Abstract
The introduced species Acacia saligna is a very promiscuous host as it can be efficiently nodulated with a wide range diversity of rhizobia taxa, including both fast and slow-growing strains. Fourteen nitrogen (N)-fixing bacteria were isolated from root nodules of wild Acacia saligna growing in distinct geographic locations in Morocco and were examined for their symbiotic efficiency and phenotypic properties. Multivariate tools, such as principal component analysis (PCA) and hierarchical clustering analysis (HCA), were used to study the correlation between phenotypic and symbiotic variables and discriminate and describe the similarities between different isolated bacteria with respect to all the phenotypic and symbiotic variables. Phenotypic characterization showed a variable response to extreme temperature, salinity and soil pH. At the plant level, the nodulation, nitrogen fixation, and the shoot and root dry weights were considered. The obtained results show that some of the tested isolates exhibit remarkable tolerances to the studied abiotic stresses while showing significant N2 fixation, indicating their usefulness as effective candidates for the inoculation of acacia trees. The PCA also allowed showing the isolates groups that present a similarity with evaluated phenotypic and symbiotic parameters. The genotypic identification of N2-fixing bacteria, carried out by the 16S rDNA approach, showed a variable genetic diversity among the 14 identified isolates, and their belonging to three different genera, namely Agrobacterium, Phyllobacterium and Rhizobium.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
The massive use of synthetic fertilizers in agriculture is nowadays subject to debates related to both environmental and public health concerns. The addition of such chemicals has induced land desertification and salinization, which reduced productivity and availability of biological resources. Desertification has become an alarming menace in many parts of the world because it leads to an unrecoverable deterioration of the vegetation cover with a consequent decrease in soil fertility (Arora 2018; Lebrazi and Fikri-Benbrahim 2022). In addition to affecting native plant communities, soil degradation also disturbs the symbioses in the plant microsphere, which play a crucial role in fostering plant growth in degraded ecosystems (Requena et al. 2001). Microorganisms associated with crops play an important role in the productivity and sustainability of several natural ecosystems and agro-ecosystems. Indeed, rhizobia are soil bacteria group from different genus that are mainly associated with plants of the legume family. These rhizobacteria can induce the development of root nodules and provide their host plants with a significant N amount through their ability to fix atmospheric dinitrogen (Suman et al. 2022).
The primordial role of trees in soil protection, litter production, and more globally in the improvement of edaphoclimatic conditions has been widely demonstrated (Ribeiro-Barros et al. 2018). Therefore, the performance of rhizobia for symbiotic nitrogen fixation plays a key role in the selection of field applications. Bacterial strain selection technology aims to improve more efficient strains to replace native soil strains. This provides an effective biotechnological tool for restoring degraded or desert ecosystems. Nodulation and symbiotic nitrogen fixation are highly correlated and related to both plant and symbiotic bacterial genotype. The introduction of tree legumes into cropping systems often involves the selection of competitive and persistent symbiotic bacteria as inoculants to improve nitrogen fixation and crop yield (Lebrazi and Fikri-Benbrahim 2022). In Morocco, the socio-economic role of Acacia is very important, and many Acacia species are widely and successfully applied in reforestation programs, for soil fertilization and dune fixation (Taoufiq et al. 2018). The ability of Acacia trees to perform symbiosis with rhizobia that fix atmospheric dinitrogen allows them to improve and maintain soil fertility levels (Bakhoum et al. 2018). By establishing bacterial and mycorrhizal symbioses, woody species have the necessary and sufficient adaptations to grow on very poor and mineral-depleted soils.
Therefore, it is possible to improve the composition of the rhizobial and mycorrhizal communities associated with the trees in order to increase the biomass production by these plants and also contribute to the mitigation of erosion.
The incorporation of Acacia species into reforestation programs based on their high resistance to various biotic and abiotic stress factors, can offer a sustainable reforestation solution in arid and semi-arid areas (Fikri Benbrahim et al. 2014). In Morocco, the Acacia woodlands cover an area of 1,128,000 ha, including Acacia mearnsii in the Mâamora region (Atlantic plain), A. gummifera and Acacia spp. in the Atlantic and Eastern Meseta (Middle Atlas), A. gummifera, A. raddiana, A. seyal and A. albida in the presahara (Souss region) as well as A. raddiana and A. seyal in the Sahara (Fikri Benbrahim et al. 2014). The increased attention to the utilization of Rhizobium as biofertilizers in agricultural systems has allowed the identification of a high number of rhizobacterial strains. To effectively exploit symbiotic N2 fixation to improve plant production, it is necessary not only to select the best host cultivar, but also to properly and sufficiently characterize the native rhizobia population. On a practical level, the selection of a suitable host plant and complementary microsymbionts depends on the edaphic environment, which is subject to several irregularities depending on the intensity and nature of crops, the geographical environment and the soil conditions (Geetha and Joshi 2013). Thus, salinity, pH and temperature perform the most important environmental constraints in plant-rhizobia symbiosis. Therefore, it is necessary to evaluate the environmental factors in order to ensure that the selected symbiotic couple has the necessary aptitudes to establish itself in a given type of soil and meet the defined requirements. Studying the diversity of rhizobia strains nodulating acacia trees and selecting efficient bacteria helps to identify the most appropriate combinations of plants and microsymbionts for use in reforestation programs (Ba et al. 2002). Therefore, the exploitation of acacias rhizosphere to promote the restoration of vegetation cover in arid and semi-arid areas seem to be interesting due to their participation in the improvement of soil stabilization and fertility through N2 transfer to the associated crops (Bakhoum et al. 2018).
Thus, the aim of this work was to study the phenotypic and genotypic parameters of rhizobia isolated from root nodules of acacia trees from different sites in Morocco with the aim of selecting efficient strains able to tolerate the different environmental variations while providing a successful N2 fixation. Multivariate tools (principal component and hierarchical clustering analyses) were used to show correlations between phenotypic and symbiotic variables and describe the similarities between the isolates’ origin sites.
Material and methods
Rhizobial strains’ isolation
Root nodules of Acacia saligna were collected from eight Moroccan geographic sites having different climates (Table 1). Bacteria were isolated according to the method recommended by Vincent (1970) and Beck et al. (1993). A digging of about 15–25 cm was conducted around the plant to a minimum depth of 20 to 50 cm to extract part of the root system (Lebrazi et al. 2018). Then, nodules surface was sterilized with 95% (v/v) ethanol for 30 s, and immersed in mercury chloride (HgCl2) solution 0.1% (w/v) for 4 min. A succession of three rinses for 10, 15 and 20 min, respectively, was carried out aseptically with sterile distilled water. Surface sterilized nodules were crushed with a few drops of 9‰ NaCl (w/v) under aseptic conditions (Beck et al. 1993). One hundred microliters of the suspension obtained was spread on Petri dishes containing YMA (Yeast, Mannitol Agar) culture medium (Vincent 1970) supplemented with 0.0025% (w/v) of Congo red. The single colonies were selected and restreaked in order to purify them (Jordan 1984). Pure cultures were stored in 20% glycerol at − 80 °C until subsequent use (Elbanna et al. 2009; Berrada et al. 2012).
Morphology of the colonies
The colony morphology of the different isolates was verified on the YMA agar medium. After incubation for 3 to 7 days at 28 °C, individual colonies were characterized for selected morphological parameters such as size, color, mucosity, transparency, shape, elevation, border and the production capacity of exopolysaccharide gum (Vincent 1970; Berrada et al. 2012).
The rhizobial isolates growth was evaluated on PGA (peptone-glucose-agar) broth medium according to Somasegaran and Hoben (1994) as a first level of selection due to weak rhizobia grow on this medium. Colonies resulting from pure cultures of the different isolates were subjected to microscopic observation after a Gram staining to allow more specific identification of these isolates and their bacterial walls to classify them into Gram positive or negative bacteria groups.
The strains' ability to alkalinize or acidify the YMA medium was evaluated by adding 0.0025% (w/v) bromothymol blue as color indicator. After incubating the inoculated plates at 28 °C for 24 h, the color change of the medium was observed. A yellow coloration indicates an acidic reaction and the rhizobia are fast growers. Dark blue coloration indicates a basic reaction and slow rhizobia growers. The isolates were divided into different morphotypes groups based on the observed morpho-cultural characteristics.
Symbiotic efficiency of bacterial isolates
The nodulation ability of the studied isolates was examined by inoculating A. saligna seedlings grown on pots containing sterile sandy-loam soil with three seeds per pot. Seeds were first surface sterilized by rinsing in 95% (v/v) ethanol, immersing for 4 min in 0.2% HgCl2 (w/v), and washing three times in sterile distilled water. They were further scarified with 95% H2SO4 and germinated on 0.9% agar in obscurity (Fikri-Benbrahim et al. 2017). One week after germination, seedlings were inoculated with approximately 1 ml of a freshly prepared bacterial suspension (108 UFC/ml) of each isolate. The plants were fed three times a week alternately with a sterilized N-free nutrient solution (Broughton and Dilworth 1971) and sterile distilled water. Non-inoculated plants were considered as N-free control (C0). Nitrogen control (NC), receiving weekly 0.5% KNO3 (w/v) as N source, was considered a non-inoculated control. Each treatment was prepared in triplicate. Six months after inoculation, plants were harvested to determine the infectivity and effectivity of isolates. Root nodules were recorded on each individual plant. The plant’s shoot and root dry weights were assessed. The average dry weight of plants inoculated with the same strain was used to estimate the relative efficiency (RE), which is expressed as the percentage of shoot dry weight of inoculated plants compared to the dry weight of nitogen control (El Hilali et al. 2007). The Kjeldahl method was then used to measure the nitrogen content of the aerial part (Nelson and Sommers 1973).
Evaluation of rhizobial tolerance to abiotic stresses
Rhizobia isolates were evaluated for their ability to grow under different environmental stresses (pH, salinity and temperature) conditions in YMA culture medium. For pH tolerance of rhizobia isolates, 10 μl of bacterial suspension of each rhizobia isolates (108 CFU/ml) were seeding on YMA culture medium adjusted at different pH values (4.8, 5.8, 6.8, 8.8, and 10.0). The plates were incubated at 28 °C for 1 week (Küçük et al. 2006). Salinity tolerance was evaluated in YMA culture medium with increased concentrations of sodium chloride (NaCl) ranging from 855 to 1710 mM (Lebrazi et al. 2018). Finally, the tolerance of rhizobial isolates to different temperatures was analyzed in YMA plates (pH 7.0) incubated at 6, 14, 28, 37, 44 and 54 °C (Hung et al. 2005). All tests were carried out in triplicate.
Amplification and sequencing of 16S rRNA gene
Genomic DNA extraction is directly performed from bacterial colonies by thermal shock technique. Universal primers 8F (5′-AGA GTT TGA TCC TGG CTC AG-3′) and 1540 R (5′-AAG GAG GTG ATC CAG CC-3′) were used for polymerase chain reaction (PCR) amplification of a 1500-bp segment of the 16S rRNA gene (Edwards et al. 1989; Lebrazi et al. 2020). Each 50 µL reaction comprised 1.0 µL of the cell lysate (approximately 20 ng DNA), 1.25 U of GoTaqR G2 DNA polymerase (Promega), 1× reaction buffer, 5% Dimethyl sulfoxide (DMSO), 0.2 mM dNTPs as well as 0.15 µM of each primer. The PCR protocol was conducted as follows: an initial cycle of denaturation at 95 °C (4 min), 35 cycles of denaturation at 95 °C (1 min each), annealing at 55 °C (1 min), extension at 72 °C (2 min), and the last extension step fixed in 7 min. Sanger sequencing was used to determine the nucleotide sequence for both strands of PCR products. DNA sequences were compared to the GenBank database by basic local alignment search tool (BLAST) requests using the blast-n algorithm and the highly similar sequence optimization (megablast) (Lebrazi et al. 2018). Phylogenetic tree was constructed using the MEGA-X program (Kumar et al. 2018).
Multivariate statistical analyses
In order to explore the possible relationships between the plant environmental stressors and symbiont characteristics, we perform the principal component analysis (PCA) to obtain a projection of variables on the factorial plane.
The considered phenotypic variables were temperature, NaCl, and pH tolerance. At the plant level, the variables of nodulation, aerial parts and root growth, as well as the nitrogen content of the aerial parts of the plant were taken into account.
The phenotypic variables considered in PCA were pH tolerance, salinity, and temperature, while, at the plant level, nodulation, shoot and root growth, as well as N content were taken into account to this analysis. The hierarchical cluster analysis (HCA) was used to better visualize the clustering of locations. Pearson’s correlation test was used to evaluate the correlation between studied variables. All statistical tests were carried out using JMP Software V 14 (Bridges 1966).
Results
Morpho-cultural characteristics of isolates
The studied isolates were identified as Gram-negative bacilli, which were unable to absorb congo red present on YMA medium and grow on PGA medium. The isolates' colonies presented the same morphology on the YMA medium: circular, convex, smooth, translucent, creamy to transparent texture. All isolates were considered fast acid producers as they changed the bromothymol blue indicator from deep green to yellow in the YMA-BTB. Moreover, 90% of the studied isolates presented a mucoid texture indicating the production of exopolysaccharides.
Evaluation of symbiotic parameters of isolates
Isolates’ efficiency was determined by examining the presence of red coloration in the nodules. Great variability in the infectivity of isolates was detected (Table 2). Even the plants receiving the same inoculum concentrations from different isolates, the number of nodules per plant was different from one plant to another. I78 was the most infective isolate and induced 89 nodules per plant, while the least infective one (I57) caused the development of only 20 nodules per plant. Some plant-rhizobia partners were superior to nitrogen control in terms of plant dry weight, with highlights for plants inoculated with I1 and I78 (Table 2). Plants inoculated with I1 exhibited a shoot dry weight of 15.32 g plant−1 and a RE of 127.49%, indicating that this isolate is the most efficient among others. The least efficient isolate was I5 with RE of the 45.25%. The RE of non-inoculated control was 10.52%. In general, the isolates I1, I28, I39 and I78 were very efficient showing a higher RE (> 101%). The total N content showed significant differences between the tested isolates (p-value < 0.0001) (Table 2). Plants inoculated with I1, I39, I70, and I78 accumulated more nitrogen and present 4.90%, 4.49%, 4,43%, and 4.37% of N content, respectively. On the other hand, plants inoculated with I57 displays the lowest N content (1.42%).
Tolerances of bacterial isolates to pH, salinity and temperature
Rhizobia isolates were able to tolerate higher salinity level (Table 3), however the maximum salinity tolerance varied greatly depending on the isolate. The ability to survive at temperatures between 14 and 28 °C was observed in all tested isolates, while a successful growth within the range of 6 to 44 °C has been recorded for six isolates (I1, I3, I5, I72, I74, and I76). Finally, the growth evaluation at different pH values showed optimum growth of all isolates at neural and slightly acidic pH (4.8 to 6.8). I1, I39, I70, I74, and I76 were able to grow in a broad pH range (4.8 to 10.0) (Table 3). I1 and I76 showed the combined ability to grow well under all evaluated stress conditions.
PCA and HCA
The PCA explain 72% of all data variability and two principal components (PC), being PC1 and PC2 explained 53.1% and 19.3%, respectively, of all data variation. A positive correlation between the four symbiotic parameters (SDW, RDW, nodulation and N content) were well explained by the first component (PC1) (Fig. 1). These results were confirmed by the Pearson correlation test showed in Table 4, which also indicated a positive correlation between tolerance to different temperature values and nodulation. The scores plotted below (Fig. 2) displays the provenances of samples that present a similarity according to the studied parameters. Thus, there was possible detect two large groups of samples. The group A was formed by Er-rachidia 2, Er-rachidia 3, Fez 1, Fez 2, Oujda 1, Oujda 3 and Oujda 4, while the group B consists from the following provenances: Berkane, Casablanca, Errachidia 1, Nador, Oujda 2, Saidia and Rabat. Hierarchical clustering analysis (HCA) was used to make an advanced classification of the isolates’ different origins (Fig. 3). The two large classes detected by PCA analysis (groups A and B) were subdivided into five subclasses by HCA, as follow (Fig. 3): (subclass 1) Er-Rachidia 2, Er-Rachidia 3, Fez 1, Oujda 1 and Oujda 4; (subclass 2) Fez 2 and Oujda 3; (subclass 3) Berkane, Er-Rachidia 1, Oujda 2 and Saidia; (subclass 4) Casablanca and Nador; and (subclass 5) Rabat. Using the biplot of scores and loadings (Fig. 4), the relationships between the individual clusters and the symbiotic parameters was detected. Thus, it was observed that rhizobia belonging to the large group A induced high values of the inoculated plants’ parameters (SDW, RDW, nodule number, N content) and possess high salinity and temperature tolerance. At the same time, those corresponding to group B are characterized by lower values of the plant’s parameters and tolerance to high pH levels.
Graph of correlations between phenotypic and symbiotic parameters according to the first two components. Vector correlation plot between the examined variables, i.e. stress tolerance (NaCl, pH and temperature tolerance) and symbiotic parameters: shoot dry weight (mg plant−1); root dry weight (mg plant−1); number of nodules (N Nod plant −1) and percentage of fixed nitrogen N (%)
Sequencing of 16S rRNA gene
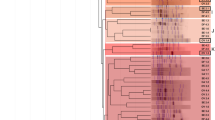
The 16S rDNA analysis showed that the sequenced isolates belonged to the genera Agrobacterium, Phyllobacterium and Rhizobium. BLAST homology searches indicated precise matches with the strain sequences present in GenBank (> 99%) for both sequences resulting from the forward and reverse sequencing reactions. Sequence analysis of the 16S rRNA gene is a rapid and precise method to identify bacterial phylogenetic position. The 16S rDNA of strain I76 (AB921256.1) was sequenced and then used to establish a phylogenetic tree (Fig. 5). Strain AB921256.1 was classified in the Rhizobium-Agrobacterium branch. It showed 99.8% similarity to Rhizobium pusense (LC208007.1) and Agrobacterium tumefaciens (KF709118.1).
Discussion
Abiotic stress is a common occurrence in the legume-rhizobium symbiosis, which significantly affects the nodulation process therefore nitrogen fixation (Basu and Kumar 2020).
Tolerance to abiotic stresses makes rhizobia very valuable inoculums to legumes grown in arid and semi-arid zones. Overall, our study revealed that the examined strains were quite tolerant to salinity in vitro, as some strains tolerated concentrations up to 1710 mM NaCl (10% NaCl). (Zahran 1999; Mohamed et al. 2000; Essendoubi et al. 2007; Diouf et al. 2007; Boukhatem et al. 2012). The range of salinity tolerance in rhizobia can be considerably different among species and even among strains of the same species (Missbah El Idrissi et al. 2021). Zerhari et al. (2000) reported that fast-growing rhizobia isolated from some Acacia species were tolerant to high NaCl concentrations compared to slow-growing strains. However, Assefa, (1993) found that some strains (slow-growing Rhizobium) from woody legumes were more tolerant to NaCl than fast-growing species. Other researches have suggested that growth rate and salt tolerance are not correlated with growth rate (Zerhari et al. 2000) but to other physiological and biochemical mechanisms (Botsford and Lewis 1990; Gouffi et al. 1999).
High soil temperature can present a serious constraint to leguminous crops. Soil surface temperatures can reach very high levels in the arid zone. Microbial inoculants are frequently exposed to high temperatures, adversely affecting rhizobia strain survival in this soil and the successful symbiotic relationship between rhizobia and legumes. High temperatures can reduce both saprophytic survival of rhizobia in the soil and effective nodulation (Patel et al. 2020). The isolated strains demonstrated relatively excellent temperature tolerance, with six strains tolerating temperatures between 6 and 44 °C, which is concordant with previous studies showing that rhizobia were able to grow within a broad temperature range (Maâtallah et al. 2002; Fikri-Benbrahim et al. 2017). Rhizobia are considered mesophilic, with optimal growth temperatures of 28 to 30 °C (Sharma et al. 2017). The maximum temperatures (Tmax) for growth of free-living rhizobia in soil are between 35 and 45 °C (Zhang et al. 1991; Zahran et al. 2012). Zerhari et al. (2000) and Assefa, (1993) also showed the ability of some rhizobia strains isolated from woody legumes to tolerate temperatures ranging from 4 to 43 °C. But, even though rhizobia grow at elevated temperatures, this does not indicate that they are efficient N2 fixers (Fentahun et al. 2013; Lebrazi et al. 2018).
The results of previous research demonstrating rhizobia's ability to grow in a diverse range of soil pH are corroborated by the evaluated strains' tolerance to different pH values ranging from (4.0–10.0), with optimal growth of all tested isolates at neutral and slightly acidic pH (4.8–6.8) (Küçük and Kıvanç 2010; Youseif et al. 2014; Lebrazi et al. 2018) even at pH 12.0 (Surange et al. 1997). Rhizobia strains contain variants that can be useful in tolerating abiotic stresses such as temperature extremes, pH and salinity. The exploration of tolerant Rhizobium strains is predicted to increase plant growth and yield, even under a combined stresses condition.
The nodules’ diagnosis highlights the presence of local native rhizobia able to nodulate Acacia, for all the fourteen samples. A variability of Acacia's nodulation and growth (shoot and root) is to be noted on all the studied bacteria. Specific variability in the efficiency of rhizobial symbiosis translates into variability in effectiveness (aerial biomass) and infectivity (nodules biomass). The criteria generally considered as determining in the selection of symbiotic bacteria concern the relationship of bacteria with plant (compatibility) or with the soil (adaptation). Strain compatibility with the host plant is defined according to infectivity and effectivity, which vary to the bacterial and plant genotype. Some rhizobial strains can be infectious and not effective but only the strains that are both infectious and effective are considered to be compatible. Using a multivariate statistical PC analysis, we were able to simultaneously evaluate the seven variables considered by condensing them into two principal components with minimum mathematical information loss. The principal components are used as axes where the data can be plotted and visualized structurally. An efficient rhizobial symbiosis is recognized by an overall statistically significant correlation between plant growth and nodulation. Significant correlations were found between dry weight of the Acacia aerial part and the number of nodules obtained by each rhizobial isolate (Fig. 1), indicating that the increased biomass recorded in plants inoculated with isolates was due to its increased nodulation potential, and not inevitably only to a greater N-fixing capacity (strain-specific). These findings proved that plants inoculated with the studied isolates (I1, I28, I39 and I78) had greater aerial biomass than the N-fertilized control plants and that these four strains resulted in an increase in root biomass compared to N-fertilized plants. This effect on root growth by inoculated rhizobia could promote plant tolerance to nutritional and drought stress through the enhancement of nutrient and water uptake by producing a more extensive root system. Significant correlations were also observed between the bacterial temperature tolerance and nodulation, as shown in Table 4. It was reported that rhizobial strains of tree legumes can nodulate and fix nitrogen at temperatures as high as 40 °C, and could represent a genetic source for nodulation at these temperatures with others species (Hungria et al. 1993). Inoculation with effective Rhizobium strains possessing high temperature tolerance and effective plant growth-promoting characteristics at higher temperatures would be required to enhance nodulation and their functioning to improve nitrogen fixation and plant growth (Patel et al. 2020).
The spatial variability of nodulation and growth observed in the different tested strains proves that it is difficult to build a model or draw conclusions by observing nodulation parameters only on a single plot. Each plot has its own specificities that influence legume nodulation. Soil, physicochemical proprieties, geomorphy, previous crops, environment, climate, plant and microorganisms are all edapho-climatic, microbiological and physiological factors that influence the interactions of legumes with the soil.
PCA was used to explore the variation in the distribution of isolates in the PC1–PC2 space according to the different studied stress tolerance phenotypes and symbiotic parameters as well as their geographic sites of origin. The phenotypic and symbiotic characteristics of strains from the same site varied considerably. Thus, as clearly demonstrated by the PCA (Fig. 4), the absence of an effect of the corresponding sampling sites on the different strains was highlighted, as no similarity could be found between the results of strains of the same geographical origin. Infact, the rhizobial community at a sample site may be more heterogeneous, and the rhizosphere of one plant may have the same rhizobial diversity as the entire sample region, hosting many rhizobial strains (Moschetti et al. 2005). Nevertheless, it was possible to detect those strains of the different geographical origins showed similar responses, as is the case for the Oujda1 and Errachidia3 strains. In the same way, the study of Boukhatem et al. (2012) on the diversity of rhizobia associated with Acacia in arid and semi-arid regions of Algeria showed the absence of correlation between the in vitro tolerances of the strains to different abiotic stress factors and their origin’s areas.
Concerning the molecular characterization, 16S rRNA gene sequencing has been widely employed to investigate rhizobial diversity associated with legume species (Cardinale et al. 2008; Muindi et al. 2017; Lebrazi et al. 2018), especially A. saligna (Zerhari et al. 2000; Amrani et al. 2010). The sequencing result indicates that the studied isolates were identified as members of Agrobacterium, Phyllobacterium and Rhizobium genera (Table 5). Earlier studies have reported that strains associated with Acacia spp. in Africa, can belong to Agrobacterium, Bradyrhizobium, Ensifer, Mesorhizobium and Rhizobium (De Lajudie et al. 1994; Odee et al. 2002; Amrani et al. 2010).
Recently, nitrogen-fixing bacteria have also been described in other genera of alphaproteobacteria, including Phyllobacterium (Zakhia et al. 2006; De Meyer et al. 2015). This finding suggests that the genes responsible for symbiosis with legumes are horizontally transmissible and function across a relatively wide range of bacterial taxa. According to previous research, several Acacia species have been nodulated by the genus Phyllobacterium. (Hoque et al. 2011; Crisóstomo et al. 2013). In some known exceptional cases, 16S rDNA sequencing cannot differentiate between some genera, as with Agrobacterium species and Rhizobium species. Although Rhizobium is the nitrogen-fixing symbiont genus and Agrobacterium is the plant pathogen genus, these two genera are phylogenetically intertwined with each other and their 16S rDNA sequences cannot separate them. Young et al. (2003) proposed that all Agrobacterium species could more correctly be considered members of Rhizobium. However, many Agrobacterium strains isolated from legume root nodules were unable to re-nodulate their original hosts (Mhamdi et al. 2005; Wang et al. 2006), but otherwise they are able to colonize pre-formed nodules. The mechanism whereby these isolates are incorporated into the nodules is currently not well understood. In contrast, the findings of this study revealed the highly symbiotic stability of our tested Agrobacterium strain for nodulating acacia roots. As reported in earlier studies (Moulin et al. 2004; García-Fraile et al. 2010), the ability of Agrobacterium to nodulate legume plants could be attributable to symbiotic genes acquisition by lateral gene transfer. Horizontal transfer can be considered a key mechanism whereby legumes can form symbioses with some bacteria that were previously classed as non-symbiotic, or unable to re-nodulate their original plants. Moreover, some Agrobacterium strains have recently been revealed that some Agrobacterium strains carry nodulating symbiosis-specific genes (e.g. nifH and nodA) identical to those of other legume symbionts (Rincón-Rosales et al. 2009; Cummings et al. 2009; Youseif et al. 2014).
It appears that genes other than those implicated in the symbiotic process should probably be involved. Transient acquisition of a symbiotic plasmid was thought to be an appropriate explanation. De Lajudie et al. (1999) showed that these Agrobacterium isolates are non-pathogenic. Mrabet et al. (2006) tested the effect of inoculation of Phaseolus vulgaris with Agrobacterium isolates on nodulation rate and vegetative yield; and showed that inoculation with these isolates negatively affected nodulation and plant growth.
Therefore, according to the results obtained during this study, it can be assumed that the selected isolates could be beneficial for their use as potential agents and promoters of plant growth and development. However, the good results obtained in vitro cannot always be reliably reproduced under field conditions. Hence, further field trials using these bacteria would be necessary to understand their potential in the agro-ecosystem as PGPRs.
Conclusion
The use of tree legumes, such as acacias, to promote canopy restoration in arid and semi-arid areas appears particularly interesting due to their potential to adapt to a long dry season and harsh ecological conditions. This work, mainly based on studying the diversity of rhizobia isolated from root nodules of A. saligna from different regions of Morocco, allowed us to select efficient rhizobia capable of fixing nitrogen and tolerating various environmental constraints. PCA was used to discriminate and describe the similarities between the different origin sites with respect to all measured phenotypic and symbiotic variables for the studied isolates. The strains' tolerance to these factors needs to be further evaluated in symbiosis with the host plant. In this perspective, further complementary investigations on tree species provenances would be interesting to further explain the responses of this symbiosis to particular edaphoclimatic conditions.
Data availability
All data generated or analyzed during this study are included in this published article.
Abbreviations
- CFU:
-
Colony Forming Unit
- HCA:
-
Hierarchical clustering analysis
- HgCl2 :
-
Mercury chloride
- NaCl:
-
Sodium chloride
- PCA:
-
Principal component analysis
- KNO3 :
-
Potassium nitrate
- PCR:
-
Polymerase chain reaction
- PGA:
-
Peptone–glucose–agar
- RDW:
-
Root dry weight
- RE:
-
Relative efficiency
- SDW:
-
Shoot dry weight
- YMA:
-
Yeast, Mannitol Agar
References
Amrani S, Noureddine NE, Bhatnagar T et al (2010) Phenotypic and genotypic characterization of rhizobia associated with Acacia saligna (Labill.) Wendl. in nurseries from Algeria. Syst Appl Microbiol 33:44–51. https://doi.org/10.1016/J.SYAPM.2009.09.003
Arora NK (2018) Bioremediation: a green approach for restoration of polluted ecosystems. Environ Sustain 14(1):305–307. https://doi.org/10.1007/S42398-018-00036-Y
Assefa F (1993) Nodulation and nitrogen fixation by Rhizobium and Bradyrhizobium spp of some indigenous tree legumes of Ethiopia—ERef Bayreuth. Universität Bayreuth, Bayreuth
Ba S, Willems A, De Lajudie P et al (2002) Symbiotic and taxonomic diversity of rhizobia isolated from Acacia tortilis subsp. raddiana in Africa. Syst Appl Microbiol 25:130–145. https://doi.org/10.1078/0723-2020-00091
Bakhoum N, Fall D, Fall F et al (2018) Senegalia senegal (synonym: Acacia senegal), its importance to sub-Saharan Africa, and its relationship with a wide range of symbiotic soil microorganisms. South African J Bot 119:362–368. https://doi.org/10.1016/J.SAJB.2018.10.007
Basu S, Kumar G (2020) Nitrogen fixation in a legume-rhizobium symbiosis: the roots of a success story. Plant Microbe Symbiosis. https://doi.org/10.1007/978-3-030-36248-5_3
Beck D, Meteron L, Afandi F (1993) Practical rhizobium legume technology manual. Manual No. 9. International Center for Agricultural Research in the Dry Areas, Aleppo
Berrada H, Nouioui I, Houssaini MI et al (2012) Phenotypic and genotypic characterizations of rhizobia isolated from root nodules of multiple legume species native of Fez, Morocco. Afr J Microbiol Res 6:5314–5324. https://doi.org/10.5897/AJMR11.1505
Botsford JL, Lewis TA (1990) Osmoregulation in Rhizobium meliloti: production of glutamic acid in response to osmotic stress. Appl Environ Microbiol 56:488–494. https://doi.org/10.1128/AEM.56.2.488-494.1990
Boukhatem ZF, Domergue O, Bekki A et al (2012) Symbiotic characterization and diversity of rhizobia associated with native and introduced acacias in arid and semi-arid regions in Algeria. FEMS Microbiol Ecol 80:534–547. https://doi.org/10.1111/J.1574-6941.2012.01315.X
Bridges CC (1966) Hierarchical cluster analysis. Psychol Rep 18:851–854. https://doi.org/10.2466/PR0.1966.18.3.851
Broughton WJ, Dilworth MJ (1971) Control of leghaemoglobin synthesis in snake beans. Biochem J 125(4):1075–1080
Cardinale M, Lanza A, Bonnì ML et al (2008) Diversity of rhizobia nodulating wild shrubs of Sicily and some neighbouring islands. Arch Microbiol 1904(190):461–470. https://doi.org/10.1007/S00203-008-0394-2
Crisóstomo JA, Rodríguez-Echeverría S, Freitas H (2013) Co-introduction of exotic rhizobia to the rhizosphere of the invasive legume Acacia saligna, an intercontinental study. Appl Soil Ecol 64:118–126. https://doi.org/10.1016/J.APSOIL.2012.10.005
Cummings SP, Gyaneshwar P, Vinuesa P et al (2009) Nodulation of Sesbania species by Rhizobium (Agrobacterium) strain IRBG74 and other rhizobia. Environ Microbiol 11:2510–2525. https://doi.org/10.1111/J.1462-2920.2009.01975.X
De Lajudie P, Willems A, Pot B et al (1994) Polyphasic taxonomy of rhizobia: Emendation of the genus Sinorhizobium and description of Sinorhizobium meliloti comb. nov., Sinorhizobium saheli sp. nov., and Sinorhizobium teranga sp. nov. Int J Syst Bacteriol 44:715–733. https://doi.org/10.1099/00207713-44-4-715/CITE/REFWORKS
De Lajudie P, Willems A, Nick G et al (1999) Agrobacterium bv. 1 Strains Isolated from Nodules of Tropical Legumes. Syst Appl Microbiol 22:119–132. https://doi.org/10.1016/S0723-2020(99)80035-6
De Meyer SE, De Beuf K, Vekeman B, Willems A (2015) A large diversity of non-rhizobial endophytes found in legume root nodules in Flanders (Belgium). Soil Biol Biochem 83:1–11. https://doi.org/10.1016/J.SOILBIO.2015.01.002
Diouf D, Samba-Mbaye R, Lesueur D et al (2007) Genetic diversity of Acacia seyal Del. rhizobial populations indigenous to Senegalese soils in relation to salinity and pH of the sampling sites. Microb Ecol 54:553–566. https://doi.org/10.1007/S00248-007-9243-0
Edwards U, Rogall T, Blockerl H et al (1989) Isolation and direct complete nucleotide determination of entire genes. Characterization of a gene coding for 16S ribosomal RNA. Nucleic Acids Res 17:7843–7853
El Hilali I, Brhada F, Thami Alami I, Filali Maltouf A (2007) Evidence of the presence of genetically different rhizobial strains in a single cluster of a lupinoid nodule, the case of Lupinus luteus. J Food, Agric Environ 5:352–359
Elbanna K, Elbadry M, Gamal-Eldin H (2009) Genotypic and phenotypic characterization of rhizobia that nodulate snap bean (Phaseolus vulgaris L.) in Egyptian soils. Syst Appl Microbiol 32:522–530. https://doi.org/10.1016/J.SYAPM.2009.07.006
Essendoubi M, Brhada F, Eljamali JE et al (2007) Osmoadaptative responses in the rhizobia nodulating Acacia isolated from south-eastern Moroccan Sahara. Environ Microbiol 9:603–611. https://doi.org/10.1111/J.1462-2920.2006.01176.X
Fentahun M, Akhtar MS, Muleta D, Lemessa F (2013) Isolation and characterization of nitrogen deficit Rhizobium isolates and their effect on growth of haricot bean. African J Agric Res 8:5942–5952. https://doi.org/10.5897/AJAR2012.6690
Fikri Benbrahim K, Berrada H, El Ghachtouli N et al (2014) Les acacias: des plantes fixatrices d’azote prometteuses pour le développement durable des zones arides et semi-arides [Acacia: Promising nitrogen fixing trees for sustainable development in arid and semi-arid areas]. Int J Innov Appl Stud 8:46–58
Fikri-Benbrahim K, Chraibi M, Lebrazi S et al (2017) Phenotypic and genotypic diversity and symbiotic effectiveness of rhizobia isolated from Acacia sp. grown in Morocco. J Agric Sci Technol 19:201–216
García-Fraile P, Mulas-García D, Peix A et al (2010) Phaseolus vulgaris is nodulated in northern Spain by Rhizobium leguminosarum strains harboring two nodC alleles present in American Rhizobium etli strains: biogeographical and evolutionary implications. Can J Microbiol 56:657–666. https://doi.org/10.1139/w10-048
Geetha SJ, Joshi SJ (2013) Engineering rhizobial bioinoculants: a strategy to improve iron nutrition. Sci World J. https://doi.org/10.1155/2013/315890
Gouffi K, Pica N, Pichereau V, Blanco C (1999) Disaccharides as a new class of nonaccumulated osmoprotectants for Sinorhizobium meliloti. Appl Environ Microbiol 65:1491–1500. https://doi.org/10.1128/AEM.65.4.1491-1500.1999/ASSET/371F56D9-CCCA-42A5-80B1-8A079398A979/ASSETS/GRAPHIC/AM0491711002.JPEG
Hoque MS, Broadhurst LM, Thrall PH (2011) Genetic characterization of root-nodule bacteria associated with Acacia salicina and A. stenophylla (Mimosaceae) across south-eastern Australia. Int J Syst Evol Microbiol 61:299–309. https://doi.org/10.1099/IJS.0.021014-0/CITE/REFWORKS
Hung M-H, Bhagwath AA, Shen F-T et al (2005) Indigenous rhizobia associated with native shrubby legumes in Taiwan. Pedobiologia (JENA) 49:577–584. https://doi.org/10.1016/j.pedobi.2005.06.002
Hungria M, Franco AA, Sprent JI (1993) New sources of high-temperature tolerant rhizobia for Phaseolus vulgaris L. Plant Soil 149:103–109. https://doi.org/10.1007/BF00010767
Jordan D (1984) Family III Rhizobiaceae Conn. 1938–254. In: Krieg NR, Holt JG (eds) Bergey’s manual of systematic bacteriology, vol 1. The Williams and Wilkins Co., Baltimore, pp 235–244
Küçük Ç, Kıvanç M (2010) Preliminary characterization of Rhizobium strains isolated from chickpea nodules. Afr J Biotechnol 7:772–775. https://doi.org/10.4314/ajb.v7i6.58526
Küçük Ç, Kivanç M, Kinaci E (2006) Characterization of Rhizobium sp. isolated from bean. Turkish J Biol 30:127–132
Kumar S, Stecher G, Li M et al (2018) MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol Biol Evol 35:1547–1549. https://doi.org/10.1093/molbev/msy096
Lebrazi S, Niehaus K, Bednarz H et al (2020) Screening and optimization of indole-3-acetic acid production and phosphate solubilization by rhizobacterial strains isolated from Acacia cyanophylla root nodules and their effects on its plant growth. J Genet Eng Biotechnol 18:1–12. https://doi.org/10.1186/s43141-020-00090-2
Lebrazi S, Fikri-Benbrahim K (2022) Potential of tree legumes in agroforestry systems and soil conservation. In: Advances in legumes for sustainable intensification. Academic Press, New York, pp 461–482
Lebrazi S, Chraibi M, Fadil M, et al (2018) Phenotypic, genotypic and symbiotic characterization of rhizobial isolates nodulating Acacia sp. in Morocco. J Pure Appl Microbiol 12:249–263. https://doi.org/10.22207/JPAM.12.1.30
Maâtallah J, Berraho EB, Muñoz S et al (2002) Phenotypic and molecular characterization of chickpea rhizobia isolated from different areas of Morocco. J Appl Microbiol 93:531–540. https://doi.org/10.1046/J.1365-2672.2002.01718.X
Mhamdi R, Mrabet M, Laguerre G et al (2005) Colonization of Phaseolus vulgaris nodules by Agrobacterium-like strains. Can J Microbiol. https://doi.org/10.1139/w04-120
Missbah El Idrissi M, Bouhnik O, ElFaik S et al (2021) Characterization of Bradyrhizobium spp. nodulating Lupinus cosentinii and L. luteus microsymbionts in Morocco. Front Agron 3:17. https://doi.org/10.3389/FAGRO.2021.661295/BIBTEX
Mohamed SH, Smouni A, Neyra M et al (2000) Phenotypic characteristics of root-nodulating bacteria isolated from Acacia spp. grown in Libya. Plant Soil 224:171–183. https://doi.org/10.1023/A:1004838218642
Moschetti G, Peluso AL, Protopapa A et al (2005) Use of nodulation pattern, stress tolerance, nodC gene amplification, RAPD-PCR and RFLP–16S rDNA analysis to discriminate genotypes of Rhizobium leguminosarum biovar viciae. Syst Appl Microbiol 28:619–631. https://doi.org/10.1016/J.SYAPM.2005.03.009
Moulin L, Béna G, Boivin-Masson C, Stȩpkowski T (2004) Phylogenetic analyses of symbiotic nodulation genes support vertical and lateral gene co-transfer within the Bradyrhizobium genus. Mol Phylogenet Evol 30:720–732. https://doi.org/10.1016/S1055-7903(03)00255-0
Mrabet M, Mnasri B, Ben RS et al (2006) Agrobacterium strains isolated from root nodules of common bean specifically reduce nodulation by Rhizobium gallicum. FEMS Microbiol Ecol 56:304–309. https://doi.org/10.1111/J.1574-6941.2006.00069.X
Muindi MM, Muthini M, Njeru EM, Maingi J (2017) Symbiotic efficiency and genetic characterization of rhizobia and non rhizobial endophytes associated with cowpea grown in semi-arid tropics of Kenya. Heliyon e06867. https://doi.org/10.1016/j.heliyon.2021.e06867
Nelson DW, Sommers LE (1973) Determination of total nitrogen in plant material. Agron J 65:109–112. https://doi.org/10.2134/agronj1973.00021962006500010033x
Odee DW, Haukka K, McInroy SG et al (2002) Genetic and symbiotic characterization of rhizobia isolated from tree and herbaceous legumes grown in soils from ecologically diverse sites in Kenya. Soil Biol Biochem 34:801–811. https://doi.org/10.1016/S0038-0717(02)00009-3
Patel K, Banjare U, Kumari A, et al (2020) Temperature tolerant Rhizobium leguminosorum bv. viciae strains with plant growth promotion traits. J Pure Appl Microbiol 14:2603–2609. https://doi.org/10.22207/JPAM.14.4.36
Requena N, Perez-Solis E, Azcón-Aguilar C et al (2001) Management of indigenous plant-microbe symbioses aids restoration of desertified ecosystems. Appl Environ Microbiol 67:495–498. https://doi.org/10.1128/AEM.67.2.495-498.2001/ASSET/2B9559B2-882D-4FFA-9FA8-5B70879ECF61/ASSETS/GRAPHIC/AM0211115003.JPEG
Ribeiro-Barros AI, Silva MJ, Moura I et al (2018) The potential of tree and shrub legumes in agroforestry systems. In: Amanullah K, Fahad S (eds) Nitrogen in agriculture—updates. InTechOpen, London, pp 223–239
Rincón-Rosales R, Lloret L, Ponce E, Martínez-Romero E (2009) Rhizobia with different symbiotic efficiencies nodulate Acaciella angustissima in Mexico, including Sinorhizobium chiapanecum sp. nov. which has common symbiotic genes with Sinorhizobium mexicanum. FEMS Microbiol Ecol 67:103–117. https://doi.org/10.1111/j.1574-6941.2008.00590.x
Sharma A, Bandamaravuri KB, Sharma A et al (2017) Phenotypic and molecular assessment of chickpea rhizobia from different chickpea cultivars of India. 3 Biotech. https://doi.org/10.1007/S13205-017-0952-X
Somasegaran P, Hoben HJ (1994) Quantifying the growth of rhizobia. In: Handbook for rhizobia. Springer, New York. https://doi.org/10.1007/978-1-4613-8375-8_5
Suman J, Rakshit A, Ogireddy SD et al (2022) Microbiome as a key player in sustainable agriculture and human health. Front Soil Sci. https://doi.org/10.3389/FSOIL.2022.821589/FULL
Surange S, Wollum AG, Kumar N, Shekhar Nautiyal C (1997) Characterization of Rhizobium from root nodules of leguminous trees growing in alkaline soils. Can J Microbiol 43:891–894. https://doi.org/10.1139/M97-130
Taoufiq K, Faghire M, Tahrouch S et al (2018) Screening and molecular identification of endophytic bacteria isolated from legumes nodules and roots cultivated in Acacia rhizosphere soils collected in an arid region, Tata-Akka in south of Morocco. Indian J Nat Sci 9:14910–14919
Vincent J. (1970) A manual for practical study of root nodule bacteria. In: IBP Hand book No. 15. Blackwell, Oxford
Wang LL, Wang ET, Liu J et al (2006) Endophytic occupation of root nodules and roots of Melilotus dentatus by Agrobacterium tumefaciens. Microb Ecol 52:436–443. https://doi.org/10.1007/S00248-006-9116-Y/METRICS
Young JM, Kuykendall LD, Martínez-Romero E et al (2003) Classification and nomenclature of Agrobacterium and Rhizobium—a reply to Farrand et al. (2003). Int J Syst Evol Microbiol 53:1689–1695. https://doi.org/10.1099/IJS.0.02762-0/CITE/REFWORKS
Youseif SH, Abd El-Megeed FH, Ageez A et al (2014) Phenotypic characteristics and genetic diversity of rhizobia nodulating soybean in Egyptian soils. Eur J Soil Biol 60:34–43. https://doi.org/10.1016/J.EJSOBI.2013.10.008
Zahran HH (1999) Rhizobium-legume symbiosis and nitrogen fixation under severe conditions and in an arid climate. Microbiol Mol Biol Rev 63:968–989. https://doi.org/10.1128/MMBR.63.4.968-989.1999
Zahran HH, Fattah A, Yasser MM et al (2012) Diversity and environmental stress responses of rhizobial bacteria from Egyptian grain legumes. Aust J Basic Appl Sci 6:571–583
Zakhia F, Jeder H, Willems A et al (2006) Diverse bacteria associated with root nodules of spontaneous legumes in Tunisia and first report for nifH-like gene within the Genera Microbacterium and Starkeya. Microb Ecol 513(51):375–393. https://doi.org/10.1007/S00248-006-9025-0
Zerhari K, Aurag J, Khbaya B et al (2000) Phenotypic characteristics of rhizobia isolates nodulating acacia species in the arid and Saharan regions of Morocco. Lett Appl Microbiol 30:351–357. https://doi.org/10.1046/J.1472-765X.2000.00730.X
Zhang X, Harper R, Karsisto M, Lindstrom K (1991) Diversity of Rhizobium bacteria isolated from the root nodules of leguminous trees. Int J Syst Bacteriol 41:104–113. https://doi.org/10.1099/00207713-41-1-104/CITE/REFWORKS
Funding
This research did not receive any specific grant from funding agencies in the public, commercial, or not-for-profit sectors.
Author information
Authors and Affiliations
Contributions
LS conducted the main experiments, designed the work, and wrote the manuscript. FM analyzed the data.CM participated in some experiments. FBK directed the work. The authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests and this work did not receive funding from any organization or institution.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Lebrazi, S., Fadil, M., Chraibi, M. et al. Phenotypic, molecular, and symbiotic characterization of the rhizobial symbionts isolated from Acacia saligna grown in different regions in Morocco: a multivariate approach. World J Microbiol Biotechnol 39, 343 (2023). https://doi.org/10.1007/s11274-023-03775-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11274-023-03775-1