Abstract
α-Galactosidase is an important exoglycosidase belonging to the hydrolase class of enzymes, which has therapeutic and industrial potential. It plays a crucial role in hydrolyzing α-1,6 linked terminal galacto-oligosaccharide residues such as melibiose, raffinose, and branched polysaccharides such as galacto-glucomannans and galactomannans. In this study, Actinoplanes utahensis B1 was explored for α-galactosidase production, yield improvement, and activity enhancement by purification. Initially, nine media components were screened using the Plackett–Burman design (PBD). Among these components, sucrose, soya bean flour, and sodium glutamate were identified as the best-supporting nutrients for the highest enzyme secretion by A. Utahensis B1. Later, the Central Composite Design (CCD) was implemented to fine-tune the optimization of these components. Based on sequential statistical optimization methodologies, a significant, 3.64-fold increase in α-galactosidase production, from 16 to 58.37 U/mL was achieved. The enzyme was purified by ultrafiltration-I followed by multimode chromatography and ultrafiltration-II. The purity of the enzyme was confirmed by Sodium Dodecyl Sulphate–Polyacrylamide Agarose Gel Electrophoresis (SDS-PAGE) which revealed a single distinctive band with a molecular weight of approximately 72 kDa. Additionally, it was determined that this process resulted in a 2.03-fold increase in purity. The purified α-galactosidase showed an activity of 2304 U/mL with a specific activity of 288 U/mg. This study demonstrates the isolation of Actinoplanes utahensis B1 and optimization of the process for the α-galactosidase production as well as single-step purification.
Graphical abstract

Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
α-Galactosidase (α-D-galactoside galactohydrolase, EC 3.2.1.22) (α-GAL) is a class of hydrolase that liberates galactose from the galacto-oligosaccharides and synthetic substrates like p-nitrophenyl-α-D-galactopyranoside (Dey and Pridham 1972). α- GAL is also known to be melibiase, which breaks the α-1,6 glycosidic bond between glucose and galactose in melibiose. Moreover, it has potential applications in biotechnological and medical industries. α-GALs are used in the animal feed industry to enhance its nutritional value (Naganagouda and Mulimani 2006; Weignerova et al. 2009) and in the paper industry, to enhance the bleaching effect in softwood pulp (Clarke et al. 2000). α-GALs are used in cane industry, for enhancement of crystallization of beet sugar and to improve the yield of sucrose (Shibuya et al. 1997; Thippeswamy and Mulimani 2002). α-GAL is employed to enhance the gelling characteristics of galactomannans, which are utilized as thickeners in food applications (Chen and Mustapha 2012). Therapeutically α-GAL is used in the treatment of Fabry disease (Kang et al. 2019) and in xenotransplantation (Liu et al. 2007; Zeyland et al. 2014). Further, trials are underway for the use of α-GAL in the conversion of ‘B’ blood group to ‘O’ blood group (Universal donor) by hydrolyzing the terminal galactose residues on the surface of B red blood cells (Balabanova et al. 2010; Liu et al. 2007).
According to the global business index, the global market for galactosidase (GAL) will increase from 1.5 to 2.35 billion USD from 2021 to 2028, with a 6.6% annual increment. (https://dataintelo.com/report/global-galactosidase-market/). This growth is attributed to the rapid expansion of the α-GAL market, which requires the latest innovations in the production and application of the enzyme. The need for novel, robust enzymes with industrial important elements such as stability and high activity under industrial processing conditions is becoming increasingly important. Further, much of the α-GAL area remains largely unexplored and there is considerable potential to develop new α-GAL with unique characteristics for industrial applications. Moreover, microbial α-GAL has gained attention from industries due to ease of cultivation, extracellular secretion, high expression levels, stability over wide range of pH and temperature, broad substrate specificity, and scope for yield improvement by molecular cloning (Aleksieva et al. 2010; Bhatia et al. 2020; Chauhan et al. 2015; Schroder et al. 2017; Stratilova et al. 2018).
The traditional approach for optimization of the medium involves altering a single independent variable while keeping all others at a fixed level which is known to be easy and simple. However, it is highly time-consuming and expensive, especially for conducting numerous experiments to determine the optimal levels. In addition, it fails to consider the interactive effects between variables (Panda et al. 2007). Statistical optimization is the preferred method as it allows for the evaluation of interactions among potential influencing factors with a limited number of experiments (Cazetta et al. 2007; Francis et al. 2003; Kennedy and Krouse 1999). This approach utilizes a specialized experimental design that minimizes errors when determining parameter effects, all while achieving cost-effectiveness. Plackett–Burman design (PBD) and response surface methodology (RSM) are two commonly employed statistical techniques for optimizing biological processes. Initially, PBD is used for screening purposes, and after finding the significant variables in this initial screening, they can be further improved using a central composite design (CCD) in RSM (Gajdhane et al. 2016; Sathish et al. 2018).
The current study deals with the isolation of high amounts of α-GAL producing Actinoplanes utahensis B1, followed by enhancement of yield by sequential optimization methods and purification of the produced enzyme. To date, there have been no reports on the use of statistical methods to optimize α-GAL production in submerged fermentation using A. utahensisB1.
Materials and methods
Chemicals and media components
Para-nitrophenyl α-D-galactopyranoside (pNPG), Para-nitrophenol and Sodium Glutamate were obtained from Sigma-Aldrich (India). Sucrose, Maltose, Glucose and galactose were purchased from Merck. Soybean meal and other chemicals were procured from commercial sources.
Bacterial isolation and growth conditions
A water sample from Shamirpet Lake, Hyderabad, Telangana, India was collected in a sterile container for the isolation of bacterial strain-producing α-GAL enzyme. From the collected water, the bacteria were isolated by serial dilution followed by spread plate technique on starch casein agar (SCA) plates (Kuster and Williams 1964) (Starch-10 g/L, Casein-0.3 g/L, K2HPO4-2 g/L, FeSO4·7H2O-0.01 g/L, MgSO4·7H2O-0.5 g/L, ZnSO4·7H2O-0.001 g/L, agar-20 g/L, pH-7.0) supplemented with raffinose 2 g/L as an inducer. The cycloheximide (40 µg/ml) was added to SCA plates to prevent the growth of fungal contaminants. The SCA plates that have been inoculated were subjected to incubation at a temperature of 28 °C for 72 h. The colonies showing maximum zone of clearance were selected and used for further experiments.
Identification of isolates by phenotypic and biochemical characterization
The colony morphology of the isolated strain, such as the shape of sporangia, mycelium color, gram staining test, margin, and elevation was determined as per Bergey’s manual of determinative bacteriology. Further, various physiochemical parameters such as temperature, sodium chloride (NaCl) and pH were also studied (Buchanan and Gibbons 1974). Biochemical identification tests such as nitrate reduction test, starch degradation test, citrate utilization test, hydrogen sulfide production, melanin formation, urease production, casein and gelatin degradation, coagulation and peptonisation of milk were conducted (Al-Dhabaan 2019; Holt 1994). Additionally, carbohydrate fermentation tests were also performed using seven carbohydrate sources for the isolated strain. Furthermore, enzymatic activities for the urease, α-GAL, β-mannosidase, β-xylanase, α-L-rhamnosidase, and lipase were studied for this strain (Table 1).
Identification of molecular methods (16 s rRNA)
The extraction of genomic DNA was carried out through the utilization of a spin column kit (HiMedia). The amplification of bacterial 16S rRNA gene (1500 bp) (Clarridge 2004) was accomplished via polymerase chain reaction and subsequently purified using Exonuclease I-Shrimp Alkaline Phosphatase (Exo-SAP) (Darby et al. 2005). The purified amplicons were subjected to sequencing through the Sanger method utilizing an ABI 3500xL genetic analyzer (Life Technologies, USA). From the obtained 16S rRNA gene sequence, the isolate was identified by the Basic Local Alignment Search Tool (BLAST) (Altschul et al. 1990).
A phylogenetic tree was reconstructed utilizing 16S rRNA gene sequences to illustrate the correlation between the isolate and related species. Multiple sequence alignment and the phylogenetic tree were constructed by using the software MEGA11.
Inoculum preparation and cultivation of microorganisms
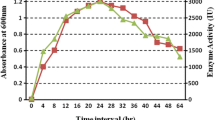
The isolated culture was cultivated in a 500 mL conical flask containing inoculum media which consists of glucose-4.0 g/L; malt extract-10.0 g/L; yeast extract-4.0 g/L and CaCO3-2.0 g/L and incubated in an orbital shaker at 28 °C, 200 rpm for 60 h. ISP 9 medium having the composition of sucrose- 1.0 g/L; ammonium Sulphate- 2.64 g/L; KH2PO4- 2.38 g/L; K2HPO4.3H2O- 5.64 g/L; MgSO4.7H2O- 1.0 g/L and trace elements- 1 mL was used as a production medium (Shirling 1966). The trace elements considered in the production media are CuSO4.5H2O-6.4 g/L; FeSO4.7H2O- 1.1 g/L; MnCl2.4H2O- 7.9 g/L; ZnSO4.7H2O- 1.5 g/L and pH of the media was adjusted to 7.0. The sterile flasks containing media are inoculated with 5% of inoculum and incubated in an orbital shaker at 28 °C, 200 rpm for 240 h. After every 24 h, samples were withdrawn and estimated for enzyme activity.
Screening of media components using Plackett–Burman design
Various carbon and nitrogen sources such as sucrose, maltose, galactose, glucose, fructose, soybean flour, yeast extract, corn steep liquor and sodium glutamate were selected for identifying the critical media components that affect the α-GAL production by isolated A. utahensis B1 strain. PBD design with twelve experiments was employed to screen the nutrients. All nine nutrients were tested at two levels. Table 2 depicts the employed PBD along with the nutrients invarious combinations.
The obtained data was analysed by a first-order regression model and the regression equation is
where Y represents the activity of α-GAL; β0 denotes the intercept; βi signifies the linear coefficient and X is the selected component. Triplet experiments were carried out and the average activity value was considered as the response variable (Y). Variables possessing confidence levels of greater than 95% were treated as significant components for α-GAL production by isolated bacteria.
Optimization of critical media components by CCD
The PBD-screened nutrients were optimized using the CCD by employing a 3-factor 5-level model. Table 3 depicts the actual and coded values of variables and the matrix of CCD with their responses of each run. The response was fitted to the second-order polynomial model to understand the relationship between the selected parameters and obtained enzyme activity. The overall polynomial equation is
where Yi represents the predicted α-GAL activity, xi and xj denote selected variables that influence the enzyme production by A. utahensis B1 strain; β0 denotes the offset term; βi represents the ith linear coefficient andβij denotes the ith quadratic coefficient. The term ‘e’ refers to the error.
Analysis of variance (ANOVA) was performed to evaluate the statistical significance of the model and model terms. The correlation coefficient (R2) was utilized to comprehend the proportion of variability of the optimization parameter that could be elucidated by the model. 3D surface plots and 2D contour plots were generated to depict the main and interactive effects of selected parameters on the production of enzyme by isolated bacteria. A statistical software Statistica version 7.0 (TIBCO Software Inc.) was used for this study.
Purification of α-GAL by chromatographic methods
The harvest culture was centrifuged at 7000 revolutions per minute (rpm) for 30 min at 4–8 °C. After centrifugation, the supernatant was collected and concentrated using a tangential flow filtration (TFF) system. A Sartocon Slice PES 50 kDa Cassette with a filter area of 0.5 m2 was used for ultrafiltration followed by diafiltration with 10 mM potassium phosphate buffer, 0.25 mM EDTA, and 3 mM NaCl for 5 diafiltration volumes (DVs). The column was pre-equilibrated with 10 mM potassium phosphate buffer, 0.25 mM EDTA, and 250 mM NaCl buffer before loading the retentate sample. The ultrafiltration and diafiltration (UFDF) retentate sample was loaded onto the Ceramic Hydroxyapatite (CHT) type II column with a flow rate of 10 mL/min. The column was washed with 8 column volumes (CVs) of wash buffer (10 mM potassium phosphate buffer, 0.25 mM EDTA, and 3 mM NaCl) to remove the unbounded protein. The enzyme bound to the column was eluted by using 8 CVs of elution buffer (50 mM potassium phosphate buffer, 0.25 mM EDTA, and 250 mM NaCl) in a step gradient manner. The elution fractions were collected and checked for α-GAL activity. Fractions having maximum activity were pooled and concentrated by ultrafiltration.
A VivaFlow 200 cassette having a pore size of 50 kDa MWCO with a membrane area of 200 cm2 was used for the UFDF experiment with a flow rate of 25 ml/min and 75 mL of fraction elute was concentrated to 25 mL to estimate the enzyme activity. In addition, the retentate samples were loaded on SDS-PAGE gel to confirm the molecular weight of the protein (Laemmli 1970).
Estimation of α-GAL activity
α-GALs activity was determined by the modified Dey and Pridham method (1969). The hydrolyzing activity of p-NPG was measured as α-GALs activity. A 50 µl of 5 mM pNPG substrate was incubated with 100 µl of enzyme sample and 850 µl of 0.1 M citrate phosphate buffer of pH 7.0 at 50 °C for 10 min. After incubation, the reaction was arrested by the addition of 2 mL of 0.5 M sodium bicarbonate. The quantity of p-nitrophenol which was released was measured using a spectrophotometer at an absorption wavelength of 405 nm. The enzyme activity was calculated by using Eq. (3) where a single unit of α-GAL activity is defined as the quantity of enzyme necessary to release 1 µMole of p-nitrophenol per minute under the assay conditions.
where AbsTest—Absorbance of the test sample at 400 nm, AbsBlank—Absorbance of blank sample at 400 nm, 18.5—Millimolar coefficient of p-nitrophenol at 405 nm, 10—Incubation minutes, 0.1—Volume of enzyme used
Specific activity
The specific activity (U/mg protein) was determined by measuring the total protein concentration of the supernatant using the Folin Lowry method, with Bovine Serum Albumin (BSA) as a reference (Waterborg et al. 2009).
SDS-PAGE(sodium dodecyl sulphate-polyacrylamide gel electrophoresis)
The Laemmli method (1970) was used to determine the molecular weight and purity of the sample through SDS-PAGE, utilizing a 3% stacking gel and 10% resolving gel. A sample buffer was prepared with 2% SDS, 10% glycerol, 0.001% bromophenol blue (dye), and 2-mercaptoethanol as a reducing agent. The protein sample and sample buffer were mixed in a 1:2 ratio and incubated at 100 °C for 90 s. The 10 μL of protein sample buffer was loaded onto the wells with the protein ladder to determine the molecular weight. Electrophoresis was conducted at 120 V power until the bromophenol marker reached the bottom of the gel. The gel was then stained with 0.1% Coomassie Brilliant Blue R-250 for an hour to stain the proteins, followed by destaining for 2 h in a solution of acetic acid (10%) and methanol (40%) to visualize the proteins.
Results
Bacterial strain isolation
The SCA plates supplemented with raffinose were examined after 72 h for the zone of clearance. The size of the zone of clearance corresponds to the degradation of raffinose in SCA medium by the extracellular α-GAL enzyme produced by the isolates. The strain with the maximum zone of clearance was selected as a potential strain for α-GAL production and it was designated as B1 strain. Mycelial suspensions of isolated B1 strain were preserved in glycerol solution (20%) and stored at − 80 °C.
Phenotypic and biochemical characterization
The colony morphology of the isolated B1 strain was found to be aerobic, motile and gram-positive filamentous bacteria. The mycelium colour of the B1 strain is orange and the sporangia are irregular in shape and digitate. The favourable temperature for the growth of the strain was observed to be between 20 to 35 °C and there was no growth above 40 °C which shows that the bacteria are not thermophilic. The B1 strain was found to grow at a neutral pH of 6 to 8 and it can withstand the salt concentration of up to 2% and showed no growth above 2%.
The biochemical tests performed for the B1 strain showed positive results for the hydrogen sulfide (H2S) gas production, nitrate reduction, melanin formation, urease production, and degradation of starch, casein and gelatin. The B1 strain also can produce enzymes such as urease, beta-mannosidase, bet-xylanase, α-Galactosidase, alpha-L-rhamnosidase and lipase. The isolated B1 strain has shown positive results for acid production by utilizing carbon sources such as fructose, galactose, glucose, maltose, sucrose, and xylose. Conversely, it has exhibited negative results concerning lactose (Table 1). Based on phenotypic and biochemical characterization it has been determined that the isolated B1 strain belongs to Actinoplanes sp.
Identification by Molecular methods (16 s rRNA)
The complete 16 s rRNA gene sequence was acquired for the isolated B1 strain. The strain showed the highest similarity with the taxon name Actinoplanes utahensis with a sequence ID of NR112105 showing a similarity of 99.86%. Figure 1 shows the phylogenetic analysis of the isolate with other Actinoplanes sps. The isolate closely relates with the Actinoplanes utahensis strains. So based on the phenotypic, biochemical and molecular methods the isolated strain was identified as Actinoplanes utahensis and it was designated as Actinoplanes utahensis B1. The 16 s rRNA sequence of the isolate has been deposited in GenBank and has been assigned the accession number OQ266884.1.
Optimization of media components
Based on preliminary studies (data not shown) on 10th day the highest titers of α-GAL production were noticed. Therefore, the fermentation time was set as 240 h. Initially, 16 U/mL of α-GAL was produced by A. utahensisB1 strain in the ISP 9 basal medium (data not shown) and found that the yields were less. To increase enzyme production, media components should be replaced or optimized for their concentrations. For this purpose, a sequential optimization process was employed to enhance the α-GAL production. Initially, PBD was used to screen the nutrients and their levels followed by CCD was employed to optimize the concentration of selected nutrients.
PBD for screening of media components
Different carbon and nitrogen sources such as sucrose, galactose, glucose, galactose, fructose, soya bean flour, corn steep liquor, yeast extract and sodium glutamate which influences α-GAL production were selected for testing. Table 2 shows PBD along with the observed enzyme produced by the A. utahensis B1.
From Table 2 it was noticed that the α-GAL production varied from 22 to 45 U/ml which indicates that the selected nutrients and their concentrations have a significant effect on the production of enzyme from A. utahensis B1. Further, the data was analyzed by regression method and each selected nutrient effect on the enzyme production was also calculated.
The regression coefficients were computed by taking the α-GAL as a response. The derived regression coefficients, effects and their corresponding p-values are depicted in Table 4. The correlation coefficient (R2) value of 0.99 designates the goodness of fit and a higher adjusted R2 value of 0.98 indicates that obtained data is flawless. The close agreement between R2 and adjusted R2 values signify to construct an equation to predict enzyme production. Equation 4, a first-order polynomial equation was constructed utilizing the computed coefficients.
The variable coefficients that have higher p-values (p > 0.05) were considered insignificant terms and the remaining were considered as significant terms. Based on coefficients p-values sucrose, soybean flour and sodium glutamate are found to be significant variables remaining all are insignificant variables. Among the three significant variables, sucrose has the highest effect (8.50) followed by soybean flour (7.66) and sodium glutamate (5.00) indicating that sucrose was an important carbon source for the production of α-GAL from the A. utahensis B1. From this data, it was observed that A. utahensis B1 strain needs sucrose a hetero disaccharide to produce a higher titer of α-GAL. Monosaccharides such as glucose, fructose, galactose, and homo disaccharide maltose were found to be insignificant. The probable reason may be mono and homo disaccharides enhance the growth of organisms rather than the production of desired enzymes. A similar type of observation was reported by Boje et al. (2011) forα-GAL production from Enterobacter dissolvens. It was also observed that with the supplement of glucose in media, the enzyme production was decreased and biomass production was increased.
Further, the levels of significant variables were determined based on the sign of the coefficient. All three significant variables have a positive sign coefficient, which indicates that for higher production of α-GAL by A. utahensis B1 needs higher amounts of sucrose, soybean flour and sodium glutamate.
Optimization of medium components by CCD method
From PBD it was observed that sucrose, soybean flour and sodium glutamate were the prominent nutrients for α-GAL production from the A. utahensis B1. Further, these components were optimized by using the CCD method, Table 3 depicts the 20 experimental designs along with the obtained α-GAL yields. From Table 3, it was observed that the α-GAL production varied from 32.5 U/ml to 56.5 U/ml, indicating the selected components and their concentrations play vital roles in the α-GAL production from the isolated A. utahensis B1 strain. The data was analyzed using the second-order polynomial regression.
The regression fit was measured by calculating the correlation coefficient (R2). In this study, the higher R2 value of 0.97, signifies the accuracy of experimental data and suitability for analysis. The adjusted R2 value of 0.95 is closer to the predicted R2 value indicating the acceptance of the constructed model (Chiranjeevi et al. 2014; Sathish et al. 2018). The lesser CV (coefficient of variation) value of 4.41% indicates that the experiments were conducted with greater precision and reliability (Mohan et al. 2014; Usman et al. 2019). To understand the relationship between the α-GAL production by A. utahensis B1 and selected nutrients, an empirical equation was constructed based on the obtained regression coefficients. Equation 5 represents the constructed polynomial regression model. This equation helps to forecast the α-GAL yields at the desired concentration of selected nutrients.
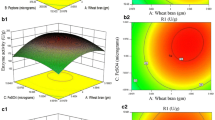
Table 5 depicts the Analysis of Variance (ANOVA) data which shows the selected three nutrients t, F and p- values. The terms that have less p-value and higher F value were considered statistically significant terms. The linear terms of soybean flour concentration and sucrose with other nutrients are not significant. The linear term of soybean flour concentration is not significant at linear term however the square term is the second highest significant, which indicates that this nutrient is highly important for α-GAL production by isolated bacteria. The interactions of nutrients were depicted as surface and contour plots. The interaction of sucrose with soybean flour and sodium glutamate is depicted in Fig. 2a–d. Both contours are circular which indicates that sucrose concentration is independent of all other nutrients. A similar trend was also noticed in soybean flour with sodium glutamate (Fig. 2e–f).
Surface plots of selected factors and their interaction (left side) A Sucrose vs Soybean flour C Sucrose vs Sodium Glutamate E Soybean flour vs Sodium Glutamate. Contour plots of selected factors and their interaction (right side) B Sucrose vs Soybean flour D Sucrose vs Sodium Glutamate F Soybean flour vs Sodium Glutamate
Based on Eq. 5, the optimum concentrations were forecasted as sucrose was 45.35 g/L, soybean flour was 30.84 g/L and sodium glutamate was 3.56 g/L, and at these concentrations the α-GAL yield predicted as 57.08 U/ml. By conducting the experiments at these conditions, 58.12 U/ml of α-GAL was obtained which indicates that the built empirical model is useful to predict the α-GAL production by A. utahensis B1.
Purification of α-GAL enzyme
The protocol for the purification of the enzyme is presented in Table 6. The α-GAL enzyme was purified to a 2.03-fold increase in activity with a yield of 32% from the initial crude enzyme extract. Initially, proteins present in the crude extract were subjected to concentration using an MWCO membrane, whereby the desired protein was retained at a 50 kDa MWCO membrane. This particular step resulted in a 0.94-fold enzyme purification, with 74% of the enzyme being recoverable. Subsequently, the retentate samples were fractionated through mixed-mode chromatography utilizing a CHT type-II column, wherein the enzyme was eluted with a step gradient of elution buffer (50 mM potassium phosphate buffer, 0.25 mM EDTA, and 250 mM NaCl). The elution profile of the purified protein is depicted in Fig. 3. The SDS-PAGE technique was executed on the eluted fractions, and the fractions that exhibited positive outcomes on SDS-PAGE (as depicted in Fig. 4) were pooled (specifically, fractions 2, 3, and 4) and concentrated to a volume of 25 mL via utilization of a 50 kDa MWCO membrane. This ultimate stage of the process yielded a 2.03-fold increase in enzyme purification, with 32% of the enzyme being retrievable.
Discussion
Several α-GAL producing microbes have been isolated, reported, and characterized over the last several decades (Bhatia et al. 2020). However, this study represents the first report of α-GAL production from A. utahensis B1 strain in submerged fermentation. Different nitrogen and carbon sources were amended in the production medium to determine their impact on α-GAL production. A PBD design was used to establish a relationship between media components and their responses to α-GAL activity. CCD was utilized to evaluate the optimum concentrations of critical media components and their interactions. Sucrose (4.53%), soybean flour (3.08%), and sodium glutamate (0.35%) were found to be the most significant variables and enhanced the yield from 16 to 58.37 U/mL. Similar studies were performed by Gajdhane et al. (2016), where they observed the highest enzyme yield from Fusarium moniliforme NCIM 1099 through the utilization of sequential statistical optimization techniques such as PBD and CCD. Similarly, Anisha et al. (2008) conducted a study on α-GAL production in Streptomyces griseoloalbus and were able to achieve a maximum yield of 50 U/mL through the implementation of PBD and BBD methods. Alvarez-Cao et al. (2019) reported the use of CCD to optimize the critical components and achieved a yield of 66 U/mL (an increase 11-fold) at 190 h of cultivation. The sequential statistical optimization of α-GAL production by A.utahensis B1 yielded a 3.64-fold increase in comparison to the unoptimized medium.
The optimum α-GAL activity and the specific activity of the purified enzyme were found to be 2304 U/mL and 288 U/mg with a purification fold of 2.03. Pure and efficient enzyme activity of α-GAL could be obtained using a single-column chromatography purification step. Previously, the purification of α-GAL was accomplished through a series of multistep purification procedures which were both laborious and time-intensive (Garro et al. 1996; Saishin et al. 2010; Sirisha et al. 2015). Nevertheless, the purification methodology utilized in this context is comparatively uncomplicated and remarkably replicable. Patil et al. (2021) achieved the purification of α-GAL from Bacillus megaterium VHM1 to homogeneity through a three-step process involving ethanol precipitation, anionic exchange chromatography, and gel filtration using G75. The yield obtained was 42.2%, with a purification fold of 94.2%. Similarly, Gote et al. (2006) employed a three-step methodology comprising ultrafiltration, alcohol precipitation and hydrophobic interaction chromatography to purify α-GAL from Bacillus stearothermophilus. The resultant yield was 44.6% and the purification fold was 369%.
The current purification study showed a discernible band that has a molecular weight of approximately 72.0 kDa, with no discernible evidence of contamination. It was observed that the molecular weight of α-GAL exhibited variation based on the origin of the enzyme. α-GALs purified from various sources such as Bacillus megaterium VHM1 (Patil et al. 2021), Bacillus stearothermophilus (Gote et al. 2006), Aspergillus fumigatus (Rezende et al. 2005), Sulfolobus solfataricusis (Brouns et al. 2006) exhibited molecular weights of 66.0, 79.9, 54.7 and 74.7 kDa respectively.
The present study showed that the isolated A. utahensis B1 strain is a potential source for α-GAL production and provided a significant scope for further studies regarding large-scale production for industrial applications.
Data availability
All data analysed in this study is included in this article.
References
Al-Dhabaan FA (2019) Morphological, biochemical and molecular identification of petroleum hydrocarbons biodegradation bacteria isolated from oil polluted soil in Dhahran. Saud Arabia Saudi J Biological Sciences 26:1247–1252
Aleksieva P, Tchorbanov B, Nacheva L (2010) High-yield production of α-galactosidase excreted from Penicillium chrysogenum and Aspergillus niger. Biotechnol Biotechnol Eq 24:1620–1623
Altschul SF, Gish W, Miller W et al (1990) Basic local alignment search tool. J Mol Biol 215(3):403–410
Alvarez-Cao ME, Cerdan ME, Gonzalez-Siso MI, Becerra M (2019) Optimization of Saccharomyces cerevisiae α-galactosidase production and application in the degradation of raffinose family oligosaccharides. Microb Cell Fact 18:172
Anisha GS, Sukumaran RK, Prema P et al (2008) Statistical Optimization of α-Galactosidase Production in Submerged Fermentation by Streptomyces griseoloalbus Using Response Surface Methodology. Food Technol Biotechnol 46(2):171–177
Balabanova LA, Bakunina IY, Nedashkovskaya OI et al (2010) Molecular characterization and therapeutic potential of a marine bacterium Pseudoalteromonas sp. KMM 701 α-galactosidase. Mar Biotechnol (NY) 12:111–120
Bhatia S, Singh A, Batra N, Singh J (2020) Microbial production and biotechnological applications of α-galactosidase. Int J Biol Macromol 150:1294–1313
Boje BG, Narasu ML, Chakravarthy BK, Savala NK (2011) Optimization of production conditions for intracellular α-galactosidase from Enterobacter dissolvens. Int J Biol Sci II:155–161
Brouns SJ, Smits N, Wu H, Snijders APL, Wright PC, de Vos WM (2006) Identification of a novel α-galactosidase from the hyperthermophilic archaeon Sulfolobus solfataricus. J Bacteriol 88:2392–2399
Buchanan RE, Gibbons NE (1974) Bergey’s manual of determinative bacteriology, 8th edn. Williams & Wilkins Co., Baltimore
Cazetta ML, Celligoi MAPC, Buzato JB, Scarmino IS (2007) Fermentation of molasses by Zymomonas mobilis: Effects of temperature and sugar concentration on ethanol production. Biores Technol 98:2824–2828
Chauhan AS, Kumar A, Siddiqi NJ, Sharma B (2015) Extracellular α-galactosidase from Trichoderma sp.(WF-3): optimization of enzyme production and biochemical characterization. Biotechnol Res Int 2015:860343
Chen M, Mustapha A (2012) Survival of freeze-dried microcapsules of α-galactosidase producing probiotics in a soybar matrix. Food Microbiol 30:68–73
Chiranjeevi PV, Pandian MR, Sathish T et al (2014) Enhancement of laccase production from Plerotus ostreatus PVcRSP-7 by altering the nutritional conditions using response surface methodology. Bioresources 9(3):4212–4225
Clarke JH, Davidson K, Rixon JE, Halstead JR, Fransen MP, Gilbert HJ, Hazlewood GP (2000) A comparison of enzyme-aided bleaching of softwood paper pulp using combinations of xylanase, mannanase and a-galactosidase. Appl Microbiol Biotechnol 53:661–667
Clarridge JE (2004) Impact of 16S rRNA gene sequence analysis for identification of bacteria on clinical microbiology and infectious diseases. Clin Microbiol Rev 4:840–862
Darby AC, Chandler SM, Welburn SC, Douglas AE (2005) Aphid-symbiotic bacteria cultured in insect cell lines. Appl Environ Microbiol 71(8):4833–4839
Dey PM, Pridham JB (1969) Purification and properties of a-galactosidases from Vicia faba seeds. Biochem J 113:49–55
Dey PM, Pridham JB (1972) Biochemistry of a-galactosidase. Adv Enzymol 36:911–930
De Rezende ST, Guimaraes VM, Rodrigues MC, Felix CR (2005) Purification and Characterization of an α-Galactosidase from Aspergillus fumigate. Braz Arch Biol Technol 48:195–202
Francis F, Sabu A, Nampoothiri KM, Ramachandran S, Ghosh S, Szakacs G, Pandey A (2003) Use of response surface methodology for optimizing process parameters for the production of a-amylase by Aspergillus oryzae. Biochem Eng J 15:107–115
Gajdhane SB, Bhagwat PK, Dandge PB (2016) Response surface methodology- based optimization of production media and purification of α-galactosidase in solid-state fermentation by Fusarium moniliforme NCIM 1099. 3 Biotech 6:1–14
Garro MS, de Valdez GF, Oliver G, de Gori GS (1996) Purification of α-galactosidase from Lactobacillus fermentum. J Biotechnol 45:103–109
Gote MM, Khan MI, Gokhale DV, Bastawde KB, Khire JM (2006) Purification, characterization and substrate specificity of thermostable α-galactosidase from Bacillus stearothermophilus (NCIM-5146). Process Biochem 41:1311–1317
Holt JG (1994) Bergey’s manual of determinative bacteriology, 9th edn. Lippincott Williams and Wilkins, Baltimore
Kang JJ, Desch KC, Kelly RJ, Shu L, Bodary PF, Shayman JA (2019) α-Galactosidase-A deficiency promotes von Willebrand factor secretion in models of Fabry disease. Kidney Int 95:149–159
Kennedy M, Krouse D (1999) Strategies for improving fermentation medium performance: a review. J Ind Microbiol Biotechnol 23:456–475
Kuster E, Williams ST (1964) Selection of media for isolation of streptomycetes. Nature 202:928–929
Laemmli UK (1970) Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227(5259):680–685
Liu QP, Sulzenbacher G, Yuan H et al (2007) Bacterial glycosidases for the production of universal red blood cells. Nat Biotechnol 25:454–464
Mohan SK, Viruthagiri T, Arunkumar C (2014) Statistical optimization of process parameters for the production of tannase by aspergillus flavus under submerged fermentation. 3 Biotech 4:159–166
Naganagouda K, Mulimani VH (2006) Gelatin blends with alginate: gel fibers for α-galactosidase immobilization and its application in reduction of non-digestible oligosaccharides in soymilk. Process Biochem 41:1903–1907
Panda BP, Ali M, Javed S (2007) Fermentation process optimization. Res J Microbiol 2:201–208
Patil AG, Kote NV, Manjula AC, Vishwanatha T (2021) Purification, characterization of α-galactosidase from a novel Bacillus megaterium VHM1, and its applications in the food industry. J Appl Biol Biotechnol 9:13–19
Saishin N, Ueta M, Wada A, Yamamoto I (2010) Purification and characterization of α-galactosidase I from Bifidobacterium longum subsp. longum JCM 7052. J Biol Micromol 10:13–22
Sathish T, Kezia D, Bramhachari PV, Prakasham RS (2018) Multi-objective based superimposed optimization method for enhancement of l-glutaminase production by Bacillus subtilis RSP-GLU. Karbala Int J of Modern Sci 4(1):50–60
Schroder C, Janzer VA, Schirrmacher G, Claren J, Antranikian G (2017) Characterization of two novel heat-active α-galactosidases from thermophilic bacteria. Extremophiles 21:85–94
Shirling G (1966) Methods of characterization of Streptomyces species. Int J Syst Bacteriol 16:313–340
Shibuya H, Kobayashi H, Sato T et al (1997) Purification, characterization and cDNA cloning of a novel α-galactosidase from Mortierella vinacea. Biosci Biotechnol Biochem 61:592–598
Sirisha E, Potumarthi R, Naveen A, Mangamoori LN (2015) Purification and characterisation of intracellular alpha-galactosidases from Acinetobacter sp. 3 Biotech 5:925–932
Stratilova B, Klaudiny J, Rehulka P et al (2018) Characterization of a long-chain α-galactosidase from Papiliotrema flavescens. World J Microbiol Biotechnol 34:19
Thippeswamy S, Mulimani VH (2002) Enzymatic degradation of raffinose family oligosaccharides in soymilk by immobilized α-galactosidase from Gibberella fujikuroi. Process Biochem 38:635–640
Usman AI, Aziz AA, Sodipo BK (2019) Application of central composite design for optimization of biosynthesized gold nanoparticles via sonochemical method. SN Applied Sciences 1:403
Waterborg JH, Walker, (2009) The Lowry method for protein quantitation. In: Walker JM (ed) The Protein Protocols Handbook. Humana Press, Totowa, NJ, pp 7–10
Weignerova L, Simerska P, Kren V (2009) α-Galactosidases and their applications in biotransformations. Biocatal Biotransform 27:79–89
Zeyland J, Wozniak A, Gawronska B et al (2014) Double Transgenic pigs with Combined expression of human α1,2-Fucosyltransferase and α-galactosidase to avoid hyperacute xenograft rejection. Arch Immunol Ther Exp 62:411–422
Acknowledgements
The authors express their gratitude to Dr. Harish Kumar Reddy, Manufacturing Science Department at Biological E Ltd for generously providing the laboratory to conduct the study. The authors would like to thank Dr. Sathish Thadikamala, Aurovaccines, Hyderabad, TS, India for his support in conducting statistical studiesas well as support during the drafting of this manuscript.
Funding
Not applicable.
Author information
Authors and Affiliations
Contributions
BK and JBD: conceptualized, designed and wrote the manuscript. VTC: revised the manuscript and also major contributor in writing the manuscript. All authors have read and approved the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
All authors declare no competing interests.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Balumahendra, K., Venkateswarulu, T.C. & Babu, D.J. Enhancement of α-galactosidase production using novel Actinoplanes utahensis B1 strain: sequential optimization and purification of enzyme. World J Microbiol Biotechnol 40, 91 (2024). https://doi.org/10.1007/s11274-023-03880-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11274-023-03880-1