Abstract
In this study, it was aimed to determine the phylogroups of Escherichia coli isolates from horse, cat, dog, sheep, cattle, and chicken feces samples and to investigate some important virulence genes of the isolates. For this purpose, a total of 600 feces samples, 100 from each animal species, were used as material. For the isolation of E.coli, feces samples were directly inoculated on MacConkey agar. The identification of the isolates was performed via phenotypic tests and species-specific multiplex Polymerase Chain Reaction (mPCR) method. PCR methods were used to phylotype E.coli isolates and to investigate virulence genes (bfpA, eaeA, LT, ST, Stx1, and Stx2). Of the total 600 E.coli isolates recovered in this study, 120 (20%), 269 (44.8%), 58 (9.7%), 19 (3.2%), 35 (5.8%), 56 (9.3%), 31 (5.2%), and 12 (2%) were identified as phylogroup A, B1, B2, C, D, E, F, and Escherichia clade I, respectively. While the virulence gene was detected in 149 (24.8%) E.coli isolates, no virulence gene was detected in 451 (75.2%) isolates. According to the analysis results, the most determined virulence gene was Stx1, while the least determined virulence gene was LT. In conclusion, in this study, when both the animal species and the number of E.coli isolates examined are considered, the data obtained are of great importance in epidemiological terms. However, the detection of virulence genes in 13.5% among phylogroup A, B1, and C isolates with commensal characteristics suggest that these isolates may show pathogenic characteristics with the virulence genes they contain.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The genus Escherichia currently consists of a total of six species (E. albertii, E. coli, E. fergusonii, E. hermannii, E. marmotae, and E. ruysiae), and E. coli is the most important species first described and known in the genus (Parte et al. 2020). E. coli is a Gram-negative, usually motile, non-spore-forming, a rod-shaped microorganism found in the normal intestinal microbiota of human and warm-blooded animals and causing intestinal or extra-intestinal infections (Moxley 2013; Gomes et al. 2016; Jang et al. 2017). Some pathogenic E. coli strains have zoonotic properties and can be transmitted to humans by direct contact with the feces of various animals or by consuming food and water contaminated with feces (Mustak et al. 2013).
Knowing the genotypic and phenotypic characteristics of E. coli recovered from animals is important for the prevention of E. coli infections. Phylotyping plays an important role in knowing the characteristic features of E. coli isolates, preventing and controlling infections, and determining new treatment methods (Mustak et al. 2013; Omerovic and Mustak 2018). In addition, the phylotyping analysis, established by Clermont et al. (2000), confers significant information about E. coli strains, because of the variety of isolates in the ecological niche, lifestyle, propensity to cause disease and phenotypic and genotypic traits regarding their phylogroup background (Amiri and Ahmadi 2019). Phylogenetic typing using PCR has 80–95% concordance with MLST analysis, showing that such testing can be used to study the genetic diversity of strains of E. coli (Gordon et al. 2008; Coura et al. 2019). The Clermont phylotyping scheme remains a popular tool for E. coli classification, as it can be performed rapidly and inexpensively in a laboratory. In addition, this classification scheme remains useful to make comparisons of newly sequenced isolates against historical literature, which contains many references to strains classified only by the Clermont scheme. The Clermont Typing Method includes PCR methods in which chuA, arpA, yjaA, trpAgpC, ArpAgpE, trpA (internal control) genes, and TspE4.C2 DNA fragment are amplified. (Clermont et al. 2013; Waters et al. 2020). Considering the data obtained from both the Clermont Typing Method and advanced molecular analyzes (Multi-Locus Sequence Typing (MLST) and Whole Genome Sequence (TGS)), currently, 8 phylogroups (A, B1, B2, C, D, E, F, and G) belonging to E. coli and five phylogroups (Escherichia clade I–V) belonging to the genus Escherichia (Clermont et al. 2013, 2019). However, it is known that the isolates responsible for the extraintestinal infection are mostly in phylogroup B2, D, E, F, and G, and commensal isolates are in phylogroups A, B1, and C (Omerovic and Mustak 2018; Clermont et al. 2019).
Pathogenic E. coli strains cause various diseases in humans and animals with their virulence factors. Investigation of genes encoding virulence factors in the strains is important in terms of determining the pathogenic character of the strains and interpreting the pathogenesis of infections (Kaper et al. 2004; Weintraub 2007). Heat-Labile Toxin (LT) is a toxin inactivating at 60 ºC, for 30 min and causing diarrhea by activating adenylate cyclase, which catalyzes cyclic AMP (cAMP). Heat-Stable Toxin (ST) is a toxin resistant to 100 ºC, 15 min, and causes diarrhea as a result of cyclic GMP (cGMP) accumulation by activating guanylate cyclase in the intestinal epithelium (Erdem 1999; Moxley 2013). Shiga toxin (Stx) is a toxin usually produced by E. coli O157:H7 and causes diarrhea, hemorrhagic colitis, and hemolytic uremic syndrome (Erdem 1999; Melton-Celsa 2014). Stx has two subgroups (Stx1 and Stx2). Stx1 is the same as the Shiga toxin produced by Shigella dysenteriae serotype 1 or differs only in one amino acid. Stx2 has more toxic effects than Stx1 (Bertin et al. 2001; Moxley 2013). Bundle-forming pilus (bfp) is fimbrial adhesin produced by typical enteropathogenic E. coli (EPEC). bfp, together with EspA (an outer membrane protein), performs the first stage of adhesion of EPEC to intestinal epithelial cells (Moxley 2013). bfp, an important virulence factor of EPEC, is encoded by the bfpA gene found in EAF (EPEC Adhesion Factor) plasmids (Kaper et al. 2004; Melo et al. 2005). lntimin (eae) is an outer membrane protein and is encoded by the eae gene located in the LEE5 (Locus of Enterocyte Effacement 5). eae is produced by pathogenic E. coli strains (EPEC and enterohemorrhagic E. coli (EHEC)) that cause attaching and effacing (A/E) lesions in host cells (Moxley 2013; Omerovic et al. 2017).
In this study, it was aimed the isolation and identification of E. coli from cattle, sheep, horse, cat, dog, and chicken feces samples, the detection of the phylotypes for E. coli isolates obtained, and the investigation of the important virulence genes present in these isolates.
Materials and methods
Feces samples
In the study, a total of 600 feces samples (100 from each animal species) collected from slaughterhouses, farms, and animal shelters visited between April and August 2019 were used as material. It was collected cattle and sheep feces samples from seven slaughterhouses in Kayseri, Turkey, chicken feces samples from four poultry slaughterhouses in Bolu, Turkey, horse feces samples from three horse farms in Kayseri, cat, and dog feces samples from three animal shelters in Kayseri and Nevşehir, Turkey. It was transported to the laboratory in the cold chain after the samples were transferred into sterile plastic containers, and analyzed within the same day.
Standard strain
Escherichia coli ATCC 25,922, E. coli NCTC 13,384, and, E. coli clinical strains (obtained from animal feces and identified by PCR) found in the culture collection of Erciyes University, Faculty of Veterinary Medicine, Department of Microbiology were used as standard strains.
DNA extraction
Commercial DNeasy UltraClean Microbial Kit (Qiagen 12224-250, USA) was used to extract DNA from E. coli isolates. The extraction procedure was carried out in accordance with the manufacturer’s instructions.
Bacterial isolation
For the isolation of E. coli, feces samples were directly inoculated on MacConkey agar (Thermo Fisher Scientific CM0007, ABD). The plates were incubated at 37 °C in an aerobic environment for 24 h. At the end of the incubation period, the E. coli-suspect colonies (bright-pink coloured colonies on MacConkey agar) were evaluated and their pure cultures were obtained on 7% sheep blood agar (Neogen NCM2013A, USA). The pure cultures of the isolates were stored at − 84 °C in Brucella Broth (Liofilchem 64,026, Italy) supplemented with 15% glycerin.
Phenotypic identification
Gram staining, motility test, oxidase, catalase, indole, methyl red (MR), hydrogen sulfide (H2S), and carbohydrate fermentation tests were performed for the phenotypic identification of the E. coli-suspect isolates.
Molecular Identification
It was performed with minor modifications in the mPCR reported by Godambe et al. (2017), and for this purpose, uidA and uspA genes were amplified. For the PCR mix prepared in a total volume of 25 µl: 2.5 µl 10X PCR Buffer (Vivantis, RB0201, Malaysia), 3 µl MgCl2 (Thermo Scientific, R0971), 0.5 µl 10 mM dNTP (Vivantis, NP2410), 0.4 µl Taq polymerase (5 U/µl) (Thermo Scientific, EP0402), 1 µl from each primer (10 pm) (Table 1) and 3 µl template DNA were used. The thermal cycling conditions were: initial denaturation at 94 °C for 5 min, 35 cycles of amplification (denaturation at 94 °C for 10 s, annealing at 55 °C for 10 s, and extension at 72 °C for 1 min), and a final extension at 72 °C for 10 min.
Clermont typing method
In order to determine the phylogroups of E. coli isolates, the Clermont Typing Method, which consists of the quadruplex PCR, phylogroup C and E-specific mPCR reported by Clermont et al. (2013), was used. Quadruplex PCR was performed with minor modifications to the PCR reported by Clermont et al. (2013), and the chuA, yjaA, arpA genes, and TspE4.C2 DNA fragments were amplified (Table 1). For the PCR mix prepared in a total volume of 25 µl: 2.5 µl of 10X PCR buffer, 3 µl of MgCl2, 0.5 µl 10 mM dNTP, 0.4 µl Taq polymerase (5 U/µl), 1 µl of each primer (10 pm) and 3 µl template DNA were used. The thermal cycling conditions were: an initial denaturation step at 94 °C for 4 min, 30 cycles of amplification (denaturation at 94 °C for 5 s, annealing at 59 °C for 20 s, and extension at 72 °C for 1 min), and a final extension at 72 °C for 5 min. As a result of quadruplex PCR, a quadruplex profile was determined for each isolate by evaluating the presence/absence of the three genes and a DNA fragment mentioned above (e.g. +−+−, arpA +, chuA −, yjaA +, TspE4.C2 -). Thus, phylogroup C specific mPCR for the differentiation of the isolates with phylogroup A/C profile (+−+−); Phylogroup E specific mPCR for differentiating of the isolates with phylogroup D/E profiles (++−− and ++−+) and phylogroup E/Escherichia clade I profile (+++−) was performed. mPCR in which trpAgpC and trpA (internal control) genes were amplified, for the determination of E. coli phylogroup C; mPCR in which ArpAgpE and trpA (internal control) genes were amplified, for determining phylogroup E was carried out. PCR mix and amplification conditions (primer annealing in phylogroup E specific mPCR: 57 °C) are the same as for the quadruplex PCR mentioned above.
Analysis of virulence genes
For the investigation of virulence genes (LT, ST, Stx1, Stx2, bfpA, eaeA) in E. coli isolates, mPCR reported by Huasai et al. (2012) was performed with minor modifications. For the PCR mix prepared in a total volume of 25 µl: 2.5 µl of 10X PCR buffer, 3 µl of MgCl2, 0.5 µl 10 mM dNTP, 0.4 µl Taq polymerase (5 U/µl), 0.2 µl of each primer (10 pm) (Table 1) and 3 µl template DNA were used. The thermal cycling conditions were: initial denaturation at 94 °C for 1 min, 30 cycles of amplification (denaturation at 94 °C for 30 s, annealing at 55 °C for 30 s, and extension at 72 °C for 1 min), and a final extension at 72 °C for 10 min.
Agarose gel electrophoresis and imaging
The PCR products obtained during the molecular identification, phylotyping, and analysis of virulence genes of E coli isolates were subjected to electrophoresis in a 1.5% agarose gel added to ethidium bromide (3µL/50mL) at 120 V, 500 mA for 50 min. Then, the agarose gel was visualized on the imaging system (Syngene G: Box F3) and evaluated.
10. Statistical analysis
All statistical analyzes were performed using the Jamovi program (version 1.6.13) in the study. The statistical significance of the difference between the phylogroups of E. coli isolates and the rates of virulence genes in the isolates was evaluated with the Pearson Chi-square test and Fisher’s exact test and the significance level was determined as P < 0.05. In addition, the statistical correlation between phylogroups and virulence genes presence in E. coli isolates was calculated by the Pearson Chi-square test and the significance level was determined as P < 0.01.
Results
Isolation and identification
All of the 600 feces samples analyzed in the current study were found to be positive for E. coli, and a total of 600 E. coli-suspect isolates, one from each sample, were obtained. All of the isolates were identified as E. coli as a result of phenotypic tests and molecular analysis (mPCR). As a result of phenotypic tests, all isolates that were Gram (−), motile, oxidase, and H2S negative, catalase, indole, MR, glucose, lactose, and sucrose positive were evaluated as E. coli. All of the isolates that formed bands both 166 bp (uidA) and 884 bp (uspA) as a result of agarose gel electrophoresis in the species-specific mPCR used in molecular identification were defined as E. coli.
Clermont typing method
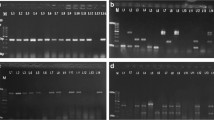
In the current study, bands of 400 bp (arpA), 288 bp (chuA), 211 bp (yjaA), and 152 bp (TspE4.C2) were detected in the agarose gel electrophoresis test performed after quadruplex PCR (Fig. 1). The phylogroups of E. coli isolates were determined according to the quadruplex profiles formed by the evaluation of the presence/absence of these three genes and a DNA fragment. In the agarose gel electrophoresis test performed after Group C specific mPCR, bands of 219 bp (trpAgpC) and 489 bp (trpA) were detected; after group E specific mPCR, bands of 301 bp (ArpAgpE) and 489 bp (trpA) were detected (Fig. 2). The isolates positive both trpAgpC and trpA in phylogroup A/C profile (+−+−) were defined as phylogroup C, while the isolates positive both ArpAgpE and trpA in phylogroup D/E profile (++−− and ++−+) or in E/Escherichia clade I profile (+++−) were defined as phylogroup E. Only the trpA positive isolates were defined as phylogroup A, D, or Escherichia clade I.
The agarose gel electrophoresis image of quadruplex PCR products. arpA (400 bp), chuA (288 bp) and yjaA (211 bp) genes, and TspE4.C2 (152 bp) DNA fragment. M: Marker (100–3000 bp), P1: Phylogroup B1 Positive Control (E. coli ATCC 25,922 (+−−+)), P2: Phylogroup B2 Positive Control (E. coli NCTC 13,384 (−++−)), N: Negative Control (Sterile deionized distilled water), 1: Phylogroup A isolate (+−−−), 2: Phylogroup A/C isolate (+−+−), 3: Phylogroup B1 isolate (+−−+), 4–6: Phylogroup B2 isolates (−++−), (−+−+), (−+++), 7–8: Phylogroup D/E isolates (++−−), (++−+), 9: Phylogroup E/Escherichia clade I isolate (+++−), 10: Phylogroup F isolate (−+−−), 11: Phylogroup Escherichia clade I isolate (−−+−)
The agarose gel electrophoresis image of phylogroup C and phylogroup E specific mPCR products. trpAgpC (219 bp), ArpAgpE (301 bp) and trpA (489 bp) (internal control) genes. M: Marker (100–3000 bp), P1: Phylogroup C Positive Control (E. coli clinical isolate), P2: Phylogroup E Positive Control (E.coli clinical isolate), N: Negative Control (Sterile deionized distilled water), 1–2: Phylogroup C isolates, 3–4: Phylogroup A isolates, 5–6: Phylogroup E isolates, 7: Phylogroup D isolate, 8: Phylogroup Escherichia clade I isolate
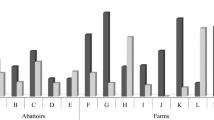
Of the total 600 E. coli isolates recovered in current study, 120 (20%), 269 (44.8%), 58 (9.7%), 19 (3.2%), 35 (5.8%), 56 (9.3%), 31 (5.2%) and 12 (2%) were identified as phylogroup A, B1, B2, C, D, E, F and Escherichia clade I, respectively. According to the analysis results, the most detected phylogroup was B1 (44.8%), while the least detected phylogroup was Escherichia clade I (2%). However, phylogroup B2, D, F, and Escherichia clade I in the isolates from horse feces, phylogroup C and Escherichia clade I in the isolates from cat feces, phylogroup C and F in the isolates from sheep feces, and, Escherichia clade I in the isolates from chicken feces was not detected. The phylogroup distribution of E. coli isolates from animal feces samples is given in Table 2. As a result of the statistical analysis, it was found a significant correlation between the animal species from which E. coli isolates were obtained and the phylogroup type detected (P < 0.05) (Table 2). Accordingly, the distribution of phylogroups varied among animal species.
Analysis of virulence genes
In the study, bands of 326 bp (bfpA), 891 bp (eaeA), 450 bp (LT), 190 bp (ST), 370 bp (Stx1), and 283 bp (Stx2) were detected in the agarose gel electrophoresis test performed after mPCR (Fig. 3). While the virulence gene was detected in 149 (24.8%) of 600 E. coli isolates, no virulence gene was detected in 451 (75.2%) of them. Of the isolates, 10 (1.6%), 5 (0.8%), 17 (2.8%), 2 (0.3%), 17 (2.8%), 54 (9%), 26 (4.3%) and 18 (3%) was found to be positive bfpA, eaeA, eaeA + Stx2, LT, ST, Stx1, Stx2 and Stx1 + Stx2, respectively (Table 3). According to the analysis results, the most determined virulence gene was Stx1, while the least determined virulence gene was LT. It was statistically found a significant difference in terms of presence rate of virulence genes in the 600 isolates of animal origin (P < 0.05) (Table 3).
The agarose gel electrophoresis image of the amplification products obtained from mPCR used to investigate the E. coli virulence genes. ST (190 bp), Stx2 (283 bp), Stx1 (370 bp), bfpA (326 bp), LT (450 bp) and eaeA (891 bp) genes. M: Marker (100–3000 bp), P1: bfpA and eaeA positive EPEC clinical isolate, P2: ST and LT positive ETEC clinical isolate, P3: Stx1 and Stx2 positive STEC clinical isolate, N: Negative Control (Sterile deionized distilled water), 1: bfpA positive isolate, 2: eaeA positive isolate, 3: eaeA and Stx2 positive isolate, 4: LT positive isolate, 5: ST positive isolate, 6: Stx1 positive isolate, 7: Stx2 positive isolate, 8: Stx1 and Stx2 positive isolate
When E. coli phylogroups and the virulence genes that they contain were compared, while the most virulence gene was detected in phylogroup A isolates, the least virulence gene was detected in Escherichia clade I isolates. The association between the phylogroups of E. coli isolates and the virulence genes detected are given in detail in Table 4. It was statistically found a significant difference in terms of phylotyping and virulence genes presence in the 600 coli isolates (P < 0.01).
Discussion
The infections caused by E. coli are one of the major causes of economic losses in the livestock sector. Therefore, knowing the genotypic and phenotypic characteristics of E. coli isolates isolated from animals is important for the prevention of these infections (Clermont et al. 2013; Mustak et al. 2013). In this study, phylogroups of 600 E. coli isolates obtained from animal feces were determined by PCR and, of the isolates, 120 (20%), 269 (44.8%), 58 (9.7%), 19 (3.2%), 35 (5.8%), 56 (9.3%), 31 (5.2%) and 12 (2%) were identified as phylogroup A, B1, B2, C, D, E, F and Escherichia clade I, respectively (Table 2).
There are a limited number of studies conducted in Turkey to determine the phylogroups of E. coli isolates recovered from animal feces. Mustak et al. (2013) reported that of 61 E. coli isolates from Wistar rats, 70.4% and 29.5% identified as phylogroup B1 and B2, respectively. The fact that the ratio of phylogroups B1 and B2 determined in this study was different from our study suggests that it may be due to the difference in animal species examined. Omerovic and Mustak (2018) reported that of 150 E. coli isolates from chicken feces, 19.32%; 35.35%; 11.33%, and 34% were defined as commensal phylogroups (A, B1, C), extraintestinal phylogroup (B2, D, E, F), Escherichia clades and unknown phylogroup, respectively. The fact that the rate of commensal phylogroups (19.32%) detected in this study was lower than the rate (57%) detected in the isolates of chicken origin in our study can be explained by the fact that the researchers used the isolates recovered from the feces of sick chickens.
When the studies on the phylotyping of E. coli isolates of animal origin worldwide were examined, Bhave et al. (2019) reported that of the 19 ExPEC strains from chicken feces, 52.63%, 36.84%, and 10.53% were identified as phylogroup B2, A and D, respectively. Goudarztalejerdi et al. (2020) defined 72% of 50 isolates obtained from chicken feces as extraintestinal phylogroups. The fact that the ratio of extraintestinal phylogroups detected in these studies was higher than the ratio (43%) detected in isolates from chicken feces in our study can be explained by the characteristics of the samples examined, nutrition, hygiene level, and the PCR protocol used these study.
In the studies performed on the phylotyping of E. coli isolates from cat and dog feces, Akhtardanesh et al. (2016) determined 66.7%, 1.2%, 13.4%, and 18.9% of 90 isolates from cats was phylogroup A, B1, B2, and D, respectively. Coura et al. (2018) reported that of 37 isolates from dogs, 1 (2.7%), 8 (21.6%), 7 (18.9%), 3 (8.1%), 8 (21.6%) and 10 (27%) identified as phylogroup A, B1, B2, D, E and unknown, respectively. While Bourne et al. (2019) detected the most phylogroup B1 (35%) in 203 isolates from dogs, they detected the most phylogroup B2 (41.3%) in 334 isolates from cats. In our study, 28% of the isolates from cat feces were commensal phylogroup (A and B1) and 72% of them were extraintestinal phylogroup (B2, D, E, and F); on the other hand, 54% of the isolates from dog feces were commensal phylogroup (A, B1, and C) and 46% of them were extraintestinal phylogroup (B2, D, E, and F). In the studies mentioned above, it was reported different results in the phylogroup distribution of the isolates from cat and dog feces. This can be associated with the character/number of the samples, geography, climate, diet, intestinal morphology, and hygiene level (Stoppe et al. 2017).
When the studies (Souto et al. 2017; Coura et al. 2017; Wang et al. 2018) performed on phylotyping of E. coli isolates from cattle and calf feces are examined, it is seen that the most detected phylogroup is B1 (> 50%). In our study, it was defined the most phylogroup B1 (75%) in the isolates from cattle feces and the results were found to be consistent with these studies (Souto et al. 2017; Coura et al. 2017; Wang et al. 2018). When the studies (Carlos et al. 2010; Johnson et al. 2017; Kennedy et al. 2018; Saei and Zavarshani 2018) performed on phylotyping of E. coli isolates from horse and sheep feces were examined, most of the isolates were defined as commensal phylogroup (A, B1) and these results were found to be consistent with our study.
In the current study, the phylogroups of 12 (2%) E. coli isolates were also determined as Escherichia clade I. In order to confirm the isolates, it is predicted to use the PCR methods used cryptic clade primers (Clermont et al. 2013).
In this study, the presence of various virulence genes in 600 E. coli isolates was investigated and while it was detected virulence genes in 149 (24.8%) isolates, it was not detected virulence genes in 451 (75.1%) isolates. The most determined virulence gene was Stx1, while the least determined virulence gene was LT. In addition, only one virulence gene in 114 isolates was detected, while two virulence genes in 35 isolates were detected together (Table 3).
There are a limited number of studies conducted in Turkey to investigate the presence of bfpA, eaeA, LT, ST, Stx1 and Stx2 virulence genes in E. coli isolates from animal feces. Aydın et al. (2010) investigated the presence of eaeA, Stx1, and Stx2 in six E. coli O157:NM strains from cattle and they detected Stx2 in 2 (33.3%) strains. Pehlivanoglu et al. (2020) examined the presence of eaeA, Stx1, and Stx2 in 17 E. coli isolates from cattle and they detected eaeA in one isolate. In these two studies, it is seen that Stx1 was not detected in E. coli isolates, positivities of eaeA and Stx2 were found between 0 and 5.9% and 0-33.3%, respectively. In our study, positivities of eaeA, Stx1, and Stx2 in the isolates of cattle origin were detected as 2%, 9%, and 9%, respectively. The minor differences in virulence gene positivity can be explained by the character of the samples examined, the number of the isolates examined or the PCR methods used.
In the studies performed on the investigation of existence of these virulence genes in worldwide, it is seen that it was eaeA the most gene detected in the E. coli isolates of cattle, horse, and cat origin (Huasai et al. 2012; Cabal et al. 2013; Chandran and Mazumder 2013; Caliman and Marin 2014; Tostes et al. 2017; Watson et al. 2017); was eaeA and Stx1 the most genes detected in the isolates of dog and sheep origin (Chandran and Mazumder 2013; Sekse et al. 2013; Ferreira et al. 2015; Torkan et al. 2016); was eaeA and Stx2 the most detected genes in the isolates of chicken origin (Ghanbarpour et al. 2011; Oh et al. 2012; Cabal et al. 2013; Chandran and Mazumder 2014), and the rate of virulence genes detected in the isolates has varied. This can be explained by the character (diarrheal/normal) of the samples, hygiene management, nutrition, or the PCR method used. The type/positivity of virulence genes detected in the E. coli isolates in our study is consistent with the studies mentioned above, except for minor differences. In addition, in the studies mentioned above, the rate of virulence genes detected in the isolates of chicken origin is between 6 and 40%. Contrary to these studies, a relatively high rate (48%) of virulence genes were detected in the isolates of chicken origin in our study. This revealed that hygienic measures in poultry slaughterhouses where samples were collected should be reviewed.
In this study, virulence genes were detected in 31 phylogroups A, 15 B1, 12 B2, 9 C, 27 D, 27 E, 26 F, and two Escherichia clade I isolates (Table 4). There are the limited number of studies (Ghanbarpour et al. 2011; Ferreira et al. 2015; Coura et al. 2017, 2018; Kennedy et al. 2018; Saei and Zavarshani 2018) in worldwide in which both phylogroups and virulence genes (bfpA, eaeA, LT, ST, Stx1, Stx2) in the E. coli isolates of animal origin (fecal) were investigated and compared. In these studies (Ghanbarpour et al. 2011; Ferreira et al. 2015; Coura et al. 2017, 2018; Kennedy et al. 2018; Saei and Zavarshani 2018), an average of 50% virulence genes presence were detected even in phylogroup A and B1 isolates, which are considered commensal. In our study, the most virulence gene was statically detected in phylogroup A isolates, and it was consistent with these studies (Table 4).
Conclusion
In the current study, the detection of virulence genes in 13.5% among phylogroup A, B1, and C isolates with commensal characteristics suggest that these isolates may show pathogenic characteristics with the virulence genes they contain. In addition, the detection of virulence genes at the rate of 48% in the E. coli isolates of chicken origin revealed that hazard analyzes and critical control points (HACCP) programs should be actively applied in poultry slaughterhouses.
References
Akhtardanesh B, Ghanbarpour R, Ganjalikhani S, Gazanfari P (2016) Determination of antibiotic resistance genes in relation to phylogenetic background in Escherichia coli isolates from fecal samples of healthy pet cats in Kerman city. Vet Res Forum 7:301–308
Amiri M, Ahmadi E (2019) A comparative phylotyping analysis of E. coli originating from avian colibacillosis based on the Clermont triplex scheme and gyrA gene sequencing. Turk J Vet Anim Sci 43:212–217. https://doi.org/10.3906/vet-1811-14
Aydın F, İça T, Yontar A (2010) Kayseri Yöresinde Süt Sığırlarında Escherichia coli O157:H7’nin Konvansiyonel ve Moleküler Yöntemlerle Araştırılması. Erciyes Univ Sağlık Bilimleri Derg 19:159–166
Bertin Y, Boukhors K, Pradel N et al (2001) Stx2 subtyping of Shiga toxin-producing Escherichia coli isolated from cattle in France: detection of a new Stx2 subtype and correlation with additional virulence factors. J Clin Microbiol 39:3060–3065. https://doi.org/10.1128/JCM.39.9.3060-3065.2001
Bhave S, Kolhe R, Mahadevaswamy R et al (2019) Phylogrouping and antimicrobial resistance analysis of extraintestinal pathogenic Escherichia coli isolated from poultry species. Turkish J Vet Anim Sci 43:117–126. https://doi.org/10.3906/vet-1808-47
Bourne JA, Chong WL, Gordon DM (2019) Genetic structure, antimicrobial resistance and frequency of human associated Escherichia coli sequence types among faecal isolates from healthy dogs and cats living in Canberra, Australia. PLoS ONE 14:1–13. https://doi.org/10.1371/journal.pone.0212867
Cabal A, Gómez-Barrero S, Porrero C et al (2013) Assessment of virulence factors characteristic of human Escherichia coli pathotypes and antimicrobial resistance in O157:H7 and non-O157:H7 isolates from livestock in Spain. Appl Environ Microbiol 79:4170–4172. https://doi.org/10.1128/AEM.00537-13
Caliman MW, Marin JM (2014) Virulence genes in Escherichia coli isolated from feces and urine of cats. Arq Bras Med Vet 66:1287–1290. https://doi.org/10.1590/1678-6694
Carlos C, Pires MM, Stoppe NC et al (2010) Escherichia coli phylogenetic group determination and its application in the identification of the major animal source of fecal contamination. BMC Microbiol 10:161. https://doi.org/10.1186/1471-2180-10-161
Chandran A, Mazumder A (2013) Prevalence of diarrhea-associated virulence genes and genetic diversity in Escherichia coli isolates from fecal material of various animal hosts. Appl Environ Microbiol 79:7371–7380. https://doi.org/10.1128/AEM.02653-13
Chandran A, Mazumder A (2014) Occurrence of diarrheagenic virulence genes and genetic diversity in Escherichia coli isolates from fecal material of various avian hosts in British Columbia, Canada. Appl Environ Microbiol 80:1933–1940. https://doi.org/10.1128/AEM.03949-13
Clermont O, Bonacorsi S, Bingen E (2000) Rapid and simple determination of the Escherichia coli phylogenetic group. Appl Environ Microbiol. 66:4555–4558. https://doi.org/10.1128/AEM.66.10.4555-4558.2000
Clermont O, Christenson JK, Denamur E, Gordon DM (2013) The Clermont Escherichia coli phylo-typing method revised: improvement of specificity and detection of new phylo-groups. Environ Microbiol Rep 5:58–65. https://doi.org/10.1111/1758-2229.12019
Clermont O, Dixit OVA, Vangchhia B et al (2019) Characterization and rapid identification of phylogroup G in Escherichia coli, a lineage with high virulence and antibiotic resistance potential. Environ Microbiol 21:3107–3117. https://doi.org/10.1111/1462-2920.14713
Coura FM, de Araújo Diniz S, Mussi JMS et al (2017) Characterization of virulence factors and phylogenetic group determination of Escherichia coli isolated from diarrheic and non-diarrheic calves from Brazil. Folia Microbiol (Praha) 62:139–144. https://doi.org/10.1007/s12223-016-0480-9
Coura FM, Diniz AN, Oliveira CA et al (2018) Detection of virulence genes and the phylogenetic groups of Escherichia coli isolated from dogs in Brazil. Ciência Rural 48:1–6. https://doi.org/10.1590/0103-8478cr20170478
Coura FM, Diniz SA, Silva MX et al (2019) Virulence factors and phylotyping of Escherichia coli isolated from non-diarrheic and diarrheic water buffalo calves. Cienc Rural 49:e20180998. https://doi.org/10.1590/0103-8478cr20180998
Erdem B (1999) Enterobacteriaceae. In: Ustaçelebi S (ed) Temel ve Klinik Mikrobiyoloji. Güneş Kitabevi, Ankara, pp 471–515
Ferreira MRA, Silva TS, Stella AE et al (2015) Detection of virulence factors and antimicrobial resistance patterns in shiga toxin-producing Escherichia coli isolates from sheep. Pesq Vet Bras 35:775–780. https://doi.org/10.1590/S0100-736X2015000900002
Ghanbarpour R, Sami M, Salehi M, Ouromiei M (2011) Phylogenetic background and virulence genes of Escherichia coli isolates from colisepticemic and healthy broiler chickens in Iran. Trop Anim Health Prod 43:153–157. https://doi.org/10.1007/s11250-010-9667-2
Godambe LP, Bandekar J, Shashidhar R (2017) Species specific PCR based detection of Escherichia coli from Indian foods. 3 Biotech 7:130. https://doi.org/10.1007/s13205-017-0784-8
Gomes TA, Elias WP, Scaletsky IC et al (2016) Diarrheagenic Escherichia coli. Braz J Microbiol 47:3–30. https://doi.org/10.1016/j.bjm.2016.10.015
Gordon DM, Clermont O, Tolley H, Denamur E (2008) Assigning Escherichia coli strains to phylogenetic groups: multi-locus sequence typing versus the PCR triplex method. Environ Microbiol 10:2484–2496. https://doi.org/10.1111/j.1462-2920.2008.01669
Goudarztalejerdi A, Mohammadzadeh A, Najafi SV et al (2020) Serogrouping, phylotyping, and virulence genotyping of commensal and avian pathogenic Escherichia coli isolated from broilers in Hamedan, Iran. Comp Immunol Microbiol Infect Dis 73:101558. https://doi.org/10.1016/j.cimid.2020.101558
Huasai S, Chen A, Wang C et al (2012) Occurrence and characteristics of virulence genes of Escherichia coli strains isolated from healthy dairy cows in Inner Mongolia, China. Braz J Microbiol 43:528–534. https://doi.org/10.1590/S1517-83822012000200013
Jang J, Hur HG, Sadowsky MJ et al (2017) Environmental Escherichia coli: ecology and public health implications-a review. J Appl Microbiol 123:570–581. https://doi.org/10.1111/jam.13468
Johnson JR, Johnston BD, Delavari P et al (2017) Phylogenetic backgrounds and virulence-associated traits of Escherichia coli isolates from surface waters and diverse animals in minnesota and wisconsin. Appl Environ Microbiol 83:e01329–e01317. https://doi.org/10.1128/AEM.01329-17
Kaper JB, Nataro JP, Mobley HL (2004) Pathogenic Escherichia coli. Nat Rev Microbiol 2:123–140. https://doi.org/10.1038/nrmicro818
Kennedy CA, Walsh C, Karczmarczyk M et al (2018) Multi-drug resistant Escherichia coli in diarrhoeagenic foals: Pulsotyping, phylotyping, serotyping, antibiotic resistance and virulence profiling. Vet Microbiol 223:144–152. https://doi.org/10.1016/j.vetmic.2018.08.009
Melo AR, Lasunskaia EB, de Almeida CM et al (2005) Expression of the virulence factor, bfpA, by enteropathogenic Escherichia coli is essential for apoptosis signalling but not for NF-kappaB activation in host cells. Scand J Immunol 61:511–519. https://doi.org/10.1111/j.1365-3083.2005.01626.x
Melton-Celsa AR (2014) Shiga toxin (Stx) classification, structure, and function. Microbiol Spectr 2:1–13. https://doi.org/10.1128/microbiolspec.EHEC-0024-2013
Moxley R (2013) Family Enterobacteriaceae and Enterobacteriaceae: Escherichia. In: McVey DS, Kennedy M, Chengappa MM (eds) Veterinary Microbiology. Wiley-Backwell, New Jersey, pp 53–74
Mustak HK, Torun E, Kaya IB et al (2013) Wistar ırkı ratlardan izole edilen Escherichia coli’lerin filogenetik analizi. Etlik Vet Mikrobiyol Derg 24:60–63
Oh JY, Kang MS, An BK et al (2012) Prevalence and characteristics of intimin-producing Escherichia coli strains isolated from healthy chickens in Korea. Poult Sci 91:2438–2443. https://doi.org/10.3382/ps.2012-02301
Omerovic M, Mustak HK (2018) Kanatlı Kökenli Escherichia coli’lerin Filogruplandırılması. Etlik Vet Mikrobiyol Derg 29:57–62. https://doi.org/10.35864/evmd.513016
Omerovic M, Mustak HK, Kaya İB (2017) Escherichia coli Patotiplerinin Virülens Faktörleri. Etlik Vet Mikrobiyol Derg 28:1–6. https://doi.org/10.35864/evmd.530084
Parte AC, Sardà Carbasse J, Meier-Kolthoff JP et al (2020) List of Prokaryotic names with Standing in Nomenclature (LPSN) moves to the DSMZ. Int J Syst Evol Microbiol 70:5607–5612. https://doi.org/10.1099/ijsem.0.004332
Pehlivanoglu F, Ozturk D, Turutoglu H (2020) Carriage of plasmidic AmpC beta-lactamase producing Escherichia coli in cattle and sheep and characterisation of the isolates in terms of antibiogram profiles, phylogeny and virulence. Kafkas Univ Vet Fak Derg 26:469–476. https://doi.org/10.9775/kvfd.2019.23541
Saei HD, Zavarshani M (2018) Phylogenetic grouping of verotoxigenic Escherichia coli (VTEC) Obtained from sheep and broiler chicken in Northwestern Iran. Acta Vet Eurasia 44:53–58. https://doi.org/10.26650/actavet.2018.010
Sekse C, Sunde M, Hopp P et al (2013) Occurrence of potentially human-pathogenic Escherichia coli O103 in Norwegian sheep. Appl Environ Microbiol 79:7502–7509. https://doi.org/10.1128/AEM.01825-13
Souto MS, Coura FM, Dorneles EM et al (2017) Antimicrobial susceptibility and phylotyping profile of pathogenic Escherichia coli and Salmonella enterica isolates from calves and pigs in Minas Gerais Brazil. Trop Anim Health Prod 49:13–23. https://doi.org/10.1007/s11250-016-1152-0
Stoppe NC, Silva JS, Carlos C et al (2017) Worldwide phylogenetic group patterns of Escherichia coli from commensal human and wastewater treatment plant isolates. Front Microbiol 8:2512. https://doi.org/10.3389/fmicb.2017.02512
Torkan S, Bahadoranian MA, Khamesipour F, Anyanwud MU (2016) Detection of virulence and antimicrobial resistance genes in Escherichia coli isolates from diarrhoiec dogs in Iran. Arch Med Vet 48:181–190. https://doi.org/10.4067/S0301-732X2016000200008
Tostes R, Goji N, Amoako K et al (2017) Subtyping Escherichia coli virulence genes isolated from feces of beef cattle and clinical cases in Alberta. Foodborne Pathog Dis 14:35–42. https://doi.org/10.1089/fpd.2016.2199
Wang LYR, Jokinen CC, Laing CR et al (2018) Multi-year persistence of verotoxigenic Escherichia coli (VTEC) in a closed Canadian beef herd: a cohort study. Front Microbiol 9:2040. https://doi.org/10.3389/fmicb.2018.02040
Waters NR, Abram F, Brennan F et al (2020) Easy phylotyping of Escherichia coli via the EzClermont web app and command-line tool. Access Microbiol 2:acmi000143. https://doi.org/10.1099/acmi.0.000143
Watson VE, Jacob ME, Flower JR et al (2017) Association of Atypical Enteropathogenic Escherichia coli with Diarrhea and Related Mortality in Kittens. J Clin Microbiol 55:2719–2735. https://doi.org/10.1128/JCM.00403-17
Weintraub A (2007) Enteroaggregative Escherichia coli: epidemiology, virulance and detection. J Med Microbiol 56:4–8. https://doi.org/10.1099/jmm.0.46930-0
Acknowledgements
The study was produced from a doctoral-PhD thesis performed by Emre KARAKAYA. Also, we would like to thank Dr. Elif CELIK for carrying out statistical analyses of our findings.
Funding
This research received no grant from any funding agency/sector.
Author information
Authors and Affiliations
Contributions
EK and FA conceived and designed the study; EK, FA, and SA contributed to the execution of analyses; EK, FA, and TK contributed to molecular analyses; EK and SA contributed to manuscript writing, drawing of figures, and editing. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interests
The authors declared that there is no conflict of interest.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Karakaya, E., Aydin, F., Kayman, T. et al. Escherichia coli in different animal feces: phylotypes and virulence genes. World J Microbiol Biotechnol 39, 14 (2023). https://doi.org/10.1007/s11274-022-03451-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11274-022-03451-w