Abstract
Genome shuffling, an efficient and practical strain improvement technology via recursive protoplasts fusion, can break through the limits of species even genus to accelerate the directed evolution of microbial strains, without requiring the comprehensively cognized genetic background and operable genetic system. Hence this technology has been widely used for many important strains to obtain the desirable industrial phenotypes. In this review, we introduce the procedure of genome shuffling, discuss the new aid strategies of genome shuffling, summarize the applications of genome shuffling for increasing metabolite yield, improving strain tolerance, enhancing substrate utilization, and put forward the outlook to the future development of this technology.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
So far, microbial strains have been widely used to produce various valuable products related to agricultural, biofuel, chemical, environmental, food and pharmaceutical industries (Zeng et al. 2020). But most of the originally isolated strains cannot be directly used for industrial production because of low productivity and weak stress tolerance, leading to an increasing interest in strain improvement over the last several decades (Zeng et al. 2020).
The strategies of strain improvement mainly include random mutagenesis, protoplast fusion, genome shuffling (GS) and rational genetic engineering approaches. Random mutagenesis followed by screening, can lead to desired mutants. As random mutagenesis is easy to operate, and not need the genetic background of microbe strains, it has succeeded in breeding many microbial strains, but it is laborious and time-consuming (Gong et al. 2009; Gu et al. 2017). Rational genetic engineering approaches, such as recombinant DNA technology, metabolic engineering, systematic engineering and genome editing, can modify the specific genes of the target strain in a rational manner (Magocha et al. 2018). But these approaches require deep understanding of the genetic background of the target strain and need necessary genetic tools, which have limited the wide application of the rational approaches.
GS, first used for strain improvement in 2002, has been applied for phenotypic improvements of many important strains (Magocha et al. 2018; Zhang et al. 2002). This practical technology has been considered as a novel whole-genome improvement method for the rapid improvement of the complex phenotypes (Gong et al. 2009; Zhang et al. 2002). GS allows for recombination between multiple parents at each generation, and through several rounds of recursive protoplast fusion, the fusants successful fused the genetic traits from multiple parental strains, significantly improving the genetic diversity of “complex progeny” and remarkably increasing the opportunity for obtaining the high-performance fusants (Gong et al. 2009). Compared with random mutagenesis, GS is more effective in obtaining desired phenotypes (Magocha et al. 2018). Compared with rational genetic engineering approaches, GS also offers more advantages. Firstly, GS is more convenient, easy to operate and can be used in most laboratories without expensive equipment (Gong et al. 2009). Furthermore, GS can break through the limits of species even genus to accelerate the directed evolution of important microbes (John et al. 2008; Luna-Flores et al. 2017), and it circumvents the essential requirements of comprehensively genetic background or metabolic network information of the target strain, therefore this technology has a wider range of application than rational approaches (Gong et al. 2009; Zhang et al. 2002). Most importantly, based on natural homologous recombination via protoplast fusion, fusants obtained by GS are not considered ‘genetically modified’, overcoming the big obstacles of the application of genetically modified organisms (Gong et al. 2009; Zhang et al. 2002).
In this review, we introduce the procedure of GS, discuss the aid strategies of GS, summarize the applications of GS for increasing metabolite yield, improving strain tolerance, enhancing substrate utilization, and put forward the outlook to the future development of this technology.
The procedure of genome shuffling
The procedure of GS mainly consists of selection of initial strain, construction of parental library, recursive protoplast fusion, screening of desired fusants and stability evaluation (Gong et al. 2009; Zhang et al. 2002) (Fig. 1).
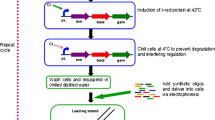
First of all, the initial strains should be selected, and then used to construct the parental library. The initial strains can be a single strain, or strains from different species, or strains from different genus (Jetti et al. 2019; John et al. 2008; Luna-Flores et al. 2017). To construct the parental library, mutagenesis is still the main choice, and in this step, the initial strains would be subjected to one or several rounds of mutagenesis. Then the selected mutants from the parental library were sent to prepare protoplasts, which would be similarly fused and regenerated. Protoplasts fusion is mainly induced by polyethylene glycol (PEG) and/or electrical pulses. The above process would be repeated several rounds, namely recursive protoplast fusion. Recursive protoplast fusion is the distinctive operation of GS, which successfully completes the efficient shuffling of diverse genes from different parental strains, ensuring the construction of desired phenotypes. The next step is to screen the desired phenotypes, which is the crucial step to ensure the success of GS, but how to efficiently screening the desired fusants is difficult and complex task. Generally, the screening methods are varied by the target of strain improvement, traditionally rely on physiological or biochemical characters of desired phenotypes. For instance, the hydrolysis zone, clear zones or inhibition zone is always employed for screening yield improved fusants, selective medium with corresponding tolerant substances is always used for screening tolerance improved fusants. Additionally, auxotroph and inactivated parental protoplast fusion were also applied in GS for screening the fusants (Gong et al. 2009). Simply, the more efficient the screening method is, the more rapidly the desired phenotypes are obtained, thus the development of the high-throughput screening (HTS) method is important for GS. The last step of GS is to evaluate the genetic stability of the selected fusants, which is always determined by evaluating the performance after continuous passage. Only the genetically stable fusants are of practical value.
Aid strategies for genome shuffling
Although GS has been successfully used for strain improvement of many microbial strains, there are still some practical needs, such as low mutant diversity, low fusion rate, low screening efficiency (Wang et al. 2016; Zhang et al. 2002). In recent years, several effective and sophisticated tools and/or methods have been developed, and introduced into GS as aid strategies to make the technology simpler, more efficient and less time consuming. Here, we describe these aid strategies to present the development potential of this practical technology.
Aid strategies for increasing the mutant efficiency
To date, random mutagenesis is still the common method for generating the mutants, however, it has some inherent limitations such as long time and low mutant efficiency. In recent years, for increasing the mutant efficiency, some novel mutation methods have been developed and applied in GS, such as atmospheric and room temperature plasma (ARTP) (Gu et al. 2017; Shi et al. 2018), atmospheric pressure non-equilibrium discharge plasma (APNEDP) (Xu et al. 2012), femtosecond laser (Liu et al. 2013), ribosome engineering (Liu et al. 2020) and low-energy ion implantation (LEII) (Xu et al. 2012) (Table 1). For example, Liu et al. applied a femtosecond laser to treat Micromonospora sagamiensis for increasing micronomicin yield (Liu et al. 2013). Under the optimized irradiation conditions of 75 mW and 180 s, a maximum of positive mutation rate of 17.8% was obtained, and maximum yield from mutant MX3004 of 263 U/ml was achieved, which was increased by 484% compared with the parent strain (Liu et al. 2013). Besides, to construct mutant libraries more rationally, the random assembly-based strategies, such as multiplex automated genome engineering (MAGE), CRISPR, and multiplex automated genome engineering for eukaryotes (EMAGE), can also be introduced into GS for generating large-scale libraries with higher diversity and different functions (Zeng et al. 2020).
Aid strategies for enhancing fusion rate
Protoplast fusion is the key event in GS, and the higher the fusion rate is, the more rapidly the desired phenotypes are obtained. However, traditional protoplasts fusion mainly relies on the presence of PEG and/or electric pulse, leading to a low fusion rate. Here we present some new technologies to enhance the fusion rate, such as femtosecond laser (Gong et al. 2008; Liu et al. 2013), nanosecond pulsed UV laser (Steubing et al. 1991), microfluidic device (Skelley et al. 2009) and optical tweezers (Steubing et al. 1991; Zhong et al. 2013). For example, femtosecond laser, possessing ultra-high temporal and spatial resolutions, was employed to induce protoplasts fusion of Phaffia rhodozyma (Gong et al. 2008). Under the optimal condition of 1.38 × 104 W and 0.25 s, the maximal fusion rate achieved 80%, providing increased efficiency in generating fusants and suggesting the great potential application of femtosecond laser in protoplasts fusion and GS (Gong et al. 2008). While Skelley et al. developed a microfluidic device that can trap and properly pair thousands of cells, and then more than 50% properly paired and fused cells were achieved (Skelley et al. 2009).
By the way, a simulation method called Monte Carlo simulation, which was used to estimate the significance level of any test statistic, was introduced into GS to simulate the simplified processes of protoplast fusion, indicating that the highest fusion rate would be achieved from 8 to 12 different parental protoplasts (Wang et al. 2016).
Aid strategies for improving screening efficiency
Recently, some developed schemes, HTS devices and/or analytical technologies have been designed for screening of desired fusants, improving the screening efficiency and making the operation of GS simpler, such as sexual and asexual reproduction (Hou 2009), drug resistance markers (Zheng et al. 2011), microplate reader (Gong et al. 2007; Liu et al. 2013), liquid chromatography-mass spectrometry (LC-MS) and fluorescence-activated cell sorting (FACS) (Fields et al. 2019; Zhang et al. 2015). To be specific, by using yeast sexual and asexual reproduction by itself instead of protoplasts fusion, a novel GS method was developed to increase ethanol production of S. cerevisiae (Hou 2009). After three rounds of GS, the obtained fusant distinctly improved ethanol resistance, but also increased ethanol yield by up to 13% compared with the control (Hou 2009). Zheng et al. introduced two drug resistance markers into GS for screening S. cerevisiae fusants with high acetic acid tolerance, with the aid of G418- and Zeocin-resistance markers, high screening efficiency was achieved (Zheng et al. 2011). While Gong et al. combined a microplate reader with 96-well microtiter plates, developing a HTS method for high epothilones-producing fusants (Gong et al. 2007). The fusants were inoculated in two parallel 96-well microtiter plates, the first plate was cultured to produce epothilone, while the second one was cultured to maintain the fusants. By using a microplate spectrophotometer, the yield of each fusant in the first plate was assayed, then the high-yield fusants were recovered from the second plate for further assay (Gong et al. 2007). Zhang et al. developed a FACS method to efficiently screen fusants of nonconventional yeast Pichia anomala for improved sugar alcohol production through GS. Parent strains were first labeled with different fluorescent stains, then fusants were selected based on their dual fluorescence by flow cytometry, achieved the efficient screening of fusants without complementary genetic markers (Zhang et al. 2015).
Application of genome shuffling for strain improvement
In 2002, GS was first applied to the improvement of tylosin production in Streptomyces fradiae (Zhang et al. 2002). Since then, GS has been extensively used to improve a variety of microbial strains for desired phenotypes (Table 1), including bacteria such as Acetobacter (Wei et al. 2012), Bacillus (Chen et al. 2020; Gu et al. 2017; John et al. 2008), Clostridium (Gu et al. 2017; Otte et al. 2009), Lactobacillus (John et al. 2008) and Zymomonas (Wang et al. 2019), actinomyces such as Streptomyces (Liu et al. 2020; Zhang et al. 2002), yeasts such as Saccharomyces (Pinel et al. 2011; Yin et al. 2016), Candida (Wei et al. 2008) and Yarrowia (Zhao et al. 2014a, b), moulds such as Aspergillus (El-Gendy et al. 2016), Aureobasidium (Kang et al. 2011) and Penicillium (Cheng et al. 2009), and microalgae such as Chlamydomonas (Fields et al. 2019) (Tables 1 and 2). Importantly, not only for strain improvement of intraspecific microbes, but GS has been also successfully applied to the improvement of interspecific microbes (Li et al. 2013; Luna-Flores et al. 2017) and intergeneric microbes (Gu et al. 2017; Jetti et al. 2019; John et al. 2008). Here, we summarize the main examples of the application of GS, especially for increasing metabolite yield, enhancing strain tolerance and improving substrate utilization, fully demonstrating the power and potential of this technology (Tables 1, 2).
Increase metabolite yield
The most important application of GS is to increase metabolite yield. For its first application, only after 2 rounds of protoplasts fusion, the generated fusants showed remarkable improvement in tylosin production, equivalent to those obtained achievement previously requiring about 20 years of effort (Zhang et al. 2002). Chen et al. used GS to increase surfactin yield in B. velezensis, after 3 rounds of GS, a high-yield fusant F34 was obtained, exhibiting a dramatic increase in surfactin yield (from 229.60 ± 7.10 mg/L to 908.15 ± 5.65 mg/L) (Chen et al. 2020). Liu et al. employed GS and ribosome engineering to improve tiancimycin-A production, then fusant CB03234-GS26 was obtained, 1.6 times higher than that of the initial Streptomyces sp. CB03234 (Liu et al. 2020). Similarly, for eucaryotic microbes, S. cerevisiae YS86, a glutathione-producing strain, was carried out 2 rounds of GS, then a 32-folds yield improvement of fusant YSF2-19 was obtained (Yin et al. 2016). P. decumbens JU-A10, a cellulase-producing strain, was also carried out 2 rounds of GS, and fusant GS2-21 achieved the maximum yield of 102.63 FPU/L/h, producing more cellulase (1.42-folds) much earlier (44 h) than that of the original strain (90 h) (Cheng et al. 2009). Furthermore, by GS, a high lovastatin-producing fusant F3/7 was obtained from A. luchuensis MERV10, which produced 57.0 mg/gds lovastatin, 6.0-folds higher than that of the starting strain (El-Gendy et al. 2016). Even for microalgae strains, GS also exhibited powerful effect, a Chlamydomonas reinhardtii fusant was obtained by GS in less than 3 months to express a green fluorescent protein (GFP) over 2% total soluble protein (TSP), which was 15-folds higher than that of the initial strain (Fields et al. 2019).
Enhance strain tolerance
Microbial cells always suffer from many environmental stresses during fermentation, such as strong acid, thermal and osmotic stresses, toxic products, and feedback inhibition, severely affecting their metabolic activity and productivity (Zhu et al. 2018). However, cell tolerance of microbial strains is regulated by distributed polygenes in the genome, thus the improvement of strain tolerance is a complex and laborious task (Zhu et al. 2018). Fortunately, GS has been reported to successfully improve stress tolerance in many industrial strains (El-Bondkly 2012), such as acetic acid-tolerant Zymomonas mobilis (Wang et al. 2019), acid-tolerant Lactobacillus (Patnaik et al. 2002; Wang et al. 2007), thermo-tolerant S. cerevisiae (Shi et al. 2009), sulphite liquor-tolerant (Pinel et al. 2011) and ethanol-tolerant S. cerevisiae (Jetti et al. 2019), glucose-tolerant L. rhamnosus (Yu et al. 2008) (Table 2).
Improve substrate utilization
Effective utilization of substrates is one of the most desired traits of industrial strains, while GS was also proved to successfully improve substrate utilization of many microbial strains (Table 2) (Jetti et al. 2019; John et al. 2008; Magocha et al. 2018). For example, Jetti et al. constructed a S. cerevisiae hybrid SP2-18 by GS, which can utilize hexose sugars as well as pentose sugars, whereas the parental strain cannot utilize xylose (Jetti et al. 2019). John et al. chose a lactic acid-producing L. delbrueckii and an amylase-producing B. amyloliquefaciens as the parental strains, after three cycles of GS, fusant F2 was obtained, which directly produced 40 g/L lactic acid from 83 g/L cassava bagasse (starch content 50% w/w) with 96% of starch conversion (John et al. 2008).
In addition, GS was proved to effectively enhance the degradation of toxic compounds for many strains (Table 2) (Dai and Copley 2004; Yi et al. 2019). For example, Pentachlorophenol (PCP) is a highly toxic pesticide, Dai and Copley used GS to obtain some high PCP-degrading fusants, which can grow on broth plates containing 6 to 8 mM PCP and completely degrade 3 mmol/L PCP in 48 h, while the original Sphingobium chlorophenolicum strain cannot grow in the presence of PCP at concentrations higher than 0.6 mmol/L (Dai and Copley 2004). Perfluorooctanoic acid (PFOA) is an emerging persistent organic pollutant, which is hard to be degraded by conventional methods because of its stable physical and chemical properties. Yi et al. successfully employed GS to improve the PFOA-degrading bacterium Pseudomonas Parafulva YAB-1, the PFOA degradation rate of fusant F3-52 was up to 58.6%, 1.8-fold higher than that of strain YAB1 (Yi et al. 2019).
Other applications
Gene clone, mutation or protoplast fusion may activate the silent genes in the genome, then the strain would produce new active metabolites (Hopwood and Chater 1980). In this respect, some researchers have proved that GS is also efficient in activating some metabolisms, thus GS could be used as a new strategy for the mining of new genes and metabolites (Cao et al. 2010; Wang et al. 2010). Wang et al. reported that fusant G-444 produced 8 new compounds, which were different in structure types and substitutions from those of the original Tubercularia sp. TF5, indicating some silent genes were activated after GS (Wang et al. 2010). Cao et al. used GS to improve the salt tolerance of Zygosaccharomyces rouxii, while the obtained fusant S3-2 not only grew well in high-salt medium, but also produced high amino acid nitrogen and ethyl acetate, and even produced a new flavor component, 4-ethylguaiacol (Cao et al. 2010). Additional, GS has also shown the powerful effects in another desired phenotypes in microbial strains, such as adhesive property in L. plantarum (Zhao et al. 2017), antimicrobial activity in Pediococcus acidilactici (Han et al. 2017) and biocontrol activity in S. bikiniensis (Zhao et al. 2014a, b). To sum up, GS is a wide range and powerful technology for strain improvement.
Future outlooks
Since the first application in 2002, GS has experienced almost two decades, and has successfully finished the improvement of many industrial strains for desired phenotypes, but it still has much room for development. Firstly, GS can combine with the rational genetic engineering techniques, for instance, with the help of rational techniques, a single gene, metabolic pathway or regulatory network with the clear genetic background can be directionally modified in advance, and then the modified phenotypes can be used for further shuffling, which will make the improvement more efficient and rational. Similarly, the fusant obtained by GS, can be sent for further directional modification in virtue of rational techniques, to harvest the better producers. Furthermore, with the development of omics and bioinformatics, the molecular mechanism of high-performance fusants can be explored, therefore GS became an important bridge to link other techniques, providing an opportunity to illustrate the metabolic networks and regulatory mechanisms.
In addition, now the availability of HTS methods is still a serious drawback for GS, while the establishment of the HTS methods is complex. In recent years, some novel devices such as biosensors, multilabel plate reader, Raman spectra, Fourier transform IR spectra, near-IR spectra and flow cytometry, have been developed and can be used for implementing the HTS methods (Zeng et al. 2020). For example, based on the spectra devices, the HTS methods for detecting a wide range of chemicals can be established, while with the introduction of flow cytometry, an HTS method can quickly analyze multiple traits of cell and rapidly classify target groups in multiple ways.
Therefore, with the development of automatic devices and rapid assay methods, the HTS method will be updated accordingly.
Conclusions
GS is a practical and effective technology for rapid improvement in microbial strains, especially for these strains with unclear genetic background. On the side, GS also offers many high-performance fusants as novel resources of rational manipulation, building an effective bridge between traditional recombination and rational manipulation. Predictably, with the development and combination of GS, it will play a much more important role in the improvement of industrial microbial strains.
References
Cao XH, Hou LH, Lu MF, Wang CL, Zeng B (2010) Genome shuffling of Zygosaccharomyces rouxii to accelerate and enhance the flavour formation of soy sauce. J Sci Food Agric 90:281–285
Chen L, Chong XY, Zhang YY, Lv YY, Hu YS (2020) Genome shuffling of Bacillus velezensis for enhanced surfactin production and variation analysis. Curr Microbiol 77:71–78
Cheng Y, Song X, Qin Y, Qu Y (2009) Genome shuffling improves production of cellulase by Penicillium decumbens JU-A10. J Appl Microbiol 107:1837–1846
Dai MH, Copley SD (2004) Genome shuffling improves degradation of the anthropogenic pesticide pentachlorophenol by Sphingobium chlorophenolicum ATCC 39723. Appl Environ Microbiol 70:2391–2397
El-Bondkly AMA (2012) Molecular identification using ITS sequences and genome shuffling to improve 2-deoxyglucose tolerance and xylanase activity of marine-derived fungus, Aspergillus sp. NRCF5. Appl Biochem Biotechnol 167:2160–2173
El-Gendy MMAA, Al-Zahrani HAA, El-Bondkly AMA (2016) Genome shuffling of mangrove endophytic Aspergillus luchuensis MERV10 for improving the cholesterol-lowering agent lovastatin under solid state fermentation. Mycobiology 44:171–179
Fields FJ, Ostrand JT, Tran M, Mayfield SP (2019) Nuclear genome shuffling significantly increases production of chloroplast-based recombinant protein in Chlamydomonas reinhardtii. Algal Res 41:101523
Gong GL, Sun X, Liu XL, Hu W, Cao WR, Liu H, Liu WF, Li YZ (2007) Mutation and a high-throughput screening method for improving the production of Epothilones of Sorangium. J Ind Microbiol Biotechnol 34:615–623
Gong JX, Zhao XM, Xing QR, Li F, Li HY, Chai L, Wang QY, Zheltikov A (2008) Femtosecond laser-induced cell fusion. Appl Phys Lett 92:093901
Gong JX, Zheng HJ, Wu ZJ, Chen T, Zhao XM (2009) Genome shuffling: progress and applications for phenotype improvement. Biotechnol Adv 27:996–1005
Gu CK, Wang GY, Mai S, Wu P, Wu J, Wang G, Liu H, Zhang J (2017) ARTP mutation and genome shuffling of ABE fermentation symbiotic system for improvement of butanol production. Appl Microbiol Biotechnol 101:2189–2199
Han GG, Song AA, Kim EB, Yoon SH, Bok JD, Cho CS, Kil DY, Kang SK, Choi YJ (2017) Improved antimicrobial activity of Pediococcus acidilactici against Salmonella Gallinarum by UV mutagenesis and genome shuffling. Appl Microbiol Biotechnol 101:5353–5363
Hopwood DA, Chater KF (1980) Fresh approaches to antibiotic production. Philos Trans R Soc Lond B Biol Sci 290:313–328
Hou L (2009) Novel methods of genome shuffling in Saccharomyces cerevisiae. Biotechnol Lett 31:671–677
Hu S, You Y, Xia F, Liu J, Dai W, Liu J, Wang Y (2019) Genome shuffling improved acid-tolerance and succinic acid production of Actinobacillus succinogenes. Food Sci Biotechnol 28:817–822
Huang QG, Zeng BD, Liang L, Wu SG, Huang JZ (2018) Genome shuffling and high-throughput screening of Brevibacterium flavum MDV1 for enhanced L-valine production. World J Microbiol Biotechnol 34:121
Jetti KD, Reddy RGNS, Garlapab D, Nammi SK (2019) Improved ethanol productivity and ethanol tolerance through genome shuffling of Saccharomyces cerevisiae and Pichia stipitis. Int Microbiol 22:247–254
John RP, Gangadharan D, Nampoothiri KM (2008) Genome shuffling of Lactobacillus delbrueckii mutant and Bacillus amyloliquefaciens through protoplasmic fusion for L-lactic acid production from starchy wastes. Bioresour Technol 99:8008–8015
Kang JX, Chen XJ, Chen WR, Li MS, Fang Y, Li DS, Ren YZ, Liu DQ (2011) Enhanced production of pullulan in Aureobasidium pullulans by a new process of genome shuffling. Process Biochem 46:792–795
Lee B-U, Cho Y-S, Park S-C, Oh K-H (2009) Enhanced degradation of TNT by genome-shuffled Stenotrophomonas maltophilia OK-5. Curr Microbiol 59:346–351
Li S, Chen X, Dong C, Zhao F, Tang L, Mao Z (2013) Combining genome shuffling and interspecific hybridization among Streptomyces improved epsilon-poly-L-lysine production. Appl Biochem Biotechnol 169:338–350
Liu P, Wen JP, Chen YL, Jia XQ (2013) Femtosecond laser-based mutagenesis strategy for micronomicin production enhancement of Micromonospora sagamiensis ATCC 21826. World J Microbiol Biotechnol 29:1121–1127
Liu H, Jiang C, Lin J, Zhuang Z, Kong W, Liu L, Huang Y, Duan Y, Zhu X (2020) Genome shuffling based on different types of ribosome engineering mutants for enhanced production of 10-membered enediyne tiancimycin-A. Appl Microbiol Biotechnol 104:4359–4369
Luna-Flores CH, Palfreyman RW, Kromer JO, Nielsen LK, Marcellin E (2017) Improved production of propionic acid using genome shuffling. Biotechnol J 12:1600120
Magocha TA, Zabed H, Yang MM, Yun JH, Zhang HH, Qi XH (2018) Improvement of industrially important microbial strains by genome shuffling: current status and future prospects. Bioresour Technol 257:281–289
Otte B, Grunwaldt E, Mahmoud O, Jennewein S (2009) Genome shuffling in Clostridium diolis DSM 15410 for improved 1,3-propanediol production. Appl Environ Microbiol 75:7610–7616
Patnaik R, Louie S, Gavrilovic V, Perry K, Stemmer WP, Ryan CM, del Cardayre S (2002) Genome shuffling of Lactobacillus for improved acid tolerance. Nat Biotechnol 20:707–712
Pinel D, D’Aoust F, del Cardayre SB, Bajwa PK, Lee H, Martin VJJ (2011) Saccharomyces cerevisiae genome shuffling through recursive population mating leads to improved tolerance to spent sulfite liquor. Appl Environ Microbiol 77:4736–4743
Shi DJ, Wang CL, Wang KM (2009) Genome shuffling to improve thermotolerance, ethanol tolerance and ethanol productivity of Saccharomyces cerevisiae. J Ind Microbiol Biotechnol 36:139–147
Shi J, Zhang M, Zhang L, Wang P, Jiang L, Deng H (2014) Xylose-fermenting Pichia stipitis by genome shuffling for improved ethanol production. Microb Biotechnol 7:90–99
Shi JR, Zhu XY, Lu YJ, Zhao HZ, Lu FX, Lu ZX (2018) Improving Iturin A production of Bacillus amyloliquefaciens by genome shuffling and its inhibition against Saccharomyces cerevisiae in orange juice. Front Microbiol 9:2683
Skelley AM, Kirak O, Suh H, Jaenisch R, Voldman J (2009) Microfluidic control of cell pairing and fusion. Nat Methods 6:147–152
Snoek T, Nicolino MP, Van den Bremt S, Mertens S, Saels V, Verplaetse A, Steensels J, Verstrepen KJ (2015) Large-scale robot-assisted genome shuffling yields industrial Saccharomyces cerevisiae yeasts with increased ethanol tolerance. Biotechnol Biofuels 8:32
Steubing RW, Cheng S, Wright WH, Numajiri Y, Berns MW (1991) Laser induced cell fusion in combination with optical tweezers: the laser cell fusion trap. Cytometry 12:505–510
Wang Y, Li Y, Pei X, Yu L, Feng Y (2007) Genome-shuffling improved acid tolerance and L-lactic acid volumetric productivity in Lactobacillus rhamnosus. J Biotechnol 129:510–515
Wang M, Liu S, Li Y, Xu R, Lu C, Shen Y (2010) Protoplast mutation and genome shuffling induce the endophytic fungus Tubercularia sp. TF5 to produce new compounds. Curr Microbiol 61:254–260
Wang H, Zhang J, Wang X, Qi W, Dai Y (2012) Genome shuffling improves production of the low-temperature alkalophilic lipase by Acinetobacter johnsonii. Biotechnol Lett 34:145–151
Wang M, Zhang W, Xu W, Shen Y, Du L (2016) Optimization of genome shuffling for high-yield production of the antitumor deacetylmycoepoxydiene in an endophytic fungus of mangrove plants. Appl Microbiol Biotechnol 100:7491–7498
Wang Y, Zhang G, Zhao X, Ling J (2017) Genome shuffling improved the nucleosides production in Cordyceps kyushuensis. J Biotechnol 260:42–47
Wang W, Wu B, Qin H, Liu P, Qin Y, Duan G, Hu G, He M (2019) Genome shuffling enhances stress tolerance of Zymomonas mobilis to two inhibitors. Biotechnol Biofuels 12:288–288
Wei P, Li Z, He P, Lin Y, Jiang N (2008) Genome shuffling in the ethanologenic yeast Candida krusei to improve acetic acid tolerance. Biotechnol Appl Biochem 49:113–120
Wei K, Cao X, Li X, Wang C, Hou L (2012) Genome shuffling to improve fermentation properties of acetic acid bacterium by the improvement of ethanol tolerance. Int J Food Sci Technol 47:2184–2189
Xu F, Jin H, Li H, Tao L, Wang J, Lv J, Chen S (2012) Genome shuffling of Trichoderma viride for enhanced cellulase production. Ann Microbiol 62:509–515
Yi L, Peng Q, Liu D, Zhou L, Tang C, Zhou Y, Chai L (2019) Enhanced degradation of perfluorooctanoic acid by a genome shuffling-modified Pseudomonas parafulva YAB-1. Environ Technol 40:3153–3161
Yin H, Ma Y, Deng Y, Xu Z, Liu J, Zhao J, Dong J, Yu J, Chang Z (2016) Genome shuffling of Saccharomyces cerevisiae for enhanced glutathione yield and relative gene expression analysis using fluorescent quantitation reverse transcription polymerase chain reaction. J Microbiol Methods 127:188–192
Yu L, Pei X, Lei T, Wang Y, Feng Y (2008) Genome shuffling enhanced L-lactic acid production by improving glucose tolerance of Lactobacillus rhamnosus. J Biotechnol 134:154–159
Zeng WZ, Guo LK, Xu S, Chen J, Zhou JW (2020) High-throughput screening technology in industrial biotechnology. Trends Biotechnol 38:888–906
Zhang YX, Perry K, Vinci VA, Powell K, Stemmer WP, Del Cardayre SB (2002) Genome shuffling leads to rapid phenotypic improvement in bacteria. Nature 415:644–646
Zhang YF, Liu SY, Du YH, Feng WJ, Liu JH, Qiao JJ (2014) Genome shuffling of Lactococcus lactis subspecies lactis YF11 for improving nisin Z production and comparative analysis. J Dairy Sci 97:2528–2541
Zhang GQ, Lin YP, Qi XN, Wang LX, He P, Wang QH, Ma YH (2015) Genome shuffling of the nonconventional yeast Pichia anomala for improved sugar alcohol production. Microb Cell Fact 14:112
Zhao S, Tian CY, Du CM (2014) Production of hybrids of Streptomyces bikiniensis strain HD-087 by genome shuffling and enhancement of its bio-control activity against Fusarium oxysporum f. sp cucumerinum. J Hortic Sci Biotechnol 89:147–152
Zhao YP, Mu XQ, Xu Y (2014) Improvement in gamma-decalactone production by Yarrowia sp. after genome shuffling. Chem Pap 68:1030–1040
Zhao YJ, Duan CC, Gao L, Yu X, Niu CH, Li SY (2017) Genome shuffling of Lactobacillus plantarum C88 improves adhesion. Biosci Biotechnol Biochem 81:184–193
Zheng DQ, Wu XC, Wang PM, Chi XQ, Tao XL, Li P, Jiang XH, Zhao YH (2011) Drug resistance marker-aided genome shuffling to improve acetic acid tolerance in Saccharomyces cerevisiae. J Ind Microbiol Biotechnol 38:415–422
Zhong MC, Gong L, Zhou JH, Wang ZQ, Li YM (2013) Optical trapping of red blood cells in living animals with a water immersion objective. Opt Lett 38:5134–5137
Zhu Z et al (2016) A new approach for breeding low-temperature-resistant Volvariella volvacea strains: genome shuffling in edible fungi. Biotechnol Appl Biochem 63:605–615
Zhu Z, Zhang J, Ji X, Fang Z, Wu Z, Chen J, Du G (2018) Evolutionary engineering of industrial microorganisms-strategies and applications. Appl Microbiol Biotechnol 102:4615–4627
Acknowledgements
This work was supported by National Key R&D Program of China (2017YFC1600803), National natural science foundation project of China (31401543), Natural science foundation project of Henan province (182300410042), Innovative Research Team in University of Henan Province (19IRTSTHN008), Henan key laboratory of cereal resource transformation and utilization (PL2017005).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest The authors declare no conflict of interest.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Chen, L., Xin, QH., Ma, LM. et al. Applications and research advance of genome shuffling for industrial microbial strains improvement. World J Microbiol Biotechnol 36, 158 (2020). https://doi.org/10.1007/s11274-020-02936-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11274-020-02936-w