pigment contents of WT and ΔMpAlkB1::MpAlkB1 strain. Nevertheless, the intracellular and extracellular yields of ΔMpAlkB1, whether red, orange or yellow pigment, were less than that of WT. In addition, the decrease degree of intracellular pigment contents of ΔMpAlkB1 strain were greater than that of its extracellular pigments. Compared with WT, the intracellular red, orange and yellow pigments of ΔMpAlkB1 decreased by about 86%, 91% and 89% on the 4th day of growth, respectively, and by approximately 18-61% in other growth stages (Fig. 5A–C), while the extracellular pigment production of ΔMpAlkB1 decreased by 12-41% during the growth stage from 4th to 14th day (Fig. 5D–F). After MpAlkB1 gene was reverted in ΔMpAlkB1::MpAlkB1, the intracellular pigment synthesis ability was restored (Fig. 5A–C).
Contents of intracellular and extracellular pigments of strains in different growth stages. (A, B, C) The yields of intracellular red, orange and yellow pigments, respectively; (D, E, F) The yields of extracellular red, orange and yellow pigments, respectively. Each value represents the mean ± SE (standard error) of three biological replicates. Asterisks represent significant differences between the transformed strain and WT strain at same growth stage (* P < 0.05, ** P < 0.01, Student’s t-test)
To analyze whether MpAlkB1 regulates citrinin formation, the intracellular citrinin contents were quantitatively detected by LC-MS. The results shown in Fig. 6, the content of citrinin in each strain increased steadily with the extension of culture time and reached the highest on the 12th day. Citrinin in ΔMpAlkB1 strain was almost undetectable before 12 days of growth, and even on the 12th day when the citrinin content was the maximum, it was only about 6% of that in the WT strain. With the resumed expression of MpAlkB1 gene, the ability of ΔMpAlkB1::MpAlkB1 strain to accumulate citrinin was restored. These results indicated that MpAlkB1 disruption downregulated the yields of Mps and severely inhibited the formation of citrinin.
The citrinin content of ΔMpAlkB1 strain decreased significantly. Each value represents the mean ± SE (standard error) of three biological replicates. Asterisks represent significant differences between the transformed strain and WT strain at same growth stage (* P < 0.05, ** P < 0.01, Student’s t-test)
The expression levels of genes related to growth and development and biosynthesis of mps and citrinin were downregulated in ΔMpAlkB1 strain
Knowing that methylation modification of nucleic acid plays an indispensable role in regulating gene expression to participate in various biological processes, the transcript levels of related genes were tested by qRT-PCR. LaeA, a putative riboprotein methyltransferase, is the first identified global regulator in filamentous fungi (Bok et al. 2004). LaeA gene knockout from Monascus ruber significantly accelerated growth, produced more aerial hyphae and conidia, and drastically reduced the production of multiple secondary metabolites, especially Mps and citrinin (Liu et al. 2016b). LaeA overexpression produced more secondary metabolites in Monascus pilosus and Monascus purpureus (Lee et al. 2013; Zhang et al. 2020a). These studies demonstrate that LaeA is a positive regulator of secondary metabolites and negatively regulates the growth in Monascus spp. Here, the level of LaeA transcript was notably reduced by more than 50% in ΔMpAlkB1 strain compared with that in WT strain throughout the growth process (Fig. 7A), supporting the results of the reduced yields of Mps and citrinin, and accelerated growth caused by MpAlkB1 knockout.
The expression levels of genes related to growth and development or biosynthesis of Mps and citrinin in ΔMpAlkB1, ΔMpAlkB1::MpAlkB1 and WT at different growth stages. The transcript levels of (A) LaeA, (B) PKS5, (C) mppR1, (D) pksCT, (E) ctnA, (F) ctnB, and (G) ctnR were detected by qRT-PCR. The error bars represent the standard deviations of three biological replicates. Asterisks represent significant differences between the transformed strain and WT strain at same growth stage (* P < 0.05, ** P < 0.01, Student’s t-test)
Since the yields of Mps and citrinin decreased in ΔMpAlkB1 (Figs. 5 and 6), the expression levels of genes belonging to the biosynthetic gene clusters (BGCs) of these two secondary metabolites were analyzed. Inactivation of the polyketide synthase gene MpPKS5 or the transcriptional regulation gene mppR1 in Monascus spp. leaded to albino mutants, confirming that these two gene in Mps BGC are crucial to pigment biosynthesis (Balakrishnan et al. 2013; Chen et al. 2019a). In ΔMpAlkB1 strain, the expression levels of MpPKS5 and mppR1 were markedly decreased at all growth stages (Fig. 7B, C), which agreed with the decrease of Mps.
In citrinin BGC, the polyketide synthase gene pksCT and ctnB (structure and function similar gene with pksCT) are two core catalytic genes, and ctnA and ctnR encode two transcriptional activators (Shimizu et al. 2007). Knockout of pksCT or ctnB leaded to the inability to produce citrinin in M. purpureus (Shimizu et al. 2005; Li et al. 2013). The ctnA deletion resulted in a decrease expression of several gene in citrinin BGC including pksCT and leaded to a significant decrease of citrinin production, suggesting that ctnA is the main transcription activator in citrinin biosynthesis pathway (Shimizu et al. 2007). The regulatory gene ctnR was also necessary for synthesis of citrinin in Monascus spp (Hong et al. 2023). Hence these four genes were selected to analyze their expression levels by qRT-PCR. As shown in Fig. 7D–G, the expressions of all four genes tested were sharply inhibited in the whole growth process of ΔMpAlkB1 strain, especially the transcripts of pksCT and ctnA were undetectable (Fig. 7D, E), and CtnB was only weakly expressed in ΔMpAlkB1 strain compared with that in WT strain (Fig. 7F). The sharp downregulation of gene expression in citrinin BGC was in line with the striking inhibition of citrinin biosynthesis in ΔMpAlkB1 strain.
Considering that MpAlkB1 belongs to the nucleic acid demethylase AlkB subfamily (Fig. 1), we speculated that MpAlkB1 plays an important role in the growth and development of the strain and the biosynthesis of secondary metabolites by mediating the expression of related genes through demethylation.
Discussion
Up to now, the epigenetic studies of nucleic acid methylation have mainly focused on mammals, followed by plants. Though as early as 1993, it was found that about 1.5% of cytosine in the genome of Neurospora crassa, a classical model organism of filamentous fungi, was methylated (Foss et al. 1993), the progress of subsequent research on nucleic acid methylation of filamentous fungi was very slow, with few reports. In recent ten years, the applications of methyltransferase inhibitors such as 5-azacytidine (5-AZA) have indirectly revealed that the epigenetics of nucleic acid methylation may play important roles in regulating gene expression, growth and development, and metabolism of filamentous fungi. For example, 5-AZA treatments could enhance the transcription of cellulase and xylanase genes of Humicola grisea grown on glucose (Manfrão-Netto et al. 2018), increase the production of diethylene glycol phthalate ester monomers and oligomers of the marine-derived fungus Cochliobolus lunatus TA26-46 (Chen et al. 2016), and affect the development and secondary metabolism of Aspergillus flavus (Lin et al. 2013). The 5-AZA even could make Aspergillus sp. SCSIOW3 and plant endophytic fungus Pestalotiopsis crassiuscula produce a new diphenylether-O-glycoside and a new coumarin, respectively (Yang et al. 2014; Li et al. 2017). Recently, overexpression of RNA m6A methyltransferase gene MTA1 in Fusarium graminearum delayed spore germination and reduced hyphal branching (Kim et al. 2024). All these indicated that epigenetics of nucleic acid methylation exists in filamentous fungi and has important functions. The present study revealed that M. purpurus MpAlkB1 belongs to nucleic acid demethylase family (Fig. 1), and MpAlkB1 knockout promoted strain growth (Fig. 3), spore formation (Fig. 4A, B), and cleistothecia development (Fig. 4C), and inhibited the biosynthesis of secondary metabolites (Figs. 5 and 6), suggesting that MpAlkB1-mediated demethylation of nucleic acid plays important roles in regulating the growth and secondary metabolism.
It is essential for maintaining normal gene expression and life process that the DNA/RNA demethylases act with methyltransferases to dynamically regulate the methylation status of nucleic acids (Luo et al. 2018). So far, the well identified nucleic acid demethylases are AlkB oxygenases, which are critical in numerous cellular processes (Perry et al. 2021). AlkB family contains multiple members with different substrate specificity, which indicates that AlkB demethylases has complex biological functions and regulatory mechanisms. For example, human AlkB family contains nine members, including ALKBH1-8 and FTO (fat mass and obesity-associated). ALKBH1 targets ssDNA/ssRNA-m3C (N3-methylcytidine) (Westbye et al. 2008), tRNA-m1A (N1-methyladenosine) (Liu et al. 2016a), and bulged DNA-m6A (N6-methyladenosine) (Zhang et al. 2020b), and is implicated in gastric cancer and glioblastoma (Xie et al. 2018; Li et al. 2019); ALKBH2 and 3 can remove m1A, m5C (5-methylcytidine) and m3C modifications, but they respectively prefer DNA and RNA, and regulate tumorigenesis (Monsen et al. 2010; Chen et al. 2019b); ALKBH4 and 5 demethylate DNA-m6A and mRNA-m6A, respectively (Tang et al. 2018; Kweon et al. 2019), and are closely related to spermatogenesis and colon cancer (Yang et al. 2020); FTO acts on m1A, m6A, m6Am (N6,2′-O-dimethyladenosine) and m3U (N3-methyluridine) in RNA, and m3T (N3-methylthymidine) in DNA (Jia et al. 2008; Wei et al. 2018), and is strongly associated with obesity, type II diabetes, Alzheimer’s disease, and nonalcoholic steatohepatitis (Keller et al. 2011; Smemo et al. 2014; Lim et al. 2016). These research experiences in mammals indicated that identifying the biological functions and substrate specificities of AlkB demethylases can effectively and deeply analyze the function and regulatory mechanism of nucleic acid methylation epigenetics in specific physiological/pathological processes. Our present study found that Monascus AlkB family contains five members (Fig. 1A), and confirmed that MpAlkB1 plays key roles in regulating the growth and development, and the formation of secondary metabolites (Figs. 2, 3, 4, 5 and 6), which can encourage to explore the physiological functions and molecular mechanism of nucleic acid methylation through further analyzing the functional divisions among AlkB members in Monascus spp.
Nucleic acid demethylase functions through demethylating its target DNA or RNA (Perry et al. 2021). Therefore, identification of the related target genes (DNA or RNA) and methylation types targeted by the AlkB demethylase is the key to reveal its regulatory mechanism in a given physiological or biosynthesis process. The downregulated expression of a global regulator gene LaeA, Mps BGC genes, and citrinin BGC genes, especially the nearly undetectable transcripts of pksCT, ctnA, and ctnB in ΔMpAlkB1 strain (Fig. 7), indicated that these genes are regulated by MpAlkB1. However, to determine whether these genes or their RNA are directly regulated by MpAlkB1, it is necessary to analyze whether these genes are methylated and their methylation types, as well as the methylation types of MpAlkB1 demethylation. Due to various types of DNA and RNA methylation modification and the functional diversity of AlkB family members, it is a very challenging research based on the current technology.
Data availability
No datasets were generated or analysed during the current study.
References
Balakrishnan B, Karki S, Chiu S et al (2013) Genetic localization and in vivo characterization of a Monascus azaphilone pigment biosynthetic gene cluster. Appl Microbiol Biot 97:6337–6345. https://doi.org/10.1007/s00253-013-4745-9
Bok JW, Keller NP (2004) LaeA, a regulator of secondary metabolism in aspergillus spp. Eukaryot Cell 3:527–535. https://doi.org/10.1128/EC.3.2.527-535.2004
Chen M, Zhang W, Shao C et al (2016) DNA methyltransferase inhibitor induced fungal biosynthetic products: diethylene glycol phthalate ester oligomers from the marine-derived fungus Cochliobolus Lunatus. Mar Biotechnol 18:409–417. https://doi.org/10.1007/s10126-016-9703-y
Chen W, Feng Y, Molnár I et al (2019a) Nature and nurture: confluence of pathway determinism with metabolic and chemical serendipity diversifies Monascus azaphilone pigments. Nat Prod Rep 36:561–572. https://doi.org/10.1039/c8np00060c
Chen Z, Qi M, Shen B et al (2019b) Transfer RNA demethylase ALKBH3 promotes cancer progression via induction of tRNA-derived small RNAs. Nucleic Acids Res 47:2533–2545. https://doi.org/10.1093/nar/gky1250
Cheng C, Pan T (2011) Protective effect of Monascus-fermented red mold rice against alcoholic liver disease by attenuating oxidative stress and inflammatory response. J Agr Food Chem 59:9950–9957. https://doi.org/10.1021/jf202577t
Cheng M, Wu M, Chen I et al (2010) Secondary metabolites from the red mould rice of Monascus Purpureus BCRC 38113. Nat Prod Res 24:1719–1725. https://doi.org/10.1080/14786410902941477
Choe D, Jung HH, Kim D et al (2020) In vivo evaluation of the anti-obesity effects ofcombinations of Monascus pigment derivatives. Rsc Adv 10:1456–1462. https://doi.org/10.1039/c9ra08036h
Delaney JC, Smeester L, Wong C et al (2005) AlkB reverses etheno DNA lesions caused by lipidoxidation in vitro and in vivo. Nat Struct Mol Biol 12:855–860. https://doi.org/10.1038/nsmb996
Falnes PØ, Johansen RF, Seeberg E (2002) AlkB-mediated oxidative demethylation reverses DNA damage in Escherichia coli. Nature 419:178–182. https://doi.org/10.1038/nature01048
Foss HM, Roberts CJ, Claeys KM et al (1993) Abnormal chromosome behavior in Neurospora mutants defective in DNA methylation. Science 262:1737–1741. https://doi.org/10.1126/science.7505062
Fu Y, Jia G, Pang X et al (2013) FTO-mediated formation of N6-hydroxymethyladenosine and N6-formyladenosine in mammalian RNA. Nat Commun 4:1798. https://doi.org/10.1038/ncomms2822
Hong X, Guo T, Xu X et al (2023) Multiplex metabolic pathway engineering of Monascus pilosus enhances lovastatin production. Appl Microbiol Biot 107:6541–6552. https://doi.org/10.1007/s00253-023-12747-2
Hsu L, Liang Y, Hsu Y et al (2013) Anti-inflammatory properties of yellow and orange pigments from Monascus purpureus NTU 568. J Agr Food Chem 61:2796–2802. https://doi.org/10.1021/jf305521v
Jia G, Yang C, Yang S et al (2008) Oxidative demethylation of 3-methylthymine and 3-methyluracil in single-stranded DNA and RNA by mouse and human FTO. Febs Lett 582:3313–3319. https://doi.org/10.1016/j.febslet.2008.08.019
Keller L, Xu W, Wang H et al (2011) The obesity related gene, FTO, interacts with APOE, and is associated with Alzheimer’s disease risk: a prospective cohort study. J Alzheimer’s Disease: JAD 23:461–469. https://doi.org/10.3233/JAD-2010-101068
Kim H, Hu J, Kang H et al (2024) Phylogenetic and functional analyses of N6-methyladenosine RNA methylation factors in the wheat scab fungus fusarium graminearum. Msphere 9:e55223. https://doi.org/10.1128/msphere.00552-23
Kweon S, Chen Y, Moon E et al (2019) An adversarial DNA N6-methyladenine-sensor network preserves polycomb silencing. Mol Cell 74:1138–1147. https://doi.org/10.1016/j.molcel.2019.03.018
Lee SS, Lee JH, Lee I (2013) Strain improvement by overexpression of the laeA gene in Monascus pilosus for the production of monascus-fermented rice. J Microbiol Biotechn 23:959–965. https://doi.org/10.4014/jmb.1303.03026
Li Y, Pan Y, Zou L et al (2013) Lower citrinin production by gene disruption of ctnB involved in citrinin biosynthesis in Monascus Aurantiacus Li AS3.4384. J Agr Food Chem 61:7397–7402. https://doi.org/10.1021/jf400879s
Li X, Xia Z, Tang J et al (2017) Identification and biological evaluation of secondary metabolites from marine derived fungi-aspergillus sp. SCSIOW3, cultivated in the presence of epigenetic modifying agents. Molecules 22. https://doi.org/10.3390/molecules22081302
Li Y, Zheng D, Wang F et al (2019) Expression of demethylase genes, FTO and ALKBH1, is associated with prognosis of gastric cancer. Digest Dis Sci 64:1503–1513. https://doi.org/10.1007/s10620-018-5452-2
Lim A, Zhou J, Sinha RA et al (2016) Hepatic FTO expression is increased in NASH and its silencing attenuates palmitic acid-induced lipotoxicity. Biochem Bioph Res Co 479:476–481. https://doi.org/10.1016/j.bbrc.2016.09.086
Lin J, Zhao X, Zhi Q et al (2013) Transcriptomic profiling of aspergillus flavus in response to 5-azacytidine. Fungal genetics and biology. Fungal Genet Biol 56:78–86. https://doi.org/10.1016/j.fgb.2013.04.007
Liu F, Clark W, Luo G et al (2016a) ALKBH1-mediated tRNA demethylation regulates translation. Cell 167:816–828. https://doi.org/10.1016/j.cell.2016.09.038
Liu Q, Cai L, Shao Y et al (2016b) Inactivation of the global regulator LaeA in Monascus ruber results in a species-dependent response in sporulation and secondarymetabolism. Fungal Biol 120:297–305. https://doi.org/10.1016/j.funbio.2015.10.008
Luo C, Hajkova P, Ecker JR (2018) Dynamic DNA methylation: in the right place at the right time. Science 361:1336–1340. https://doi.org/10.1126/science.aat6806
Manfrão-Netto JHC, Mello-De-Sousa TM, Mach-Aigner AR et al (2018) The DNA-methyltransferase inhibitor 5-aza-2-deoxycytidine affects Humicola grisea enzyme activities and the glucose-mediated gene repression. J Basic Microb 58:144–153. https://doi.org/10.1002/jobm.201700415
Michalak EM, Burr ML, Bannister AJ et al (2019) The roles of DNA, RNA and histone methylation in ageing and cancer. Nat Rev Mol Cell Biol 20:573–589. https://doi.org/10.1038/s41580-019-0143-1
Monsen VT, Sundheim O, Aas PA et al (2010) Divergent ß-hairpins determine double-strand versus single-strand substrate recognition of human AlkB-homologueues 2 and 3. Nucleic Acids Res 38:6447–6455. https://doi.org/10.1093/nar/gkq518
Patakova P (2013) Monascus secondary metabolites: production and biological activity. J Ind Microbiol Biot 40:169–181. https://doi.org/10.1007/s10295-012-1216-8
Perry GS, Das M, Woon ECY (2021) Inhibition of AlkB nucleic acid demethylases: promising new epigenetic targets. J Med Chem 64:16974–17003. https://doi.org/10.1021/acs.jmedchem.1c01694
Shao Y, Ding Y, Zhao Y et al (2009) Characteristic analysis of transformants in T-DNA mutation library of Monascus ruber. World J Microbiol Biotechnol 25:989–995. https://doi.org/10.1007/s11274-009-9977-6
Shen L, Song C, He C et al (2014) Mechanism and function of oxidative reversal of DNA and RNA methylation. Annu Rev Biochem 83:585–614. https://doi.org/10.1146/annurev-biochem-060713-035513
Shi Y, Pan T (2011) Beneficial effects of Monascus purpureus NTU 568-fermented products: a review. Appl Microbiol Biot 90:1207–1217. https://doi.org/10.1007/s00253-011-3202-x
Shimizu T, Kinoshita H, Ishihara S et al (2005) Polyketide synthase gene responsible for citrinin biosynthesis in Monascus Purpureus. Appl Environ Microb 71:3453–3457. https://doi.org/10.1016/j.funbio.2015.10.008
Shimizu T, Kinoshita H, Nihira T (2007) Identification and in vivo functional analysis by gene disruption of ctnA, an activator gene involved in citrinin biosynthesis in Monascus Purpureus. Appl Environ Microb 73:5097–5103. https://doi.org/10.1128/AEM.01979-06
Smemo S, Tena JJ, Kim K et al (2014) Obesity-associated variants within FTO form longrange functional connections with IRX3. Nature 507:371–375. https://doi.org/10.1038/nature13138
Tan H, Xing Z, Chen G et al (2018) Evaluating antitumor and antioxidant activities of yellow Monascus pigments from Monascus ruber fermentation. Molecules 23. https://doi.org/10.3390/molecules23123242
Tang C, Klukovich R, Peng H et al (2018) ALKBH5-dependent m6A demethylation demethylation controls splicing and stability of long 3′-UTR mRNAs in male germ cells. P Natl Acad Sci Usa 115:E325–E333. https://doi.org/10.1073/pnas.1717794115
Trewick SC, Henshaw TF, Hausinger RP et al (2002) Oxidative demethylation by Escherichia coli AlkB directly reverts DNA base damage. Nature 419:174–178. https://doi.org/10.1038/nature00908
Wei J, Liu F, Lu Z et al (2018) Differential m6A, m6Am, and m1A demethylation mediated by FTO in the cell nucleus and cytoplasm. Mol Cell 71:973–985. https://doi.org/10.1016/j.molcel.2018.08.011
Westbye MP, Feyzi E, Aas PA et al (2008) Human AlkB homolog 1 is a mitochondrial protein that demethylates 3-methylcytosine in DNA and RNA. J Biol Chem 283:25046–25056. https://doi.org/10.1074/jbc.M803776200
Xie Q, Wu TP, Gimple RC et al (2018) N6-methyladenine DNA modification in glioblastoma. Cell 175:1228–1243. https://doi.org/10.1016/j.cell.2018.10.006
Xie L, Liu L, Cheng L (2020) Selective inhibitors of AlkB family of nucleic acid demethylases. Biochemistry 59:230–239. https://doi.org/10.1021/acs.biochem.9b00774
Xu N, Li L, Chen F (2022) Construction of gene modification system with highly efficient and markerless for Monascus ruber M7. Front Microbiol 13:952323. https://doi.org/10.3389/fmicb.2022.952323
Yang X, Huang L, Ruan X (2014) Epigenetic modifiers alter the secondary metabolite composition of a plant endophytic fungus, Pestalotiopsis Crassiuscula obtained fromthe leaves of Fragaria chiloensis. J Asian Nat Prod Res 16:412–417. https://doi.org/10.1080/10286020.2014.881356
Yang P, Wang Q, Liu A et al (2020) ALKBH5 holds prognostic values and inhibits the metastasis of colon cancer. Pathol Oncol Research: POR 26:1615–1623. https://doi.org/10.1007/s12253-019-00737-7
Zhang Z, Ali Z, Khan SI et al (2016) Cytotoxic monacolins from red yeast rice, a Chinese medicine and food. Food Chem 202:262–268. https://doi.org/10.1016/j.foodchem.2015.12.039
Zhang C, Zhang H, Zhu Q et al (2020a) Overexpression of global regulator LaeA increases secondary metabolite production in Monascus Purpureus. Appl Microbiol Biot 104:3049–3060. https://doi.org/10.1007/s00253-020-10379-4
Zhang M, Yang S, Nelakanti R et al (2020b) Mammalian ALKBH1 serves as an N6-mA demethylase of unpairing DNA. Cell Res 30:197–210. https://doi.org/10.1038/s41422-019-0237-5
Acknowledgements
We gratefully acknowledge Professor Gang Liu (State Key Laboratory of Mycology, Institute of Microbiology, Chinese Academy of Sciences) for providing pAg1-H3 plasmid.
Funding
This study was supported by the National Key Research and Development Program of China (No. 2022YFC2106201).
Author information
Authors and Affiliations
Contributions
YY and TQ conceived the experiments and the project. TQ, LZ and YC performed the experiments and analyzed the data. TQ and YY wrote the manuscript. All authors have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Qiu, T., Zeng, L., Chen, Y. et al. Nucleic acid demethylase MpAlkB1 regulates the growth, development, and secondary metabolite biosynthesis in Monascus purpureus. World J Microbiol Biotechnol 40, 282 (2024). https://doi.org/10.1007/s11274-024-04094-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11274-024-04094-9