Abstract
Lactate are proved to be attractive electron donor for the production of n-caproic acid (CA) that is a high value-added fuel precursor and chemical feedstock, but little is known about molecular mechanism of lactate transformation. In the present study, the gene for L-lactate dehydrogenase (LDH, EC.1.1.1.27) from a Ruminococcaceae strain CPB6 was cloned and expressed in Escherichia coli BL21 (DE3) with plasmid pET28a. The recombinant LDH exhibited molecular weight of 36–38 kDa in SDS-PAGE. The purified LDH was found to have the maximal oxidation activity of 29.6 U/mg from lactate to pyruvate at pH 6.5, and the maximal reduction activity of 10.4 U/mg from pyruvate to lactate at pH 8.5, respectively. Strikingly, its oxidative activity predominates over reductive activity, leading to a 17-fold increase for the utilization of lactate in E. coli/pET28a-LDH than E. coli/pET28a. The CPB6 LDH gene encodes a 315 amino acid protein sharing 42.19% similarity with Clostridium beijerinckii LDH, and lower similarity with LDHs of other organisms. Significant difference were observed between the CPB6 LDH and C. beijerinckii and C. acetobutylicum LDH in the predicted tertiary structure and active center. Further, X-ray crystal structure analysis need to be performed to verify the specific active center of the CPB6 LDH and its role in the conversion of lactate into CA.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Medium chain fatty acids (MCFAs) are gaining increasing attention due to their broad use. MCFAs can be synthesized by chain-elongating microbes from various substrates (Wu et al. 2019), such as ethanol (Agler et al. 2012), methanol (Chen et al. 2016), and D-galactitol (Andersen et al. 2017). Recently, researchers have also described the production of MCFAs (e.g., n-caproic acid) from lactate-containing food wastewater in reactor microbiome systems (Kucek et al. 2016; Zhu et al. 2015), and lactate is regarded as an attractive alternative to ethanol for the production of MCFAs (Zhu et al. 2017).
Ruminococcaceae are generally the producers of short-chain fatty acids (Zhu et al. 2017). As it turns out, Ruminococcaceae play critical roles in degrading a variety of polysaccharides and fibres (Louis and Flint 2009; Scott et al. 2008). Recently, we reported a new Ruminococcaceae bacterium, CPB6, with the capability of producing a high concentration of n-caproic acid (CA) from lactate (Wang et al. 2018; Zhu et al. 2017). More recently, complete genomic sequencing and annotation showed that the Ruminococcaceae species encodes most genes related to lactate utilization and CA production (Hahnke et al. 2018; Tao et al. 2017), and two genes encoding L-lactate dehydrogenase (L-LDH) and D-lactate dehydrogenase (D-LDH) were identified in the CPB6 genome. In the process of lactate conversion into CA, the oxidation of lactate is assumed to provide energy and acetyl-CoA for chain elongation (Cavalcante et al. 2017), but the enzymatic mechanism of lactate conversion remains unknown.
LDH (EC 1.1.1.27) is a key enzyme in glycolysis (Dong et al. 2014; Nadeem et al. 2018), catalysing the mutual conversion of pyruvate and lactic acid in the presence of nicotinamide adenine dinucleotide coenzyme (Özkan et al. 2004; Zhang et al. 2014). The enzyme plays a crucial role in food industries fermenting lactic acid (Collins et al. 1991), wine and yoghurt (Gaspar et al. 2004; Sun et al. 2016). Some studies have described the cloning, expression and characterization of LDH from various organisms (Lu et al. 2006; Singh et al. 2012), such as Lactobacillus (Lee et al. 2004), Selenomonas (Evans and Martin 2002), Pediococcus (Zhou et al. 2003), and Clostridium (Contag et al. 1990; Stevenson and Weimer 2005). These methods are effective for understanding the structure of function of LDH and further improving the yield of targeted compounds catalysed by LDH (Kim et al. 2018; Kouassi et al. 2007; Wang et al. 2019). In order to investigate the enzymatic mechanism of lactate conversion into CA by Ruminococcaceae bacterium CPB6, an L-LDH gene from strain CPB6 was cloned into plasmid pET28a to construct recombinant pET28a-LDH, which was expressed in E. coli BL21. The oxidative and reductive activities of the recombinant LDH protein were determined, and its effect on the utilization of lactic acid by E. coli BL21 was assayed. Finally, bioinformatics analysis of LDH was performed to analyse its structure and function. This study is likely to be valuable for revealing the mechanism of action of lactate conversion into CA and genetically modifying strain CPB6 to further improve CA productivity.
Materials and methods
Bacterial strains and plasmids
Ruminococcaceae strain CPB6 was cultivated as described previously (Zhu et al. 2017). The plasmid pET28a was used as an LDH expression vector. E. coli BL21 (DE3) (Biomed, Beijing, China) was used as a host for gene cloning and expression of the recombinant plasmid. According to the predicted LDH gene from the CPB6 genome (GenBank: CP020705.1) (Tao et al. 2017), the primers YT45 (5′-AGCAAATGGGTCGCGGATCCATGAAAACAAGAAAAATCGGCGTCA-3′) and YT42 (5′-GCGGCCGCAAGCTTGTCGACGCAAAGGTCTTTTAGCTGTTCCATATTTTTACG-3′) (bold underscore letters match with the sequence of pET28a for seamless cloning) were designed to amplify the ORF of the CPB6 LDH protein. PCR was performed in a 50.0 μL system, which contained DNA template 2.5 μL, primers 2.5 μL, DMSO 1.5 μL, high fidelity Taq enzyme mix 25.0 μL (Thermo, Waltham, USA), and the required amount of deionized water. The reaction was initially heated at 98 °C for 2 min, followed by 30 cycles of amplification. Each cycle consisted of incubation at 98 °C for 10 s, 52 °C for 30 s, and 72 °C for 30 s. Finally, the reaction was incubated at 72 °C for 5 min for the final extension. The PCR products were verified by agarose electrophoresis and recovered using the PCR purification kit (Fuji, Chengdu, China).
Plasmid construction and gibson assembly
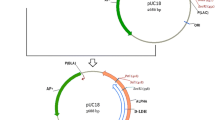
The plasmid pET28a was used as the parent vector for the construction of the recombinant plasmid. The PCR product was seamlessly inserted into plasmid pET28a double-digested with Sall I and BamH I (Thermo, Waltham, USA) to construct the recombinant plasmid pET28a-LDH, which was transformed into E. coli BL21. The positive clones with the recombinant plasmid pET28a-LDH were preliminarily screened by Kana-resistance (50 µg/mL) LB plates. The recombinant plasmid was extracted from the culture of the positive clone using the plasmid extraction kit (Tiangen, Beijing, China) and was verified by enzyme digestion analysis and sequencing.
Prokaryotic expression and purification
The coding region of the recombinant plasmid pET28a-LDH was positioned in frame with the 6 × His-tag framework, and the plasmid was transformed into E. coli BL21 (DE3); the resulting strain was cultured in LB medium with Kana (50 µg/mL) overnight at 37 °C with shaking at 200 rpm. When the OD600 of the culture reached 0.4–0.6, isopropyl-β-D-thiogalactopyranoside (IPTG) was added to a final concentration of 0.5 mM to induce the expression of the recombinant His-tagged LDH fusion protein. After induction at 18 °C for 12 h, cells were collected by centrifugation at 12,000 rpm for 15 min at 4 °C. The enzyme was purified with Ni-IDA Resin (Genscript, Nanjing, China) and eluted with 50 mM sodium phosphate (pH 8.0) containing 300 mM NaCl, 200 mM imidazole. The samples were analysed by SDS-PAGE with 5% stacking and 10% separating gels after the samples were boiled in water at 100 °C for 5 min. Protein bands were visualized by Coomassie Brilliant Blue G-250 (Kelong, Chengdu, China). The protein concentrations were measured by the BCA protein assay kit (Solarbio, Beijing, China).
In addition, the purified recombinant LDH protein was verified by Western blot analysis with anti-His monoclonal antibody. The purified enzyme was analyzed by SDS-PAGE, followed by transferring of the protein onto a polyvinylidene difluoride (PVDF) membrane (Bio-Rad, California, USA). The membrane was cut into vertical strips prior to incubation with blocking solution of 5% nonfat dry milk in Tris-buffered saline for 1 h at room temperature with constant shaking and then probed with diluted Anti-His mouse monoclonal antibody (1:5000) for 2 h. The membrane strips were washed three times with TBST and treated for an additional 1 h with biotinylated goat antihuman IgM/IgG (1:2500), the secondary antibody. After additional three washes with TBST, the membranes were stained with Immobilon TM Western Chemiluminescent HRP Substrate (Millipore) and detected using an Image Quant LAS4000 system (GE Healthcare).
Enzyme assays
LDH activity was assayed spectrophotometrically by measuring the first-order change in absorbance at 340 nm as a result of oxidation of NADH or reduction of NAD + , following the method in the literature(Lu et al. 2006). For preparation of crude enzyme solution, first, cell pellets were collected after centrifugation at 10,000 rpm and 4 °C for 10 min. Then, the cells were suspended at a 1000:1 ratio of bacterial population (104) to the extracting solution (mL), and the cells were lysed by sonication in an ice bath with a power of 200 W every 4 s with an interval of 6 s repeated 100 times. Finally, the cells were centrifuged for 10 min at 8000 g and 4 °C, and the supernatant was collected and stored on ice. The reduction reaction of pyruvate to lactate was determined in a reaction mixture containing 100 mmol/L sodium phosphate buffer (pH 5.0–10.0), 80 mmol/L pyruvate and 4 mmol/L NADH. For the oxidation of lactate to pyruvate, the activity was measured in a reaction mixture containing 100 mmol/L sodium phosphate buffer (pH 5.0–10.0), 50 mmol/L sodium lactate and 1 mmol/L NAD + . All reactions were started after adding the crude enzyme or the purified enzyme solution at 25 °C. Every 0.5 min for a total of 3–5 min after the beginning of the reaction, the change in absorbance value was measured by using a TU-1810 UV/visible (UV–Vis) spectrophotometer (Puxi, Beijing, China). Each data point represents the mean value of three biological duplicates. One unit of activity was defined as 1 μmol of product (NAD +) formed from the reduction of pyruvate to lactate and 1 μmol of product (NADH) formed from the oxidation of lactate to pyruvate per minute per milligram of protein. The activity of LDH (U/mg of protein) = (A340nm/min × dilution factors)/(the amount of enzyme solution added (10 µL) × 10–1 × the protein concentration). All assays were performed in triplicate.
Utilization of lactic acid by E. coli
To verify the effect of the LDH gene on the utilization of lactic acid in living E. coli, E. coli/pET28a-LDH and E. coli/pET28a were inoculated into LB medium with 10 g/L lactic acid and incubated at 37 °C with shaking at 200 rpm. IPTG (0.5 mM final concentration) was added when cells reached the logarithmic phase (OD600nm 0.4–0.6), and all cells were incubated for a total of 72 h. Residual lactic acid in the medium was determined by HPLC (Agilent 1260 Infinity, California, USA) equipped with a differential refraction detector (RID) and a HiPlex H column (300 × 6.5 mm) (Wang et al. 2018). For HPLC sample preparation, 1 mL of the sample was centrifuged at 12,000 rpm for 3 min; the supernatant was diluted 20 times with distilled water and then filtered through a 0.22 μm filter (Wandao, Chengdu, China). The resulting filtrate was injected into the HPLC for lactic acid analysis. Culture growth was measured by following the optical density (OD) in the fermentation broth at 600 nm using a TU-1810 UV/Vis Spectrophotometer (Puxi, Beijing, China).
Bioinformatics analysis
Sequence alignment was performed using ESPript (Cuff and Barton 2000). Representative LDH sequences, including those from Clostridium, Ruminococcaceae and Lactobacillus, were retrieved from the NCBI database. MEGA-X software was used to perform sequence alignment and construct a phylogenetic tree (Tamura et al. 2011). The active sites of LDH were predicted by ScanProsite, and the three-dimensional LDH structure was simulated by NCBI-CDD to search for templates in SWISS-MODEL and was embellished and labelled by PyMOL 2.3.3 software (Arnold et al. 2006; Marchler-Bauer et al. 2009).
Results and discussion
Cloning and plasmid construction
According to the CPB6 genome sequence, primers targeting the CPB6 LDH gene were designed and synthesized. DNA from strain CPB6 was prepared with a DNA extraction kit (Tiangen, Beijing, China) and used as a template for PCR amplification. Agarose gel electrophoresis showed that the size of the PCR product was approximately 1000 bp (Fig. S1A), consistent with the expected size of the CPB6 LDH gene (948 bp). The recombinant plasmid, which was confirmed by agarose electrophoresis of the double digestion product (Fig. S1B), was sequenced by the Sanger method. Sequence analysis demonstrated that the cloned gene had 100% similarity with the predicted LDH gene of strain CPB6 (Fig. S2), indicating that the recombinant E. coli/pET28a-LDH was successfully constructed.
Expression and purification of the recombinant protein
E. coli/pET28a-LDH was incubated in LB medium until OD600nm 0.4–0.6, and the culture was induced at 18 °C for 12 h by adding 0.5 mM IPTG. A series of controls were also performed simultaneously (Fig. S3). SDS-PAGE analysis showed that after induction with IPTG, a new significant protein band (approximately 36–38 kDa) appeared compared to the controls (Fig. S4). The expressed protein band was roughly the same size as the expected CPB6 LDH band (36 kDa), indicating that the CPB6 LDH gene was expressed successfully in the E. coli host. After affinity chromatography purification, the purified enzyme showed a single band after SDS-PAGE, corresponding to a molecular weight of 36 kDa (Fig. 1), which agreed well with the predicted LDH molecular masses. Moreover, Western blot analysis also confirmed the successful expression of the purified recombinant protein (Fig. 1b). The purified LDH was added 50% glycerin and stored at − 80 °C and used when needed.
SDS-PAGE and Western blot analysis. a Polyacrylamide gel electrophoresis (SDS-PAGE); b Western blot probed with the purified recombinant protein. M, protein Marker; Lane 1, crude cell extract of E. coli BL21/pET28a by IPTG induction; Lane 2, crude cell extract of E. coli BL21/pET28a-LDH without IPTG induction; Lane 3, crude cell extract of E. coli BL21/pET28a-LDH by IPTG induction; Lane 4, the purified E. coli BL21/pET28a-LDH by affinity chromatography eluate
Enzyme assay
The enzymatic assay for the crude enzyme showed that the recombinant protein has a maximal oxidation activity of 5.40 U/mg from lactate to pyruvate at pH 6.5 and a maximal reduction activity of 1.15 U/mg from pyruvate to lactate at pH 8.5 (Fig. 2). This result is consistent with previous studies that LDH has oxidation activity under acidic conditions and reduction activity under alkaline conditions (Sharkey et al. 2011; Skory 2000). As shown in Table 1, the LDH reduction activity of E. coli/pET28a-LDH was remarkably higher than that of the control E. coli/pET28a (1.76 vs 0.61 U/mg of protein). Likewise, the oxidative activity of E. coli/pET28a-LDH was markedly higher than that of the control (6.21 vs 0.81 U/mg of protein). After purification, the maximum oxidative and reductive activities of LDH were 29.6 U/mg and 10.4 U/mg, respectively, further indicating that the oxidative activity of the enzyme predominates over the reductive activity. Thus, it is estimated that the direction of the main metabolic flux in E. coli/pET28a-LDH is from lactate to pyruvate (Fig. S5). Compared with other literatures, such as the LDH of Clostridium acetobutylicum (Contag et al. 1990), which was reported to only have reductive activity from pyruvate to lactate. The LDH of Rhizopus oryzae (Skory 2000) has higher reductive activity (2.2 U/mg of protein) related to oxidative activity (0.18 U/mg of protein). Likewise, similar properties have been observed in LDHs of Clostridium thermocellum (Özkan et al. 2004) and Corynebacterium glutamicum (Sharkey et al. 2011). Up to now, there are few microbes is reported to have so high LDH oxidative activity (29.6 U/mg) like the CPB6 strain. In recent years, the production mid-chain fatty acids from lactic acid is becoming an interesting study (Tao et al. 2016; Wang et al. 2018). The oxidation of lactate to pyruvate is the first step for the conversion of lactate to mid-chain fatty acids, and thus the LDH with high oxidation activity needs to focus on.
Changes in the activity of the recombinant LDH (the crude enzyme) under different pH values. The left y-axis shows the specific activity, which is the measure of E. coli/pET28a-LDH minus E. coli/pET28a. The x-axis denotes pH values. The buffer for adjusting the pH value is 1 M HCl or 1 M NaOH solution. Meanwhile, solid dot curve shows the oxidizing activity of LDH, and solid diamond curve shows the reducing activity of LDH. Unstable activity is not shown in the figure. Both sets of reactions were carried out in a mixed buffer at 25 °C as described in the “Materials and methods” section. All data are presented as the means ± standard deviations (n = 3)
Cell growth and utilization of lactic acid by E. coli
As illustrated in Fig. 3, both E. coli/pET28a-LDH and E. coli/pET28a displayed similar growth rates and growth kinetics. All cells grew rapidly within 12 h and then gradually entered the stationary phase (Fig. 3a). However, E. coli/pET28a-LDH consumed 3.606 g/L lactic acid after induction for 24 h, while E. coli/pET28a consumed only 0.2 g/L lactic acid (Fig. 3b). The consumption of lactic acid by E. coli/pET28a-LDH was increased by 17-fold compared with E. coli/pET28a, indicating that CPB6 LDH improves the utilization of lactic acid by E. coli under weak acid or neutral conditions. This supports the estimation that expression of the CPB6 LDH gene leads to the direction of the metabolic flux from lactate to pyruvate in E. coli.
Curve of cell growth and lactic acid utilization by E. coli BL21/pET28a-LDH and E. coli BL21/pET28a. The x-axis represents culture time (h), and the time starts after IPTG (0.5 mM) addition. a The left y‑axis shows the absorbance value of bacterial solution at 600 nm, and the initial absorbance value was measured when cells reached the logarithmic phase and after IPTG (0.5 mM) was added. b The left y-axis represents the utilization of lactic acid (g/L) by E. coli BL21/pET28a-LDH and E. coli BL21/pET28a. The error bars indicate standard deviations of measurements from three independent cultures
In general, LDH catalyses not only the reduction of pyruvate to lactate but also the conversion of lactate to pyruvate (Skory 2000). To date, there are many reports on the exogenous cloning and expression of LDH from different organisms, but most of the reports focus on the reductive activity of LDH and the high yield of lactic acid by expressing LDH in E. coli (Fu et al. 2018; Li 2008; Singhvi et al. 2017). Here, the CPB6 LDH protein showed higher oxidative activity than reductive activity, indicating that the enzyme is different from other LDHs. The crude LDH exhibited the maximum oxidative activity at pH 6.5, consistent with our previous study, which showed that strain CPB6 efficiently produced CA from lactate at pH 6.5 and achieved the maximum yield of CA (Wang et al. 2018). Additionally, the kinetics of lactic acid utilization were increased by 17-fold in E. coli/pET28a-LDH compared with E. coli/pET28a, demonstrating that the expression of LDH significantly improved the utilization of lactic acid by E. coli. However, the utilization of lactic acid by E. coli/pET28a-LDH was lower than that by CPB6. This was probably because the recombinant LDH protein contains a small fusion tag, which may lead to lower activity of the fusion LDH protein (Gokarn et al. 2000; Tebianian et al. 2011).
Multiple sequence alignments of the LDH
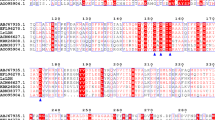
Amino acid sequence similarity is an important parameter reflecting gene evolutionary status (Hannenhalli and Russell 2000). LDH is widely distributed in a variety of organisms, including Lactobacillus, Clostridium and Pediococcus, and its biochemical properties have been characterized (Dietrich et al. 2009; Li 2008; Ozkan et al. 2009; Zhou and Shao 2010). In order to further analyse the characteristics of the CPB6 LDH protein, thirteen LDH sequences of other organisms were retrieved from the National Center for Biotechnology Information (https://www.ncbi.nlm.nih.gov), which are the same species as CPB6 and the LDH function are similar or opposite. Multiple alignment of amino acid sequences showed that the CPB6 LDH protein had low similarity with LDHs from other organisms (Fig. 4), such as C. beijerinckii (42.19%), C. difficile (35.53%), C. acetobutylicum (26.33%), Ruminococcus flavefaciens (17.45%), and L. acidophilus (14.24%). The phylogenetic analysis showed that the CPB6 LDH protein was clustered with the C. beijerinckii and C. difficile LDH proteins (Fig. S6), implying close phylogenetic relationships between them.
Multiple amino acid sequence alignments of LDHs from different strains. The amino acid residues in the red shaded boxes are the same, with 100% of the residues identical and no gaps. The consistency of amino acid residues in the blue box is more than 75%, and the red characters indicate that the amino acid residues belong to the same class and have similar structures and functions. Although some amino acids are different, the overall protein effect may not be affected
Prediction of active sites and 3-dimensional (3D) structure
The analysis of the active site showed that most LDH active sites (except LDH of R. flavefaciens) consist of seven amino acids, located in the range of amino acids 171 to 186. Among them, there are four highly conserved amino acids (“GEHG”) (Table 2). This is the site of the enzyme that the substrate binds and that performs the catalytic activity. However, the active site of the CPB6 LDH protein could not be predicted using ScanProsite software, demonstrating the difference in the sequence and structure of the CPB6 LDH protein compared with other LDH proteins. Combining the result of the multiple sequence alignments (Fig. 4), the active site of the CPB6 LDH located in the range of amino acids 175–181, which is “MGEHGVK” (Table 2). It can be seen that its position showed a shift in the CPB6 LDH.
Both C. beijerinckii and C. acetobutylicum are solventogenic and butyrate-producing bacteria (Wang et al. 2013; Xiao et al. 2011); these bacteria have significant features of genomic organization and unique adaptation strategies (Toth et al. 1999; Nolling et al. 2001). The LDH gene of C. acetobutylicum was cloned and expressed in E. coli; the resulting strain had improved growth and could produce lactic acid under anaerobic conditions, and the crude cell extract of this strain had higher LDH activity, which was demonstrated by the reduction of pyruvate to lactic acid (Contag et al. 1990). In this study, we compared the 3D structures and the active centres of the CPB6 LDH protein and C. acetobutylicum LDH protein (Fig. 5). Although the active site of the CPB6 LDH protein could not be predicted with ScanProsite software, we preliminarily identified its possible position and amino acid sequence according to the sequence alignment of LDHs from various organisms, including strain CPB6 (Fig. 4). The 3D structure of the CPB6 LDH protein was similar to that of the C. acetobutylicum LDH protein, but there was a difference between them in the location and structure of the active centre (Fig. 5). Obviously, the active centre of the CPB6 LDH protein, located between amino acids 175 and 181 (MGEHGVK), formed a circular structure that may be the binding site of the enzyme and substrate. However, the amino acid sequence from 173 aa-179 aa (MGEHGDS) of the C. acetobutylicum LDH protein did not form a loop in the active centre. Additionally, the active centre of the CPB6 LDH protein was different from that of the C. beijerinckii LDH protein, whose active centre was located between amino acids 175 and 181 (MGEHGDS). There were significant differences between the two proteins in the structures of the active centres (Fig. S7).
The 3-dimensional (3D) structures and active centres of LDHs from Ruminococcaceae CPB6 and C. acetobutylicum. The 3D structures of the LDHs are on the left. The structures of the active centre are on the right, and the red represents one hydrogen, in which one amino acid is attached to each two hydrogens. a The 3D structure and active centre of LDH from Ruminococcaceae CPB6 (CP020705.1), and its active site is from amino acids 175–181 (MGEHGVK). b The 3D structure and active centre of LDH from C. acetobutylicum (WP 010963590.1), and its active site is from amino acids 173–179 (MGEHGDS)
Strong interactions between adjacent domains of tetramers are considered to be the key to subunit interactions, and the stability of the amino acid spatial structure is also the key to achieving high catalytic efficiency (Zoraghi et al. 2010). The structure of the CPB6 LDH protein is similar to that of the C. acetobutylicum LDH protein, but there are great distinctions between the amino acid sequences and 3D structures of their active centres, which may affect their catalytic activity and binding of substrates. Perhaps this particular structure is responsible for the higher oxidation activity of the CPB6 LDH protein compared to its reduction activity. However, the exact active centre and function of the CPB6 LDH protein at the molecular level certainly remains to be determined by X-ray crystal structure analysis.
Concluding remarks
The gene encoding L-lactate dehydrogenase (LDH; EC 1.1.1.27) from strain CPB6 was successfully cloned and expressed in E. coli BL21 (DE3) with the plasmid pET28a. The purified enzyme of the CPB6 LDH had higher oxidative activity relative to its reductive activity, implying that it catalyses the conversion of lactate to pyruvate with the formation of energy and acetyl-CoA. This explains the increased utilization of lactic acid by recombinant E. coli. Additionally, the CPB6 LDH protein had low amino acid sequence similarity with other LDH proteins. This explains why the active centre and 3D structure of the CPB6 LDH protein were different from those of other LDH proteins. These results will help us deeply understand the molecular mechanism of lactate conversion into CA.
References
Agler MT, Spirito CM, Usack JG, Werner JJ, Angenent LT (2012) Chain elongation with reactor microbiomes: upgrading dilute ethanol to medium-chain carboxylates. Energy Environ Sci 5:8189–8192
Andersen SJ, De Groof V, Khor WC, Roume H, Props R, Coma M, Rabaey K (2017) A Clostridium Group IV species dominates and suppresses a mixed culture fermentation by tolerance to medium chain fatty acids products. Front Bioeng Biotechnol 5:1–10
Arnold K, Bordoli L, Kopp J, Schwede T (2006) The SWISS-MODEL workspace: a web-based environment for protein structure homology modelling. Bioinformatics 22:195–201
Cavalcante WDA, Leitão RC, Gehring TA, Angenent LT, Santaella ST (2017) Anaerobic fermentation for n-caproic acid production: a review. Process Biochem 54:106–119
Chen WS, Ye Y, Steinbusch KJJ, Strik DPBTB, Buisman CJN (2016) Methanol as an alternative electron donor in chain elongation for butyrate and caproate formation. Biomass Bioenerg 93:201–208
Collins SS, Keeton JT, Smith SB (1991) Lactate dehydrogenase enzyme activity in raw, cured, and heated porcine muscle. J Agric Food Chem 39:1294–1297
Contag PR, Williams MG, Rogers P (1990) Cloning of a lactate dehydrogenase gene from Clostridium acetobutylicum B643 and expression in Escherichia coli. Appl Environ Microbiol 56:3760–3765
Cuff JA, Barton GJ (2000) Application of multiple sequence alignment profiles to improve protein secondary structure prediction. Proteins 40:502–511
Dietrich C, Nato A, Bost B, Le Marechal P, Guyonvarch A (2009) Regulation of ldh expression during biotin-limited growth of Corynebacterium glutamicum. Microbiology 155:1360–1375
Dong H, Wang Y, Zhao Q, Han H, Zhu S, Li L, Wu Y, Huang B (2014) Molecular cloning and characterization of lactate dehydrogenase gene from Eimeria tenella. Parasitol Res 113:2915–2923
Evans JD, Martin SA (2002) Cloning of the L-lactate dehydrogenase gene from the ruminal bacterium Selenomonasruminantium HD4. Curr Microbiol 44:155–160
Fu Y, Sun X, Zhu H, Jiang R, Luo X, Yin L (2018) An optimized fed-batch culture strategy integrated with a one-step fermentation improves L-lactic acid production by Rhizopusoryzae. World J Microbiol Biotechnol 34:74–82
Gaspar P, Neves AR, Ramos A, Gasson MJ, Shearman CA, Santos H (2004) Engineering Lactococcuslactis for production of mannitol: high yields from food-grade strains deficient in lactate dehydrogenase and the mannitol transport system. Appl Environ Microbiol 70:1466–1474
Gokarn RR, Eiteman MA, Altman E (2000) Metabolic analysis of Escherichiacoli in the presence and absence of the carboxylating enzymes phosphoenolpyruvate carboxylase and pyruvate carboxylase. Appl Environ Microbiol 66:1844–1850
Hahnke S, Abendroth C, Langer T, Codoner FM, Ramm P, Porcar M, Luschnig O, Klocke M (2018) Complete genome sequence of a new ruminococcaceae bacterium isolated from anaerobic biomass hydrolysis. Genome Announc 6:1–2
Hannenhalli SS, Russell RB (2000) Analysis and prediction of functional sub-types from protein sequence alignments. J Mol Biol 303:61–76
Kim SG, Jang S, Lim JH, Jeon BS, Kim J, Kim KH, Sang BI, Jung GY (2018) Optimization of hexanoic acid production in recombinant Escherichiacoli by precise flux rebalancing. Bioresour Technol 247:1253–1257
Kouassi GK, Anantheswaran RC, Knabel SJ, Floros JD (2007) Effect of high-pressure processing on activity and structure of alkaline phosphatase and lactate dehydrogenase in buffer and milk. J Agric Food Chem 55:9520–9529
Kucek LA, Nguyen M, Angenent LT (2016) Conversion of L-lactate into n-caproate by a continuously fed reactor microbiome. Water Res 93:163–171
Lee JH, Choi MH, Park JY, Kang HK, Ryu HW, Sunwo CS, Wee YJ, Park KD, Kim DW, Kim D (2004) Cloning and characterization of the lactate dehydrogenase genes from Lactobacillus sp. RKY2. Biotechnol Bioprocess Eng 9:318–322
Li X (2008) Purification and partial characterization of Lactobacillus species SK007 lactate dehydrogenase (LDH) catalyzing phenylpyruvic acid (PPA) conversion into phenyllactic acid (PLA). J Agric Food Chem 56:2392–2399
Louis P, Flint HJ (2009) Diversity, metabolism and microbial ecology of butyrate-producing bacteria from the human large intestine. FEMS Microbiol Lett 294:1–8
Lu G, Hu X, Peng Z, Xie H, Li Y, Wu Z, Yu X (2006) Expression and characterization of lactate dehydrogenase from Schistosomajaponicum. Parasitol Res 99:593–596
Marchler-Bauer A, Anderson JB, Chitsaz F, Derbyshire MK, DeWeese-Scott C, Fong JH, Geer LY, Geer RC, Gonzales NR, Gwadz M, He S, Hurwitz DI, Jackson JD, Ke Z, Lanczycki CJ, Liebert CA, Liu C, Lu F, Lu S, Marchler GH, Mullokandov M, Song JS, Tasneem A, Thanki N, Yamashita RA, Zhang D, Zhang N, Bryant SH (2009) CDD: specific functional annotation with the Conserved Domain Database. Nucleic Acids Res 37:D205-210
Nadeem MS, Al-Ghamdi MA, Khan JA, Sadath S, Al-Malki A (2018) Recombinant production and biochemical and in silico characterization of lactate dehydrogenase from Geobacillus thermodenitrificans DSM-465. Electron J Biotechnol 35:18–24
Nolling J, Breton G, Omelchenko MV, Makarova KS, Zeng Q, Gibson R, Lee HM, Dubois J, Qiu D, Hitti J, Wolf YI, Tatusov RL, Sabathe F, Doucette-Stamm L, Soucaille P, Daly MJ, Bennett GN, Koonin EV, Smith DR (2001) Genome sequence and comparative analysis of the solvent-producing bacterium Clostridiumacetobutylicum. J Bacteriol 183:4823–4838
Özkan M, Yllmaz EI, Lynd LR, Özcengiz G (2004) Cloning and expression of the Clostridiumthermocellum L-lactate dehydrogenase gene in Escherichiacoli and enzyme characterization1. Can J Microbiol 50:845–851
Ozkan M, Erhan E, Terzi O, Tan I, Ozoner SK (2009) Thermostable amperometric lactate biosensor with Clostridiumthermocellum L-LDH for the measurement of blood lactate. Talanta 79:1412–1417
Scott KP, Martin JC, Mrazek J, Flint HJ (2008) Transfer of conjugative elements from rumen and human Firmicutes bacteria to Roseburiainulinivorans. Appl Environ Microbiol 74:3915–3917
Sharkey MA, Maher MA, Guyonvarch A, Engel PC (2011) Kinetic characterisation of recombinant Corynebacterium glutamicum NAD+-dependent LDH over-expressed in E. coli and its rescue of an lldD- phenotype in C. glutamicum: the issue of reversibility re-examined. Arch Microbiol 193:731–740
Singh V, Kaushal DC, Rathaur S, Kumar N, Kaushal NA (2012) Cloning, overexpression, purification and characterization of Plasmodium knowlesi lactate dehydrogenase. Protein Expr Purif 84:195–203
Singhvi M, Zendo T, Iida H, Gokhale D, Sonomoto K (2017) Stimulation of d- and l-lactate dehydrogenases transcriptional levels in presence of diammonium hydrogen phosphate resulting to enhanced lactic acid production by Lactobacillus strain. J Biosci Bioeng 124:674–679
Skory CD (2000) Isolation and expression of lactate dehydrogenase genes from Rhizopusoryzae. Appl Environ Microbiol 66:2343–2348
Stevenson DM, Weimer PJ (2005) Expression of 17 genes in Clostridiumthermocellum ATCC 27405 during fermentation of cellulose or cellobiose in continuous culture. Appl Environ Microbiol 71:4672–4678
Sun L, Zhang C, Lyu P, Wang Y, Wang L, Yu B (2016) Contributory roles of two l-lactate dehydrogenases for l-lactic acid production in thermotolerant Bacilluscoagulans. Sci Rep 6:37916–37923
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731–2739
Tao Y, Hu X, Zhu X, Jin H, Xu Z, Tang Q, Li X (2016) Production of butyrate from lactate by a newly isolated Clostridium sp. BPY5. Appl Biochem Biotechnol 179:361–374
Tao Y, Zhu X, Wang H, Wang Y, Li X, Jin H, Ru J (2017) Complete genome sequence of Ruminococcaceae bacterium CPB6: a newly isolated culture for efficient n-caproic acid production from lactate. J Biotechnol 259:91–94
Tebianian M, Hoseini AZ, Ebrahimi SM, Memarnejadian A, Mokarram AR, Mahdavi M, Sohrabi N, Taghizadeh M (2011) Cloning, expression, and immunogenicity of novel fusion protein of Mycobacterium tuberculosis based on ESAT-6 and truncated C-terminal fragment of HSP70. Biologicals 39:143–148
Toth J, Ismaiel AA, Chen JS (1999) The ald gene, encoding a coenzyme A-acylating aldehyde dehydrogenase, distinguishes Clostridiumbeijerinckii and two other solvent-producing Clostridia from Clostridiumacetobutylicum. Appl Environ Microbiol 65:4973–4980
Wang Y, Li X, Milne CB, Janssen H, Lin W, Phan G, Huiying Hu, Jin Y-S (2013) Development of a gene knockout system using mobile Group II Introns (Targetron) and genetic disruption of acid production pathways in Clostridiumbeijerinckii. Appl Environ Microbiol 79:5853–5863
Wang H, Li X, Wang Y, Tao Y, Lu S, Zhu X, Li D (2018) Improvement of n-caproic acid production with Ruminococcaceae bacterium CPB6: selection of electron acceptors and carbon sources and optimization of the culture medium. Microb Cell Fact 17:99–108
Wang P, Zhang J, Feng J, Wang S, Guo L, Wang Y, Lee YY, Taylor S, McDonald T, Wang Y (2019) Enhancement of acid re-assimilation and biosolvent production in Clostridiumsaccharoperbutylacetonicum through metabolic engineering for efficient biofuel production from lignocellulosic biomass. Bioresour Technol 281:217–225
Wu Q, Bao X, Guo W, Wang B, Li Y, Luo H, Wang H, Ren N (2019) Medium chain carboxylic acids production from waste biomass: current advances and perspectives. Biotechnol Adv 37:599–615
Xiao Y, Francke C, Abee T, Wells-Bennik MHJ (2011) Clostridial spore germination versus bacilli: genome mining and current insights. Food Microbiol 28:266–274
Zhang QL, Yang M, Zhao YY, Zhang SZ, He QH, Meng XY, Tan WR (2014) Cloning and characterization of lactate dehydrogenase C4 from Ochotonacurzoniae. Mol Biol 48:105–112
Zhou Q, Shao WL (2010) Molecular genetic characterization of the thermostable L-lactate dehydrogenase gene (ldhL) of Thermoanaerobacterethanolicus JW200 and biochemical characterization of the enzyme. Biochemistry (Mosc) 75:526–530
Zhou S, Shanmugam KT, Ingram LO (2003) Functional replacement of the Escherichiacoli D-(-)-lactate dehydrogenase gene (ldhA) with the L-(+)-lactate dehydrogenase gene (ldhL) from Pediococcusacidilactici. Appl Environ Microbiol 69:2237–2244
Zhu X, Tao Y, Liang C, Li X, Wei N, Zhang W, Zhou Y, Yang Y, Bo T (2015) The synthesis of n-caproate from lactate: a new efficient process for medium-chain carboxylates production. Sci Rep 5:14360–14369
Zhu X, Zhou Y, Wang Y, Wu T, Li X, Li D, Tao Y (2017) Production of high-concentration n-caproic acid from lactate through fermentation using a newly isolated Ruminococcaceae bacterium CPB6. Biotechnol Biofuels 10:102–114
Zoraghi R, See RH, Gong H, Lian T, Swayze R, Finlay BB, Brunham RC, McMaster WR, Reiner NE (2010) Functional analysis, overexpression, and kinetic characterization of pyruvate kinase from methicillin-resistant Staphylococcusaureus. Biochemistry 49:7733–7747
Acknowledgements
This work was financially supported by the Natural Science Foundation of China (Grant No. 31770090), the Open-foundation project of CAS Key Laboratory of Environmental and Applied Microbiology (KLCAS-2017-01).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Yang, Q., Wei, C., Guo, S. et al. Cloning and characterization of a l-lactate dehydrogenase gene from Ruminococcaceae bacterium CPB6. World J Microbiol Biotechnol 36, 182 (2020). https://doi.org/10.1007/s11274-020-02958-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11274-020-02958-4