Abstract
Stable and efficient hydrocarbon degrading microbial consortia were developed from a refinery sludge through nitrate amendment for their application in enhanced bioremediation of petroleum contaminated waste. Nitrate induced biostimulation of refinery sludge resulted in increased abundance of hydrocarbon degrading Rhodocyclaceae, Xanthomonadaceae, Syntrophaceae and Comamonadaceae members. Repeated subculturing of nitrate stimulated communities in crude oil supplemented basal medium was done under aerobic and anaerobic conditions. Aerobically enriched consortia (composed of Pseudomonadaceae, Pseudoxanthomonadaceae and unclassified Comamonadaceae) showed their ability to utilize alkanes, aromatics and crude oil as growth substrates. Anaerobically enriched consortium was predominated by Bacillaceae, Pseudomonadaceae, Xanthomonadaceae, Porphyromonadaceae and Comamonadaceae members. Anaerobic consortium was found to be relatively less efficient in terms of TPH (total petroleum hydrocarbons) degradation compared to its aerobic counterpart. These enriched microbial consortia were finally tested for their biodegradation performance and stability during bioremediation of highly contaminated refinery sludge using different strategies. A 30 days microcosm based bioremediation trial showed that bioaugmentation of aerobic cultures with refinery sludge was more effective in TPH degradation (~ 65% degradation) compared to the anaerobic consortium (only 36% TPH degradation) and a combination of bioaugmentation and nitrate amendment with sludge resulted in enhanced hydrocarbon attenuation (up to 86% TPH degradation). Subsequent community analysis at the end of bioremediation trial confirmed the stability of the added microbial populations. Thus, the strategy of bioaugmentation of specially enriched native microbial populations in combination with nitrate amendment was successfully used for the enhanced bioremediation of petroleum hydrocarbon contaminated refinery waste.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Management of billions of tons of hazardous petroleum hydrocarbon containing waste sludge generated by oil refineries all over the world has posed severe environmental challenge for oil industries, whose improper handling could pose serious threat to the environment (Liu et al. 2015; Roy et al. 2018a; Kuyukina et al. 2020). Remediation approaches vary from cost intensive physicochemical ones to eco-friendly biological ones, with advantages and limitations of both the strategies (Das and Kazy 2014; Davoodi et al. 2020; Kuyukina et al. 2020). Eco-friendly and cost-effective biological treatment strategies have been adopted in many cases in order to achieve complete removal of complex hydrocarbon pollutants. These strategies may rely either on improvement of local environmental conditions so that native catabolically relevant microorganisms can perform efficiently (biostimulation) or by augmenting the contaminated sites with potent hydrocarbon degrading microorganisms (bioaugmentation) to achieve enhanced bioremediation (Guerra et al. 2018; Naeem and Qazi 2020; Mishra et al. 2020). The complexity of microbial hydrocarbon degradation processes was found to be dependent on (a) nature and levels of toxic hydrocarbons (b) catabolic potential of native microbiota, (c) availability of the essential nutrients in sufficient quantity for catabolically relevant groups of microorganisms and (d) cellular uptake mechanisms of hydrophobic substrates (Sarkar et al. 2016; Roy et al. 2018b; Xu et al. 2018; Ławniczak et al. 2020). It has been well reported that different types of petroleum hydrocarbon degrading microbes prevail under harsh conditions of hydrocarbon impacted environments (Hazen et al. 2010; Tan et al. 2015; Varjani 2017; Katarína et al. 2018; Poursat et al. 2019). Past bioremediation studies have clearly indicated that designing a bioremediation strategy must entail a thorough understanding of the indigenous microbial community composition and function (De’Lorenzo 2008; Bamforth and Singleton 2005; Roy et al. 2018a). In this context, inability of the augmented exogenous microorganisms to survive and perform under field conditions have indulged researchers to focus on utilizing the potency of native microbial community members (endogenous) to sustain and perform in contaminated sites (Shankar et al. 2014; Poi et al. 2017). Taxonomically and metabolically diverse types of microorganisms including both who can oxidize the hydrocarbons directly, as well as those who are involved only in catalyzing the peripheral biodegradation pathways have been reported to work in syntrophic associations towards biodegradation of petroleum waste. Such microbial consortia have been found to be more effective as bioaugmenting agents for remediation of highly complex petroleum contaminated sludges (Szulc et al. 2014; Wang et al. 2016; Tao et al. 2017).
Enrichment studies have been conducted by previous investigators to understand the response of microbial communities towards different types of organic pollutants (aliphatics, aromatics, etc.) or terminal electron acceptors (sulfate, nitrate, iron, etc.) (Tan et al. 2015; Xu et al. 2016; Sarkar et al. 2017; Guerra et al. 2018). Several hydrocarbon specific enrichments’ under nitrate- and sulfate-reducing conditions have been studied extensively and diverse culturable as well as non-culturable microbial communities have been identified (Dou et al. 2008; Hidaka et al. 2018; Sperfeld et al. 2018; Roy et al. 2018a; Shin et al. 2019). Nitrate enriched microbial community was found to be composed of γ-Proteobacteria and Actinobacteria members, which could degrade PAHs (Polycyclic aromatic hydrocarbons) such as naphthalene (Martirani et al. 2017; Révész et al. 2020), phenanthrene and biphenyl (Rockne and Strand 2001) effectively (Sarkar et al. 2016; Roy et al. 2018a). Efficiency of sulfate-enriched communities in degradation of alicyclic aliphatics and aromatics has also been reported previously (Rios-Hernandez et al. 2003; Dou et al. 2008). Compared to all these enrichment attempts that have established the usefulness of biostimulation in bioremediation, very few studies have explored the scope for obtaining the enriched, hydrocarbon metabolizing microbial populations as active culture and use the same as inoculant for more efficient hydrocarbon remediation (Dueholm et al. 2015; Spini et al. 2018). Concerted metabolic regimes established by native microbial consortia have been reported to be effective in management of hydrocarbon contaminated environments (Cerqueira et al. 2011; Roy et al. 2018a). Our previous studies on nitrate biostimulation and bioaugmentation of native microorganisms in hydrocarbon containing sludge indicated towards effective strategies for the bioremediation of refinery sludge (Sarkar et al. 2017; Roy et al. 2018a). However, improvement in efficiency in both the cases was further required for complete degradation of high TPH containing sludge.
The present study aimed to (a) obtain specialized, hydrocarbon-degrading microbial cultures (as consortia) through biostimulation based enrichment approach and (b) characterize and test the efficacy of such cultured consortia in bioaugmentation agent for more efficient bioremediation of petroleum hydrocarbons. For this study a high TPH containing refinery waste sludge was used. The nitrate based biostimulation approach was followed by development of stable aerobic and anaerobic, hydrocarbon degrading consortia. Metagenome based high throughput sequencing and analysis of 16S rRNA genes, coupled with TPH degradation by gas chromatography were used to characterize the cultures. The consortia were finally tested for their bioremediation efficacy through bioaugmentation and a combined approach of bioaugmentation plus biostimulation.
Materials and methods
Experimental steps for the development of specialized, hydrocarbon degrading microbial consortia from refinery sludge and their bioremediation performance assessment have been summarized in Fig. 1.
Overall workflow of the study. Step 1, nitrate biostimulation of refinery sludge microorganisms, Step 2, enrichment of hydrocarbon degrading microbial consortia through repeated subculturing of previously stimulated sludge microorganisms (from step 1) in crude oil and nitrate supplemented basal medium under different conditions, and Step 3, assessment of refinery sludge bioremediation performance and stability of the enriched microbial consortia (from step 2) by adopting different strategies. Nomenclature of different microcosm set-ups of this study has been shown in the figure and description of each set-ups has been mentioned in Table 1. Microbial community compositions were determined from the samples of the sets marked with asterisk (*)
Biostimulation of native refinery sludge microorganisms by nitrate amendment
Petroleum refinery sludge (20 g) was amended with two different nitrate concentrations (10 mM; for low nitrate, designated as LN and 50 mM; for high nitrate and designated as HN) in 100 mL glass serum vial microcosms to biostimulate native hydrocarbon degrading sludge microorganisms. One killed (using 1% (w/v) sodium azide) and one unamended control sets were also maintained (to ascertain abiotic losses of hydrocarbons as well as effect of stimulation on sludge microorganisms). Periodic sampling upto 30 days followed by analysis of TPH and nitrate levels were done. TPH estimation was done by gravimetry and GC-FID (elaborated below). Nitrate levels were estimated through spectrophometric method (Cataldo et al. 1975; Sarkar et al. 2016; Roy et al. 2018a). Total DNA from the native sludge (J0d) and experimental microcosms following 21 days incubation (as maximum TPH degradation was achieved after 21 days) were extracted and analysed through amplicon sequencing and qPCR (described below) to elucidate microbial community composition and load in native sludge as well as in nitrate amended sludge.
Enrichment of hydrocarbon-degrading microbial consortia from nitrate stimulated sludge
Enrichment of cultivable hydrocarbon degrading microbial populations was done through repeated sub-culturing of microorganisms from LN and HN sets under both aerobic and anaerobic conditions in crude oil and nitrate supplemented basal medium. The composition of basal medium was as follows (g L−1):K2HPO4, 0.8; KH2PO4, 0.2; NaCl, 1; Na2SO4, 0.43; MgCl2·2H2O, 0.2; CaCl2·2H2O 0.03 and 2.5 mL of trace element solution (Das and Kazy 2014). For the enrichment of hydrocarbon degrading microbial communities, 10 mL sludge slurry from each microcosm set-ups of step 1 (i.e. from LN, HN after 21 days) was inoculated in the basal medium amended with appropriate (low/high) nitrate concentrations and 1 mL crude oil using 100 mL glass serum vials. Four sets of microcosms were thus prepared and incubated under aerobic as well as anaerobic conditions. The details of aerobic (XLN and XHN) and anaerobic (ALN and AHN) enrichment set-ups have been described in Table 1. Each of these enrichment cultures were sub-cultured under appropriate conditions following seven days of incubation for aerobic and 15 days for anaerobic enrichments. Following three sub-culturing of aerobic and anaerobic enrichment cultures, the hydrocarbon enriched microbial populations from XLN, XHN, ALN, and AHN sets were assessed in terms of their hydrocarbon degradation potential. Each of these four enriched microbial consortia was tested for their hydrocarbon degradation ability using mixture of alkanes (K), aromatics (R), alkanes + aromatics (K + R) and crude oil (CO) as substrates. Alkane mixture (K) consisted of dodecane, nonadecane and docosane at an initial concentration of 450 ppm, aromatic compound mixture (R) consisted of benzene, toluene, xylene and naphthalene (450 ppm). For K + R mixture, concentrations of the alkane and aromatic compounds were adjusted to obtain 450 ppm initial concentration, while crude oil was added as 10% (w/v) of the medium for the CO experimental set. Total petroleum hydrocarbon (TPH) degradation and nitrate consumption by aerobic consortia were monitored till 15 days, while that of anaerobic set were measured after 30 days because of their slow growth rate. TPH was measured by gas chromatography equipped with flame ionization detector (GC-FID, Clarus 580, Perkin Elmer, USA) based procedure developed by our laboratory (Sarkar et al. 2016; Roy et al. 2018a). Details of the TPH and nitrate estimation methods are presented separately. Microbial community compositions of all four enriched cultures (XLN, XHN, ALN, and AHN) were determined through 16S rRNA gene amplicon sequencing. Total DNA of the cultures was extracted following manufacturer’s protocol using DNeasy® PowerSoil® kit (Qiagen 12888-50) (Sarkar et al. 2016). Real time qPCR based quantification of total (bacterial and archaeal 16S rRNA genes) and selected (β-Proteobacteria and Firmicutes) community members were also ascertained (Gupta et al. 2018).
Assessment of refinery sludge bioremediation performance of enriched microbial consortia
The enriched microbial cultures were further assessed in terms of their refinery sludge bioremediation potential by using them as bioaugmenting agents. For evaluating the TPH removal performance of each of these microbial consortia, 10 mL culture inoculum was added to 20 g refinery sludge along with 70 mL basal medium in a 100 mL glass serum vial. Microbial TPH degradation was tested under aerobic and anaerobic conditions. Apart from killed and unamended native sludge controls, seven different sets of refinery sludge were incubated under following conditions: (i) biostimulation only (with 30 mM NaNO3; no added culture; set designated as BS), (ii) bioaugmentation only (addition of XLN, XHN and AHN cultures with the sludge at 10% (v/v) concentration; sets designated as BAXLN, BAXHN, and BAAHN), and (iii) combination of nitrate biostimulation and bioaugmentation of enriched cultures with sludge (sets designated as BSBAXLN, BSBAXHN, and BSBAAHN) (Table 1). All experimental set-ups were incubated till 30 days. TPH and nitrate concentrations were monitored through periodical sampling. Total DNA from each set-up was extracted after 30 days of incubation, followed by qPCR and 16S rRNA gene amplicon sequencing.
Measurement of TPH removal
Hydrocarbon removal from each of the set-ups was estimated using gas chromatography equipped with flame ionization detector (GC-FID, Clarus 580, Perkin Elmer, USA). Extraction of hydrocarbon was performed using n-Hexane from 1 g sludge in triplicate with 1:10 (m/v) solvent ratio, following USEPA method 3530, followed by 30 min speed vortexing and centrifugation at 10,000 rpm for 10 min. TPH content was estimated by injecting 2 µL of extract in GC using the method as described by Wallisch et al (2014). Quantification of the amount of TPH present in each sample was achieved by adding all unresolved and resolved components of the hexane extract between retention times 5 to 30 min (for C10H22 to C40H82). Degradation efficiency was measured as percentage TPH reduction using the following equation:
For individual components, n-hexane for alkanes, acetone for aromatics and 1:1 mixture of acetone and n-hexane for mixture and crude oil were used as extraction solvent in ratio of 1:5 (v/v) at the beginning and at various intervals of incubation.GC peaks corresponding to added aliphatics or aromatics were ascertained and degradation calculated accordingly. However, for crude oil, same method as TPH degradation was applied due to observation of multiple peaks. To estimate the rate of bioremediation process, the kinetics of degradation for individual components as well as TPH reduction, was fitted into a first order kinetic model as has been previously suggested (Sarkar et al. 2017) and shown in Eq. 2 below:
where C0 is the initial and Ct is the residual TPH content of sludge (g kg−1 at time t (day) and k is the biodegradation rate constant (day−1). Biodegradation rate was calculated for each treatment by plotting logarithm of residual TPH concentration versus time.
Estimation of microbial load using quantitative PCR
Quantification of microbial load (bacterial as well as archaeal) and abundance of β-Proteobacteria and Firmicutes members in nitrate stimulated sludge (LN and HN), enriched microbial consortia (XLN, XHN and AHN), and selected sludge bioremediation set-ups (BSBAXLN, BSBAXHN and BSBAAHN) were estimated using quantitative real time PCR (qPCR). Primer sets and annealing temperatures used for each group were mentioned in Table 2. qPCR (Quant Studio 5 Real-Time PCR System, Thermo Fisher Scientific) was performed using Power SYBR green PCR mastermix (Invitrogen) (5 μL), 5 pM of each primer set and 2 μL of template metagenome (total reaction volume 10 μL) in triplicate. The amplification protocol followed: 95 °C for 10 min, 40 cycles of 95 °C for 15 s, annealing temperature for 30 s and 72 °C for 30 s, followed by melt curve analysis. Annealing temperatures of 55 °C, 61 °C and 57 °C were used for bacterial and archaeal 16S, β-Proteobacteria and Firmicutes, respectively (Muyzer 1993; Mühling et al. 2008; Purkamo et al. 2016). Melting curve analysis was run after each assay to check PCR specificity. Copy number of genes were determined against standard curve (R2 > 0.993) generated using known concentrations (102 to 108 copies μL−1) of plasmid DNA containing cloned target gene as per the protocol outlined in Lee et al. (2008).
Amplicon sequencing and data analysis
Microbial community compositions of samples from nitrate stimulated sludge(LN and HN), enriched microbial consortia (XLN, XHN and AHN), and selected sludge bioremediation set-ups (BSBAXLN, BSBAXHN and BSBAAHN) were determined using amplicon sequencing of 16S rRNA genes targeting V4 regions. V4 region specific primers 515F and 806R were used for PCR amplification using AmpliTaq GoldTM 360 Master Mix and the following amplification conditions: 95 °C for 5 min, 35 cycles of 95 °C for 40 s, 50 °C for 45 s and 72 °C for 40 s with final extension at 72 °C for 7 min. Amplified products were extracted using 2% E-gel (E-Gel SizeSelect II Agarose Gel, Thermo Fisher Scientific) and sequenced in Ion S5 platform (Thermo Fischer Scientific) with Ion 530 chip using manufacturer’s protocol. Taxonomic compositions of microbial communities were analyzed using QIIME (Quantitative Insights Into Microbial Ecology) pipeline (Caporaso et al. 2010). Raw reads were filtered (for length between 250 and 300 bps), clustered into Operational taxonomic units (OTUs) at 97% identity and assigned taxonomy using SILVA 128 database. The raw reads were deposited to the NCBI Sequence Read Archive database (SRA) under Bioproject ID PRJNA515418.
Statistical analysis
The data were subjected to one-way analysis of variance (ANOVA) at 5% probability. Mean of the different treatments were tested for level of significant differences at p < 0.05 by Tukey (Honestly Significant Difference) test. Associations between variables were calculated by Pearson’s correlation.
Results
Biostimulation of native refinery sludge microorganisms by nitrate amendment
Biostimulation of native sludge microorganisms by the addition of two different levels of nitrate concentrations (LN, 10 mM and HN, 50 mM) resulted in considerable amount (up to 55%) of sludge TPH degradation (i.e., from 400 to 180 g kg−1) within three weeks of incubation, which is much higher than that of the unamended control set (Fig. 2a). TPH degradation in biostimulation set-ups increased steadily over the incubation period with concurrent reduction in nitrate concentrations and increment in bacterial load within the microcosms (Fig. 2b, c), which confirmed the effectiveness of biostimulation with nitrate amendment as a strategy for the bioremediation of oil containing sludge. The degradation of sludge TPH was comparable between low nitrate (LN) and high nitrate (HN) sets. However, the nitrate utilization profile indicated depletion of nitrate in LN set within 30 days, while presence of sufficient nitrate was observed in HN set (2b). qPCR analysis indicated a significant increase in bacterial 16S rRNA gene copy numbers with biostimulation within two weeks (Fig. 2c).
Effect of nitrate amendment (biostimulation) on hydrocarbon degradation and microbial community composition in oil refinery sludge. a TPH degradation in sludge microcosms with/without nitrate amendment (low, LN, 10 mM nitrate and high, HN, 50 mM), b Nitrate utilization profile of the biostimulated (LN, HN) and non-stimulated (Unamended) sludge microcosms, c qPCR analysis of bacterial cell counts based on copy number of bacteria specific 16S rRNA genes in biostimulated and unamended sludge samples, and d Taxonomic distribution of major families (cumulative abundance 0.5%) in native unamended sludge control (J0d) and biostimulated (LN and HN) sludge samples
The microbial community compositions of the native unamended sludge (J0d) and that after 3 weeks (21 days) of nitrate stimulation (both LN and HN) were assessed using 16S rRNA genes-V4 targeted amplicon sequencing. Taxonomic assignment of sequence reads showed distinct shift in community composition with nitrate amendment at family level. Dominant families (cumulative abundance > 1%) detected in three samples and their relative abundances were represented in Fig. 2d. In the native sludge sample (J0d), 56% of total reads were assigned to the members of following 10 families, Pseudomonadaceae (29%), Sphingomonadaceae (6.3%), Comamonadaceae (4.4%), Xanthomonadaceae (4.4%), Rhodocyclaceae (4.3%), Cytophagaceae (2.5%), Moraxellaceae (1.7%), Chitinophagaceae (1.4%), Verrucomicrobiaceae (1.2%) and Microbacteriaceae (1%). Incubation of sludge under low nitrate condition for 21 days (LN) resulted in a microbial community, which was majorly composed of Rhodocyclaceae (21%), Syntrophaceae (19%), Xanthomonadaceae (16%), Comamonadaceae (8.4%), Geobacteraceae (4.5%), Anaerolineaceae (3.7%), Porphyromonadaceae (3.2%), Coriobacteraceae (3%), Methanoregulaceae (2%), Bacteriovoradaceae (2%), Methanosetaceae (1.7%), Enterobacteraceae (1.2%), Thermodesulfobiaceae (1.1%), Spirochaetaceae (1%) and Ruminococcaceae (1%) members. In this case, noticeable increment was observed in the abundance of Syntrophaceae, Anaerolineaceae, Porphyromonadaceae, Coriobacteraceae, Methanoregulaceae, Bacteriovoradaceae, Thermodesulfobiaceae, Ruminococcaceae, Caldisericaceae, Desulfobulbaceae, Synergistaceae, Syntrophobacteraceae and Vellionellaceae members. Some minor groups (abundance < 0.5%) like Peptococcaceae, Oxalobacteraceae, Syntrophomonadaceae and Crenarchaeotal member Desulfurococcaceae also showed considerable hike in relative abundance within low nitrate stimulated sludge (Fig. 2d). With high nitrate incubation of sludge (HN), similar type of microbial community composition was noticed along with increase in the abundance of Hydrogenophilaceae, Hyphomonadaceae, Bradyrhizobiaceae, Thermodesulfobacteraceae and Microbacteriaceae members. However, marginal increment in the abundance of members of Rhodocyclaceae, Caldiseica_TTA-B1, Desulfurococcaceae, Porphyromonadaceae, Ruminococcaceae, Caldiserica_TTA-B15 was observed in later case.
At OTU level, 3766 OTUs were shared among LN and HN sets covering almost (36%) and (61%) of reads, while 6633 and 2341 OTUs were unique in each sample, respectively (Fig. 3a). Figure 3b showed that most abundant (top 30) OTUs of LN and HN sets are common. This indicated stimulation of a core community with nitrate amendment, irrespective of its concentration. Rank abundance plot (Fig. 3c) indicated that the most abundant OTU of LN and HN sets (denovo29972, covering almost 12% and 19% reads of LN and HN) showed closest BLAST match to hydrocarbon degrading Rhodocyclaceae members. The next abundant OTU of both the sets (denovo27926) could be assigned to the well known hydrocarbon degrading Pseudoxanthomonas sp. Third abundant OTU (covering 4.2% of LN, 2.6% of HN reads) was assigned to the syntrophic, alkane degrading Smithella sp. Fourth abundant OTU (covering 4% LN, 2.3% HN reads) was affiliated to hydrocarbon degrading Comamonadaceae member. Other common OTUs of both the LN and HN sets could be assigned to organic pollutant degrading Geobacter, Proteiniphilum, etc.
Distribution of OTUs among native sludge (J0d) and biostimulated sludge (LN and HN): a Venn diagram showing common and unique OTUs among the samples; b Relative read distribution of most abundant top 30 OTUs of J0d, LN and HN in each of the three samples, c Rank abundance plot showing read distribution of most abundant and common OTUs of LN and HN sets
Enrichment of hydrocarbon-degrading microbial consortia from nitrate stimulated sludge and assessment of their hydrocarbon degradation potential
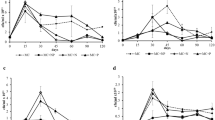
After three subculturing of nitrate biostimulated sludge microbial communities in crude oil amended basal medium under both aerobic and anaerobic conditions, four sets of enriched microbial consortia (aerobic XLN and XHN; anaerobic ALN and AHN) were obtained. To assess their hydrocarbon degradation ability, each of these consortia was tested for alkanes (K), aromatics (R), mixture of K + R and crude oil (CO) degradation. Figure 4a represented the results of aerobic biodegradation after 7 and 14 days incubation. It was observed that alkanes (K) were the most preferred carbon source for the enriched consortia followed by K + R mixture, crude oil and aromatics as evident from their rate and extent of biodegradation. However, biodegradation performance of XLN and XHN sets were found to be comparable indicating towards the positive effect of nitrate amendment in biodegradation even at low concentration. A marginal loss (10–15%) of hydrocarbons in control sets could be attributed to abiotic oxidation over the incubation period. Rate of degradation of each set was calculated assuming first order degradation kinetics and the result was presented in Table 3. Degradation rate constant k was maximum for alkanes in both XLN (k = 0.11 day−1) and XHN (k = 0.13 day−1) sets. Slower rate of degradation was found for aromatics in which case, XHN (56% TPH degradation, k = 0.06 day−1) consortium performed marginally better than XLN (50% TPH degradation, k = 0.05 day−1). In a similar comparison, anaerobic consortia ALN and AHN also showed higher degradation of alkanes over aromatics and other compounds tested albeit consuming longer time (Fig. 5a). It was observed that AHN consortium was a better performer in terms of TPH degradation, which could degrade 54% of K, 45% of R, 48% of K + R and 39% of CO compounds after 30 days of incubation. Although ALN consortium also showed competent degradation rates, unfortunate contamination in culture impeded its further growth and this consortium was thus not used for subsequent experiments.
Hydrocarbon degradation ability and taxonomic composition of aerobic enrichments: a Hydrocarbon degradation efficiency and nitrate utilization of aerobically enriched consortia (XLN and XHN) with aliphatic (K), aromatic (R), aliphatic and aromatics mix (K + R) and crude oil (CO); b Taxonomic composition (at family level) of initial (LN, HN) and aerobically enriched communities (XLN, XHN) with cumulative relative read abundance of 1%
Hydrocarbon degradation ability and taxonomic composition of anaerobic enrichments: a Hydrocarbon degradation and nitrate consumption at the end of 30 days for anaerobic enriched cultures (ALN and AHN), b Taxonomic composition of initial sludge (J0d), starter culture (HN) and enriched community under high nitrate (AHN) at family level with 1% cumulative abundance
Taxonomic characterization of enriched microbial populations in basal medium
Repeated subculturing of nitrate stimulated sludge microorganisms in crude oil amended basal medium for the enrichment of hydrocarbon degrading microbial populations under aerobic and anaerobic conditions, showed a prominent effect on the composition of microbial communities obtained therein. Amplicon sequencing of aerobic enrichments XLN and XHN generated an average of 0.2 million quality filtered reads distributed over 1017 OTUs. Compared to the native or nitrated stimulated microbial communities, decrease in microbial diversity was observed in the XLN, XHN and ALN consortia (Figs. 4b, 5b). Taxonomic assignments of the reads indicated preponderance of certain taxa, viz., Pseudomonadaceae, Comamonadaceae and Xanthomonadaceae in XLN as well as in XHN set. However, Pseudomonadaceae members were dominant in low nitrate XLN community, while Comamonadaceae members were most abundant in high nitrate XHN community. In both the consortium most abundant genera were assigned to Pseudomonas, Pseudoxanthomonas and Comamonadaceae members, Brachymonas, Ottowia, Comamonas, Extensimonas, Acidovorax and Paracoccus (Table 4). On the other hand, taxonomic data of anaerobic AHN community showed that 97% of the total quality filtered reads belonged to Bacillaceae (Bacillus, 64%, most abundant), Pseudomonadaceae (Pseudomonas, 12.5%), Porphyromonadaceae (Proteiniphilum, 10%), Hydrogenophilaceae (Tepidiphilus, 1.8%) and Comamonadaceae (Brachymonas, Ottowia, etc.) (Fig. 5b; Table 4). OTU level analysis indicated that most abundant 23 OTUs of XLN community covered almost 89% of its total reads and 88% of XHN community reads, indicating similar effect of nitrate levels (Fig. 6). The most abundant OTU of XLN (covering 60% reads) was assigned to Pseudomonas, while in XHN community the dominant OTU was assigned to Comamonadaceae member. The second most abundant OTU of XLN community (covering 10% reads), which was also the most abundant OTU of XHN, was assigned to Comamonadaceae family and showed similarity to Alicycliphilus member. Other dominant OTUs in both the consortium was related to Pseudoxanthomonas and Candidate division TM6 (Dependentiaceae) members. Increase in betaproteobacteria members have been confirmed through both amplicon and quantitative real time PCR study (Fig. 7).
Quantitative estimation of microbial abundance in samples using qPCR. Quantitative real time based estimation of bacteria, archaea, β-Proteobacteria and Firmicutes members in the native sludge (J0d), stimulated sludge (LN, HN), enriched consortia (XLN, XHN, AHN) and selected sludge samples after bioremediation experiment (BSBAXLN, BSBAXHN, BSBAAHN)
Assessment of sludge bioremediation performance of enriched microbial consortia
The enriched hydrocarbon degrading microbial consortia (XLN, XHN and AHN) were further assessed in terms of their refinery sludge bioremediation potential by using them as bioaugmenting agents. Problems in culturing ALN consortium led us to eliminate it for further study. TPH degradation performance of each of these cultures were monitored by adopting different bioremediation strategies that include (i) their bioaugmentation with sludge (bioremediation set-ups were designated as BAXLN, BAXHN and BAAHN) and (ii) a combination of their bioaugmentation + nitrate biostimulation (BSBAXLN, BSBAXHN and BSBAAHN). The results of TPH biodegradation during bioremediation experiment have been presented in Fig. 8a. Abiotic loses accounted for 8% TPH reduction, while unamended control (native sludge) accounted for 30% TPH degradation in 30 days. Within the same timeframe, aerobic consortia (XLN and XHN) performed better than anaerobic ones (AHN). Nitrate biostimulation (BS only) of native sludge could achieve up to 59% TPH degradation in 30 days, while only bioaugmentation strategy could achieve 38–58% TPH reduction (varying for BAXLN, BAXHN and BAAHN set-ups). Maximum TPH degradation could be observed in the set amended with XHN consortium and 30 mM nitrate (BSBAXHN). The initial TPH of 320 ± 25 g kg−1 was reduced to 46 g kg−1 in BSBAXHN set-up and 112 g kg−1in BSBAXLN. In anaerobic bioremediation set-ups (BAAHN and BSBAAHN), rate and extent of TPH degradation was inhibited (36% in 30 days). Compared to other bioremediation strategies, combined biostimulation and bioaugmentation set-up showed faster and efficient degradation kinetics (Table 5). Survivability of the added microbial populations as well as the microbial community composition after 30 days of bioremediation treatment, were monitored using 16SrRNA gene amplicon sequencing of selected sludge microbiome (aerobic bioremediation sets BSBAXLN and BSBAXHN; anaerobic BSBAAHN) (Fig. 8b). The aerobic sets generated an average of 0.2 million reads distributed over 1000 OTUs. Taxonomically BSBAXLN was predominated by γ-(83%) [Pseudomonadaceae, Xanthomonadaceae] and β- (16%) [Comamonadaceae] Proteobacteria, members as also observed in the added XLN community (Fig. 8b). Only four genera, viz., Pseudomonas, Pseudoxanthomonas, Paracoccus and Brachymonas constituted majority (covering > 80% reads) in the BSBAXLN community (Table 4). Similarly, after bioremediation, BSBAXHN community was also composed of the members of γ-[Xanthomonadaceae, Pseudomonadaceae] and β-[Comamonadaceae] Proteobacteria, covering 89% of quality filtered reads in the library, which is in line with the similar community composition of the added XHN populations in this set-up (Fig. 8b). Pseudoxanthomonas, Pseudomonas and Brachymonas were also found to be the most prevalent genera in BSBAXHN community (Table 4). The anaerobic set-up (BSBAAHN) also showed the survival of the amended microbial populations (AHN). After 30 days, BSBAAHN set-up was found to be composed of the members of Firmicutes (Bacillaceae), γ-Proteobacteria (Pseudomonadaceae, Xanthomonadaceae), β-Proteobacteria (Comamonadaceae, Hydrogenophilaceae), and Bacteoidetes (Porphyromonadaceae) (Fig. 8b). Sludge derived community after 30 days treatment, BSBAAHN indicated presence of similar groups as AHN community with marginal increase in abundance of Comamonadaceae members Brachymonas, Comamonas, Ottowia, Aquabacterium etc. Presence of β-Proteobacteria and Firmicutes members in the sludge samples before and after bioremediation has also been confirmed through qPCR study (Fig. 7). Overall observation showed the survivability and stability of the added microbial consortia during bioremediation of refinery sludge while performing very well in sludge TPH degradation.
Assessment of refinery sludge bioremediation potential and stability of enriched microbial consortia during bioremediation trial: a TPH degradation during refinery sludge bioremediation using nitrate biostimulation, bioaugmentation with XLN, XHN or AHN and a combination of nitrate biostimulation + bioaugmentation (XLN, XHN or AHN) over 30 days; b Microbial community composition in microcosms after 30 days of bioremediation experiment using combined biostimulation + bioaugmentation strategy (BSBAXLN, BSBAXHN and BSBAAHN) and comparison with starter cultures (XLN, XHN and AHN) at family level covering > 90% of reads in each library
Discussion
Biostimulation of oil refinery sludge microorganisms by the addition of nitrate showed enhanced TPH degradation with concomitant nitrate utilization and enhanced bacterial abundance within the sample. It has been shown earlier that the degradation of petroleum hydrocarbons by the indigenous community can be enhanced by the extraneous supply of the required nutrients in the contaminated site (Delille et al. 2004; Tyagi et al. 2011; Sarkar et al. 2016). Diverse types of hydrocarbon residues present in the refinery waste sludge represent a substantial source of carbon and electrons for the indigenous microorganisms, whereas, the presence/bioavailability of nitrogen and phosphorous is still very limited. The refinery sludge used in this study was deprived of nitrogen and as soon as available nitrate was provided, the resident microorganisms started utilization of the same rapidly, which was confirmed by the depletion of the nitrate concentration. Availability of nitrogen (as nitrate) favorably alleviates the N deficient state of high organic carbon-rich sludge and helps in stimulating a plethora of biogeochemical transformations thereby allowing efficient metabolism and cell growth which could be evidenced by steep rise in bacterial cell numbers, coupled with TPH reduction (Mason et al. 2014; Zhang et al. 2015; Xu et al. 2018). With respect to the different nitrate levels used in this study, we found that concentration of nitrate had a marginal role on the amount of TPH degradation as it was slightly higher in HN set compared to the LN set, which indicated that even the addition of low amount of nitrate could stimulate the growth of resident microorganisms within the sludge thus resulting in enhanced TPH degradation (Hazen et al. 2016). It has been reported that nitrate has two distinct functions with respect to microorganisms of hydrocarbon rich environments, which could facilitate hydrocarbon degradation: (a) nitrate mediated hydrocarbon activation with concomitant growth of mainly β-Proteobacteria members, and (b) nitrate can act as terminal electron acceptor by dissimilatory nitrate reducing bacteria during oxidation of organic compounds (Zedelius et al. 2011; Dashti et al. 2015; Sarkar et al. 2016).
Previous studies on petroleum contaminated samples have demonstrated the efficacy of biostimulation as a remediation strategy (Atlas and Hazen 2011; Wu et al. 2016; Liu et al. 2020). Taxonomic composition of these biostimulated communities were ascertained to identify the members, which have been active in achieving enhanced degradation as compared to native sludge community (Lamendella et al. 2014; Singleton et al. 2016; Roy et al. 2018a). Among the abundant families detected in the unamended native sludge (J0d) of the present study, Pseudomonadaceae members have been ubiquitously reported from other hydrocarbon contaminated environments, while members of Sphingomonadaceae have been found to be oligotrophic and able to utilize a wide range of recalcitrant aromatics under a wide range of temperature and nutrient regimes (Yang et al. 2014; Wang et al. 2016; Zhou et al. 2016; Lee et al. 2019). Betaproteobacteria members Rhodocyclaceae and Comamonadaceae, which constitute a part of the native community, have also been previously reported from petroleum affected environments and their function in nitrogen cycle was noteworthy (Yergeau et al. 2012; Bell et al. 2013; Singleton et al. 2016). Chitiniphagaceae members, detected in our sample (Sedimentibacterium), have been reported to degrade recalcitrant crude oil components like tar (Llado et al. 2015). Thus, our observation suggested presence of potent hydrocarbon degrading microbial community in the native refinery sludge, which was comparable to that reported from such site previously (Roy et al. 2018b). Biostimulation with low nitrate level led to proliferation of metabolically diverse microorganisms known to be involved in hydrocarbon degradation, nitrate and sulfate reduction, syntrophic and methanogenic activities (Rhodocyclaceae, Xanthomonadaceae, Comamonadaceae, Anaerolineaceae, Geobacteraceae, Syntrophaceae, Porphyromonadaceae, Coriobacteraceae, Methanoregulaceae members (Gray et al. 2011; Zedelius et al. 2011; An et al. 2013; Dashti et al. 2015; Tan et al. 2015; Fowler et al. 2016). With high nitrate incubation (HN), similar trend was also noticed along with the increase in abundance of other organisms including Hydrogenophilaceae, Hyphomonadaceae, Bradyrhizobiaceae, Thermodesulfobacteraceae and Microbacteriaceae members. Phylogenetic interpretation of the abundant OTUs of nitrate stimulated communities (LN and HN) clearly indicated close phylogenetic relation of the members with previously reported hydrocarbon degrading microorganisms. The phylogenetic study in particular revealed the close lineage of biostimulated members to organisms not only inhabiting/degrading hydrocarbons, but also creating physical/chemical ambience to other members of the community, thus, favoring an overall enhanced community performance. For example, the most abundant OTU of both the samples indicated its close similarity to Rhodocyclales member Azovibrio as well as to Dechloromonas, which have also been reported to create niches for aerobic hydrocarbon degrading bacteria to function (Wolterink et al. 2005). Detection of syntrophic groups such as Syntrophaceae, Anaerolineaceae, Porphyromonadaceae and Coriobacteriaceae indicated that they could provide organic acid intermediates like acetate or formate, which in turn could act as substrates for methanogenic archaeal members (Sutton et al. 2013; Noguchi et al. 2014; Fowler et al. 2016). An organism reported to have diverse hydrocarbon degradation capacity, Pseudoxanthomonas, was detected as the second most abundant OTU confirming their presence in the nitrate stimulated sludge samples. These organisms have also been previously reported from nitrate associated hydrocarbon contaminated environments (Young et al. 2007; Al-Mailem et al. 2019). Presence of anaerobic, acetogenic, syntrophic organisms related to Smithella indicated towards the presence of anaerobic niches in the sludge. Abundance of Comamonadaceae members, which have been previously reported to be involved in hydrocarbon degradation under nitrate stimulated conditions, ascertained their possible role as major players in the hydrocarbon degradation process (Qiu et al. 2015). The shift in community composition after nitrate stimulation of native sludge corroborated very well with OTU level analysis. OTU level analysis of biostimulated samples (both LN and HN) clearly indicated stimulation of a specific microbial community with nitrate amendment in sludge. The data reaffirmed the effect of nitrate biostimulation on shifting of microbial community composition, which could be linked to corresponding enhanced TPH degradation as has also been suggested by previous investigators (Roy et al. 2018a; Siles and Margesin 2018). Overall observation also suggested that aerobic and anaerobic microbes could work together to facilitate hydrocarbon degradation under biostimulated condition.
Enrichment of hydrocarbon degrading microbial populations through repeated subculturing of these nitrate biostimulated communities in crude oil amended basal medium under aerobic and anaerobic conditions resulted in four microbial communities (aerobic: XLN and XHN; anaerobic: ALN and AHN). Biodegradation of a range of hydrocarbon compounds by the four enriched consortia indicated that aerobic consortia (XLN, XHN) were superior to anaerobic ones. Under both the conditions, alkanes were preferred over aromatics. Microbial hydrocarbon degradation ability has been reported to be affected by the chemical structure of the hydrocarbons and several studies have reported the order of preference as follows: linear alkane > branched alkane > aromatics > cyclic compounds > PAHs (Varjani 2017). Aerobic degradation of hydrocarbons has been reported to be faster and more metabolically efficient than anaerobic degradation (Rojo 2009; Brzeszcz and Kaszycki 2018; Xu et al. 2018). Repeated sub-culturing of hydrocarbon degrading populations in the basal medium with nitrate showed a prominent effect on the composition of microbial communities obtained from original biostimulation. Aerobic enrichments (XLN, XHN) showed preponderance of certain taxons, viz., γ- and β-Proteobacteria and Candidate division TM6, specifically Pseudomonas, Pseudoxanthomonas and Comamonadaceae members, Brachymonas, Ottowia, Comamonas, Extensimonas, Acidovorax and Paracoccus, all of them have been known to be capable of hydrocarbon degradation and nitrate reduction (Rotaru et al. 2010; Al-Mailem et al. 2019; Ali et al. 2020). Low nitrate levels stimulated Pseudomonadaceae members (Pseudomonas), while high nitrate stimulated Comamonadaceae members (Comamonas, Brachymonas). OTU level analysis of the enriched aerobic hydrocarbon degrading communities reiterated the effect of nitrate enrichment, irrespective of the concentration, wherein most abundant OTUs in both the samples were shared covering majority of reads in each of the libraries. The most abundant OTU of XLN showed closest taxonomic match with Pseudomonas, which is well known for its hydrocarbon degradation capability in diverse environments. The second most abundant OTU of XLN and the most abundant OTU of XHN was assigned to Comamonadaceae family member, which showed similarity to Alicycliphilus member reported to be capable of degrading hydrocarbons under both aerobic and anaerobic conditions (Mechichi et al. 2003). Members of Betaproteobacteria have been implicated in hydrocarbon removal under nitrate reducing conditions (Zedelius et al. 2011; Dashti et al. 2015), which was also observed in this study and a significant increase was confirmed through both amplicon and quantitative PCR analysis. Thus, this study reaffirms the role of β- and γ-Proteobacteria in rapid aerobic degradation of hydrocarbons in presence of nitrate. Anaerobic enrichment (AHN) resulted in a distinct community with preponderance of Firmicutes (Bacillus) members along with few common members as also observed in aerobic sets (Pseudomonadaceae, Porphyromonadaceae, Hydrogenophilaceae snd Comamonadaceae). Bacillus members have been reported ubiquitously in diverse hydrocarbon contaminated environments and are known to have strong role in hydrocarbon attenuation (Das and Kazy 2014; Tao et al. 2017; Roy et al. 2018a) under anaerobic condition. Several members are capable of hydrocarbon degradation followed by fermentation thus producing intermediates useful for other organisms.
Sludge bioremediation experiments were set up to investigate the TPH biodegradation efficiency of the enriched microbial consortia. Compared to other bioremediation strategies tested, a combined nitrate biostimulation and bioaugmentation (addition of enriched microbial consortia) approach was observed to be most effective emphasizing the importance of nutrient availability as well as presence of capable degraders in order to achieve efficient bioremediation of refinery sludge. However, the sludge bioremediation performance of aerobically enriched microbial consortia (XLN and XHN) was superior over the anaerobic populations (AHN), which corroborated with very well-known fact about the efficacy of aerobic system on hydrocarbon degradation. It should be noted that the performance of anaerobically enriched consortium (AHN) was tested for its sludge bioremediation ability under anaerobic condition only. Although anaerobically enriched community performed poorly compared to the aerobic enrichments, anaerobic regimes deep inside the sludge pit, might support the activities of anaerobic populations in subsequent degradation of hydrocarbons. In order to validate the maintenance of added microbial consortia (through bioaugmentation), we assessed the taxonomic composition of the three bioremediation set-ups (BSBAXLN, BSBAXHN and BSBAAHN) after the completion of sludge TPH biodegradation experiment. It was clearly evident that the major bacterial taxa, augmented to the refinery sludge with XLN and XHN communities, could survive and flourish satisfactorily as the same OTUs, as found in the XLN, XHN and AHN communities before treatment, constituted the major proportions in the sludge bioremediation set-ups after the completion of experiment. Predominance of gamma (Pseudomonadaceae, Xanthomonadaceae) and beta- (Comamonadaceae) proteobacterial members emphasized on their role in efficient TPH biodegradation in refinery sludge under aerobic conditions (Bargiela et al. 2015; Singleton et al. 2016; Duarte et al. 2017). It was noticeable that augmented microbial communities could successfully survive and establish a stable and efficient hydrocarbon degrading community within refinery sludge. Although, anaerobic bioremediation experiment could not yield satisfactory result in terms of efficient TPH degradation within 30 days, taxonomic analysis, however, revealed the survival of the added microbial community (AHN) after the treatment period, verifying the members as active participants of hydrocarbon degradation process. Although anaerobic enrichment could not perform comparably to aerobic consortia but adaptability within sludge bioremediation environment was comparable, which strongly supported the use of native microbial populations as bioaugmenting agents for accelerating refinery sludge bioremediation under aerobic as well as anaerobic conditions. The survival, stability and performance of enriched microbial community members within sludge bioremediation set-ups could be attributed to their capability to survive and flourish within their native environment, wherein these microbes have been already adjusted with the harsh conditions. Thus the approach of bioaugmentation of autochthonous microbial populations could be successfully used for the enhanced bioremediation of petroleum hydrocarbon contaminated refinery waste.
Conclusion
This study allowed enrichment and characterization of hydrocarbon degrading microbial consortia from a highly contaminated refinery waste sludge. The enriched hydrocarbon degrading populations were predominantly composed of bacteria and these cultures were successfully tested for their application as bioaugmentation agent for refinery waste remediation. Our work confirmed that catabolically superior, hydrocarbon degrading mixed bacterial culture can be enriched from refinery waste sludge. Nitrate reducing, hydrocarbonoclastic aerobic genera Comamonas, Brachymonas, Pseudomonas and Pseudoxanthomonas were obtained as aerobic consortia under both high and low nitrate condition. The anaerobic consortium obtained through high nitrate amendment was constituted by anaerobic, fermentative, members affiliated to Bacillus, Proteiniphilum, etc. The study successfully demonstrated that nitrate utilizaing, hydrocarbon degrading microbial consortia obtained from refinery sludge can be used as potent bioaugmentation agent for efficient remediation of such complex waste. An advanced bioremediation strategy was developed by combining biostimulation and bioaugmentation approaches for superior hydrocarbon degradation. This microbial consortia obtained and advanced bioremediation strategy developed could be implemented for attenuation of petroleum hydrocarbon pollution.
Data availability
The raw sequence reads obtained were submitted to the NCBI Sequence Read Archive (SRA) database under Bioproject ID PRJNA515418.
References
Ali N, Dashti N, Khanafer M, Al-Awadhi H, Radwan S (2020) Bioremediation of soils saturated with spilled crude oil. Sci Rep 10:1116. https://doi.org/10.1038/s41598-019-57224-x
Al-Mailem DM, Kansour MK, Radwan SS (2019) Cross-bioaugmentation among four remote soil samples contaminated with oil exerted just inconsistent effects on oil-bioremediation. Front Microbiol 10:2827. https://doi.org/10.3389/fmicb.2019.02827
An D, Caffrey SM, Soh J, Agrawal A, Brown D, Budwill K, Dong X, Dunfield PF, Foght J, Gieg LM, Hallam SJ, Hanson NW, He Z, Jack TR, Klassen J, Konwar KM, Kuatsjah E, Li C, Larter S, Leopatra V, Nesbø CL, Oldenburg T, Page AP, Padron ER, Rochman FF, Mehrabad AS, Sensen CW, Sipahimalani P, Song YC, Wilson S, Wolbring G, Wong ML, Voordouw G (2013) Metagenomics of hydrocarbon resource environments indicates aerobic taxa and genes to be unexpectedly common. Environ Sci Technol 47:10708–10717. https://doi.org/10.1021/es4020184
Atlas RM, Hazen TC (2011) Oil biodegradation and bioremediation: a tale of the two worst spills in U.S. history. Environ Sci Technol 45:6709–6715. https://doi.org/10.1021/es2013227
Bamforth SM, Singleton I (2005) Bioremediation of polycyclic aromatic hydrocarbons: current knowledge and future directions. J Chem Technol Biotechnol 80:723–736. https://doi.org/10.1002/jctb.1276
Bargiela R, Mapelli F, Rojo D, Chouaia B, Tornés J, Borin S, Richter M, Del Pozo MV, Cappello S, Gertler C, Genovese M (2015) Bacterial population and biodegradation potential in chronically crude oil-contaminated marine sediments are strongly linked to temperature. Sci Rep 5:11651. https://doi.org/10.1038/srep11651
Bell TH, Yergeau E, Juck FD, Whyte GL, Greer WC (2013) Alteration of microbial community structure affects diesel biodegradation in an Arctic soil. FEMS Microb Ecol 85:51–61. https://doi.org/10.1111/1574-6941.12102
Brzeszcz J, Kaszycki P (2018) Aerobic bacteria degrading both n-alkanes and aromatic hydrocarbons: an undervalued strategy for metabolic diversity and flexibility. Biodegradation 29:359–407. https://doi.org/10.1007/s10532-018-9837-x
Caporaso JG, Kuczynski J, Stombaugh J, Bittinger K, Bushman FD, Costello EK, Fierer N, Pena AG, Goodrich JK, Gordon JI, Huttley GA, Kelley ST, Knights D, Koenig JE, Ley RE, Lozupone CA, McDonald D, Muegge BD, Pirrung M, Reeder J, Sevinsky JR, Turnbaugh PJ, William AW, Widmann J, Yatsunenko T, Zaneveld J, Knight R (2010) QIIME allows analysis of high throughput community sequencing data. Nat Methods 7:335–336. https://doi.org/10.1038/nmeth.f.303
Cataldo DA, Maroon M, Schrader LE, Youngs VL (1975) Rapid colorimetric determination of nitrate in plant tissues by nitration of salicylic acid. Commun Soil Sci Plant Anal 6:71–80. https://doi.org/10.1080/00103627509366547
Cerqueira VS, Hollenbach EB, Maboni F, Vainstein MH, Camargo FA, Maria do Carmo RP, Bento FM (2011) Biodegradation potential of oily sludge by pure and mixed bacterial cultures. Bioresour Technol 102:11003–11010. https://doi.org/10.1016/j.biortech.2011.09.074
Das R, Kazy SK (2014) Microbial diversity, community composition and metabolic potential in hydrocarbon contaminated oily sludge: prospects for in situ bioremediation. Environ Sci Pollut Res 21:7369–7389. https://doi.org/10.1007/s11356-014-2640-2
Dashti N, Ali N, Eliyas M, Khanafer M, Sorkhoh NA, Radwan SS (2015) Most hydrocarbonoclastic bacteria in the total environment are diazotrophic, which highlights their value in the bioremediation of hydrocarbon contaminants. Microbes Environ 30:70–75. https://doi.org/10.1264/jsme2.ME14090
Davoodi SM, Miri S, Taheran M, Brar SK, Galvez-Cloutier R, Martel R (2020) Bioremediation of unconventional oil contaminated ecosystems under natural and assisted conditions: a review. Environ Sci Technol 54:2054–2067. https://doi.org/10.1021/acs.est.9b00906
Delille D, Coulon F, Pelletier E (2004) Effects of temperature warming during a bioremediation study of natural and nutrient-amended hydrocarbon-contaminated sub-Antarctic soils. Cold Reg Sci Technol 40:61–70. https://doi.org/10.1016/j.coldregions.2004.05.005
De’Lorenzo V, (2008) Systems biology approaches to bioremediation. Curr Opin Biotech 19:579–589. https://doi.org/10.1016/j.copbio.2008.10.004
Dou J, Liu X, Hu Z, Deng D (2008) Anaerobic BTEX biodegradation linked to nitrate and sulfate reduction. J Hazard Mater 151:720–729. https://doi.org/10.1016/j.jhazmat.2007.06.043
Duarte M, Nielsen A, Camarinha Silva A, Vilchez Vargas R, Bruls T, Wos Oxley ML, Jauregui R, Pieper DH (2017) Functional soil metagenomics: elucidation of polycyclic aromatic hydrocarbon degradation potential following 12 years of in situ bioremediation. Environ Microbiol 19:2992–3011. https://doi.org/10.1111/1462-2920.13756
Dueholm MS, Marques IG, Karst SM, D’Imperio S, Tale VP, Lewis D, Nielsen PH, Nielsen JL (2015) Survival and activity of individual bioaugmentation strains. Bioresour Technol 186:192–199. https://doi.org/10.1016/j.biortech.2015.02.111
Fowler SJ, Toth CRA, Gieg LM (2016) Community structure in methanogenic enrichments provides insight into syntrophic interactions in hydrocarbon-impacted environments. Front Microbiol 7:562. https://doi.org/10.3389/fmicb.2016.00562
Gray ND, Sherry A, Grant RJ, Rowan AK, Hubert CR, Callbeck CM, Aitken CM, Jones DM, Adams JJ, Larter SR, Head IM (2011) The quantitative significance of Syntrophaceae and syntrophic partnerships in methanogenic degradation of crude oil alkanes. Environ Microbiol 13:2957–2975. https://doi.org/10.1111/j.1462-2920.2011.02570.x
Guerra AB, Oliveira JS, Silva-Portela RC, Araújo W, Carlos AC, Vasconcelos AT, Freitas AT, Domingos YS, de Farias MF, Fernandes GJ, Agnez-Lima LF (2018) Metagenome enrichment approach used for selection of oil-degrading bacteria consortia for drill cutting residue bioremediation. Environ Pollut 235:869–880. https://doi.org/10.1016/j.envpol.2018.01.014
Gupta A, Dutta A, Sarkar J, Panigrahi MK, Sar P (2018) Low-abundance members of the Firmicutes facilitate bioremediation of soil impacted by highly acidic mine drainage from the Malanjkhand copper project India. Front Microbiol 9:2882. https://doi.org/10.3389/fmicb.2018.02882
Hazen TC, Dubinsky EA, DeSantis TZ, Andersen GL, Piceno YM, Singh N, Jansson JK, Probst A, Borglin SE, Fortney JL, Stringfellow WT (2010) Deep-sea oil plume enriches indigenous oil-degrading bacteria. Science 330:204–208. https://doi.org/10.1126/science.1195979
Hazen TC, Prince RC, Mahmoudi N (2016) Marine oil biodegradation. Environ Sci Technol 50:2121–2129. https://doi.org/10.1021/acs.est.5b03333
Hidaka K, Miyanaga K, Tanji Y (2018) The presence of nitrate-and sulfate-reducing bacteria contributes to ineffectiveness souring control by nitrate injection. Int Biodeter Biodegr 129:81–88. https://doi.org/10.1016/j.ibiod.2018.01.007
Katarína D, Slavomíra M, Hana D, Katarína L, Hana H (2018) The adaptation mechanisms of bacteria applied in bioremediation of hydrophobic toxic environmental pollutants: how indigenous and introduced bacteria can respond to persistent organic pollutants-induced stress? In: Donyinah SK (ed) Persistent organic pollutants. IntechOpen, London, pp 71–97
Kuyukina MS, Krivoruchko AV, Ivshina IB (2020) Advanced bioreactor treatments of hydrocarbon-containing wastewater. Appl Sci 10:831–849. https://doi.org/10.3390/app10030831
Lamendella R, Strutt S, Borglin SE, Chakraborty R, Tas N, Mason OU, Hultman J, Prestat E, Hazen TC, Jansson J (2014) Assessment of the deepwater horizon oil spill impact on gulf coast microbial communities. Front Microbiol 5:130. https://doi.org/10.3389/fmicb.2014.00130
Ławniczak Ł, Woźniak-Karczewska M, Loibner AP, Heipieper HJ, Chrzanowski Ł (2020) Microbial degradation of hydrocarbons—Basic principles for bioremediation: a review. Molecules 25:856. https://doi.org/10.3390/molecules25040856
Lee C, Lee S, Shin SG, Hwang S (2008) Real-time PCR determination of rRNA gene copy number: absolute and relative quantification assays with Escherichia coli. Appl Microbiol Biotechnol 78:371–376. https://doi.org/10.1007/s00253-007-1300-6
Lee Y, Lee Y, Jeon CO (2019) Biodegradation of naphthalene, BTEX, and aliphatic hydrocarbons by Paraburkholderia aromaticivorans BN5 isolated from petroleum-contaminated soil. Sci Rep 9:860. https://doi.org/10.1038/s41598-018-36165-x
Liu Q, Tang J, Bai Z, Hecker M, Giesy JP (2015) Distribution of petroleum degrading genes and factor analysis of petroleum-contaminated soil from the Dagang oil field, China. Sci Rep 5:11068. https://doi.org/10.1038/srep11068
Liu H, Tan X, Guo J, Liang X, Xie Q, Chen S (2020) Bioremediation of oil-contaminated soil by combination of soil conditioner and microorganism. J Soils Sediment 20:2121–2129. https://doi.org/10.1007/s11368-020-02591-6
Lladó S, Covino S, Solanas AM, Petruccioli M, D’annibale A, Viñas M, (2015) Pyrosequencing reveals the effect of mobilizing agents and lignocellulosic substrate amendment on microbial community composition in a real industrial PAH-polluted soil. J Hazard Mater 283:35–43. https://doi.org/10.1016/j.jhazmat.2014.08.065
Mason OU, Scott NM, Gonzalez A, Robbins-Pianka A, Bælum J, Kimbrel J, Bouskill NJ, Prestat E, Borglin S, Joyner DC, Fortney JL (2014) Metagenomics reveals sediment microbial community response to Deepwater Horizon oil spill. ISME J 8:1464–1475. https://doi.org/10.1038/ismej.2013.254
Martirani-Von Abercron SM, Marín P, Solsona-Ferraz M, Castañeda-Cataña MA, Marqués S (2017) Naphthalene biodegradation under oxygen-limiting conditions: community dynamics and the relevance of biofilm-forming capacity. Microb Biotechnol 10:1781–1796. https://doi.org/10.1111/1751-7915.12842
Mechichi T, Stackebrandt E, Fuchs G (2003) Alicycliphilus denitrificans gen. nov., sp. nov., a cyclohexanol-degrading, nitrate-reducing β-proteobacterium. Int J Syst Evol Microbiol 53:147–152. https://doi.org/10.1099/ijs.0.02276-0
Mishra B, Varjani S, Kumar G, Awasthi MK, Awasthi SK, Sindhu R, Binod P, Rene ER, Zhang Z (2020) Microbial approaches for remediation of pollutants: innovations, future outlook, and challenges. Energy Environ. https://doi.org/10.1177/0958305X19896781
Mühling M, Woolven-Allen J, Murrell JC, Joint I (2008) Improved group-specific PCR primers for denaturing gradient gel electrophoresis analysis of the genetic diversity of complex microbial communities. ISME J 2:379–392. https://doi.org/10.1038/ismej.2007.97
Muyzer G (1993) Profiling of complex microbial populations by denaturing gradient gel electrophoresis analysis of polymerase chain reaction-amplified genes coding for 16S rRNA. Appl Environ Microbiol 59:695–700. https://doi.org/10.1128/AEM.59.3.695-700.1993
Naeem U, Qazi MA (2020) Leading edges in bioremediation technologies for removal of petroleum hydrocarbons. Environ Sci Pollut Res 27:27370–27382. https://doi.org/10.1007/s11356-019-06124-8
Noguchi M, Kurisu F, Kasuga I, Furumai H (2014) Time-resolved DNA stable isotope probing links Desulfobacterales-and Coriobacteriaceae-related bacteria to anaerobic degradation of benzene under methanogenic conditions. Microbes Environ 29:191–199. https://doi.org/10.1264/jsme2.ME13104
Poi G, Aburto-Medina A, Mok PC, Ball AS, Shahsavari E (2017) Large scale bioaugmentation of soil contaminated with petroleum hydrocarbons using a mixed microbial consortium. Ecol Eng 102:64–71. https://doi.org/10.1016/j.ecoleng.2017.01.048
Poursat BA, van Spanning RJ, de Voogt P, Parsons JR (2019) Implications of microbial adaptation for the assessment of environmental persistence of chemicals. Crit Rev Environ Sci Technol 49:2220–2255. https://doi.org/10.1080/10643389.2019.1607687
Purkamo L, Bomberg M, Kietäväinen R, Salavirta H, Nyyssönen M, Nuppunen-Puputti M, Ahonen L, Kukkonen I, Itävaara M (2016) Microbial co-occurrence patterns in deep Precambrian bedrock fracture fluids. Biogeosciences 13:3091–3108. https://doi.org/10.5194/bg-13-3091-2016
Qiu T, Zuo Z, Gao J, Gao M, Han M, Sun L, Zhang L, Wang X (2015) Diaphorobacter polyhydroxybutyrativorans sp. nov., a novel poly (3-hydroxybutyrate-co-3-hydroxyvalerate)-degrading bacterium isolated from biofilms. Int J Syst Evol Microbiol 65:2913–2918. https://doi.org/10.1099/ijs.0.000353
Révész F, Figueroa-Gonzalez PA, Probst AJ, Kriszt B, Banerjee S, Szoboszlay S, Maróti G, Táncsics A (2020) Microaerobic conditions caused the overwhelming dominance of Acinetobacter spp. and the marginalization of Rhodococcus spp. in diesel fuel/crude oil mixture-amended enrichment cultures. Arch Microbiol 202:329–342. https://doi.org/10.1007/s00203-019-01749-2
Rios-Hernandez LA, Gieg LM, Suflita JM (2003) Biodegradation of an alicyclic hydrocarbon by a sulfate-reducing enrichment from a gas condensate-contaminated aquifer. Appl Environ Microbiol 69:434–443. https://doi.org/10.1128/AEM.69.1.434-443.2003
Rockne KJ, Strand SE (2001) Anaerobic biodegradation of naphthalene, phenanthrene, and biphenyl by a denitrifying enrichment culture. Water Res 35:291–299. https://doi.org/10.1016/S0043-1354(00)00246-3
Rojo F (2009) Degradation of alkanes by bacteria. Environ Microbiol 11:2477–2490. https://doi.org/10.1111/j.1462-2920.2009.01948.x
Rotaru AE, Probian C, Wilkes H, Harder J (2010) Highly enriched Betaproteobacteria growing anaerobically with p-xylene and nitrate. FEMS Microbiol Ecol 71:460–468. https://doi.org/10.1111/j.1574-6941.2009.00814.x
Roy A, Dutta A, Pal S, Gupta A, Sarkar J, Chatterjee A, Saha A, Sarkar P, Sar P, Kazy SK (2018a) Biostimulation and bioaugmentation of native microbial community accelerated bioremediation of oil refinery sludge. Bioresour Technol 253:22–32. https://doi.org/10.1016/j.biortech.2018.01.004
Roy A, Sar P, Sarkar J, Dutta A, Sarkar P, Gupta A, Mohapatra B, Pal S, Kazy SK (2018b) Petroleum hydrocarbon rich oil refinery sludge of North-East India harbours anaerobic, fermentative, sulfate-reducing, syntrophic and methanogenic microbial populations. BMC Microbiol 18:151. https://doi.org/10.1186/s12866-018-1275-8
Sarkar J, Kazy SK, Gupta A, Dutta A, Mohapatra B, Roy A, Bera P, Mitra A, Sar P (2016) Biostimulation of indigenous microbial community for bioremediation of petroleum refinery sludge. Front Microbiol 7:1407. https://doi.org/10.3389/fmicb.2016.01407
Sarkar P, Roy A, Pal S, Mohapatra B, Kazy SK, Maiti MK, Sar P (2017) Enrichment and characterization of hydrocarbon-degrading bacteria from petroleum refinery waste as potent bioaugmentation agent for in situ bioremediation. Bioresour Technol 242:15–27. https://doi.org/10.1016/j.biortech.2017.05.010
Shankar S, Kansrajh C, Dinesh MG, Satyan RS, Kiruthika S, Tharanipriya A (2014) Application of indigenous microbial consortia in bioremediation of oil-contaminated soils. Int J Environ Sci Technol 11:367–376. https://doi.org/10.1007/s13762-013-0366-1
Shin B, Kim M, Zengler K, Chin KJ, Overholt WA, Gieg LM, Konstantinidis KT, Kostka JE (2019) Anaerobic degradation of hexadecane and phenanthrene coupled to sulfate reduction by enriched consortia from northern Gulf of Mexico seafloor sediment. Sci Rep 9:1239. https://doi.org/10.1038/s41598-018-36567-x
Siles JA, Margesin R (2018) Insights into microbial communities mediating the bioremediation of hydrocarbon-contaminated soil from an Alpine former military site. Appl Microbiol Biotechnol 102:4409–4421. https://doi.org/10.1007/s00253-018-8932-6
Singleton DR, Adrion AC, Aitken MD (2016) Surfactant-induced bacterial community changes correlated with increased polycyclic aromatic hydrocarbon degradation in contaminated soil. Appl Environ Microbiol 100:10165–10177. https://doi.org/10.1007/s00253-016-7867-z
Sperfeld M, Rauschenbach C, Diekert G, Studenik S (2018) Microbial community of a gasworks aquifer and identification of nitrate-reducing Azoarcus and Georgfuchsia as key players in BTEX degradation. Water Res 132:146–157. https://doi.org/10.1016/j.watres.2017.12.040
Spini G, Spina F, Poli A, Blieux AL, Regnier T, Gramellini C, Varese GC, Puglisi E (2018) Molecular and microbiological insights on the enrichment procedures for the isolation of petroleum degrading bacteria and fungi. Front Microbiol 9:2543. https://doi.org/10.3389/fmicb.2018.02543
Sutton NB, Maphosa F, Morillo JA, Al-Soud WA, Langenhoff AA, Grotenhuis T, Rijnaarts HH, Smidt H (2013) Impact of long-term diesel contamination on soil microbial community structure. Appl Environ Microbiol 79:619–630. https://doi.org/10.1128/AEM.02747-12
Szulc A, Ambrożewicz D, Sydow M, Ławniczak Ł, Piotrowska-Cyplik A, Marecik R, Chrzanowski Ł (2014) The influence of bioaugmentation and biosurfactant addition on bioremediation efficiency of diesel-oil contaminated soil: feasibility during field studies. J Environ Manag 132:121–128. https://doi.org/10.1016/j.jenvman.2013.11.006
Tan B, Fowler SJ, Laban NA, Dong X, Sensen CW, Foght J, Gieg LM (2015) Comparative analysis of metagenomes from three methanogenic hydrocarbon-degrading enrichment cultures with 41 environmental samples. ISME J 9:2028–2045. https://doi.org/10.1038/ismej.2015.22
Tao K, Liu X, Chen X, Hu X, Cao L, Yuan X (2017) Biodegradation of crude oil by a defined co-culture of indigenous bacterial consortium and exogenous Bacillus subtilis. Bioresour Technol 224:327–332. https://doi.org/10.1016/j.biortech.2016.10.073
Tyagi M, da Fonseca MM, de Carvalho CC (2011) Bioaugmentation and biostimulation strategies to improve the effectiveness of bioremediation processes. Biodegradation 22:231–241. https://doi.org/10.1007/s10532-010-9394-4
Varjani SJ (2017) Microbial degradation of petroleum hydrocarbons. Bioresour Technol 223:277–286. https://doi.org/10.1016/j.biortech.2016.10.037
Wang H, Wang B, Dong W, Hu X (2016) Co-acclimation of bacterial communities under stresses of hydrocarbons with different structures. Sci Rep 6:34588. https://doi.org/10.1038/srep34588
Wallisch S, Gril T, Dong X, Welzl G, Burns C, Heath E, Marion E, Suhadolc M, Schloeter M (2014) Effects of different compost amendments on the abundance and composition of alkB harboring bacterial communities in a soil under industrial use contaminated with hydrocarbons. Front Microbiol 5:96. https://doi.org/10.3389/fmicb.2014.00096
Wolterink A, Kim S, Muusse M, Kim IS, Roholl PJ, van Ginkel CG, Stams AJ, Kengen SW (2005) Dechloromonas hortensis sp. nov. and strain ASK-1, two novel (per) chlorate-reducing bacteria, and taxonomic description of strain GR-1. Int J Syst Evol Microbiol 55:2063–2068. https://doi.org/10.1099/ijs.0.63404-0
Wu M, Dick WA, Li W, Wang X, Yang Q, Wang T, Xu L, Zhang M, Chen L (2016) Bioaugmentation and biostimulation of hydrocarbon degradation and the microbial community in a petroleum-contaminated soil. Int Biodeterior Biodegr 107:158–164. https://doi.org/10.1016/j.ibiod.2015.11.019
Xu GL, Liu H, Li MJ, Li ZM, Peng ZH, Zuo LM, He X, Liu WW, Cai LG (2016) In situ bioremediation of crude oil contaminated site: a case study in Jianghan oil field, China. Petrol Sci Technol 34:63–70. https://doi.org/10.1080/10916466.2015.1115873
Xu X, Liu W, Tian S, Wang W, Qi Q, Jiang P, Gao X, Li F, Li H, Yu H (2018) Petroleum hydrocarbon-degrading bacteria for the remediation of oil pollution under aerobic conditions: a perspective analysis. Front Microbiol 9:2885. https://doi.org/10.3389/fmicb.2018.02885
Yang S, Wen X, Zhao L, Shi Y, Jin H (2014) Crude oil treatment leads to shift of bacterial communities in soils from the deep active layer and upper permafrost along the China-Russia crude oil pipeline route. PLoS ONE 9:e96552. https://doi.org/10.1371/journal.pone.0096552
Yergeau E, Lawrence JR, Sanschagrin S, Waiser MJ, Korber DR, Greer CW (2012) Next-generation sequencing of microbial communities in the Athabasca River and its tributaries in relation to oil sands mining activities. Appl Environ Microb 78:7626–7637. https://doi.org/10.1128/AEM.02036-12
Young CC, Ho MJ, Arun AB, Chen WM, Lai WA, Shen FT, Rekha PD, Yassin AF (2007) Pseudoxanthomonas spadix sp. nov., isolated from oil-contaminated soil. Int J Syst Evol Microbiol 57:1823–1827. https://doi.org/10.1099/ijs.0.65053-0
Zedelius J, Rabus R, Grundmann O, Werner I, Brodkorb D, Schreiber F, Ehrenreich P, Behrends A, Wilkes H, Kube M, Reinhardt R (2011) Alkane degradation under anoxic conditions by a nitrate reducing bacterium with possible involvement of the electron acceptor in substrate activation. Environ Microbiol Rep 3:125–135. https://doi.org/10.1111/j.1758-2229.2010.00198.x
Zhang Z, Lo IM, Yan DY (2015) An integrated bioremediation process for petroleum hydrocarbons removal and odor mitigation from contaminated marine sediment. Water Res 83:21–30. https://doi.org/10.1016/j.watres.2015.06.022
Zhou L, Li H, Zhang Y, Han S, Xu H (2016) Sphingomonas from petroleum-contaminated soils in Shenfu, China and their PAHs degradation abilities. Braz J Microbiol 47:271–278. https://doi.org/10.1016/j.bjm.2016.01.001
Acknowledgements
Authors acknowledge the financial support provided by Department of Biotechnology, Government of India under NER Twinning project (BT/226/NE/TBP/2011). The generous help from Noonmati Refinery (Indian Oil Corporation Limited), Guwahati, Assam, India for providing refinery waste sludge is gratefully acknowledged.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Sarkar, J., Saha, A., Roy, A. et al. Development of nitrate stimulated hydrocarbon degrading microbial consortia from refinery sludge as potent bioaugmenting agent for enhanced bioremediation of petroleum contaminated waste. World J Microbiol Biotechnol 36, 156 (2020). https://doi.org/10.1007/s11274-020-02925-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11274-020-02925-z